
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MDM2-BRI3BP (FusionGDB2 ID:52391) |
Fusion Gene Summary for MDM2-BRI3BP |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MDM2-BRI3BP | Fusion gene ID: 52391 | Hgene | Tgene | Gene symbol | MDM2 | BRI3BP | Gene ID | 4193 | 140707 |
| Gene name | MDM2 proto-oncogene | BRI3 binding protein | |
| Synonyms | ACTFS|HDMX|LSKB|hdm2 | BNAS1|HCCR-1|HCCR-2|HCCRBP-1|HCCRBP-3|KG19 | |
| Cytomap | 12q15 | 12q24.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | E3 ubiquitin-protein ligase Mdm2MDM2 oncogene, E3 ubiquitin protein ligaseMDM2 proto-oncogene, E3 ubiquitin protein ligaseMdm2, p53 E3 ubiquitin protein ligase homologMdm2, transformed 3T3 cell double minute 2, p53 binding proteindouble minute 2, hum | BRI3-binding proteinI3-binding proteincervical cancer 1 proto-oncogene-binding protein KG19cervical cancer oncogene binding protein | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | . | Q8WY22 | |
| Ensembl transtripts involved in fusion gene | ENST00000478070, ENST00000258148, ENST00000258149, ENST00000350057, ENST00000462284, ENST00000299252, ENST00000348801, ENST00000356290, ENST00000360430, ENST00000393410, ENST00000393412, ENST00000393413, ENST00000428863, ENST00000517852, ENST00000540827, ENST00000544125, ENST00000544561, ENST00000545204, | ENST00000341446, | |
| Fusion gene scores | * DoF score | 39 X 20 X 11=8580 | 8 X 8 X 4=256 |
| # samples | 47 | 9 | |
| ** MAII score | log2(47/8580*10)=-4.19024498582191 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/256*10)=-1.50814690367033 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MDM2 [Title/Abstract] AND BRI3BP [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MDM2(69214154)-BRI3BP(125509890), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | MDM2-BRI3BP seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. MDM2-BRI3BP seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MDM2 | GO:0000122 | negative regulation of transcription by RNA polymerase II | 9271120|17310983 |
| Hgene | MDM2 | GO:0006511 | ubiquitin-dependent protein catabolic process | 11278372|15314173|16173922|17310983 |
| Hgene | MDM2 | GO:0016567 | protein ubiquitination | 9450543|15878855|19656744|20153724 |
| Hgene | MDM2 | GO:0031648 | protein destabilization | 9529249|10360174|15314173 |
| Hgene | MDM2 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process | 11278372 |
| Hgene | MDM2 | GO:0034504 | protein localization to nucleus | 10360174 |
| Hgene | MDM2 | GO:0042176 | regulation of protein catabolic process | 9153395 |
| Hgene | MDM2 | GO:0043518 | negative regulation of DNA damage response, signal transduction by p53 class mediator | 9529249|10360174 |
| Hgene | MDM2 | GO:0045184 | establishment of protein localization | 10360174 |
| Hgene | MDM2 | GO:0045892 | negative regulation of transcription, DNA-templated | 9271120 |
| Hgene | MDM2 | GO:0065003 | protein-containing complex assembly | 10608892|12915590 |
| Hgene | MDM2 | GO:0071157 | negative regulation of cell cycle arrest | 9529249 |
| Hgene | MDM2 | GO:0071480 | cellular response to gamma radiation | 16213212 |
| Hgene | MDM2 | GO:0072717 | cellular response to actinomycin D | 15314173 |
| Hgene | MDM2 | GO:1901797 | negative regulation of signal transduction by p53 class mediator | 16173922 |
 Fusion gene breakpoints across MDM2 (5'-gene) Fusion gene breakpoints across MDM2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
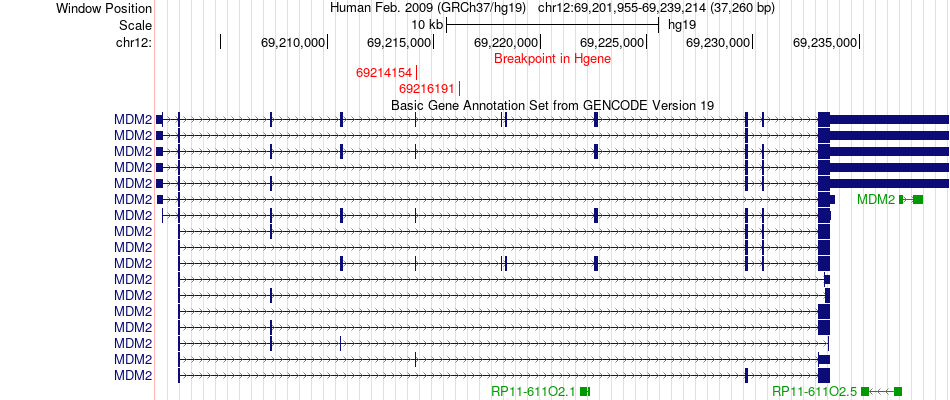 |
 Fusion gene breakpoints across BRI3BP (3'-gene) Fusion gene breakpoints across BRI3BP (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
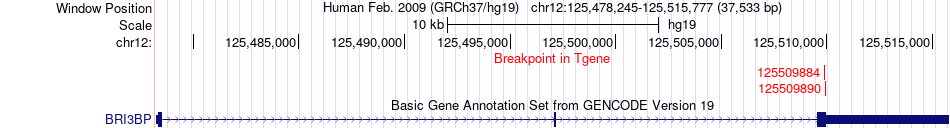 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PRAD | TCGA-KK-A7B2-01A | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| ChimerDB4 | PRAD | TCGA-KK-A7B2 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| ChimerDB4 | PRAD | TCGA-KK-A7B2 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
Top |
Fusion Gene ORF analysis for MDM2-BRI3BP |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000478070 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| In-frame | ENST00000258148 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| In-frame | ENST00000258149 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| In-frame | ENST00000350057 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| In-frame | ENST00000462284 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000258148 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000258149 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000299252 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000299252 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000348801 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000348801 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000350057 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000356290 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000356290 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000360430 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000360430 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000393410 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000393410 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000393412 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000393412 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000393413 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000393413 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000428863 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000428863 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000462284 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000478070 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000517852 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000517852 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000540827 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000540827 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000544125 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000544125 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000544561 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000544561 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
| intron-3CDS | ENST00000545204 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + |
| intron-3CDS | ENST00000545204 | ENST00000341446 | MDM2 | chr12 | 69216191 | + | BRI3BP | chr12 | 125509884 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000462284 | MDM2 | chr12 | 69214154 | + | ENST00000341446 | BRI3BP | chr12 | 125509890 | + | 6547 | 660 | 266 | 745 | 159 |
| ENST00000258149 | MDM2 | chr12 | 69214154 | + | ENST00000341446 | BRI3BP | chr12 | 125509890 | + | 6532 | 645 | 251 | 730 | 159 |
| ENST00000258148 | MDM2 | chr12 | 69214154 | + | ENST00000341446 | BRI3BP | chr12 | 125509890 | + | 6259 | 372 | 14 | 457 | 147 |
| ENST00000350057 | MDM2 | chr12 | 69214154 | + | ENST00000341446 | BRI3BP | chr12 | 125509890 | + | 6152 | 265 | 2591 | 2232 | 119 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000462284 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + | 0.006561065 | 0.9934389 |
| ENST00000258149 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + | 0.006466644 | 0.9935334 |
| ENST00000258148 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + | 0.005607127 | 0.9943929 |
| ENST00000350057 | ENST00000341446 | MDM2 | chr12 | 69214154 | + | BRI3BP | chr12 | 125509890 | + | 0.021022916 | 0.9789771 |
Top |
Fusion Genomic Features for MDM2-BRI3BP |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
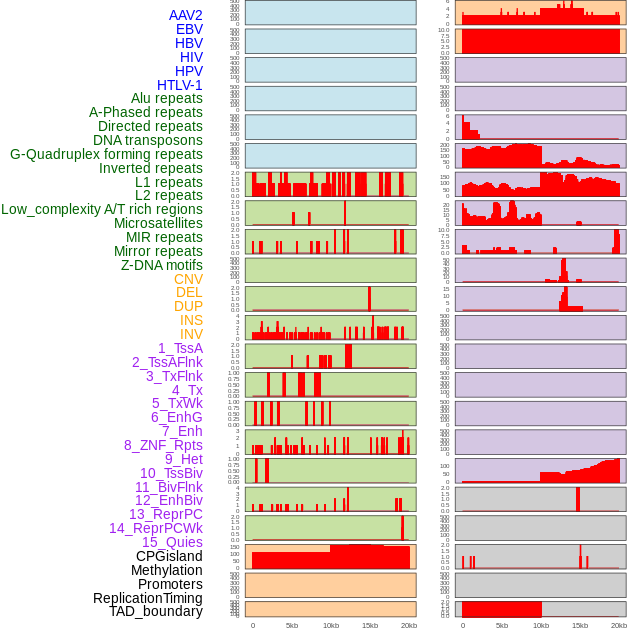 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for MDM2-BRI3BP |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:69214154/chr12:125509890) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | BRI3BP |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Involved in tumorigenesis and may function by stabilizing p53/TP53. {ECO:0000269|PubMed:17943721}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 26_109 | 113 | 437.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 26_109 | 119 | 498.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 1_101 | 113 | 437.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 1_101 | 119 | 498.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Tgene | BRI3BP | chr12:69214154 | chr12:125509890 | ENST00000341446 | 0 | 3 | 217_247 | 0 | 252.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | BRI3BP | chr12:69214154 | chr12:125509890 | ENST00000341446 | 0 | 3 | 125_145 | 0 | 252.0 | Transmembrane | Helical | |
| Tgene | BRI3BP | chr12:69214154 | chr12:125509890 | ENST00000341446 | 0 | 3 | 13_33 | 0 | 252.0 | Transmembrane | Helical | |
| Tgene | BRI3BP | chr12:69214154 | chr12:125509890 | ENST00000341446 | 0 | 3 | 146_166 | 0 | 252.0 | Transmembrane | Helical | |
| Tgene | BRI3BP | chr12:69214154 | chr12:125509890 | ENST00000341446 | 0 | 3 | 185_205 | 0 | 252.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 210_215 | 113 | 437.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 243_301 | 113 | 437.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 210_215 | 0 | 322.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 243_301 | 0 | 322.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 210_215 | 0 | 266.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 243_301 | 0 | 266.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 210_215 | 0 | 322.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 243_301 | 0 | 322.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 210_215 | 0 | 297.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 243_301 | 0 | 297.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 210_215 | 0 | 219.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 243_301 | 0 | 219.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 210_215 | 0 | 219.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 243_301 | 0 | 219.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 210_215 | 0 | 271.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 243_301 | 0 | 271.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 210_215 | 119 | 498.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 243_301 | 119 | 498.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 210_215 | 0 | 297.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 243_301 | 0 | 297.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 26_109 | 0 | 322.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 26_109 | 0 | 266.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 26_109 | 0 | 322.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 26_109 | 0 | 297.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 26_109 | 0 | 219.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 26_109 | 0 | 219.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 26_109 | 0 | 271.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 26_109 | 0 | 297.0 | Domain | SWIB/MDM2 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 179_185 | 113 | 437.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 190_202 | 113 | 437.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 466_473 | 113 | 437.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 179_185 | 0 | 322.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 190_202 | 0 | 322.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 466_473 | 0 | 322.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 179_185 | 0 | 266.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 190_202 | 0 | 266.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 466_473 | 0 | 266.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 179_185 | 0 | 322.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 190_202 | 0 | 322.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 466_473 | 0 | 322.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 179_185 | 0 | 297.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 190_202 | 0 | 297.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 466_473 | 0 | 297.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 179_185 | 0 | 219.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 190_202 | 0 | 219.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 466_473 | 0 | 219.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 179_185 | 0 | 219.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 190_202 | 0 | 219.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 466_473 | 0 | 219.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 179_185 | 0 | 271.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 190_202 | 0 | 271.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 466_473 | 0 | 271.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 179_185 | 119 | 498.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 190_202 | 119 | 498.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 466_473 | 119 | 498.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 179_185 | 0 | 297.0 | Motif | Nuclear localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 190_202 | 0 | 297.0 | Motif | Note=Nuclear export signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 466_473 | 0 | 297.0 | Motif | Nucleolar localization signal |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 210_304 | 113 | 437.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 242_331 | 113 | 437.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 1_101 | 0 | 322.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 210_304 | 0 | 322.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 242_331 | 0 | 322.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 1_101 | 0 | 266.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 210_304 | 0 | 266.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 242_331 | 0 | 266.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 1_101 | 0 | 322.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 210_304 | 0 | 322.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 242_331 | 0 | 322.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 1_101 | 0 | 297.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 210_304 | 0 | 297.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 242_331 | 0 | 297.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 1_101 | 0 | 219.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 210_304 | 0 | 219.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 242_331 | 0 | 219.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 1_101 | 0 | 219.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 210_304 | 0 | 219.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 242_331 | 0 | 219.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 1_101 | 0 | 271.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 210_304 | 0 | 271.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 242_331 | 0 | 271.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 210_304 | 119 | 498.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 242_331 | 119 | 498.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 1_101 | 0 | 297.0 | Region | Sufficient to promote the mitochondrial pathway of apoptosis |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 210_304 | 0 | 297.0 | Region | Note=ARF-binding |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 242_331 | 0 | 297.0 | Region | Note=Region II |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 299_328 | 113 | 437.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 438_479 | 113 | 437.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 299_328 | 0 | 322.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 438_479 | 0 | 322.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 299_328 | 0 | 266.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 438_479 | 0 | 266.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 299_328 | 0 | 322.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 438_479 | 0 | 322.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 299_328 | 0 | 297.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 438_479 | 0 | 297.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 299_328 | 0 | 219.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 438_479 | 0 | 219.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 299_328 | 0 | 219.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 438_479 | 0 | 219.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 299_328 | 0 | 271.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 438_479 | 0 | 271.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 299_328 | 119 | 498.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 438_479 | 119 | 498.0 | Zinc finger | RING-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 299_328 | 0 | 297.0 | Zinc finger | RanBP2-type |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 438_479 | 0 | 297.0 | Zinc finger | RING-type |
Top |
Fusion Gene Sequence for MDM2-BRI3BP |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >52391_52391_1_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000258148_BRI3BP_chr12_125509890_ENST00000341446_length(transcript)=6259nt_BP=372nt CGCGAAAACCCCGGATGGTGAGGAGCAGGCAAATGTGCAATACCAACATGTCTGTACCTACTGATGGTGCTGTAACCACCTCACAGATTC CAGCTTCGGAACAAGAGACCCTGGTTAGACCAAAGCCATTGCTTTTGAAGTTATTAAAGTCTGTTGGTGCACAAAAAGACACTTATACTA TGAAAGAGGTTCTTTTTTATCTTGGCCAGTATATTATGACTAAACGATTATATGATGAGAAGCAACAACATATTGTATATTGTTCAAATG ATCTTCTAGGAGATTTGTTTGGCGTGCCAAGCTTCTCTGTGAAAGAGCACAGGAAAATATATACCATGATCTACAGGAACTTGGTAGTAG TCAATCAGCAGGTGGAGCACCTGGAGAAGCAGGTCAGACTGCTCAACATCCGTCTCAACCGGGTGCTCGAGAGCCTGGACCGCTCCAAGG ACAAGTGAAGGTCAGCCGGCCGGGCGGGTCCACAGTTACCAGCACGCTGTCTCAGAAAACGAAAACGGAGGAAAAAAACCCCAAACCCCA AACAATCTTAATAAACACGACTGAGCAAGAAAGTGGCGCTGTGTAGGGCTATTTCCACCCACCCGGCAGCTCTTAGGACACATTCCCAGA AGAGCGGAAAGATCATTGACGTGGAACTACACACGAAGTGTAATTAGTGGGGGAAAAAATATTTTTTAAACAAAGGATATAACCATATTT AGTTGTACAGTAAGAGAAATTTATCTGTGCATAGAGCATAAAGTTAATTTTTTCAAGCATTTAAATACATCTTTTGTAAGGTTTTTTAAT AAAGGCAGATTGAGTCAAGTTTATTAAAACTTTACATGGAGGGGAAGAATATGCTAATTTGACATTTGAGAAAAGTTTAAATGCGAGCGT TTGTTTTCACAGCTTAGGGATTGGTGGGTGAGGTCTGTGTGCGTTATTTTCATGTTCCCAAACTGTGAGCTTTATTTCATGTTATTTTTG GTGGATGTTGACTTAGTTCATGAATGTTGTCACTTCTAAATGTGGACAGTTGGGCTGCAGGATGCAACTCAACCTCTTTTGGATCCTAGA GACCCCTGCCAAGCCCCCATCAGTCCTATTTTTAAATGAAGACATCGCCATTTCTAGAAGACCTGAGATTTTCTTCCCTCTGTGCTTCGA TGTATTTTAACCTCAACATTTTGAAACGGAAAATGCAAGGAGGGGCCAAGAAATGTCCTGGGGATTTGATGCCAGGAAGTACCTGGTAAG GATTGGAGATTGCTCCAGAATCTACTGCATGTCCCAATTTGCAGTTTGCTCACTAGGGCTATATACAAAAGCTATACCCTTGAGGGTTTT TTTTTTTTTCTTTTGATTTCCTCAGGACCTTAGAGGGAAAACAAACAGTAGCAGCTAATATTCTCAAGTATATTGCTGCTTAGAAAGATC CTCAGGAACAATTAGCAGCAATAAGCAAGCCTTTGAAAAGATCTGAATTCTTTTTCCTGAAATATTTACGATACACAGGTGCTTTTTATC TGAAATCTGTTGTGTCCTCCTTTTAGGCAGTCTCTGTGGGCAGAGAGTGGGACTTGCGAGGTGGACAGCTGTGGGGATCCTGGGCAAGGG AGTTTCAGAAGGTGTGGCTCAGGGGCTGGTAGAAACCCCCTTTTGTTAGGACTTGTTTTTTAAAAAAGAAAGCCCTAGGCCAGGCGCAGT GGCTGATGCCTGTAATCCCAGCGCTTTGGGAGGCCGAGGAGTATGGATCACCTGAGGTCAGGAGTTCAAGACCAGCCTGGCCAACATGGT GAAACCCCCGTCTCTACTAAAAATACAAAAATTAGTCAGGCATGGTGGTGGGTGCCTGTAGTCCAGCTACTCAGGAGGCTGAGGCAGGAG AATTGCTTGAACCTGGGAGGTGGAGGTTGCGGTGAGCCGAGATCACGCCACTGCACTCCAGCCTAGGCAACAGAGCAAGACTTCATCTTA AAAAAAAAAAAAAAAGCCCAGCATTTTGCAGCTTTGAAGCTGCTGGGGATCACGACTGTCTCTGTGGCCCTTACCTGGCCACAGATGTAC CCTGGCAACCCGCTCACAGTGCCCTGCGTGCAGGTTTATAGATAACCCCTGCCAGACCTGCTACAACCAAGACCTTATGGTCCAGGTCTC AGTTTGGAGAAGATTTAGCTGTCTACCACTTCTTGTGTAGTGTTCCTGTGAAGTTTTATTTTTGGTTGCAAGTATCTGGAAAACAGATGC AGATGTTTTTGTGGATGTTGTTGAGACTTTTGAGACTTCTGTTTCTAAGGTACACATGATGGGACCCAAACTCTCCTGGAGAGTTCCATT CACTTCCTGAAACGTGCACCTGTTTCAGGTGCATGTTTATTCAGTGTGCATGGATATTTAAGATTTGCTGGAGCAGGCTGGGCAGGGTGG CTCACCCCTATAATCCCAACACTTTGGGAGGCCAAGGCAGGTGGATCACCTGAGGTCAGGAGTTTGAGACCAGCCCGGCCAACAGAGTGA AACCCCGTCTCTACTAAAAATACAAAAATTAGCCAGGCCTGGTGGTGGGCGCCTGTGATCCCAGCTACTTGGGAGGCTGAGGCTGGAGAA TCGCTTGAACATGGGAGGCGGAGGTTGCAGTCAGCTGAGATCATACCACCACACTCTAGCCTCAGCAACAGAGTGAGACTCCGTCTCAAA AAAAATTATTTGCTGGATCGAATCTTTTTTTTTTTGAGATGGAGTCTCACTGTCAACCAGGCTGGAGTGCAGTGGCACCATCTCGGCTCA CTGCAAGCTCCGCCTCCTGGGTTCACGCCATTCTCCTACCTCAGCCTCCCAAGTAGCCGGGACTACAGGCGCCCACCATCATGCCCGGCT AATTTTTTATATTTTTAGTAGAGACGGGGTTTCACCGTGTTAGCCAGGATGGTCTCGATCTCCTGACCTCGTGATCCGCCCGCCTCGGCC TCCCAAAGTGCTGGGATTACAGGCGTGAGCCACCGCGCTGGGCCTAGATCAAATCTTTATCCATGCACATTGGAACACAGGATTACTGGG TTGAAATCATTCTAGTTTTGTCATTTAGATACTTGTAGATGAATCTATTTTAGCACAAGGTATAAATAACTCGGGAGGTCATCTCTATCT TCTTTCCTTTTGTGCATTTGGCTATACCACGTTTAGGTACTAAAACAGCTTTGCTTATGTTGGCCAGGGGAAAACATGGCATTCTGTGCG CAAAGCTAATGATCGCCAGCCCTGCCTTGGCCCCTCCCTTGTTTATGGTCATTGTAAGATGCCCGCATGTTAAGGCTTAAGCTGTCACTG GGCTGGGTGTAATACCCGCTTCATTCCTCCTCCCACCCTCTTACCCGAAACATGAAGGGCACTGTGCTCTATTGAGATCTCGATAAGATC ATCATTTTAACTTGTATTCAACTGAGGGAGGTAACATGATATCTGGGTGCTGGTGGATTGAGACCTCAAGCATCAATTCAAAATTGCTGT CGAAAATATGCACTATTTTTTTTTTCCACTCTGGCTAAGTGTAGGCTCTGAAGCCAGTTTTATAGCTTGATACATTCTGGTGAAAGATGT CTTCATTTTTAATGAAAGAACCTACAGTTAGTAGGTGTCACCTCAGGGTAGAAATTCTGATAGAAATTCTGATTATTTGAAACTCTGATA GAAATCCTAGATGCTAGATTTGAAGAAAACCTGGGTTTTATTGCATAGATTTCACCTTTTCTGGGGCACATCTTACCCCAAAATAATTGG GGATTTTTTTGTTCCTTCTCCATTCCATGGAACCTTGTAGGGTGTTGGGATTGGGCCTCCCCTCGCCACCCGCCACATGTCTTTGTTTTT TTCTCTCTTTTGTATGTGGAAAAATGTATTTTTGTACCATATCTCATACCTGGAGAGAAACCTTTTTTCTATCTTCAGGGCTGTAGAATA TATTAGCGTAACACTTCACTGAATGTGACGTAAAAGCATTCTGATGAAAACTGGTTCTGTTATAATTAAGACTCAGTGTTTAAAATATCT TAATCAGTGTTAATTTCCCAGTGTTTCAGTATCACAGTAAATACTTGTTTTACTGAGAACAGAAAATAATCCAGAAAAGGAAAAGGACCA GCCCTCTCTAGATTGCTGGAATTGTATGGATGTGTTGGGGTTGGTTTAGTGATCTCTTCCTTGACACCCTAGCAAAGATGCTTTTGTGGA GTGCAAACCTTTCATAAATATACTTTCCAGGCTGGGTGCAGTGGCTCAAGCTTGTAATCCCAGCACTTTGGGAGGCCAAGGCGGGTGGAT CACTTGCGGTCGGGAGTTCGAGACCAGCCTGGCCAACATAGTGAAATCCCGTCTCTACCAAAAAATACAAAAAAAAAAAAAAAAAAAAAA ATAGCCAGGTGTGGTGGTGGGTTCCTGTAGTCCCAGCTACTCAGGAGGCTGAGGTGGGAGAATTGTATGAACCTGGGAGGCAGAGGTTGC AGTGAACTGAGATCACACCACTGTACTCCAGTTGAGGCGACAGAGTGAGACCCAGTCTCAAAAAAAAAAAAAAGTGTGTGTGTGTGTATA TATATGTATATATATATACACACACACACACTTTCCAATACCAGTCGGCATTACCCGCCCTTTGGTTTCTTTTGCTTTGGATTTCAGCCC CCATATCATTTCTGGTAACAGAGCACAGCAATAGAATTTCTGGGAAGCAGACCTTGTGAATCGTGCCTGTGTCTTAATTCACAAAGGGTA AGTGAATGCCGTTGAAAGGTGTTTCATACACTTACCCCAGAAATGAGAGCCTGTCTCATGCAGGAGTCGGGAGGCAGCGATGTCAGGGAA CTGGGACTATGTCTTTATGACTGTGATGGTGGTAACCAAGTGGGGTGGTCCCCTCTGGGGAGCTGAGTAAGAACGCAAAGGTTTGGGTCA GTGATTAGGAAGACTATAATATGACTTTGTGCATGCCCGGGAGGGCTGCCTTGTAGAGAGGATGTGAGCAGCTTAGTCGCTCATCTGGCC CTGTGATTCAGGCTTATGGAGCGTTAAGAATAACAGCTGTCAAATGGCCTAGACATGGTTAATGCAATTTGTTGCTAGTGGAAATCCTGA ATTGCTTCCTTTCTGTGATCACTGCTACTTCTTAAGATGCTTTTGATGAATGTCATCTGCCTTACAAGTTGACACCTGATAACTTCTCCC TGATGGGTTTCCGAACTGGCTGACTTAACCAAAAAGCCAGCTCTTGCCATCTATCTTGCATTAAAAGGAATTCCTGAGGCTCCTAAGGGG TCAGCTGCCCCACTCCTGACTTTTTTATTTTTAATGGTCTATACCTTCTGCAACATTTTTGTTTATGGCCATTTGAATAGTTGGACTTTG ACTCCTCACTTGTGAATAATAGGAATATATTTTGCAGAATCTAACATAATACCCTTAAAATTCATACTGGACAACCATCAGTGTGATGTA TAGTATCTGGTGTAAACAAATTTTATTCAGCATATTAAATTATTCTGTGTTTTGCTTTTCCTTGATATGTAGGAGGTGCACCAGTACCAG TTTTTCTCTTGTGGTGGCTTAAAACGCTGATTGCCATTTGCATTGTCTACTGAAAAGCTGTGGCAGCGAGTGGTATTATTGACCATGTGT TCTCATGTCTTTTTGCAGATTTAAAGGTTTTATTGTAGTGTCCATTTAATATCTGCATGTGTGTTTAATCTCTGAATACCATGTAATTAA ACTGATTAACTTGTGATGGTGATGGTGATTAGCACACACTAAAAACCCAATGAAGCAATTGTCTCCCTCTTCTTAACTGGAAATCACACT TGTGAGGTATGGTATCCATTATCAACAGTGGATTCATGATTGACTTCATATTTTGTGTATCTGGTTATAAATTTTGTATAGCATTTAATT TGGCACTTAATATAAATCCATTTGATTTGAATAGTGTGTGTGTGTACATGGTGTGTGTGTGTGTGTGCACTTTAGTTTCGTGAGCCTTGG GAAACTGATTCTACAAATATCTTATATAGAGATAAATGTATCAGGCATTTTTTTGCAAAGCGATATATACATTCCTCTTATTTATTAAAT CACTGGCTTTGGTGTTATGTAGAGATATTTGTAGATCTCAGAACAGAATTAAAATATTGTGAAAATTACAGCTTAATAAATGTTTATTGT >52391_52391_1_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000258148_BRI3BP_chr12_125509890_ENST00000341446_length(amino acids)=147AA_BP=120 MVRSRQMCNTNMSVPTDGAVTTSQIPASEQETLVRPKPLLLKLLKSVGAQKDTYTMKEVLFYLGQYIMTKRLYDEKQQHIVYCSNDLLGD -------------------------------------------------------------- >52391_52391_2_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000258149_BRI3BP_chr12_125509890_ENST00000341446_length(transcript)=6532nt_BP=645nt GCACCGCGGCGAGCTTGGCTGCTTCTGGGGCCTGTGTGGCCCTGTGTGTCGGAAAGATGGAGCAAGAAGCCGAGCCCGAGGGGCGGCCGC GACCCCTCTGACCGAGATCCTGCTGCTTTCGCAGCCAGGAGCACCGTCCCTCCCCGGATTAGTGCGTACGAGCGCCCAGTGCCCTGGCCC GGAGAGTGGAATGATCCCCGAGGCCCAGGGCGTCGTGCTTCCGCGCGCCCCGTGAAGGAAACTGGGGAGTCTTGAGGGACCCCCGACTCC AAGCGCGAAAACCCCGGATGGTGAGGAGCAGGCAAATGTGCAATACCAACATGTCTGTACCTACTGATGGTGCTGTAACCACCTCACAGA TTCCAGCTTCGGAACAAGAGACCCTGGTTAGACCAAAGCCATTGCTTTTGAAGTTATTAAAGTCTGTTGGTGCACAAAAAGACACTTATA CTATGAAAGAGGTTCTTTTTTATCTTGGCCAGTATATTATGACTAAACGATTATATGATGAGAAGCAACAACATATTGTATATTGTTCAA ATGATCTTCTAGGAGATTTGTTTGGCGTGCCAAGCTTCTCTGTGAAAGAGCACAGGAAAATATATACCATGATCTACAGGAACTTGGTAG TAGTCAATCAGCAGGTGGAGCACCTGGAGAAGCAGGTCAGACTGCTCAACATCCGTCTCAACCGGGTGCTCGAGAGCCTGGACCGCTCCA AGGACAAGTGAAGGTCAGCCGGCCGGGCGGGTCCACAGTTACCAGCACGCTGTCTCAGAAAACGAAAACGGAGGAAAAAAACCCCAAACC CCAAACAATCTTAATAAACACGACTGAGCAAGAAAGTGGCGCTGTGTAGGGCTATTTCCACCCACCCGGCAGCTCTTAGGACACATTCCC AGAAGAGCGGAAAGATCATTGACGTGGAACTACACACGAAGTGTAATTAGTGGGGGAAAAAATATTTTTTAAACAAAGGATATAACCATA TTTAGTTGTACAGTAAGAGAAATTTATCTGTGCATAGAGCATAAAGTTAATTTTTTCAAGCATTTAAATACATCTTTTGTAAGGTTTTTT AATAAAGGCAGATTGAGTCAAGTTTATTAAAACTTTACATGGAGGGGAAGAATATGCTAATTTGACATTTGAGAAAAGTTTAAATGCGAG CGTTTGTTTTCACAGCTTAGGGATTGGTGGGTGAGGTCTGTGTGCGTTATTTTCATGTTCCCAAACTGTGAGCTTTATTTCATGTTATTT TTGGTGGATGTTGACTTAGTTCATGAATGTTGTCACTTCTAAATGTGGACAGTTGGGCTGCAGGATGCAACTCAACCTCTTTTGGATCCT AGAGACCCCTGCCAAGCCCCCATCAGTCCTATTTTTAAATGAAGACATCGCCATTTCTAGAAGACCTGAGATTTTCTTCCCTCTGTGCTT CGATGTATTTTAACCTCAACATTTTGAAACGGAAAATGCAAGGAGGGGCCAAGAAATGTCCTGGGGATTTGATGCCAGGAAGTACCTGGT AAGGATTGGAGATTGCTCCAGAATCTACTGCATGTCCCAATTTGCAGTTTGCTCACTAGGGCTATATACAAAAGCTATACCCTTGAGGGT TTTTTTTTTTTTCTTTTGATTTCCTCAGGACCTTAGAGGGAAAACAAACAGTAGCAGCTAATATTCTCAAGTATATTGCTGCTTAGAAAG ATCCTCAGGAACAATTAGCAGCAATAAGCAAGCCTTTGAAAAGATCTGAATTCTTTTTCCTGAAATATTTACGATACACAGGTGCTTTTT ATCTGAAATCTGTTGTGTCCTCCTTTTAGGCAGTCTCTGTGGGCAGAGAGTGGGACTTGCGAGGTGGACAGCTGTGGGGATCCTGGGCAA GGGAGTTTCAGAAGGTGTGGCTCAGGGGCTGGTAGAAACCCCCTTTTGTTAGGACTTGTTTTTTAAAAAAGAAAGCCCTAGGCCAGGCGC AGTGGCTGATGCCTGTAATCCCAGCGCTTTGGGAGGCCGAGGAGTATGGATCACCTGAGGTCAGGAGTTCAAGACCAGCCTGGCCAACAT GGTGAAACCCCCGTCTCTACTAAAAATACAAAAATTAGTCAGGCATGGTGGTGGGTGCCTGTAGTCCAGCTACTCAGGAGGCTGAGGCAG GAGAATTGCTTGAACCTGGGAGGTGGAGGTTGCGGTGAGCCGAGATCACGCCACTGCACTCCAGCCTAGGCAACAGAGCAAGACTTCATC TTAAAAAAAAAAAAAAAAGCCCAGCATTTTGCAGCTTTGAAGCTGCTGGGGATCACGACTGTCTCTGTGGCCCTTACCTGGCCACAGATG TACCCTGGCAACCCGCTCACAGTGCCCTGCGTGCAGGTTTATAGATAACCCCTGCCAGACCTGCTACAACCAAGACCTTATGGTCCAGGT CTCAGTTTGGAGAAGATTTAGCTGTCTACCACTTCTTGTGTAGTGTTCCTGTGAAGTTTTATTTTTGGTTGCAAGTATCTGGAAAACAGA TGCAGATGTTTTTGTGGATGTTGTTGAGACTTTTGAGACTTCTGTTTCTAAGGTACACATGATGGGACCCAAACTCTCCTGGAGAGTTCC ATTCACTTCCTGAAACGTGCACCTGTTTCAGGTGCATGTTTATTCAGTGTGCATGGATATTTAAGATTTGCTGGAGCAGGCTGGGCAGGG TGGCTCACCCCTATAATCCCAACACTTTGGGAGGCCAAGGCAGGTGGATCACCTGAGGTCAGGAGTTTGAGACCAGCCCGGCCAACAGAG TGAAACCCCGTCTCTACTAAAAATACAAAAATTAGCCAGGCCTGGTGGTGGGCGCCTGTGATCCCAGCTACTTGGGAGGCTGAGGCTGGA GAATCGCTTGAACATGGGAGGCGGAGGTTGCAGTCAGCTGAGATCATACCACCACACTCTAGCCTCAGCAACAGAGTGAGACTCCGTCTC AAAAAAAATTATTTGCTGGATCGAATCTTTTTTTTTTTGAGATGGAGTCTCACTGTCAACCAGGCTGGAGTGCAGTGGCACCATCTCGGC TCACTGCAAGCTCCGCCTCCTGGGTTCACGCCATTCTCCTACCTCAGCCTCCCAAGTAGCCGGGACTACAGGCGCCCACCATCATGCCCG GCTAATTTTTTATATTTTTAGTAGAGACGGGGTTTCACCGTGTTAGCCAGGATGGTCTCGATCTCCTGACCTCGTGATCCGCCCGCCTCG GCCTCCCAAAGTGCTGGGATTACAGGCGTGAGCCACCGCGCTGGGCCTAGATCAAATCTTTATCCATGCACATTGGAACACAGGATTACT GGGTTGAAATCATTCTAGTTTTGTCATTTAGATACTTGTAGATGAATCTATTTTAGCACAAGGTATAAATAACTCGGGAGGTCATCTCTA TCTTCTTTCCTTTTGTGCATTTGGCTATACCACGTTTAGGTACTAAAACAGCTTTGCTTATGTTGGCCAGGGGAAAACATGGCATTCTGT GCGCAAAGCTAATGATCGCCAGCCCTGCCTTGGCCCCTCCCTTGTTTATGGTCATTGTAAGATGCCCGCATGTTAAGGCTTAAGCTGTCA CTGGGCTGGGTGTAATACCCGCTTCATTCCTCCTCCCACCCTCTTACCCGAAACATGAAGGGCACTGTGCTCTATTGAGATCTCGATAAG ATCATCATTTTAACTTGTATTCAACTGAGGGAGGTAACATGATATCTGGGTGCTGGTGGATTGAGACCTCAAGCATCAATTCAAAATTGC TGTCGAAAATATGCACTATTTTTTTTTTCCACTCTGGCTAAGTGTAGGCTCTGAAGCCAGTTTTATAGCTTGATACATTCTGGTGAAAGA TGTCTTCATTTTTAATGAAAGAACCTACAGTTAGTAGGTGTCACCTCAGGGTAGAAATTCTGATAGAAATTCTGATTATTTGAAACTCTG ATAGAAATCCTAGATGCTAGATTTGAAGAAAACCTGGGTTTTATTGCATAGATTTCACCTTTTCTGGGGCACATCTTACCCCAAAATAAT TGGGGATTTTTTTGTTCCTTCTCCATTCCATGGAACCTTGTAGGGTGTTGGGATTGGGCCTCCCCTCGCCACCCGCCACATGTCTTTGTT TTTTTCTCTCTTTTGTATGTGGAAAAATGTATTTTTGTACCATATCTCATACCTGGAGAGAAACCTTTTTTCTATCTTCAGGGCTGTAGA ATATATTAGCGTAACACTTCACTGAATGTGACGTAAAAGCATTCTGATGAAAACTGGTTCTGTTATAATTAAGACTCAGTGTTTAAAATA TCTTAATCAGTGTTAATTTCCCAGTGTTTCAGTATCACAGTAAATACTTGTTTTACTGAGAACAGAAAATAATCCAGAAAAGGAAAAGGA CCAGCCCTCTCTAGATTGCTGGAATTGTATGGATGTGTTGGGGTTGGTTTAGTGATCTCTTCCTTGACACCCTAGCAAAGATGCTTTTGT GGAGTGCAAACCTTTCATAAATATACTTTCCAGGCTGGGTGCAGTGGCTCAAGCTTGTAATCCCAGCACTTTGGGAGGCCAAGGCGGGTG GATCACTTGCGGTCGGGAGTTCGAGACCAGCCTGGCCAACATAGTGAAATCCCGTCTCTACCAAAAAATACAAAAAAAAAAAAAAAAAAA AAAATAGCCAGGTGTGGTGGTGGGTTCCTGTAGTCCCAGCTACTCAGGAGGCTGAGGTGGGAGAATTGTATGAACCTGGGAGGCAGAGGT TGCAGTGAACTGAGATCACACCACTGTACTCCAGTTGAGGCGACAGAGTGAGACCCAGTCTCAAAAAAAAAAAAAAGTGTGTGTGTGTGT ATATATATGTATATATATATACACACACACACACTTTCCAATACCAGTCGGCATTACCCGCCCTTTGGTTTCTTTTGCTTTGGATTTCAG CCCCCATATCATTTCTGGTAACAGAGCACAGCAATAGAATTTCTGGGAAGCAGACCTTGTGAATCGTGCCTGTGTCTTAATTCACAAAGG GTAAGTGAATGCCGTTGAAAGGTGTTTCATACACTTACCCCAGAAATGAGAGCCTGTCTCATGCAGGAGTCGGGAGGCAGCGATGTCAGG GAACTGGGACTATGTCTTTATGACTGTGATGGTGGTAACCAAGTGGGGTGGTCCCCTCTGGGGAGCTGAGTAAGAACGCAAAGGTTTGGG TCAGTGATTAGGAAGACTATAATATGACTTTGTGCATGCCCGGGAGGGCTGCCTTGTAGAGAGGATGTGAGCAGCTTAGTCGCTCATCTG GCCCTGTGATTCAGGCTTATGGAGCGTTAAGAATAACAGCTGTCAAATGGCCTAGACATGGTTAATGCAATTTGTTGCTAGTGGAAATCC TGAATTGCTTCCTTTCTGTGATCACTGCTACTTCTTAAGATGCTTTTGATGAATGTCATCTGCCTTACAAGTTGACACCTGATAACTTCT CCCTGATGGGTTTCCGAACTGGCTGACTTAACCAAAAAGCCAGCTCTTGCCATCTATCTTGCATTAAAAGGAATTCCTGAGGCTCCTAAG GGGTCAGCTGCCCCACTCCTGACTTTTTTATTTTTAATGGTCTATACCTTCTGCAACATTTTTGTTTATGGCCATTTGAATAGTTGGACT TTGACTCCTCACTTGTGAATAATAGGAATATATTTTGCAGAATCTAACATAATACCCTTAAAATTCATACTGGACAACCATCAGTGTGAT GTATAGTATCTGGTGTAAACAAATTTTATTCAGCATATTAAATTATTCTGTGTTTTGCTTTTCCTTGATATGTAGGAGGTGCACCAGTAC CAGTTTTTCTCTTGTGGTGGCTTAAAACGCTGATTGCCATTTGCATTGTCTACTGAAAAGCTGTGGCAGCGAGTGGTATTATTGACCATG TGTTCTCATGTCTTTTTGCAGATTTAAAGGTTTTATTGTAGTGTCCATTTAATATCTGCATGTGTGTTTAATCTCTGAATACCATGTAAT TAAACTGATTAACTTGTGATGGTGATGGTGATTAGCACACACTAAAAACCCAATGAAGCAATTGTCTCCCTCTTCTTAACTGGAAATCAC ACTTGTGAGGTATGGTATCCATTATCAACAGTGGATTCATGATTGACTTCATATTTTGTGTATCTGGTTATAAATTTTGTATAGCATTTA ATTTGGCACTTAATATAAATCCATTTGATTTGAATAGTGTGTGTGTGTACATGGTGTGTGTGTGTGTGTGCACTTTAGTTTCGTGAGCCT TGGGAAACTGATTCTACAAATATCTTATATAGAGATAAATGTATCAGGCATTTTTTTGCAAAGCGATATATACATTCCTCTTATTTATTA AATCACTGGCTTTGGTGTTATGTAGAGATATTTGTAGATCTCAGAACAGAATTAAAATATTGTGAAAATTACAGCTTAATAAATGTTTAT >52391_52391_2_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000258149_BRI3BP_chr12_125509890_ENST00000341446_length(amino acids)=159AA_BP=132 MRDPRLQARKPRMVRSRQMCNTNMSVPTDGAVTTSQIPASEQETLVRPKPLLLKLLKSVGAQKDTYTMKEVLFYLGQYIMTKRLYDEKQQ -------------------------------------------------------------- >52391_52391_3_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000350057_BRI3BP_chr12_125509890_ENST00000341446_length(transcript)=6152nt_BP=265nt ATGTGCAATACCAACATGTCTGTACCTACTGATGGTGCTGTAACCACCTCACAGATTCCAGCTTCGGAACAAGAGACCCTGGTTCTTTTT TATCTTGGCCAGTATATTATGACTAAACGATTATATGATGAGAAGCAACAACATATTGTATATTGTTCAAATGATCTTCTAGGAGATTTG TTTGGCGTGCCAAGCTTCTCTGTGAAAGAGCACAGGAAAATATATACCATGATCTACAGGAACTTGGTAGTAGTCAATCAGCAGGTGGAG CACCTGGAGAAGCAGGTCAGACTGCTCAACATCCGTCTCAACCGGGTGCTCGAGAGCCTGGACCGCTCCAAGGACAAGTGAAGGTCAGCC GGCCGGGCGGGTCCACAGTTACCAGCACGCTGTCTCAGAAAACGAAAACGGAGGAAAAAAACCCCAAACCCCAAACAATCTTAATAAACA CGACTGAGCAAGAAAGTGGCGCTGTGTAGGGCTATTTCCACCCACCCGGCAGCTCTTAGGACACATTCCCAGAAGAGCGGAAAGATCATT GACGTGGAACTACACACGAAGTGTAATTAGTGGGGGAAAAAATATTTTTTAAACAAAGGATATAACCATATTTAGTTGTACAGTAAGAGA AATTTATCTGTGCATAGAGCATAAAGTTAATTTTTTCAAGCATTTAAATACATCTTTTGTAAGGTTTTTTAATAAAGGCAGATTGAGTCA AGTTTATTAAAACTTTACATGGAGGGGAAGAATATGCTAATTTGACATTTGAGAAAAGTTTAAATGCGAGCGTTTGTTTTCACAGCTTAG GGATTGGTGGGTGAGGTCTGTGTGCGTTATTTTCATGTTCCCAAACTGTGAGCTTTATTTCATGTTATTTTTGGTGGATGTTGACTTAGT TCATGAATGTTGTCACTTCTAAATGTGGACAGTTGGGCTGCAGGATGCAACTCAACCTCTTTTGGATCCTAGAGACCCCTGCCAAGCCCC CATCAGTCCTATTTTTAAATGAAGACATCGCCATTTCTAGAAGACCTGAGATTTTCTTCCCTCTGTGCTTCGATGTATTTTAACCTCAAC ATTTTGAAACGGAAAATGCAAGGAGGGGCCAAGAAATGTCCTGGGGATTTGATGCCAGGAAGTACCTGGTAAGGATTGGAGATTGCTCCA GAATCTACTGCATGTCCCAATTTGCAGTTTGCTCACTAGGGCTATATACAAAAGCTATACCCTTGAGGGTTTTTTTTTTTTTCTTTTGAT TTCCTCAGGACCTTAGAGGGAAAACAAACAGTAGCAGCTAATATTCTCAAGTATATTGCTGCTTAGAAAGATCCTCAGGAACAATTAGCA GCAATAAGCAAGCCTTTGAAAAGATCTGAATTCTTTTTCCTGAAATATTTACGATACACAGGTGCTTTTTATCTGAAATCTGTTGTGTCC TCCTTTTAGGCAGTCTCTGTGGGCAGAGAGTGGGACTTGCGAGGTGGACAGCTGTGGGGATCCTGGGCAAGGGAGTTTCAGAAGGTGTGG CTCAGGGGCTGGTAGAAACCCCCTTTTGTTAGGACTTGTTTTTTAAAAAAGAAAGCCCTAGGCCAGGCGCAGTGGCTGATGCCTGTAATC CCAGCGCTTTGGGAGGCCGAGGAGTATGGATCACCTGAGGTCAGGAGTTCAAGACCAGCCTGGCCAACATGGTGAAACCCCCGTCTCTAC TAAAAATACAAAAATTAGTCAGGCATGGTGGTGGGTGCCTGTAGTCCAGCTACTCAGGAGGCTGAGGCAGGAGAATTGCTTGAACCTGGG AGGTGGAGGTTGCGGTGAGCCGAGATCACGCCACTGCACTCCAGCCTAGGCAACAGAGCAAGACTTCATCTTAAAAAAAAAAAAAAAAGC CCAGCATTTTGCAGCTTTGAAGCTGCTGGGGATCACGACTGTCTCTGTGGCCCTTACCTGGCCACAGATGTACCCTGGCAACCCGCTCAC AGTGCCCTGCGTGCAGGTTTATAGATAACCCCTGCCAGACCTGCTACAACCAAGACCTTATGGTCCAGGTCTCAGTTTGGAGAAGATTTA GCTGTCTACCACTTCTTGTGTAGTGTTCCTGTGAAGTTTTATTTTTGGTTGCAAGTATCTGGAAAACAGATGCAGATGTTTTTGTGGATG TTGTTGAGACTTTTGAGACTTCTGTTTCTAAGGTACACATGATGGGACCCAAACTCTCCTGGAGAGTTCCATTCACTTCCTGAAACGTGC ACCTGTTTCAGGTGCATGTTTATTCAGTGTGCATGGATATTTAAGATTTGCTGGAGCAGGCTGGGCAGGGTGGCTCACCCCTATAATCCC AACACTTTGGGAGGCCAAGGCAGGTGGATCACCTGAGGTCAGGAGTTTGAGACCAGCCCGGCCAACAGAGTGAAACCCCGTCTCTACTAA AAATACAAAAATTAGCCAGGCCTGGTGGTGGGCGCCTGTGATCCCAGCTACTTGGGAGGCTGAGGCTGGAGAATCGCTTGAACATGGGAG GCGGAGGTTGCAGTCAGCTGAGATCATACCACCACACTCTAGCCTCAGCAACAGAGTGAGACTCCGTCTCAAAAAAAATTATTTGCTGGA TCGAATCTTTTTTTTTTTGAGATGGAGTCTCACTGTCAACCAGGCTGGAGTGCAGTGGCACCATCTCGGCTCACTGCAAGCTCCGCCTCC TGGGTTCACGCCATTCTCCTACCTCAGCCTCCCAAGTAGCCGGGACTACAGGCGCCCACCATCATGCCCGGCTAATTTTTTATATTTTTA GTAGAGACGGGGTTTCACCGTGTTAGCCAGGATGGTCTCGATCTCCTGACCTCGTGATCCGCCCGCCTCGGCCTCCCAAAGTGCTGGGAT TACAGGCGTGAGCCACCGCGCTGGGCCTAGATCAAATCTTTATCCATGCACATTGGAACACAGGATTACTGGGTTGAAATCATTCTAGTT TTGTCATTTAGATACTTGTAGATGAATCTATTTTAGCACAAGGTATAAATAACTCGGGAGGTCATCTCTATCTTCTTTCCTTTTGTGCAT TTGGCTATACCACGTTTAGGTACTAAAACAGCTTTGCTTATGTTGGCCAGGGGAAAACATGGCATTCTGTGCGCAAAGCTAATGATCGCC AGCCCTGCCTTGGCCCCTCCCTTGTTTATGGTCATTGTAAGATGCCCGCATGTTAAGGCTTAAGCTGTCACTGGGCTGGGTGTAATACCC GCTTCATTCCTCCTCCCACCCTCTTACCCGAAACATGAAGGGCACTGTGCTCTATTGAGATCTCGATAAGATCATCATTTTAACTTGTAT TCAACTGAGGGAGGTAACATGATATCTGGGTGCTGGTGGATTGAGACCTCAAGCATCAATTCAAAATTGCTGTCGAAAATATGCACTATT TTTTTTTTCCACTCTGGCTAAGTGTAGGCTCTGAAGCCAGTTTTATAGCTTGATACATTCTGGTGAAAGATGTCTTCATTTTTAATGAAA GAACCTACAGTTAGTAGGTGTCACCTCAGGGTAGAAATTCTGATAGAAATTCTGATTATTTGAAACTCTGATAGAAATCCTAGATGCTAG ATTTGAAGAAAACCTGGGTTTTATTGCATAGATTTCACCTTTTCTGGGGCACATCTTACCCCAAAATAATTGGGGATTTTTTTGTTCCTT CTCCATTCCATGGAACCTTGTAGGGTGTTGGGATTGGGCCTCCCCTCGCCACCCGCCACATGTCTTTGTTTTTTTCTCTCTTTTGTATGT GGAAAAATGTATTTTTGTACCATATCTCATACCTGGAGAGAAACCTTTTTTCTATCTTCAGGGCTGTAGAATATATTAGCGTAACACTTC ACTGAATGTGACGTAAAAGCATTCTGATGAAAACTGGTTCTGTTATAATTAAGACTCAGTGTTTAAAATATCTTAATCAGTGTTAATTTC CCAGTGTTTCAGTATCACAGTAAATACTTGTTTTACTGAGAACAGAAAATAATCCAGAAAAGGAAAAGGACCAGCCCTCTCTAGATTGCT GGAATTGTATGGATGTGTTGGGGTTGGTTTAGTGATCTCTTCCTTGACACCCTAGCAAAGATGCTTTTGTGGAGTGCAAACCTTTCATAA ATATACTTTCCAGGCTGGGTGCAGTGGCTCAAGCTTGTAATCCCAGCACTTTGGGAGGCCAAGGCGGGTGGATCACTTGCGGTCGGGAGT TCGAGACCAGCCTGGCCAACATAGTGAAATCCCGTCTCTACCAAAAAATACAAAAAAAAAAAAAAAAAAAAAAATAGCCAGGTGTGGTGG TGGGTTCCTGTAGTCCCAGCTACTCAGGAGGCTGAGGTGGGAGAATTGTATGAACCTGGGAGGCAGAGGTTGCAGTGAACTGAGATCACA CCACTGTACTCCAGTTGAGGCGACAGAGTGAGACCCAGTCTCAAAAAAAAAAAAAAGTGTGTGTGTGTGTATATATATGTATATATATAT ACACACACACACACTTTCCAATACCAGTCGGCATTACCCGCCCTTTGGTTTCTTTTGCTTTGGATTTCAGCCCCCATATCATTTCTGGTA ACAGAGCACAGCAATAGAATTTCTGGGAAGCAGACCTTGTGAATCGTGCCTGTGTCTTAATTCACAAAGGGTAAGTGAATGCCGTTGAAA GGTGTTTCATACACTTACCCCAGAAATGAGAGCCTGTCTCATGCAGGAGTCGGGAGGCAGCGATGTCAGGGAACTGGGACTATGTCTTTA TGACTGTGATGGTGGTAACCAAGTGGGGTGGTCCCCTCTGGGGAGCTGAGTAAGAACGCAAAGGTTTGGGTCAGTGATTAGGAAGACTAT AATATGACTTTGTGCATGCCCGGGAGGGCTGCCTTGTAGAGAGGATGTGAGCAGCTTAGTCGCTCATCTGGCCCTGTGATTCAGGCTTAT GGAGCGTTAAGAATAACAGCTGTCAAATGGCCTAGACATGGTTAATGCAATTTGTTGCTAGTGGAAATCCTGAATTGCTTCCTTTCTGTG ATCACTGCTACTTCTTAAGATGCTTTTGATGAATGTCATCTGCCTTACAAGTTGACACCTGATAACTTCTCCCTGATGGGTTTCCGAACT GGCTGACTTAACCAAAAAGCCAGCTCTTGCCATCTATCTTGCATTAAAAGGAATTCCTGAGGCTCCTAAGGGGTCAGCTGCCCCACTCCT GACTTTTTTATTTTTAATGGTCTATACCTTCTGCAACATTTTTGTTTATGGCCATTTGAATAGTTGGACTTTGACTCCTCACTTGTGAAT AATAGGAATATATTTTGCAGAATCTAACATAATACCCTTAAAATTCATACTGGACAACCATCAGTGTGATGTATAGTATCTGGTGTAAAC AAATTTTATTCAGCATATTAAATTATTCTGTGTTTTGCTTTTCCTTGATATGTAGGAGGTGCACCAGTACCAGTTTTTCTCTTGTGGTGG CTTAAAACGCTGATTGCCATTTGCATTGTCTACTGAAAAGCTGTGGCAGCGAGTGGTATTATTGACCATGTGTTCTCATGTCTTTTTGCA GATTTAAAGGTTTTATTGTAGTGTCCATTTAATATCTGCATGTGTGTTTAATCTCTGAATACCATGTAATTAAACTGATTAACTTGTGAT GGTGATGGTGATTAGCACACACTAAAAACCCAATGAAGCAATTGTCTCCCTCTTCTTAACTGGAAATCACACTTGTGAGGTATGGTATCC ATTATCAACAGTGGATTCATGATTGACTTCATATTTTGTGTATCTGGTTATAAATTTTGTATAGCATTTAATTTGGCACTTAATATAAAT CCATTTGATTTGAATAGTGTGTGTGTGTACATGGTGTGTGTGTGTGTGTGCACTTTAGTTTCGTGAGCCTTGGGAAACTGATTCTACAAA TATCTTATATAGAGATAAATGTATCAGGCATTTTTTTGCAAAGCGATATATACATTCCTCTTATTTATTAAATCACTGGCTTTGGTGTTA TGTAGAGATATTTGTAGATCTCAGAACAGAATTAAAATATTGTGAAAATTACAGCTTAATAAATGTTTATTGTAATTTTGTTGTATGGAC >52391_52391_3_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000350057_BRI3BP_chr12_125509890_ENST00000341446_length(amino acids)=119AA_BP= MRRSLTLLLRLECGGMISADCNLRLPCSSDSPASASQVAGITGAHHQAWLIFVFLVETGFHSVGRAGLKLLTSGDPPALASQSVGIIGVS -------------------------------------------------------------- >52391_52391_4_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000462284_BRI3BP_chr12_125509890_ENST00000341446_length(transcript)=6547nt_BP=660nt GGGGCGCGCACCGAGGCACCGCGGCGAGCTTGGCTGCTTCTGGGGCCTGTGTGGCCCTGTGTGTCGGAAAGATGGAGCAAGAAGCCGAGC CCGAGGGGCGGCCGCGACCCCTCTGACCGAGATCCTGCTGCTTTCGCAGCCAGGAGCACCGTCCCTCCCCGGATTAGTGCGTACGAGCGC CCAGTGCCCTGGCCCGGAGAGTGGAATGATCCCCGAGGCCCAGGGCGTCGTGCTTCCGCGCGCCCCGTGAAGGAAACTGGGGAGTCTTGA GGGACCCCCGACTCCAAGCGCGAAAACCCCGGATGGTGAGGAGCAGGCAAATGTGCAATACCAACATGTCTGTACCTACTGATGGTGCTG TAACCACCTCACAGATTCCAGCTTCGGAACAAGAGACCCTGGTTAGACCAAAGCCATTGCTTTTGAAGTTATTAAAGTCTGTTGGTGCAC AAAAAGACACTTATACTATGAAAGAGGTTCTTTTTTATCTTGGCCAGTATATTATGACTAAACGATTATATGATGAGAAGCAACAACATA TTGTATATTGTTCAAATGATCTTCTAGGAGATTTGTTTGGCGTGCCAAGCTTCTCTGTGAAAGAGCACAGGAAAATATATACCATGATCT ACAGGAACTTGGTAGTAGTCAATCAGCAGGTGGAGCACCTGGAGAAGCAGGTCAGACTGCTCAACATCCGTCTCAACCGGGTGCTCGAGA GCCTGGACCGCTCCAAGGACAAGTGAAGGTCAGCCGGCCGGGCGGGTCCACAGTTACCAGCACGCTGTCTCAGAAAACGAAAACGGAGGA AAAAAACCCCAAACCCCAAACAATCTTAATAAACACGACTGAGCAAGAAAGTGGCGCTGTGTAGGGCTATTTCCACCCACCCGGCAGCTC TTAGGACACATTCCCAGAAGAGCGGAAAGATCATTGACGTGGAACTACACACGAAGTGTAATTAGTGGGGGAAAAAATATTTTTTAAACA AAGGATATAACCATATTTAGTTGTACAGTAAGAGAAATTTATCTGTGCATAGAGCATAAAGTTAATTTTTTCAAGCATTTAAATACATCT TTTGTAAGGTTTTTTAATAAAGGCAGATTGAGTCAAGTTTATTAAAACTTTACATGGAGGGGAAGAATATGCTAATTTGACATTTGAGAA AAGTTTAAATGCGAGCGTTTGTTTTCACAGCTTAGGGATTGGTGGGTGAGGTCTGTGTGCGTTATTTTCATGTTCCCAAACTGTGAGCTT TATTTCATGTTATTTTTGGTGGATGTTGACTTAGTTCATGAATGTTGTCACTTCTAAATGTGGACAGTTGGGCTGCAGGATGCAACTCAA CCTCTTTTGGATCCTAGAGACCCCTGCCAAGCCCCCATCAGTCCTATTTTTAAATGAAGACATCGCCATTTCTAGAAGACCTGAGATTTT CTTCCCTCTGTGCTTCGATGTATTTTAACCTCAACATTTTGAAACGGAAAATGCAAGGAGGGGCCAAGAAATGTCCTGGGGATTTGATGC CAGGAAGTACCTGGTAAGGATTGGAGATTGCTCCAGAATCTACTGCATGTCCCAATTTGCAGTTTGCTCACTAGGGCTATATACAAAAGC TATACCCTTGAGGGTTTTTTTTTTTTTCTTTTGATTTCCTCAGGACCTTAGAGGGAAAACAAACAGTAGCAGCTAATATTCTCAAGTATA TTGCTGCTTAGAAAGATCCTCAGGAACAATTAGCAGCAATAAGCAAGCCTTTGAAAAGATCTGAATTCTTTTTCCTGAAATATTTACGAT ACACAGGTGCTTTTTATCTGAAATCTGTTGTGTCCTCCTTTTAGGCAGTCTCTGTGGGCAGAGAGTGGGACTTGCGAGGTGGACAGCTGT GGGGATCCTGGGCAAGGGAGTTTCAGAAGGTGTGGCTCAGGGGCTGGTAGAAACCCCCTTTTGTTAGGACTTGTTTTTTAAAAAAGAAAG CCCTAGGCCAGGCGCAGTGGCTGATGCCTGTAATCCCAGCGCTTTGGGAGGCCGAGGAGTATGGATCACCTGAGGTCAGGAGTTCAAGAC CAGCCTGGCCAACATGGTGAAACCCCCGTCTCTACTAAAAATACAAAAATTAGTCAGGCATGGTGGTGGGTGCCTGTAGTCCAGCTACTC AGGAGGCTGAGGCAGGAGAATTGCTTGAACCTGGGAGGTGGAGGTTGCGGTGAGCCGAGATCACGCCACTGCACTCCAGCCTAGGCAACA GAGCAAGACTTCATCTTAAAAAAAAAAAAAAAAGCCCAGCATTTTGCAGCTTTGAAGCTGCTGGGGATCACGACTGTCTCTGTGGCCCTT ACCTGGCCACAGATGTACCCTGGCAACCCGCTCACAGTGCCCTGCGTGCAGGTTTATAGATAACCCCTGCCAGACCTGCTACAACCAAGA CCTTATGGTCCAGGTCTCAGTTTGGAGAAGATTTAGCTGTCTACCACTTCTTGTGTAGTGTTCCTGTGAAGTTTTATTTTTGGTTGCAAG TATCTGGAAAACAGATGCAGATGTTTTTGTGGATGTTGTTGAGACTTTTGAGACTTCTGTTTCTAAGGTACACATGATGGGACCCAAACT CTCCTGGAGAGTTCCATTCACTTCCTGAAACGTGCACCTGTTTCAGGTGCATGTTTATTCAGTGTGCATGGATATTTAAGATTTGCTGGA GCAGGCTGGGCAGGGTGGCTCACCCCTATAATCCCAACACTTTGGGAGGCCAAGGCAGGTGGATCACCTGAGGTCAGGAGTTTGAGACCA GCCCGGCCAACAGAGTGAAACCCCGTCTCTACTAAAAATACAAAAATTAGCCAGGCCTGGTGGTGGGCGCCTGTGATCCCAGCTACTTGG GAGGCTGAGGCTGGAGAATCGCTTGAACATGGGAGGCGGAGGTTGCAGTCAGCTGAGATCATACCACCACACTCTAGCCTCAGCAACAGA GTGAGACTCCGTCTCAAAAAAAATTATTTGCTGGATCGAATCTTTTTTTTTTTGAGATGGAGTCTCACTGTCAACCAGGCTGGAGTGCAG TGGCACCATCTCGGCTCACTGCAAGCTCCGCCTCCTGGGTTCACGCCATTCTCCTACCTCAGCCTCCCAAGTAGCCGGGACTACAGGCGC CCACCATCATGCCCGGCTAATTTTTTATATTTTTAGTAGAGACGGGGTTTCACCGTGTTAGCCAGGATGGTCTCGATCTCCTGACCTCGT GATCCGCCCGCCTCGGCCTCCCAAAGTGCTGGGATTACAGGCGTGAGCCACCGCGCTGGGCCTAGATCAAATCTTTATCCATGCACATTG GAACACAGGATTACTGGGTTGAAATCATTCTAGTTTTGTCATTTAGATACTTGTAGATGAATCTATTTTAGCACAAGGTATAAATAACTC GGGAGGTCATCTCTATCTTCTTTCCTTTTGTGCATTTGGCTATACCACGTTTAGGTACTAAAACAGCTTTGCTTATGTTGGCCAGGGGAA AACATGGCATTCTGTGCGCAAAGCTAATGATCGCCAGCCCTGCCTTGGCCCCTCCCTTGTTTATGGTCATTGTAAGATGCCCGCATGTTA AGGCTTAAGCTGTCACTGGGCTGGGTGTAATACCCGCTTCATTCCTCCTCCCACCCTCTTACCCGAAACATGAAGGGCACTGTGCTCTAT TGAGATCTCGATAAGATCATCATTTTAACTTGTATTCAACTGAGGGAGGTAACATGATATCTGGGTGCTGGTGGATTGAGACCTCAAGCA TCAATTCAAAATTGCTGTCGAAAATATGCACTATTTTTTTTTTCCACTCTGGCTAAGTGTAGGCTCTGAAGCCAGTTTTATAGCTTGATA CATTCTGGTGAAAGATGTCTTCATTTTTAATGAAAGAACCTACAGTTAGTAGGTGTCACCTCAGGGTAGAAATTCTGATAGAAATTCTGA TTATTTGAAACTCTGATAGAAATCCTAGATGCTAGATTTGAAGAAAACCTGGGTTTTATTGCATAGATTTCACCTTTTCTGGGGCACATC TTACCCCAAAATAATTGGGGATTTTTTTGTTCCTTCTCCATTCCATGGAACCTTGTAGGGTGTTGGGATTGGGCCTCCCCTCGCCACCCG CCACATGTCTTTGTTTTTTTCTCTCTTTTGTATGTGGAAAAATGTATTTTTGTACCATATCTCATACCTGGAGAGAAACCTTTTTTCTAT CTTCAGGGCTGTAGAATATATTAGCGTAACACTTCACTGAATGTGACGTAAAAGCATTCTGATGAAAACTGGTTCTGTTATAATTAAGAC TCAGTGTTTAAAATATCTTAATCAGTGTTAATTTCCCAGTGTTTCAGTATCACAGTAAATACTTGTTTTACTGAGAACAGAAAATAATCC AGAAAAGGAAAAGGACCAGCCCTCTCTAGATTGCTGGAATTGTATGGATGTGTTGGGGTTGGTTTAGTGATCTCTTCCTTGACACCCTAG CAAAGATGCTTTTGTGGAGTGCAAACCTTTCATAAATATACTTTCCAGGCTGGGTGCAGTGGCTCAAGCTTGTAATCCCAGCACTTTGGG AGGCCAAGGCGGGTGGATCACTTGCGGTCGGGAGTTCGAGACCAGCCTGGCCAACATAGTGAAATCCCGTCTCTACCAAAAAATACAAAA AAAAAAAAAAAAAAAAAAATAGCCAGGTGTGGTGGTGGGTTCCTGTAGTCCCAGCTACTCAGGAGGCTGAGGTGGGAGAATTGTATGAAC CTGGGAGGCAGAGGTTGCAGTGAACTGAGATCACACCACTGTACTCCAGTTGAGGCGACAGAGTGAGACCCAGTCTCAAAAAAAAAAAAA AGTGTGTGTGTGTGTATATATATGTATATATATATACACACACACACACTTTCCAATACCAGTCGGCATTACCCGCCCTTTGGTTTCTTT TGCTTTGGATTTCAGCCCCCATATCATTTCTGGTAACAGAGCACAGCAATAGAATTTCTGGGAAGCAGACCTTGTGAATCGTGCCTGTGT CTTAATTCACAAAGGGTAAGTGAATGCCGTTGAAAGGTGTTTCATACACTTACCCCAGAAATGAGAGCCTGTCTCATGCAGGAGTCGGGA GGCAGCGATGTCAGGGAACTGGGACTATGTCTTTATGACTGTGATGGTGGTAACCAAGTGGGGTGGTCCCCTCTGGGGAGCTGAGTAAGA ACGCAAAGGTTTGGGTCAGTGATTAGGAAGACTATAATATGACTTTGTGCATGCCCGGGAGGGCTGCCTTGTAGAGAGGATGTGAGCAGC TTAGTCGCTCATCTGGCCCTGTGATTCAGGCTTATGGAGCGTTAAGAATAACAGCTGTCAAATGGCCTAGACATGGTTAATGCAATTTGT TGCTAGTGGAAATCCTGAATTGCTTCCTTTCTGTGATCACTGCTACTTCTTAAGATGCTTTTGATGAATGTCATCTGCCTTACAAGTTGA CACCTGATAACTTCTCCCTGATGGGTTTCCGAACTGGCTGACTTAACCAAAAAGCCAGCTCTTGCCATCTATCTTGCATTAAAAGGAATT CCTGAGGCTCCTAAGGGGTCAGCTGCCCCACTCCTGACTTTTTTATTTTTAATGGTCTATACCTTCTGCAACATTTTTGTTTATGGCCAT TTGAATAGTTGGACTTTGACTCCTCACTTGTGAATAATAGGAATATATTTTGCAGAATCTAACATAATACCCTTAAAATTCATACTGGAC AACCATCAGTGTGATGTATAGTATCTGGTGTAAACAAATTTTATTCAGCATATTAAATTATTCTGTGTTTTGCTTTTCCTTGATATGTAG GAGGTGCACCAGTACCAGTTTTTCTCTTGTGGTGGCTTAAAACGCTGATTGCCATTTGCATTGTCTACTGAAAAGCTGTGGCAGCGAGTG GTATTATTGACCATGTGTTCTCATGTCTTTTTGCAGATTTAAAGGTTTTATTGTAGTGTCCATTTAATATCTGCATGTGTGTTTAATCTC TGAATACCATGTAATTAAACTGATTAACTTGTGATGGTGATGGTGATTAGCACACACTAAAAACCCAATGAAGCAATTGTCTCCCTCTTC TTAACTGGAAATCACACTTGTGAGGTATGGTATCCATTATCAACAGTGGATTCATGATTGACTTCATATTTTGTGTATCTGGTTATAAAT TTTGTATAGCATTTAATTTGGCACTTAATATAAATCCATTTGATTTGAATAGTGTGTGTGTGTACATGGTGTGTGTGTGTGTGTGCACTT TAGTTTCGTGAGCCTTGGGAAACTGATTCTACAAATATCTTATATAGAGATAAATGTATCAGGCATTTTTTTGCAAAGCGATATATACAT TCCTCTTATTTATTAAATCACTGGCTTTGGTGTTATGTAGAGATATTTGTAGATCTCAGAACAGAATTAAAATATTGTGAAAATTACAGC >52391_52391_4_MDM2-BRI3BP_MDM2_chr12_69214154_ENST00000462284_BRI3BP_chr12_125509890_ENST00000341446_length(amino acids)=159AA_BP=132 MRDPRLQARKPRMVRSRQMCNTNMSVPTDGAVTTSQIPASEQETLVRPKPLLLKLLKSVGAQKDTYTMKEVLFYLGQYIMTKRLYDEKQQ -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MDM2-BRI3BP |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 170_306 | 113.33333333333333 | 437.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 170_306 | 0.0 | 322.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 170_306 | 0.0 | 266.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 170_306 | 0 | 322.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 170_306 | 0.0 | 297.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 170_306 | 0 | 219.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 170_306 | 0.0 | 219.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 170_306 | 0 | 271.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 170_306 | 119.33333333333333 | 498.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 170_306 | 0 | 297.0 | MTBP |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 150_230 | 113.33333333333333 | 437.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 150_230 | 0.0 | 322.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 150_230 | 0.0 | 266.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 150_230 | 0 | 322.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 150_230 | 0.0 | 297.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 150_230 | 0 | 219.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 150_230 | 0.0 | 219.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 150_230 | 0 | 271.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 150_230 | 119.33333333333333 | 498.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 150_230 | 0 | 297.0 | PYHIN1 and necessary for interaction with RFFL and RNF34 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000258149 | + | 5 | 9 | 223_232 | 113.33333333333333 | 437.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000299252 | + | 1 | 5 | 223_232 | 0.0 | 322.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000348801 | + | 1 | 3 | 223_232 | 0.0 | 266.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000356290 | + | 1 | 6 | 223_232 | 0 | 322.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000360430 | + | 1 | 4 | 223_232 | 0.0 | 297.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393412 | + | 1 | 3 | 223_232 | 0 | 219.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000393413 | + | 1 | 2 | 223_232 | 0.0 | 219.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000428863 | + | 1 | 4 | 223_232 | 0 | 271.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000462284 | + | 5 | 11 | 223_232 | 119.33333333333333 | 498.0 | USP7 |
| Hgene | MDM2 | chr12:69214154 | chr12:125509890 | ENST00000540827 | + | 1 | 5 | 223_232 | 0 | 297.0 | USP7 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MDM2-BRI3BP |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MDM2-BRI3BP |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
