
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MED1-GDPD1 (FusionGDB2 ID:52651) |
Fusion Gene Summary for MED1-GDPD1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MED1-GDPD1 | Fusion gene ID: 52651 | Hgene | Tgene | Gene symbol | MED1 | GDPD1 | Gene ID | 8930 | 284161 |
| Gene name | methyl-CpG binding domain 4, DNA glycosylase | glycerophosphodiester phosphodiesterase domain containing 1 | |
| Synonyms | MED1 | GDE4 | |
| Cytomap | 3q21.3 | 17q22 | |
| Type of gene | protein-coding | protein-coding | |
| Description | methyl-CpG-binding domain protein 43,N(4)-ethenocytosine glycosylaseG/5-fluorouracil mismatch glycosylase with biphasic kineticsG/T mismatch glycosylaseG/U mismatch glycosylasemethyl-CpG binding domain protein 4methyl-CpG-binding endonuclease 1meth | lysophospholipase D GDPD1glycerophosphodiester phosphodiesterase 4glycerophosphodiester phosphodiesterase domain-containing protein 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q15648 | Q8N9F7 | |
| Ensembl transtripts involved in fusion gene | ENST00000300651, ENST00000394287, | ENST00000284116, ENST00000581140, ENST00000581276, | |
| Fusion gene scores | * DoF score | 42 X 23 X 11=10626 | 5 X 5 X 3=75 |
| # samples | 41 | 7 | |
| ** MAII score | log2(41/10626*10)=-4.69583089796762 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/75*10)=-0.0995356735509144 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MED1 [Title/Abstract] AND GDPD1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MED1(37579580)-GDPD1(57335112), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across MED1 (5'-gene) Fusion gene breakpoints across MED1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
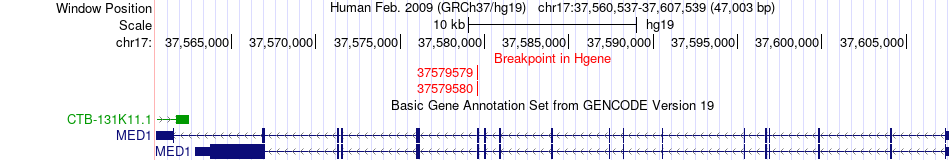 |
 Fusion gene breakpoints across GDPD1 (3'-gene) Fusion gene breakpoints across GDPD1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
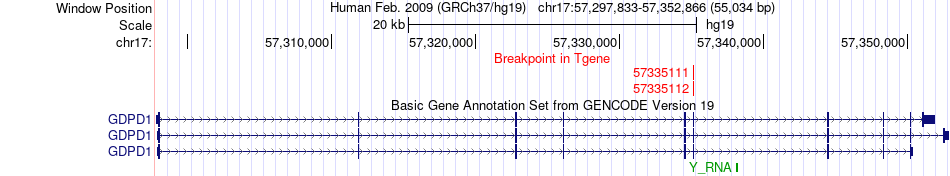 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-A8-A09I-01A | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| ChimerDB4 | BRCA | TCGA-A8-A09I | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
Top |
Fusion Gene ORF analysis for MED1-GDPD1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000300651 | ENST00000284116 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| In-frame | ENST00000300651 | ENST00000284116 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
| In-frame | ENST00000300651 | ENST00000581140 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| In-frame | ENST00000300651 | ENST00000581140 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
| In-frame | ENST00000300651 | ENST00000581276 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| In-frame | ENST00000300651 | ENST00000581276 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
| In-frame | ENST00000394287 | ENST00000284116 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| In-frame | ENST00000394287 | ENST00000284116 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
| In-frame | ENST00000394287 | ENST00000581140 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| In-frame | ENST00000394287 | ENST00000581140 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
| In-frame | ENST00000394287 | ENST00000581276 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + |
| In-frame | ENST00000394287 | ENST00000581276 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000394287 | MED1 | chr17 | 37579580 | - | ENST00000284116 | GDPD1 | chr17 | 57335112 | + | 2499 | 1301 | 206 | 1759 | 517 |
| ENST00000394287 | MED1 | chr17 | 37579580 | - | ENST00000581140 | GDPD1 | chr17 | 57335112 | + | 1992 | 1301 | 206 | 1684 | 492 |
| ENST00000394287 | MED1 | chr17 | 37579580 | - | ENST00000581276 | GDPD1 | chr17 | 57335112 | + | 1822 | 1301 | 206 | 1687 | 493 |
| ENST00000300651 | MED1 | chr17 | 37579580 | - | ENST00000284116 | GDPD1 | chr17 | 57335112 | + | 2517 | 1319 | 224 | 1777 | 517 |
| ENST00000300651 | MED1 | chr17 | 37579580 | - | ENST00000581140 | GDPD1 | chr17 | 57335112 | + | 2010 | 1319 | 224 | 1702 | 492 |
| ENST00000300651 | MED1 | chr17 | 37579580 | - | ENST00000581276 | GDPD1 | chr17 | 57335112 | + | 1840 | 1319 | 224 | 1705 | 493 |
| ENST00000394287 | MED1 | chr17 | 37579579 | - | ENST00000284116 | GDPD1 | chr17 | 57335111 | + | 2499 | 1301 | 206 | 1759 | 517 |
| ENST00000394287 | MED1 | chr17 | 37579579 | - | ENST00000581140 | GDPD1 | chr17 | 57335111 | + | 1992 | 1301 | 206 | 1684 | 492 |
| ENST00000394287 | MED1 | chr17 | 37579579 | - | ENST00000581276 | GDPD1 | chr17 | 57335111 | + | 1822 | 1301 | 206 | 1687 | 493 |
| ENST00000300651 | MED1 | chr17 | 37579579 | - | ENST00000284116 | GDPD1 | chr17 | 57335111 | + | 2517 | 1319 | 224 | 1777 | 517 |
| ENST00000300651 | MED1 | chr17 | 37579579 | - | ENST00000581140 | GDPD1 | chr17 | 57335111 | + | 2010 | 1319 | 224 | 1702 | 492 |
| ENST00000300651 | MED1 | chr17 | 37579579 | - | ENST00000581276 | GDPD1 | chr17 | 57335111 | + | 1840 | 1319 | 224 | 1705 | 493 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000394287 | ENST00000284116 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + | 0.000835761 | 0.9991642 |
| ENST00000394287 | ENST00000581140 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + | 0.001145928 | 0.99885404 |
| ENST00000394287 | ENST00000581276 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + | 0.002009511 | 0.9979905 |
| ENST00000300651 | ENST00000284116 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + | 0.000817874 | 0.99918216 |
| ENST00000300651 | ENST00000581140 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + | 0.00113621 | 0.9988638 |
| ENST00000300651 | ENST00000581276 | MED1 | chr17 | 37579580 | - | GDPD1 | chr17 | 57335112 | + | 0.001946389 | 0.99805367 |
| ENST00000394287 | ENST00000284116 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + | 0.000835761 | 0.9991642 |
| ENST00000394287 | ENST00000581140 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + | 0.001145928 | 0.99885404 |
| ENST00000394287 | ENST00000581276 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + | 0.002009511 | 0.9979905 |
| ENST00000300651 | ENST00000284116 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + | 0.000817874 | 0.99918216 |
| ENST00000300651 | ENST00000581140 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + | 0.00113621 | 0.9988638 |
| ENST00000300651 | ENST00000581276 | MED1 | chr17 | 37579579 | - | GDPD1 | chr17 | 57335111 | + | 0.001946389 | 0.99805367 |
Top |
Fusion Genomic Features for MED1-GDPD1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
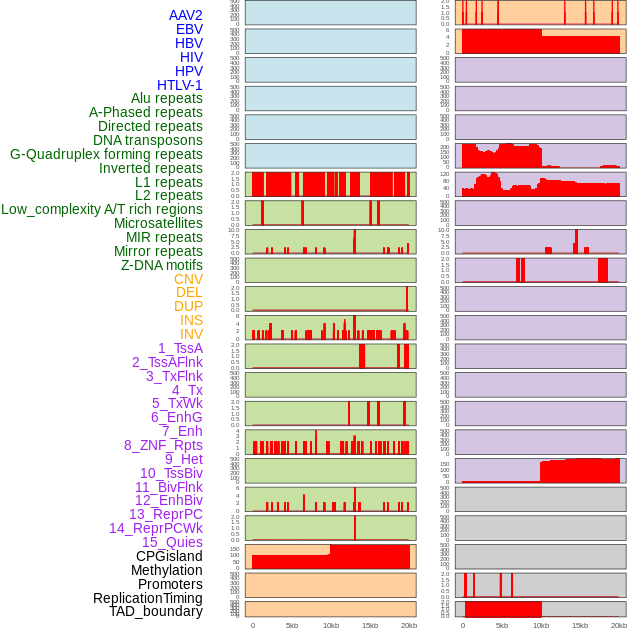 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
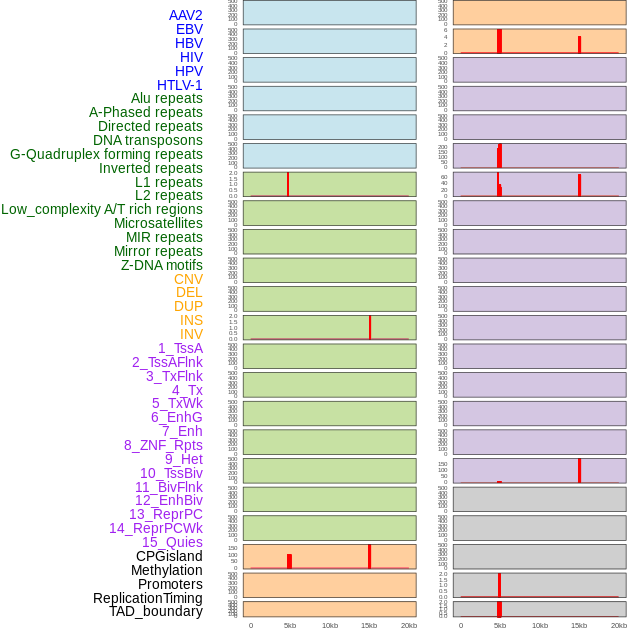 |
Top |
Fusion Protein Features for MED1-GDPD1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr17:37579580/chr17:57335112) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MED1 | GDPD1 |
| FUNCTION: Component of the Mediator complex, a coactivator involved in the regulated transcription of nearly all RNA polymerase II-dependent genes. Mediator functions as a bridge to convey information from gene-specific regulatory proteins to the basal RNA polymerase II transcription machinery. Mediator is recruited to promoters by direct interactions with regulatory proteins and serves as a scaffold for the assembly of a functional preinitiation complex with RNA polymerase II and the general transcription factors (PubMed:10406464, PubMed:11867769, PubMed:12037571, PubMed:12218053, PubMed:12556447, PubMed:14636573, PubMed:15340084, PubMed:15471764, PubMed:15989967, PubMed:16574658, PubMed:9653119). Acts as a coactivator for GATA1-mediated transcriptional activation during erythroid differentiation of K562 erythroleukemia cells (PubMed:24245781). {ECO:0000269|PubMed:10406464, ECO:0000269|PubMed:11867769, ECO:0000269|PubMed:12037571, ECO:0000269|PubMed:12218053, ECO:0000269|PubMed:12556447, ECO:0000269|PubMed:14636573, ECO:0000269|PubMed:15340084, ECO:0000269|PubMed:15471764, ECO:0000269|PubMed:15989967, ECO:0000269|PubMed:16574658, ECO:0000269|PubMed:24245781, ECO:0000269|PubMed:9653119}. | FUNCTION: Hydrolyzes lysoglycerophospholipids to produce lysophosphatidic acid (LPA) and the corresponding amines (PubMed:27637550, PubMed:25596343). Shows a preference for 1-O-alkyl-sn-glycero-3-phosphocholine (lyso-PAF), lysophosphatidylethanolamine (lyso-PE) and lysophosphatidylcholine (lyso-PC) (PubMed:27637550, PubMed:25596343). May be involved in bioactive N-acylethanolamine biosynthesis from both N-acyl-lysoplasmenylethanolamin (N-acyl-lysoPlsEt) and N-acyl-lysophosphatidylethanolamin (N-acyl-lysoPE) (PubMed:27637550, PubMed:25596343). In addition, hydrolyzes glycerophospho-N-acylethanolamine to N-acylethanolamine (PubMed:27637550). Does not display glycerophosphodiester phosphodiesterase activity, since it cannot hydrolyze either glycerophosphoinositol or glycerophosphocholine (By similarity). {ECO:0000250|UniProtKB:Q9CRY7, ECO:0000269|PubMed:25596343, ECO:0000269|PubMed:27637550}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000284116 | 4 | 10 | 217_314 | 162 | 315.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581140 | 4 | 10 | 217_314 | 162 | 290.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581276 | 4 | 9 | 217_314 | 162 | 291.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000284116 | 4 | 10 | 217_314 | 162 | 315.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581140 | 4 | 10 | 217_314 | 162 | 290.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581276 | 4 | 9 | 217_314 | 162 | 291.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000284116 | 4 | 10 | 196_216 | 162 | 315.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581140 | 4 | 10 | 196_216 | 162 | 290.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581276 | 4 | 9 | 196_216 | 162 | 291.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000284116 | 4 | 10 | 196_216 | 162 | 315.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581140 | 4 | 10 | 196_216 | 162 | 290.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581276 | 4 | 9 | 196_216 | 162 | 291.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 1078_1482 | 365 | 1582.0 | Compositional bias | Note=Ser-rich |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 1496_1529 | 365 | 1582.0 | Compositional bias | Note=Lys-rich |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 1078_1482 | 365 | 557.0 | Compositional bias | Note=Ser-rich |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 1496_1529 | 365 | 557.0 | Compositional bias | Note=Lys-rich |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 1078_1482 | 365 | 1582.0 | Compositional bias | Note=Ser-rich |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 1496_1529 | 365 | 1582.0 | Compositional bias | Note=Lys-rich |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 1078_1482 | 365 | 557.0 | Compositional bias | Note=Ser-rich |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 1496_1529 | 365 | 557.0 | Compositional bias | Note=Lys-rich |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 604_608 | 365 | 1582.0 | Motif | Note=LXXLL motif 1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 645_649 | 365 | 1582.0 | Motif | Note=LXXLL motif 2 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 875_902 | 365 | 1582.0 | Motif | Integrase domain-binding motif (IBM) |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 604_608 | 365 | 557.0 | Motif | Note=LXXLL motif 1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 645_649 | 365 | 557.0 | Motif | Note=LXXLL motif 2 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 875_902 | 365 | 557.0 | Motif | Integrase domain-binding motif (IBM) |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 604_608 | 365 | 1582.0 | Motif | Note=LXXLL motif 1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 645_649 | 365 | 1582.0 | Motif | Note=LXXLL motif 2 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 875_902 | 365 | 1582.0 | Motif | Integrase domain-binding motif (IBM) |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 604_608 | 365 | 557.0 | Motif | Note=LXXLL motif 1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 645_649 | 365 | 557.0 | Motif | Note=LXXLL motif 2 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 875_902 | 365 | 557.0 | Motif | Integrase domain-binding motif (IBM) |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000284116 | 4 | 10 | 40_309 | 162 | 315.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581140 | 4 | 10 | 40_309 | 162 | 290.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581276 | 4 | 9 | 40_309 | 162 | 291.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000284116 | 4 | 10 | 40_309 | 162 | 315.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581140 | 4 | 10 | 40_309 | 162 | 290.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581276 | 4 | 9 | 40_309 | 162 | 291.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000284116 | 4 | 10 | 1_3 | 162 | 315.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000284116 | 4 | 10 | 25_195 | 162 | 315.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581140 | 4 | 10 | 1_3 | 162 | 290.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581140 | 4 | 10 | 25_195 | 162 | 290.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581276 | 4 | 9 | 1_3 | 162 | 291.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581276 | 4 | 9 | 25_195 | 162 | 291.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000284116 | 4 | 10 | 1_3 | 162 | 315.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000284116 | 4 | 10 | 25_195 | 162 | 315.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581140 | 4 | 10 | 1_3 | 162 | 290.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581140 | 4 | 10 | 25_195 | 162 | 290.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581276 | 4 | 9 | 1_3 | 162 | 291.0 | Topological domain | Extracellular | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581276 | 4 | 9 | 25_195 | 162 | 291.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000284116 | 4 | 10 | 4_24 | 162 | 315.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581140 | 4 | 10 | 4_24 | 162 | 290.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579579 | chr17:57335111 | ENST00000581276 | 4 | 9 | 4_24 | 162 | 291.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000284116 | 4 | 10 | 4_24 | 162 | 315.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581140 | 4 | 10 | 4_24 | 162 | 290.0 | Transmembrane | Helical | |
| Tgene | GDPD1 | chr17:37579580 | chr17:57335112 | ENST00000581276 | 4 | 9 | 4_24 | 162 | 291.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for MED1-GDPD1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >52651_52651_1_MED1-GDPD1_MED1_chr17_37579579_ENST00000300651_GDPD1_chr17_57335111_ENST00000284116_length(transcript)=2517nt_BP=1319nt ACGGTTTATCTCGCGAATTTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAG ATTGATCCTTCCCGGGAAGAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAG GGCTTATCCTTTTGTGGCGGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGA TGAGTTCTCTCCTGGAACGGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGA AGAGGGTTGTGATGAGTTCTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAG CAATGACTGATCGTTTGGAGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAG ATATGTTCTATGTGGAAGTGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTC CGGAGCTTGTACAGCAGCTAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGG ACAACAAACTGAAGACTAAAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTA ATGCTGGTCCCTTGGATAAGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATG TCTCTCCTTCTGACCTACTGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCAT CAGTGACAATTGAAGGAACATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCC CTTCCTTCTCCTCAATCACCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAG CATTTGTTCAGAAACTGCAGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGT TTGAGCTATCAAAGGACCCTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAG AACACTTAACAGTGTGGGGTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTAC AACGTGTCCTGCTCATTCTTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTT CTATTATACTGAAGCTAAAAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAG CTTTGTTTGACCACCTAACTGCTCGAGGCATTCAAGTGTATATTTGGGTATTAAATGAAGAACAAGAATACAAAAGAGCTTTTGATTTGG GAGCAACTGGGGTGATGACAGACTATCCAACAAAGCTTAGGGATTTTTTACATAACTTTTCAGCATAGAAAAAGAGGTACTTAGAAGTAT TGAAGGAAAAAATGAAGACCTAAGAAAAAAATATTTCATGATCATTTCCCTAAGCCATTTCCAGAATGGTAAAAGGTTTAATCAGTTTTT ATTACCTCATTTTTAAGCCTGTATGAGAATGTAGAAACTATATATTATATGTATATTTATTTTAAATAATATTGTATATTTTATGTTTGT AAATTGTTTAGAAAGATAATTGGTTATGAGATGTAAGTTTTAATTTCTTAATGTGCATTTTTGTTTCTAGATCTTATACAGAAATCTTGA TTAATAACATACACAGAAATGTACATACTACATCATCTACAGAAATCTTGATCAATAACCTAGAAACTAGGTTATCTAGGTTATTGATCA AGATTTCTGTAGATGATGCAGTGTTCTATCATAAGTAATTCTGGAATCAAAGACTATTGGATACATTTGGCATTGGGCTGAGTGTGGTGG CTCATGCCTGTAATCCCAGCACTTTGGGAGGCTGAGACAGGCGGATCATCTGAAGTCAGGAGTTAAAGACCAGCCTGGCCAACATGGCAA AACCCCATCTCTACCAAAAATACAAAAATTAACCATGCGTGGTGGTACACGTCTGTCATCCCAGCAGCTCTTAAGGCTGAGGCACAAGAA >52651_52651_1_MED1-GDPD1_MED1_chr17_37579579_ENST00000300651_GDPD1_chr17_57335111_ENST00000284116_length(amino acids)=517AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_2_MED1-GDPD1_MED1_chr17_37579579_ENST00000300651_GDPD1_chr17_57335111_ENST00000581140_length(transcript)=2010nt_BP=1319nt ACGGTTTATCTCGCGAATTTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAG ATTGATCCTTCCCGGGAAGAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAG GGCTTATCCTTTTGTGGCGGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGA TGAGTTCTCTCCTGGAACGGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGA AGAGGGTTGTGATGAGTTCTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAG CAATGACTGATCGTTTGGAGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAG ATATGTTCTATGTGGAAGTGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTC CGGAGCTTGTACAGCAGCTAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGG ACAACAAACTGAAGACTAAAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTA ATGCTGGTCCCTTGGATAAGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATG TCTCTCCTTCTGACCTACTGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCAT CAGTGACAATTGAAGGAACATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCC CTTCCTTCTCCTCAATCACCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAG CATTTGTTCAGAAACTGCAGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGT TTGAGCTATCAAAGGACCCTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAG AACACTTAACAGTGTGGGGTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTAC AACGTGTCCTGCTCATTCTTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTT CTATTATACTGAAGCTAAAAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAG CTTTGTTTGAACCACTGCACCCGGCCAGTAAAAGAAATTTTGAAGGCCATTGCAGCTATTTGGTAGTGTCTTGTTATTTCTAGGTGTACC TTAGTTAAAGAGGAAAAATAAAACGGAAAAAAGCTTGGAAATCAGTGATGTGTAGTTATTTGGCAAGTTATACATAATCAGCAGCAGCCA GGCTCAAGAAAATAAAAGTTGATTAGTTGATCAGAAATAAAATCTGTAGAGTGAATTAGATTTCTGAGTTGTTGTTGTTAATGGAACATT CTATTTGAGACCTTTTTCAGGTGTGTAGCAATTCTACCATGTCCATTTTTTTAAGCATTAAAAAGGAACTTACCAGTTGTAAATTAAGAC >52651_52651_2_MED1-GDPD1_MED1_chr17_37579579_ENST00000300651_GDPD1_chr17_57335111_ENST00000581140_length(amino acids)=492AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_3_MED1-GDPD1_MED1_chr17_37579579_ENST00000300651_GDPD1_chr17_57335111_ENST00000581276_length(transcript)=1840nt_BP=1319nt ACGGTTTATCTCGCGAATTTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAG ATTGATCCTTCCCGGGAAGAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAG GGCTTATCCTTTTGTGGCGGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGA TGAGTTCTCTCCTGGAACGGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGA AGAGGGTTGTGATGAGTTCTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAG CAATGACTGATCGTTTGGAGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAG ATATGTTCTATGTGGAAGTGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTC CGGAGCTTGTACAGCAGCTAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGG ACAACAAACTGAAGACTAAAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTA ATGCTGGTCCCTTGGATAAGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATG TCTCTCCTTCTGACCTACTGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCAT CAGTGACAATTGAAGGAACATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCC CTTCCTTCTCCTCAATCACCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAG CATTTGTTCAGAAACTGCAGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGT TTGAGCTATCAAAGGACCCTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAG AACACTTAACAGTGTGGGGTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTAC AACGTGTCCTGCTCATTCTTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTT CTATTATACTGAAGCTAAAAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAG CTTTGTTTGACCACCTAACTGCTCGAGGCATTCAAGTAAGTTTCTGGAATGATGCATTTTGGAAGCAGCATAGCTCACCAGTTTAACACA AATATAAAGGGCAAGAACTTTTAACTGACTTAGTTTGGCCCTTGACTCTGATGCTGCTGCTTTACTGATCCACAAATGAGGACCTTAAGC >52651_52651_3_MED1-GDPD1_MED1_chr17_37579579_ENST00000300651_GDPD1_chr17_57335111_ENST00000581276_length(amino acids)=493AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_4_MED1-GDPD1_MED1_chr17_37579579_ENST00000394287_GDPD1_chr17_57335111_ENST00000284116_length(transcript)=2499nt_BP=1301nt TTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAGATTGATCCTTCCCGGGAA GAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAGGGCTTATCCTTTTGTGGC GGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGATGAGTTCTCTCCTGGAAC GGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGAAGAGGGTTGTGATGAGTT CTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAGCAATGACTGATCGTTTGG AGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAGATATGTTCTATGTGGAAG TGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTCCGGAGCTTGTACAGCAGC TAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGGACAACAAACTGAAGACTA AAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTAATGCTGGTCCCTTGGATA AGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATGTCTCTCCTTCTGACCTAC TGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCATCAGTGACAATTGAAGGAA CATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCCCTTCCTTCTCCTCAATCA CCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAGCATTTGTTCAGAAACTGC AGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGTTTGAGCTATCAAAGGACC CTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAGAACACTTAACAGTGTGGG GTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTACAACGTGTCCTGCTCATTC TTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTTCTATTATACTGAAGCTAA AAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAGCTTTGTTTGACCACCTAA CTGCTCGAGGCATTCAAGTGTATATTTGGGTATTAAATGAAGAACAAGAATACAAAAGAGCTTTTGATTTGGGAGCAACTGGGGTGATGA CAGACTATCCAACAAAGCTTAGGGATTTTTTACATAACTTTTCAGCATAGAAAAAGAGGTACTTAGAAGTATTGAAGGAAAAAATGAAGA CCTAAGAAAAAAATATTTCATGATCATTTCCCTAAGCCATTTCCAGAATGGTAAAAGGTTTAATCAGTTTTTATTACCTCATTTTTAAGC CTGTATGAGAATGTAGAAACTATATATTATATGTATATTTATTTTAAATAATATTGTATATTTTATGTTTGTAAATTGTTTAGAAAGATA ATTGGTTATGAGATGTAAGTTTTAATTTCTTAATGTGCATTTTTGTTTCTAGATCTTATACAGAAATCTTGATTAATAACATACACAGAA ATGTACATACTACATCATCTACAGAAATCTTGATCAATAACCTAGAAACTAGGTTATCTAGGTTATTGATCAAGATTTCTGTAGATGATG CAGTGTTCTATCATAAGTAATTCTGGAATCAAAGACTATTGGATACATTTGGCATTGGGCTGAGTGTGGTGGCTCATGCCTGTAATCCCA GCACTTTGGGAGGCTGAGACAGGCGGATCATCTGAAGTCAGGAGTTAAAGACCAGCCTGGCCAACATGGCAAAACCCCATCTCTACCAAA AATACAAAAATTAACCATGCGTGGTGGTACACGTCTGTCATCCCAGCAGCTCTTAAGGCTGAGGCACAAGAATTGCTTGAACCCGGGAGG >52651_52651_4_MED1-GDPD1_MED1_chr17_37579579_ENST00000394287_GDPD1_chr17_57335111_ENST00000284116_length(amino acids)=517AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_5_MED1-GDPD1_MED1_chr17_37579579_ENST00000394287_GDPD1_chr17_57335111_ENST00000581140_length(transcript)=1992nt_BP=1301nt TTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAGATTGATCCTTCCCGGGAA GAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAGGGCTTATCCTTTTGTGGC GGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGATGAGTTCTCTCCTGGAAC GGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGAAGAGGGTTGTGATGAGTT CTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAGCAATGACTGATCGTTTGG AGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAGATATGTTCTATGTGGAAG TGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTCCGGAGCTTGTACAGCAGC TAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGGACAACAAACTGAAGACTA AAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTAATGCTGGTCCCTTGGATA AGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATGTCTCTCCTTCTGACCTAC TGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCATCAGTGACAATTGAAGGAA CATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCCCTTCCTTCTCCTCAATCA CCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAGCATTTGTTCAGAAACTGC AGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGTTTGAGCTATCAAAGGACC CTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAGAACACTTAACAGTGTGGG GTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTACAACGTGTCCTGCTCATTC TTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTTCTATTATACTGAAGCTAA AAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAGCTTTGTTTGAACCACTGC ACCCGGCCAGTAAAAGAAATTTTGAAGGCCATTGCAGCTATTTGGTAGTGTCTTGTTATTTCTAGGTGTACCTTAGTTAAAGAGGAAAAA TAAAACGGAAAAAAGCTTGGAAATCAGTGATGTGTAGTTATTTGGCAAGTTATACATAATCAGCAGCAGCCAGGCTCAAGAAAATAAAAG TTGATTAGTTGATCAGAAATAAAATCTGTAGAGTGAATTAGATTTCTGAGTTGTTGTTGTTAATGGAACATTCTATTTGAGACCTTTTTC AGGTGTGTAGCAATTCTACCATGTCCATTTTTTTAAGCATTAAAAAGGAACTTACCAGTTGTAAATTAAGACAAGATCCAAATAGTCATA >52651_52651_5_MED1-GDPD1_MED1_chr17_37579579_ENST00000394287_GDPD1_chr17_57335111_ENST00000581140_length(amino acids)=492AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_6_MED1-GDPD1_MED1_chr17_37579579_ENST00000394287_GDPD1_chr17_57335111_ENST00000581276_length(transcript)=1822nt_BP=1301nt TTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAGATTGATCCTTCCCGGGAA GAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAGGGCTTATCCTTTTGTGGC GGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGATGAGTTCTCTCCTGGAAC GGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGAAGAGGGTTGTGATGAGTT CTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAGCAATGACTGATCGTTTGG AGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAGATATGTTCTATGTGGAAG TGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTCCGGAGCTTGTACAGCAGC TAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGGACAACAAACTGAAGACTA AAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTAATGCTGGTCCCTTGGATA AGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATGTCTCTCCTTCTGACCTAC TGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCATCAGTGACAATTGAAGGAA CATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCCCTTCCTTCTCCTCAATCA CCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAGCATTTGTTCAGAAACTGC AGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGTTTGAGCTATCAAAGGACC CTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAGAACACTTAACAGTGTGGG GTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTACAACGTGTCCTGCTCATTC TTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTTCTATTATACTGAAGCTAA AAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAGCTTTGTTTGACCACCTAA CTGCTCGAGGCATTCAAGTAAGTTTCTGGAATGATGCATTTTGGAAGCAGCATAGCTCACCAGTTTAACACAAATATAAAGGGCAAGAAC TTTTAACTGACTTAGTTTGGCCCTTGACTCTGATGCTGCTGCTTTACTGATCCACAAATGAGGACCTTAAGCTCTTCCTTACACTTAGCA >52651_52651_6_MED1-GDPD1_MED1_chr17_37579579_ENST00000394287_GDPD1_chr17_57335111_ENST00000581276_length(amino acids)=493AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_7_MED1-GDPD1_MED1_chr17_37579580_ENST00000300651_GDPD1_chr17_57335112_ENST00000284116_length(transcript)=2517nt_BP=1319nt ACGGTTTATCTCGCGAATTTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAG ATTGATCCTTCCCGGGAAGAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAG GGCTTATCCTTTTGTGGCGGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGA TGAGTTCTCTCCTGGAACGGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGA AGAGGGTTGTGATGAGTTCTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAG CAATGACTGATCGTTTGGAGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAG ATATGTTCTATGTGGAAGTGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTC CGGAGCTTGTACAGCAGCTAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGG ACAACAAACTGAAGACTAAAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTA ATGCTGGTCCCTTGGATAAGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATG TCTCTCCTTCTGACCTACTGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCAT CAGTGACAATTGAAGGAACATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCC CTTCCTTCTCCTCAATCACCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAG CATTTGTTCAGAAACTGCAGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGT TTGAGCTATCAAAGGACCCTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAG AACACTTAACAGTGTGGGGTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTAC AACGTGTCCTGCTCATTCTTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTT CTATTATACTGAAGCTAAAAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAG CTTTGTTTGACCACCTAACTGCTCGAGGCATTCAAGTGTATATTTGGGTATTAAATGAAGAACAAGAATACAAAAGAGCTTTTGATTTGG GAGCAACTGGGGTGATGACAGACTATCCAACAAAGCTTAGGGATTTTTTACATAACTTTTCAGCATAGAAAAAGAGGTACTTAGAAGTAT TGAAGGAAAAAATGAAGACCTAAGAAAAAAATATTTCATGATCATTTCCCTAAGCCATTTCCAGAATGGTAAAAGGTTTAATCAGTTTTT ATTACCTCATTTTTAAGCCTGTATGAGAATGTAGAAACTATATATTATATGTATATTTATTTTAAATAATATTGTATATTTTATGTTTGT AAATTGTTTAGAAAGATAATTGGTTATGAGATGTAAGTTTTAATTTCTTAATGTGCATTTTTGTTTCTAGATCTTATACAGAAATCTTGA TTAATAACATACACAGAAATGTACATACTACATCATCTACAGAAATCTTGATCAATAACCTAGAAACTAGGTTATCTAGGTTATTGATCA AGATTTCTGTAGATGATGCAGTGTTCTATCATAAGTAATTCTGGAATCAAAGACTATTGGATACATTTGGCATTGGGCTGAGTGTGGTGG CTCATGCCTGTAATCCCAGCACTTTGGGAGGCTGAGACAGGCGGATCATCTGAAGTCAGGAGTTAAAGACCAGCCTGGCCAACATGGCAA AACCCCATCTCTACCAAAAATACAAAAATTAACCATGCGTGGTGGTACACGTCTGTCATCCCAGCAGCTCTTAAGGCTGAGGCACAAGAA >52651_52651_7_MED1-GDPD1_MED1_chr17_37579580_ENST00000300651_GDPD1_chr17_57335112_ENST00000284116_length(amino acids)=517AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_8_MED1-GDPD1_MED1_chr17_37579580_ENST00000300651_GDPD1_chr17_57335112_ENST00000581140_length(transcript)=2010nt_BP=1319nt ACGGTTTATCTCGCGAATTTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAG ATTGATCCTTCCCGGGAAGAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAG GGCTTATCCTTTTGTGGCGGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGA TGAGTTCTCTCCTGGAACGGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGA AGAGGGTTGTGATGAGTTCTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAG CAATGACTGATCGTTTGGAGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAG ATATGTTCTATGTGGAAGTGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTC CGGAGCTTGTACAGCAGCTAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGG ACAACAAACTGAAGACTAAAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTA ATGCTGGTCCCTTGGATAAGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATG TCTCTCCTTCTGACCTACTGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCAT CAGTGACAATTGAAGGAACATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCC CTTCCTTCTCCTCAATCACCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAG CATTTGTTCAGAAACTGCAGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGT TTGAGCTATCAAAGGACCCTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAG AACACTTAACAGTGTGGGGTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTAC AACGTGTCCTGCTCATTCTTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTT CTATTATACTGAAGCTAAAAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAG CTTTGTTTGAACCACTGCACCCGGCCAGTAAAAGAAATTTTGAAGGCCATTGCAGCTATTTGGTAGTGTCTTGTTATTTCTAGGTGTACC TTAGTTAAAGAGGAAAAATAAAACGGAAAAAAGCTTGGAAATCAGTGATGTGTAGTTATTTGGCAAGTTATACATAATCAGCAGCAGCCA GGCTCAAGAAAATAAAAGTTGATTAGTTGATCAGAAATAAAATCTGTAGAGTGAATTAGATTTCTGAGTTGTTGTTGTTAATGGAACATT CTATTTGAGACCTTTTTCAGGTGTGTAGCAATTCTACCATGTCCATTTTTTTAAGCATTAAAAAGGAACTTACCAGTTGTAAATTAAGAC >52651_52651_8_MED1-GDPD1_MED1_chr17_37579580_ENST00000300651_GDPD1_chr17_57335112_ENST00000581140_length(amino acids)=492AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_9_MED1-GDPD1_MED1_chr17_37579580_ENST00000300651_GDPD1_chr17_57335112_ENST00000581276_length(transcript)=1840nt_BP=1319nt ACGGTTTATCTCGCGAATTTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAG ATTGATCCTTCCCGGGAAGAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAG GGCTTATCCTTTTGTGGCGGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGA TGAGTTCTCTCCTGGAACGGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGA AGAGGGTTGTGATGAGTTCTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAG CAATGACTGATCGTTTGGAGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAG ATATGTTCTATGTGGAAGTGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTC CGGAGCTTGTACAGCAGCTAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGG ACAACAAACTGAAGACTAAAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTA ATGCTGGTCCCTTGGATAAGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATG TCTCTCCTTCTGACCTACTGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCAT CAGTGACAATTGAAGGAACATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCC CTTCCTTCTCCTCAATCACCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAG CATTTGTTCAGAAACTGCAGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGT TTGAGCTATCAAAGGACCCTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAG AACACTTAACAGTGTGGGGTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTAC AACGTGTCCTGCTCATTCTTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTT CTATTATACTGAAGCTAAAAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAG CTTTGTTTGACCACCTAACTGCTCGAGGCATTCAAGTAAGTTTCTGGAATGATGCATTTTGGAAGCAGCATAGCTCACCAGTTTAACACA AATATAAAGGGCAAGAACTTTTAACTGACTTAGTTTGGCCCTTGACTCTGATGCTGCTGCTTTACTGATCCACAAATGAGGACCTTAAGC >52651_52651_9_MED1-GDPD1_MED1_chr17_37579580_ENST00000300651_GDPD1_chr17_57335112_ENST00000581276_length(amino acids)=493AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_10_MED1-GDPD1_MED1_chr17_37579580_ENST00000394287_GDPD1_chr17_57335112_ENST00000284116_length(transcript)=2499nt_BP=1301nt TTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAGATTGATCCTTCCCGGGAA GAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAGGGCTTATCCTTTTGTGGC GGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGATGAGTTCTCTCCTGGAAC GGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGAAGAGGGTTGTGATGAGTT CTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAGCAATGACTGATCGTTTGG AGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAGATATGTTCTATGTGGAAG TGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTCCGGAGCTTGTACAGCAGC TAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGGACAACAAACTGAAGACTA AAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTAATGCTGGTCCCTTGGATA AGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATGTCTCTCCTTCTGACCTAC TGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCATCAGTGACAATTGAAGGAA CATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCCCTTCCTTCTCCTCAATCA CCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAGCATTTGTTCAGAAACTGC AGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGTTTGAGCTATCAAAGGACC CTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAGAACACTTAACAGTGTGGG GTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTACAACGTGTCCTGCTCATTC TTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTTCTATTATACTGAAGCTAA AAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAGCTTTGTTTGACCACCTAA CTGCTCGAGGCATTCAAGTGTATATTTGGGTATTAAATGAAGAACAAGAATACAAAAGAGCTTTTGATTTGGGAGCAACTGGGGTGATGA CAGACTATCCAACAAAGCTTAGGGATTTTTTACATAACTTTTCAGCATAGAAAAAGAGGTACTTAGAAGTATTGAAGGAAAAAATGAAGA CCTAAGAAAAAAATATTTCATGATCATTTCCCTAAGCCATTTCCAGAATGGTAAAAGGTTTAATCAGTTTTTATTACCTCATTTTTAAGC CTGTATGAGAATGTAGAAACTATATATTATATGTATATTTATTTTAAATAATATTGTATATTTTATGTTTGTAAATTGTTTAGAAAGATA ATTGGTTATGAGATGTAAGTTTTAATTTCTTAATGTGCATTTTTGTTTCTAGATCTTATACAGAAATCTTGATTAATAACATACACAGAA ATGTACATACTACATCATCTACAGAAATCTTGATCAATAACCTAGAAACTAGGTTATCTAGGTTATTGATCAAGATTTCTGTAGATGATG CAGTGTTCTATCATAAGTAATTCTGGAATCAAAGACTATTGGATACATTTGGCATTGGGCTGAGTGTGGTGGCTCATGCCTGTAATCCCA GCACTTTGGGAGGCTGAGACAGGCGGATCATCTGAAGTCAGGAGTTAAAGACCAGCCTGGCCAACATGGCAAAACCCCATCTCTACCAAA AATACAAAAATTAACCATGCGTGGTGGTACACGTCTGTCATCCCAGCAGCTCTTAAGGCTGAGGCACAAGAATTGCTTGAACCCGGGAGG >52651_52651_10_MED1-GDPD1_MED1_chr17_37579580_ENST00000394287_GDPD1_chr17_57335112_ENST00000284116_length(amino acids)=517AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_11_MED1-GDPD1_MED1_chr17_37579580_ENST00000394287_GDPD1_chr17_57335112_ENST00000581140_length(transcript)=1992nt_BP=1301nt TTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAGATTGATCCTTCCCGGGAA GAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAGGGCTTATCCTTTTGTGGC GGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGATGAGTTCTCTCCTGGAAC GGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGAAGAGGGTTGTGATGAGTT CTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAGCAATGACTGATCGTTTGG AGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAGATATGTTCTATGTGGAAG TGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTCCGGAGCTTGTACAGCAGC TAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGGACAACAAACTGAAGACTA AAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTAATGCTGGTCCCTTGGATA AGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATGTCTCTCCTTCTGACCTAC TGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCATCAGTGACAATTGAAGGAA CATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCCCTTCCTTCTCCTCAATCA CCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAGCATTTGTTCAGAAACTGC AGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGTTTGAGCTATCAAAGGACC CTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAGAACACTTAACAGTGTGGG GTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTACAACGTGTCCTGCTCATTC TTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTTCTATTATACTGAAGCTAA AAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAGCTTTGTTTGAACCACTGC ACCCGGCCAGTAAAAGAAATTTTGAAGGCCATTGCAGCTATTTGGTAGTGTCTTGTTATTTCTAGGTGTACCTTAGTTAAAGAGGAAAAA TAAAACGGAAAAAAGCTTGGAAATCAGTGATGTGTAGTTATTTGGCAAGTTATACATAATCAGCAGCAGCCAGGCTCAAGAAAATAAAAG TTGATTAGTTGATCAGAAATAAAATCTGTAGAGTGAATTAGATTTCTGAGTTGTTGTTGTTAATGGAACATTCTATTTGAGACCTTTTTC AGGTGTGTAGCAATTCTACCATGTCCATTTTTTTAAGCATTAAAAAGGAACTTACCAGTTGTAAATTAAGACAAGATCCAAATAGTCATA >52651_52651_11_MED1-GDPD1_MED1_chr17_37579580_ENST00000394287_GDPD1_chr17_57335112_ENST00000581140_length(amino acids)=492AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- >52651_52651_12_MED1-GDPD1_MED1_chr17_37579580_ENST00000394287_GDPD1_chr17_57335112_ENST00000581276_length(transcript)=1822nt_BP=1301nt TTTGGGAAGTTCCGTTGGGGAAGATGGCGGCGGCCTCGAGCACCCTTCTCTTCTTGCCGCCGGGGACTTCAGATTGATCCTTCCCGGGAA GAGTAGGGACTGCTGGTGCCCTGCGTCCCGGGATCCCGAGCCAACTTGTTTCCTCCGTTAGTGGTGGGGAAGGGCTTATCCTTTTGTGGC GGATCTAGCTTCTCCTCGCCTTCAGGATGAAAGCTCAGGGGGAAACCGAGGAGTCAGAAAAGCTGAGTAAGATGAGTTCTCTCCTGGAAC GGCTCCATGCAAAATTTAACCAAAATAGACCCTGGAGTGAAACCATTAAGCTTGTGCGTCAAGTCATGGAGAAGAGGGTTGTGATGAGTT CTGGAGGGCATCAACATTTGGTCAGCTGTTTGGAGACATTGCAGAAGGCTCTCAAAGTAACATCTTTACCAGCAATGACTGATCGTTTGG AGTCCATAGCAAGACAGAATGGACTGGGCTCTCATCTCAGTGCCAGTGGCACTGAATGTTACATCACGTCAGATATGTTCTATGTGGAAG TGCAGTTAGATCCTGCAGGACAGCTTTGTGATGTAAAAGTGGCTCACCATGGGGAGAATCCTGTGAGCTGTCCGGAGCTTGTACAGCAGC TAAGGGAAAAAAATTTTGATGAATTTTCTAAGCACCTTAAGGGCCTTGTTAATCTGTATAACCTTCCAGGGGACAACAAACTGAAGACTA AAATGTACTTGGCTCTCCAATCCTTAGAACAAGATCTTTCTAAAATGGCAATTATGTACTGGAAAGCAACTAATGCTGGTCCCTTGGATA AGATTCTTCATGGAAGTGTTGGCTATCTCACACCAAGGAGTGGGGGTCATTTAATGAACCTGAAGTACTATGTCTCTCCTTCTGACCTAC TGGATGACAAGACTGCATCTCCCATCATTTTGCATGAGAATAATGTTTCTCGATCTTTGGGCATGAATGCATCAGTGACAATTGAAGGAA CATCTGCTGTGTACAAACTCCCAATTGCACCATTAATTATGGGGTCACATCCAGTTGACAATAAATGGACCCCTTCCTTCTCCTCAATCA CCAGTGCCAACAGTGTTGATCTTCCTGCCTGTTTCTTCTTGAAATTTCCCCAGCCAATCCCAGTATCTAGAGCATTTGTTCAGAAACTGC AGAACTGCACAGGAATTCCATTGTTTGAAACTCAACCAACTTATGCACCCCTGTATGAACTGATCACTCAGTTTGAGCTATCAAAGGACC CTGACCCCATACCTTTGAATCACAACATGAGATTTTATGCTGTTTCAGAGTTGGTGAAGCGGTATAATCGAGAACACTTAACAGTGTGGG GTAATGCCAATTATGAAATTGTAGAAAAGTGCTACAAAGAGAATTCAGATATTCCTATACTCTTCAGTCTACAACGTGTCCTGCTCATTC TTGGCCTTTTCTTCACTGGCCTCTTGCCCTTTGTGCCCATTCGAGAACAGTTTTTTGAAATCCCAATGCCTTCTATTATACTGAAGCTAA AAGAACCACACACCATGTCCAGAAGTCAAAAGTTTCTCATCTGGCTTTCTGATCTCTTACTAATGAGGAAAGCTTTGTTTGACCACCTAA CTGCTCGAGGCATTCAAGTAAGTTTCTGGAATGATGCATTTTGGAAGCAGCATAGCTCACCAGTTTAACACAAATATAAAGGGCAAGAAC TTTTAACTGACTTAGTTTGGCCCTTGACTCTGATGCTGCTGCTTTACTGATCCACAAATGAGGACCTTAAGCTCTTCCTTACACTTAGCA >52651_52651_12_MED1-GDPD1_MED1_chr17_37579580_ENST00000394287_GDPD1_chr17_57335112_ENST00000581276_length(amino acids)=493AA_BP=365 MKAQGETEESEKLSKMSSLLERLHAKFNQNRPWSETIKLVRQVMEKRVVMSSGGHQHLVSCLETLQKALKVTSLPAMTDRLESIARQNGL GSHLSASGTECYITSDMFYVEVQLDPAGQLCDVKVAHHGENPVSCPELVQQLREKNFDEFSKHLKGLVNLYNLPGDNKLKTKMYLALQSL EQDLSKMAIMYWKATNAGPLDKILHGSVGYLTPRSGGHLMNLKYYVSPSDLLDDKTASPIILHENNVSRSLGMNASVTIEGTSAVYKLPI APLIMGSHPVDNKWTPSFSSITSANSVDLPACFFLKFPQPIPVSRAFVQKLQNCTGIPLFETQPTYAPLYELITQFELSKDPDPIPLNHN MRFYAVSELVKRYNREHLTVWGNANYEIVEKCYKENSDIPILFSLQRVLLILGLFFTGLLPFVPIREQFFEIPMPSIILKLKEPHTMSRS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MED1-GDPD1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 108_212 | 365.0 | 1582.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 108_212 | 365.0 | 557.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 108_212 | 365.0 | 1582.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 108_212 | 365.0 | 557.0 | the Mediator complex |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 16_590 | 365.0 | 1582.0 | ESR1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 656_1066 | 365.0 | 1582.0 | ESR1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 16_590 | 365.0 | 557.0 | ESR1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 656_1066 | 365.0 | 557.0 | ESR1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 16_590 | 365.0 | 1582.0 | ESR1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 656_1066 | 365.0 | 1582.0 | ESR1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 16_590 | 365.0 | 557.0 | ESR1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 656_1066 | 365.0 | 557.0 | ESR1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 681_715 | 365.0 | 1582.0 | GATA1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 681_715 | 365.0 | 557.0 | GATA1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 681_715 | 365.0 | 1582.0 | GATA1 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 681_715 | 365.0 | 557.0 | GATA1 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 622_701 | 365.0 | 1582.0 | PPARGC1A and THRA |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 622_701 | 365.0 | 557.0 | PPARGC1A and THRA |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 622_701 | 365.0 | 1582.0 | PPARGC1A and THRA |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 622_701 | 365.0 | 557.0 | PPARGC1A and THRA |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 215_390 | 365.0 | 1582.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 215_390 | 365.0 | 557.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 215_390 | 365.0 | 1582.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 215_390 | 365.0 | 557.0 | the Mediator complex |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 1_670 | 365.0 | 1582.0 | the Mediator complex and THRA |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 1_670 | 365.0 | 557.0 | the Mediator complex and THRA |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 1_670 | 365.0 | 1582.0 | the Mediator complex and THRA |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 1_670 | 365.0 | 557.0 | the Mediator complex and THRA |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 405_644 | 365.0 | 1582.0 | THRA |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 405_644 | 365.0 | 557.0 | THRA |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 405_644 | 365.0 | 1582.0 | THRA |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 405_644 | 365.0 | 557.0 | THRA |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 1249_1421 | 365.0 | 1582.0 | TP53 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 1249_1421 | 365.0 | 557.0 | TP53 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 1249_1421 | 365.0 | 1582.0 | TP53 |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 1249_1421 | 365.0 | 557.0 | TP53 |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000300651 | - | 13 | 17 | 542_789 | 365.0 | 1582.0 | VDR |
| Hgene | MED1 | chr17:37579579 | chr17:57335111 | ENST00000394287 | - | 13 | 18 | 542_789 | 365.0 | 557.0 | VDR |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000300651 | - | 13 | 17 | 542_789 | 365.0 | 1582.0 | VDR |
| Hgene | MED1 | chr17:37579580 | chr17:57335112 | ENST00000394287 | - | 13 | 18 | 542_789 | 365.0 | 557.0 | VDR |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MED1-GDPD1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MED1-GDPD1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
