
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:AP3B1-IPO11 (FusionGDB2 ID:5267) |
Fusion Gene Summary for AP3B1-IPO11 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: AP3B1-IPO11 | Fusion gene ID: 5267 | Hgene | Tgene | Gene symbol | AP3B1 | IPO11 | Gene ID | 8546 | 51194 |
| Gene name | adaptor related protein complex 3 subunit beta 1 | importin 11 | |
| Synonyms | ADTB3|ADTB3A|HPS|HPS2|PE | RanBP11 | |
| Cytomap | 5q14.1 | 5q12.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | AP-3 complex subunit beta-1AP-3 complex beta-3A subunitadaptor protein complex AP-3 subunit beta-1adaptor related protein complex 3 beta 1 subunitbeta-3A-adaptinclathrin assembly protein complex 3 beta-1 large chain | importin-11Ran binding protein 11imp11ran-binding protein 11 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | O00203 | Q9UI26 | |
| Ensembl transtripts involved in fusion gene | ENST00000255194, ENST00000519295, ENST00000523204, | ENST00000409534, ENST00000512177, ENST00000325324, ENST00000409296, | |
| Fusion gene scores | * DoF score | 10 X 12 X 6=720 | 6 X 6 X 7=252 |
| # samples | 12 | 7 | |
| ** MAII score | log2(12/720*10)=-2.58496250072116 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/252*10)=-1.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: AP3B1 [Title/Abstract] AND IPO11 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | AP3B1(77590276)-IPO11(61762958), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across AP3B1 (5'-gene) Fusion gene breakpoints across AP3B1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
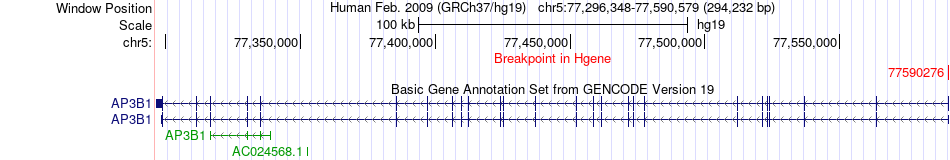 |
 Fusion gene breakpoints across IPO11 (3'-gene) Fusion gene breakpoints across IPO11 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
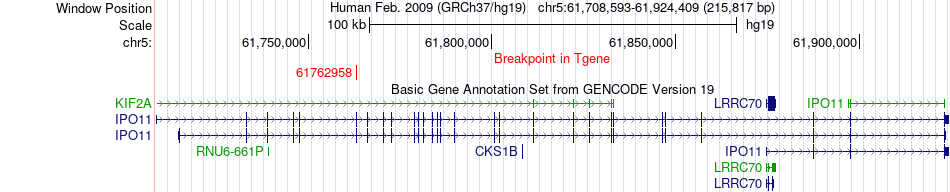 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SKCM | TCGA-D3-A51N-06A | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
Top |
Fusion Gene ORF analysis for AP3B1-IPO11 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000255194 | ENST00000409534 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| 5CDS-intron | ENST00000255194 | ENST00000512177 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| In-frame | ENST00000255194 | ENST00000325324 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| In-frame | ENST00000255194 | ENST00000409296 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-3CDS | ENST00000519295 | ENST00000325324 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-3CDS | ENST00000519295 | ENST00000409296 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-3CDS | ENST00000523204 | ENST00000325324 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-3CDS | ENST00000523204 | ENST00000409296 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-intron | ENST00000519295 | ENST00000409534 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-intron | ENST00000519295 | ENST00000512177 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-intron | ENST00000523204 | ENST00000409534 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
| intron-intron | ENST00000523204 | ENST00000512177 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000255194 | AP3B1 | chr5 | 77590276 | - | ENST00000325324 | IPO11 | chr5 | 61762958 | + | 3975 | 304 | 337 | 2715 | 792 |
| ENST00000255194 | AP3B1 | chr5 | 77590276 | - | ENST00000409296 | IPO11 | chr5 | 61762958 | + | 3102 | 304 | 337 | 2715 | 792 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000255194 | ENST00000325324 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + | 0.000530878 | 0.9994691 |
| ENST00000255194 | ENST00000409296 | AP3B1 | chr5 | 77590276 | - | IPO11 | chr5 | 61762958 | + | 0.000819178 | 0.99918085 |
Top |
Fusion Genomic Features for AP3B1-IPO11 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| AP3B1 | chr5 | 77590275 | - | IPO11 | chr5 | 61762957 | + | 4.19E-05 | 0.99995816 |
| AP3B1 | chr5 | 77590275 | - | IPO11 | chr5 | 61762957 | + | 4.19E-05 | 0.99995816 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
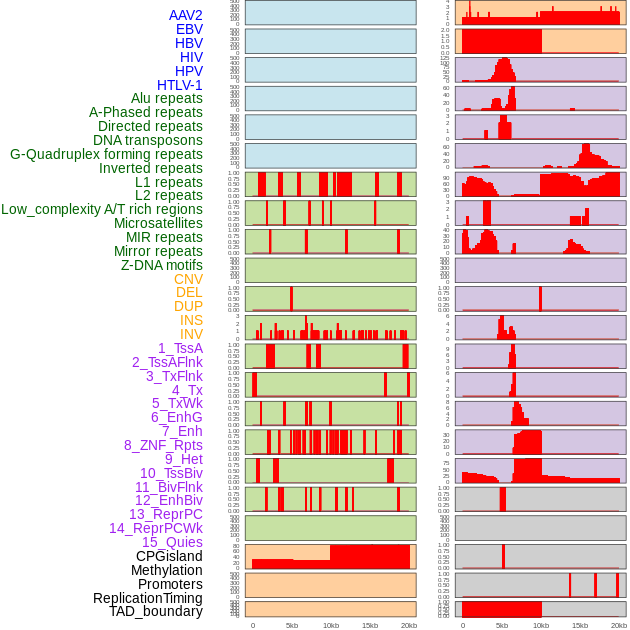 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
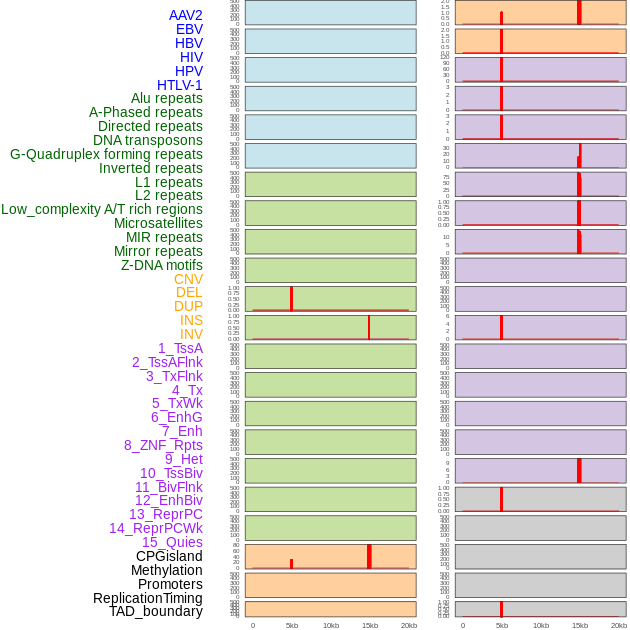 |
Top |
Fusion Protein Features for AP3B1-IPO11 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr5:77590276/chr5:61762958) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| AP3B1 | IPO11 |
| FUNCTION: Subunit of non-clathrin- and clathrin-associated adaptor protein complex 3 (AP-3) that plays a role in protein sorting in the late-Golgi/trans-Golgi network (TGN) and/or endosomes. The AP complexes mediate both the recruitment of clathrin to membranes and the recognition of sorting signals within the cytosolic tails of transmembrane cargo molecules. AP-3 appears to be involved in the sorting of a subset of transmembrane proteins targeted to lysosomes and lysosome-related organelles. In concert with the BLOC-1 complex, AP-3 is required to target cargos into vesicles assembled at cell bodies for delivery into neurites and nerve terminals. {ECO:0000305|PubMed:9151686}. | FUNCTION: Functions in nuclear protein import as nuclear transport receptor. Serves as receptor for nuclear localization signals (NLS) in cargo substrates. Is thought to mediate docking of the importin/substrate complex to the nuclear pore complex (NPC) through binding to nucleoporin and the complex is subsequently translocated through the pore by an energy requiring, Ran-dependent mechanism. At the nucleoplasmic side of the NPC, Ran binds to the importin, the importin/substrate complex dissociates and importin is re-exported from the nucleus to the cytoplasm where GTP hydrolysis releases Ran. The directionality of nuclear import is thought to be conferred by an asymmetric distribution of the GTP- and GDP-bound forms of Ran between the cytoplasm and nucleus (By similarity). Mediates the nuclear import of UBE2E3, and of RPL12 (By similarity). {ECO:0000250, ECO:0000269|PubMed:11032817}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 209_248 | 172 | 976.0 | Repeat | Note=HEAT 2 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 318_356 | 172 | 976.0 | Repeat | Note=HEAT 3 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 422_459 | 172 | 976.0 | Repeat | Note=HEAT 4 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 473_509 | 172 | 976.0 | Repeat | Note=HEAT 5 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 511_548 | 172 | 976.0 | Repeat | Note=HEAT 6 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 555_593 | 172 | 976.0 | Repeat | Note=HEAT 7 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 600_636 | 172 | 976.0 | Repeat | Note=HEAT 8 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 640_677 | 172 | 976.0 | Repeat | Note=HEAT 9 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 683_720 | 172 | 976.0 | Repeat | Note=HEAT 10 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 743_764 | 172 | 976.0 | Repeat | Note=HEAT 11 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 765_804 | 172 | 976.0 | Repeat | Note=HEAT 12 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 819_849 | 172 | 976.0 | Repeat | Note=HEAT 13 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 850_887 | 172 | 976.0 | Repeat | Note=HEAT 14 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 957_974 | 172 | 976.0 | Repeat | Note=HEAT 15 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 209_248 | 212 | 1016.0 | Repeat | Note=HEAT 2 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 318_356 | 212 | 1016.0 | Repeat | Note=HEAT 3 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 422_459 | 212 | 1016.0 | Repeat | Note=HEAT 4 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 473_509 | 212 | 1016.0 | Repeat | Note=HEAT 5 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 511_548 | 212 | 1016.0 | Repeat | Note=HEAT 6 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 555_593 | 212 | 1016.0 | Repeat | Note=HEAT 7 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 600_636 | 212 | 1016.0 | Repeat | Note=HEAT 8 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 640_677 | 212 | 1016.0 | Repeat | Note=HEAT 9 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 683_720 | 212 | 1016.0 | Repeat | Note=HEAT 10 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 743_764 | 212 | 1016.0 | Repeat | Note=HEAT 11 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 765_804 | 212 | 1016.0 | Repeat | Note=HEAT 12 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 819_849 | 212 | 1016.0 | Repeat | Note=HEAT 13 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 850_887 | 212 | 1016.0 | Repeat | Note=HEAT 14 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 957_974 | 212 | 1016.0 | Repeat | Note=HEAT 15 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | AP3B1 | chr5:77590276 | chr5:61762958 | ENST00000255194 | - | 1 | 27 | 677_802 | 42 | 1095.0 | Compositional bias | Note=Glu/Ser-rich |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 28_100 | 172 | 976.0 | Domain | Importin N-terminal | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 28_100 | 212 | 1016.0 | Domain | Importin N-terminal | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000325324 | 4 | 30 | 123_160 | 172 | 976.0 | Repeat | Note=HEAT 1 | |
| Tgene | IPO11 | chr5:77590276 | chr5:61762958 | ENST00000409296 | 4 | 30 | 123_160 | 212 | 1016.0 | Repeat | Note=HEAT 1 |
Top |
Fusion Gene Sequence for AP3B1-IPO11 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >5267_5267_1_AP3B1-IPO11_AP3B1_chr5_77590276_ENST00000255194_IPO11_chr5_61762958_ENST00000325324_length(transcript)=3975nt_BP=304nt ACTGCGCATGCGCAGGGGGTGGACTGCCAGGTCGGCTCAGGGAGCCGTGACGAGTCCGGAAGCGCCTGCGCGCGCTCCTCCGTACGAGAA CTAGTTTTGTTCCGTGCCCTCTGGACTGGAACCTTTTGGAGAGAACCCCCGGCAGGACCAACCCCGCACCCGCCAGCACCGCGGCAATGT CCAGCAATAGTTTTCCTTACAATGAGCAGTCCGGAGGAGGGGAGGCGACGGAGCTGGGTCAGGAGGCGACCTCAACCATTTCCCCCTCGG GGGCCTTCGGCCTCTTTAGCAGCGATTTGAAGAATTAGCTTCTGGAATTTATAATTTTGCCTGCTCTCTGTGGAATCACCACACAGACAC ATTCCTGCAAGAAGTTTCTTCTGGCAATGAAGCTGCAATTTTGAGTTCACTAGAACGAACACTGCTATCATTGAAAGTGCTGCGTAAGTT AACTGTTAATGGATTTGTGGAACCTCATAAGAATATGGAGGTGATGGGTTTTTTACATGGAATATTTGAACGTCTAAAACAGTTTCTGGA ATGCAGTAGAAGTATAGGTACAGATAATGTGTGTAGAGATAGACTGGAAAAGACCATCATTCTTTTTACTAAAGTGCTTTTGGACTTCTT GGATCAGCATCCTTTTTCATTTACTCCTCTAATTCAGAGATCACTGGAATTTTCTGTAAGCTATGTTTTTACAGAAGTTGGTGAAGGCGT TACATTTGAACGATTCATTGTCCAATGTATGAATCTTATTAAGATGATTGTCAAAAATTATGCTTATAAGCCATCCAAAAATTTTGAAGA TAGCAGCCCTGAAACTCTTGAAGCCCATAAGATTAAGATGGCATTCTTCACATATCCTACTTTGACAGAGATATGTAGAAGATTAGTCTC TCATTATTTCCTATTAACTGAAGAAGAACTGACAATGTGGGAAGAAGACCCAGAAGGCTTTACAGTGGAAGAAACAGGAGGAGATTCTTG GAAATATAGTTTGAGGCCATGCACTGAAGTATTATTTATAGATATATTCCATGAATATAATCAGACTCTTACTCCTGTACTTCTAGAAAT GATGCAAACACTTCAAGGACCCACAAATGTGGAAGATATGAATGCACTGTTAATCAAAGATGCTGTGTATAATGCTGTTGGATTAGCTGC TTATGAGCTCTTTGACAGTGTTGATTTTGATCAGTGGTTTAAAAACCAGCTTCTTCCAGAATTACAAGTCATTCACAATAGGTATAAGCC ATTGCGACGCAGGGTGATTTGGCTCATCGGTCAGTGGATTTCTGTGAAATTCAAGTCTGACTTAAGACCCATGCTTTATGAAGCAATCTG TAACTTGCTTCAAGATCAAGATTTAGTGGTCCGTATTGAAACAGCTACAACTTTGAAGTTAACTGTTGATGATTTTGAATTTAGAACAGA TCAGTTTCTACCGTATTTGGAAACCATGTTCACACTACTTTTTCAGTTACTGCAGCAAGTTACAGAATGTGACACAAAGATGCATGTTTT GCATGTCCTTTCTTGTGTGATCGAAAGAGTCAACATGCAGATACGACCATATGTGGGATGTTTGGTACAATATTTGCCCCTCCTTTGGAA GCAGAGTGAAGAACACAATATGTTGAGATGTGCTATTTTGACAACACTTATTCATCTTGTTCAGGGATTAGGAGCAGACAGCAAGAACCT GTACCCTTTCCTGCTCCCAGTTATTCAACTGAGTACAGATGTTTCACAGCCTCCACATGTTTATCTTCTGGAAGATGGTTTAGAATTATG GTTAGTAACTTTGGAAAACAGTCCATGTATTACACCAGAGTTGCTTCGTATATTTCAGAATATGTCACCACTTCTTGAACTAAGTTCAGA AAATCTTAGAACTTGCTTTAAGATCATCAATGGTTATATCTTTTTATCATCAACAGAATTTTTACAGACATACGCAGTAGGTCTATGCCA GTCCTTTTGTGAACTTTTAAAGGAAATTACTACAGAAGGTCAAGTTCAGGTGCTCAAGGTTGTGGAAAATGCCCTTAAAGTGAACCCAAT ACTAGGTCCACAAATGTTTCAACCGATTTTACCCTATGTTTTCAAGGGTATTATAGAAGGGGAGAGGTATCCTGTAGTGATGTCCACGTA TCTTGGAGTTATGGGTCGAGTTCTACTACAAAACACTAGTTTTTTTTCTTCACTACTTAATGAGATGGCCCATAAATTTAATCAGGAGAT GGACCAGCTTTTGGGAAATATGATTGAAATGTGGGTTGATCGAATGGACAACATTACCCAGCCTGAAAGAAGAAAACTTTCAGCTTTGGC TTTGCTCTCTCTTCTGCCATCTGATAATAGTGTTATCCAAGATAAATTCTGTGGGATTATAAACATTTCAGTAGAAGGCCTGCATGATGT CATGACGGAAGATCCTGAAACAGGAACTTATAAAGACTGTATGTTGATGTCTCATCTTGAGGAACCAAAAGTAACAGAAGATGAAGAACC ACCCACAGAACAAGATAAGAGGAAAAAGATGCTGGCCCTGAAGGACCCTGTTCATACAGTGTCACTGCAGCAGTTCATCTACGAGAAGCT CAAGGCACAGCAGGAGATGCTAGGAGAACAAGGTTTCCAGTCCCTCATGGAAACAGTGGATACGGAGATTGTCACCCAGCTACAGGAGTT TTTGCAAGGATTCTAAGAGCACATGACATGTGGCTGCCTCCCCTTTCAGAAACAAGCTGAGTAACCCAGCCTGCCGTTTGTATGTGAGAG CCTGCTGAGATGAAGAAATCACTTCATGAAAATAAGCAAAGACCACACATTTTTTACTACAAAATGTAAAGGATAAATGTAAATCCTGCA TAACTAAAATCACAAACCTATTCCTCAAAAGAATTTAATTTTATATTTATGAGGGGGCCCTTCACTAAAAAGTACATGTAAAAGTACATT TGATGACAATAGCTGCTTAGTTTCCTGTTAAGAGAAGAAACTTTATCTTTTAATTATGTGCTCTTAATATTTGAAGATGAGAGTTAATAC CTGAGATGTTTTTCTGCAACCAAAATTCATTAAATTTGGCTGCCTTATCCTTTTTTTAAGCTAATGAAACTACAGGTTTGAAAAATGACA AAGCTGTTCAGATGATGCTATTAAAGAAATGTGTGTACTAAGCAAAAATATATAAATAGTGACAAATACACATTACCAAGCTTATCTTGC AAGGGAGTTATTTTCATCTAACATAGAAAGTGTGTTTTATCAGACAAATGCTTTTATTTTCATTCTAATAATTTGATACAGAAATTAGTA AAGGCATTTTTTTCTTTTTTTTTCCAGTAAATACATTGGGTCTATAAATGTGCATTTGTAAGGGCCACAAAAGTGAACGTGTGGTACTGT AGTACCACGTGGGAGACCTCTGGTTATGGTTTAGTCCTAGTTCCTTTGTTACTCCTGTGAGCACCGAGAAGAACTGGGCGACTCCCAGTC CCACCTGTGCTGTGACAGTCCCACGTGGCTATGACAGACTGTTTAGTACTTACCCTTCTCAGGTTCCTCAGTGCAGGGGTGCATCAGGGC CTCAATAATAGGGGTATACCTGGGAGGATCCAGCAGTAATCCCCAGGGTACTAGGATTACTAGTACTCTGATGGAACTAGTCTTCCTTCC TTATTCCTCGAACATGCAGTACATAAAAAGGGGAAAAGGAGAAAAAAAAAGCCTTACTTTGTTTTACTTGCCATTTATTGTAAGGAAACT TTAAAGCATTTTTTAGGAAATACTCAAAAGCAAGGTTGGAAAATGTTTTATCTTTCTATAGAAAGTTGGGTACAGTATGTAACTGCGGGA AACCCACTGCCCCTTTGTAAGCTGTGGAACCCAAACTGTATGGGGATATTTGATGTTTTCAGAAAGAGGAAGAAAATATGGTCCAAATTA >5267_5267_1_AP3B1-IPO11_AP3B1_chr5_77590276_ENST00000255194_IPO11_chr5_61762958_ENST00000325324_length(amino acids)=792AA_BP= MWNHHTDTFLQEVSSGNEAAILSSLERTLLSLKVLRKLTVNGFVEPHKNMEVMGFLHGIFERLKQFLECSRSIGTDNVCRDRLEKTIILF TKVLLDFLDQHPFSFTPLIQRSLEFSVSYVFTEVGEGVTFERFIVQCMNLIKMIVKNYAYKPSKNFEDSSPETLEAHKIKMAFFTYPTLT EICRRLVSHYFLLTEEELTMWEEDPEGFTVEETGGDSWKYSLRPCTEVLFIDIFHEYNQTLTPVLLEMMQTLQGPTNVEDMNALLIKDAV YNAVGLAAYELFDSVDFDQWFKNQLLPELQVIHNRYKPLRRRVIWLIGQWISVKFKSDLRPMLYEAICNLLQDQDLVVRIETATTLKLTV DDFEFRTDQFLPYLETMFTLLFQLLQQVTECDTKMHVLHVLSCVIERVNMQIRPYVGCLVQYLPLLWKQSEEHNMLRCAILTTLIHLVQG LGADSKNLYPFLLPVIQLSTDVSQPPHVYLLEDGLELWLVTLENSPCITPELLRIFQNMSPLLELSSENLRTCFKIINGYIFLSSTEFLQ TYAVGLCQSFCELLKEITTEGQVQVLKVVENALKVNPILGPQMFQPILPYVFKGIIEGERYPVVMSTYLGVMGRVLLQNTSFFSSLLNEM AHKFNQEMDQLLGNMIEMWVDRMDNITQPERRKLSALALLSLLPSDNSVIQDKFCGIINISVEGLHDVMTEDPETGTYKDCMLMSHLEEP -------------------------------------------------------------- >5267_5267_2_AP3B1-IPO11_AP3B1_chr5_77590276_ENST00000255194_IPO11_chr5_61762958_ENST00000409296_length(transcript)=3102nt_BP=304nt ACTGCGCATGCGCAGGGGGTGGACTGCCAGGTCGGCTCAGGGAGCCGTGACGAGTCCGGAAGCGCCTGCGCGCGCTCCTCCGTACGAGAA CTAGTTTTGTTCCGTGCCCTCTGGACTGGAACCTTTTGGAGAGAACCCCCGGCAGGACCAACCCCGCACCCGCCAGCACCGCGGCAATGT CCAGCAATAGTTTTCCTTACAATGAGCAGTCCGGAGGAGGGGAGGCGACGGAGCTGGGTCAGGAGGCGACCTCAACCATTTCCCCCTCGG GGGCCTTCGGCCTCTTTAGCAGCGATTTGAAGAATTAGCTTCTGGAATTTATAATTTTGCCTGCTCTCTGTGGAATCACCACACAGACAC ATTCCTGCAAGAAGTTTCTTCTGGCAATGAAGCTGCAATTTTGAGTTCACTAGAACGAACACTGCTATCATTGAAAGTGCTGCGTAAGTT AACTGTTAATGGATTTGTGGAACCTCATAAGAATATGGAGGTGATGGGTTTTTTACATGGAATATTTGAACGTCTAAAACAGTTTCTGGA ATGCAGTAGAAGTATAGGTACAGATAATGTGTGTAGAGATAGACTGGAAAAGACCATCATTCTTTTTACTAAAGTGCTTTTGGACTTCTT GGATCAGCATCCTTTTTCATTTACTCCTCTAATTCAGAGATCACTGGAATTTTCTGTAAGCTATGTTTTTACAGAAGTTGGTGAAGGCGT TACATTTGAACGATTCATTGTCCAATGTATGAATCTTATTAAGATGATTGTCAAAAATTATGCTTATAAGCCATCCAAAAATTTTGAAGA TAGCAGCCCTGAAACTCTTGAAGCCCATAAGATTAAGATGGCATTCTTCACATATCCTACTTTGACAGAGATATGTAGAAGATTAGTCTC TCATTATTTCCTATTAACTGAAGAAGAACTGACAATGTGGGAAGAAGACCCAGAAGGCTTTACAGTGGAAGAAACAGGAGGAGATTCTTG GAAATATAGTTTGAGGCCATGCACTGAAGTATTATTTATAGATATATTCCATGAATATAATCAGACTCTTACTCCTGTACTTCTAGAAAT GATGCAAACACTTCAAGGACCCACAAATGTGGAAGATATGAATGCACTGTTAATCAAAGATGCTGTGTATAATGCTGTTGGATTAGCTGC TTATGAGCTCTTTGACAGTGTTGATTTTGATCAGTGGTTTAAAAACCAGCTTCTTCCAGAATTACAAGTCATTCACAATAGGTATAAGCC ATTGCGACGCAGGGTGATTTGGCTCATCGGTCAGTGGATTTCTGTGAAATTCAAGTCTGACTTAAGACCCATGCTTTATGAAGCAATCTG TAACTTGCTTCAAGATCAAGATTTAGTGGTCCGTATTGAAACAGCTACAACTTTGAAGTTAACTGTTGATGATTTTGAATTTAGAACAGA TCAGTTTCTACCGTATTTGGAAACCATGTTCACACTACTTTTTCAGTTACTGCAGCAAGTTACAGAATGTGACACAAAGATGCATGTTTT GCATGTCCTTTCTTGTGTGATCGAAAGAGTCAACATGCAGATACGACCATATGTGGGATGTTTGGTACAATATTTGCCCCTCCTTTGGAA GCAGAGTGAAGAACACAATATGTTGAGATGTGCTATTTTGACAACACTTATTCATCTTGTTCAGGGATTAGGAGCAGACAGCAAGAACCT GTACCCTTTCCTGCTCCCAGTTATTCAACTGAGTACAGATGTTTCACAGCCTCCACATGTTTATCTTCTGGAAGATGGTTTAGAATTATG GTTAGTAACTTTGGAAAACAGTCCATGTATTACACCAGAGTTGCTTCGTATATTTCAGAATATGTCACCACTTCTTGAACTAAGTTCAGA AAATCTTAGAACTTGCTTTAAGATCATCAATGGTTATATCTTTTTATCATCAACAGAATTTTTACAGACATACGCAGTAGGTCTATGCCA GTCCTTTTGTGAACTTTTAAAGGAAATTACTACAGAAGGTCAAGTTCAGGTGCTCAAGGTTGTGGAAAATGCCCTTAAAGTGAACCCAAT ACTAGGTCCACAAATGTTTCAACCGATTTTACCCTATGTTTTCAAGGGTATTATAGAAGGGGAGAGGTATCCTGTAGTGATGTCCACGTA TCTTGGAGTTATGGGTCGAGTTCTACTACAAAACACTAGTTTTTTTTCTTCACTACTTAATGAGATGGCCCATAAATTTAATCAGGAGAT GGACCAGCTTTTGGGAAATATGATTGAAATGTGGGTTGATCGAATGGACAACATTACCCAGCCTGAAAGAAGAAAACTTTCAGCTTTGGC TTTGCTCTCTCTTCTGCCATCTGATAATAGTGTTATCCAAGATAAATTCTGTGGGATTATAAACATTTCAGTAGAAGGCCTGCATGATGT CATGACGGAAGATCCTGAAACAGGAACTTATAAAGACTGTATGTTGATGTCTCATCTTGAGGAACCAAAAGTAACAGAAGATGAAGAACC ACCCACAGAACAAGATAAGAGGAAAAAGATGCTGGCCCTGAAGGACCCTGTTCATACAGTGTCACTGCAGCAGTTCATCTACGAGAAGCT CAAGGCACAGCAGGAGATGCTAGGAGAACAAGGTTTCCAGTCCCTCATGGAAACAGTGGATACGGAGATTGTCACCCAGCTACAGGAGTT TTTGCAAGGATTCTAAGAGCACATGACATGTGGCTGCCTCCCCTTTCAGAAACAAGCTGAGTAACCCAGCCTGCCGTTTGTATGTGAGAG CCTGCTGAGATGAAGAAATCACTTCATGAAAATAAGCAAAGACCACACATTTTTTACTACAAAATGTAAAGGATAAATGTAAATCCTGCA TAACTAAAATCACAAACCTATTCCTCAAAAGAATTTAATTTTATATTTATGAGGGGGCCCTTCACTAAAAAGTACATGTAAAAGTACATT TGATGACAATAGCTGCTTAGTTTCCTGTTAAGAGAAGAAACTTTATCTTTTAATTATGTGCTCTTAATATTTGAAGATGAGAGTTAATAC >5267_5267_2_AP3B1-IPO11_AP3B1_chr5_77590276_ENST00000255194_IPO11_chr5_61762958_ENST00000409296_length(amino acids)=792AA_BP= MWNHHTDTFLQEVSSGNEAAILSSLERTLLSLKVLRKLTVNGFVEPHKNMEVMGFLHGIFERLKQFLECSRSIGTDNVCRDRLEKTIILF TKVLLDFLDQHPFSFTPLIQRSLEFSVSYVFTEVGEGVTFERFIVQCMNLIKMIVKNYAYKPSKNFEDSSPETLEAHKIKMAFFTYPTLT EICRRLVSHYFLLTEEELTMWEEDPEGFTVEETGGDSWKYSLRPCTEVLFIDIFHEYNQTLTPVLLEMMQTLQGPTNVEDMNALLIKDAV YNAVGLAAYELFDSVDFDQWFKNQLLPELQVIHNRYKPLRRRVIWLIGQWISVKFKSDLRPMLYEAICNLLQDQDLVVRIETATTLKLTV DDFEFRTDQFLPYLETMFTLLFQLLQQVTECDTKMHVLHVLSCVIERVNMQIRPYVGCLVQYLPLLWKQSEEHNMLRCAILTTLIHLVQG LGADSKNLYPFLLPVIQLSTDVSQPPHVYLLEDGLELWLVTLENSPCITPELLRIFQNMSPLLELSSENLRTCFKIINGYIFLSSTEFLQ TYAVGLCQSFCELLKEITTEGQVQVLKVVENALKVNPILGPQMFQPILPYVFKGIIEGERYPVVMSTYLGVMGRVLLQNTSFFSSLLNEM AHKFNQEMDQLLGNMIEMWVDRMDNITQPERRKLSALALLSLLPSDNSVIQDKFCGIINISVEGLHDVMTEDPETGTYKDCMLMSHLEEP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for AP3B1-IPO11 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for AP3B1-IPO11 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for AP3B1-IPO11 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
