
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MED26-SLC1A6 (FusionGDB2 ID:52750) |
Fusion Gene Summary for MED26-SLC1A6 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MED26-SLC1A6 | Fusion gene ID: 52750 | Hgene | Tgene | Gene symbol | MED26 | SLC1A6 | Gene ID | 9441 | 6511 |
| Gene name | mediator complex subunit 26 | solute carrier family 1 member 6 | |
| Synonyms | CRSP7|CRSP70 | EAAT4 | |
| Cytomap | 19p13.11 | 19p13.12 | |
| Type of gene | protein-coding | protein-coding | |
| Description | mediator of RNA polymerase II transcription subunit 26ARC70CRSP complex subunit 7activator-recruited cofactor 70 kDa componentcofactor required for Sp1 transcriptional activation subunit 7cofactor required for Sp1 transcriptional activation, subunit | excitatory amino acid transporter 4sodium-dependent glutamate/aspartate transportersolute carrier family 1 (high affinity aspartate/glutamate transporter), member 6 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | O95402 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000263390, | ENST00000544886, ENST00000598504, ENST00000600144, ENST00000221742, ENST00000430939, | |
| Fusion gene scores | * DoF score | 16 X 4 X 12=768 | 7 X 6 X 7=294 |
| # samples | 18 | 9 | |
| ** MAII score | log2(18/768*10)=-2.09310940439148 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/294*10)=-1.70781924850669 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MED26 [Title/Abstract] AND SLC1A6 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MED26(16738683)-SLC1A6(15083729), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | MED26-SLC1A6 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. MED26-SLC1A6 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. MED26-SLC1A6 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. MED26-SLC1A6 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MED26 | GO:0006357 | regulation of transcription by RNA polymerase II | 15989967 |
 Fusion gene breakpoints across MED26 (5'-gene) Fusion gene breakpoints across MED26 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
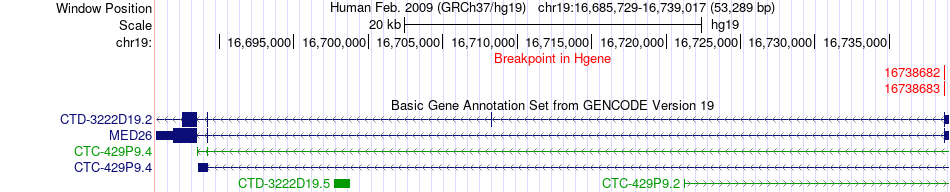 |
 Fusion gene breakpoints across SLC1A6 (3'-gene) Fusion gene breakpoints across SLC1A6 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
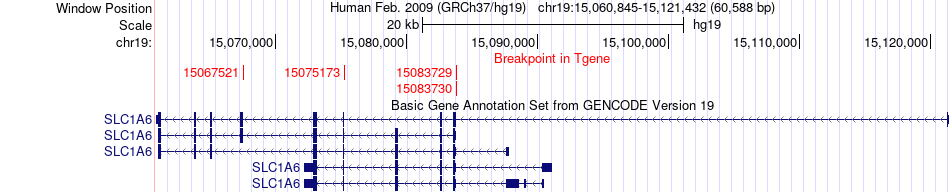 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | OV | TCGA-61-1728-01A | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15067521 | - |
| ChimerDB4 | OV | TCGA-61-1728-01A | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15075173 | - |
| ChimerDB4 | OV | TCGA-61-1728-01A | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - |
| ChimerDB4 | OV | TCGA-61-1728 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15067521 | - |
| ChimerDB4 | OV | TCGA-61-1728 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15075173 | - |
| ChimerDB4 | OV | TCGA-61-1728 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - |
Top |
Fusion Gene ORF analysis for MED26-SLC1A6 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000263390 | ENST00000544886 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - |
| 5CDS-5UTR | ENST00000263390 | ENST00000544886 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - |
| 5CDS-5UTR | ENST00000263390 | ENST00000598504 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - |
| 5CDS-5UTR | ENST00000263390 | ENST00000598504 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - |
| 5CDS-5UTR | ENST00000263390 | ENST00000600144 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - |
| 5CDS-5UTR | ENST00000263390 | ENST00000600144 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - |
| 5CDS-intron | ENST00000263390 | ENST00000544886 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15067521 | - |
| 5CDS-intron | ENST00000263390 | ENST00000544886 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15067521 | - |
| 5CDS-intron | ENST00000263390 | ENST00000598504 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15067521 | - |
| 5CDS-intron | ENST00000263390 | ENST00000598504 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15067521 | - |
| 5CDS-intron | ENST00000263390 | ENST00000600144 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15067521 | - |
| 5CDS-intron | ENST00000263390 | ENST00000600144 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15067521 | - |
| Frame-shift | ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15067521 | - |
| Frame-shift | ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15067521 | - |
| Frame-shift | ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15067521 | - |
| Frame-shift | ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15067521 | - |
| Frame-shift | ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000544886 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000544886 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000598504 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000598504 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000600144 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15075173 | - |
| Frame-shift | ENST00000263390 | ENST00000600144 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15075173 | - |
| In-frame | ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - |
| In-frame | ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - |
| In-frame | ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - |
| In-frame | ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000263390 | MED26 | chr19 | 16738683 | - | ENST00000430939 | SLC1A6 | chr19 | 15083729 | - | 1993 | 335 | 263 | 1831 | 522 |
| ENST00000263390 | MED26 | chr19 | 16738683 | - | ENST00000221742 | SLC1A6 | chr19 | 15083729 | - | 2054 | 335 | 343 | 2037 | 564 |
| ENST00000263390 | MED26 | chr19 | 16738682 | - | ENST00000430939 | SLC1A6 | chr19 | 15083730 | - | 1993 | 335 | 263 | 1831 | 522 |
| ENST00000263390 | MED26 | chr19 | 16738682 | - | ENST00000221742 | SLC1A6 | chr19 | 15083730 | - | 2054 | 335 | 343 | 2037 | 564 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - | 0.01632094 | 0.983679 |
| ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738683 | - | SLC1A6 | chr19 | 15083729 | - | 0.01355212 | 0.98644793 |
| ENST00000263390 | ENST00000430939 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - | 0.01632094 | 0.983679 |
| ENST00000263390 | ENST00000221742 | MED26 | chr19 | 16738682 | - | SLC1A6 | chr19 | 15083730 | - | 0.01355212 | 0.98644793 |
Top |
Fusion Genomic Features for MED26-SLC1A6 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
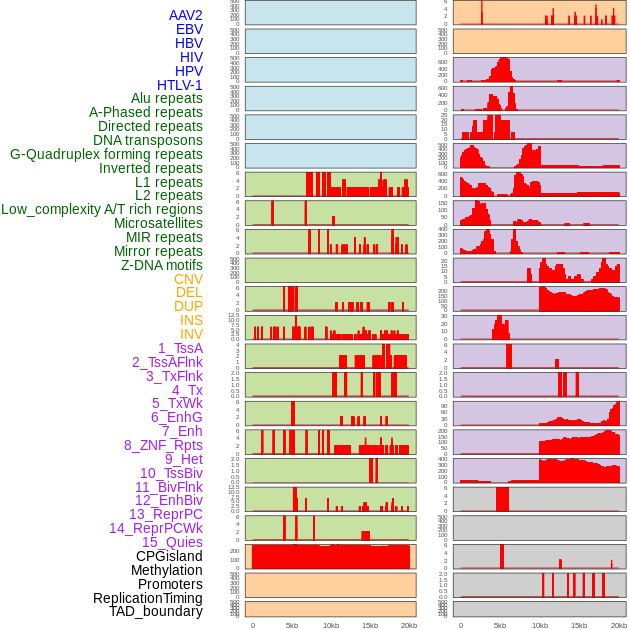 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for MED26-SLC1A6 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:16738683/chr19:15083729) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MED26 | . |
| FUNCTION: Component of the Mediator complex, a coactivator involved in the regulated transcription of nearly all RNA polymerase II-dependent genes. Mediator functions as a bridge to convey information from gene-specific regulatory proteins to the basal RNA polymerase II transcription machinery. Mediator is recruited to promoters by direct interactions with regulatory proteins and serves as a scaffold for the assembly of a functional pre-initiation complex with RNA polymerase II and the general transcription factors. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 371_401 | 68 | 565.0 | Intramembrane | Discontinuously helical | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 451_484 | 68 | 565.0 | Intramembrane | Discontinuously helical | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 371_401 | 68 | 565.0 | Intramembrane | Discontinuously helical | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 451_484 | 68 | 565.0 | Intramembrane | Discontinuously helical | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 388_390 | 68 | 565.0 | Region | Aspartate binding | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 468_472 | 68 | 565.0 | Region | Aspartate binding | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 388_390 | 68 | 565.0 | Region | Aspartate binding | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 468_472 | 68 | 565.0 | Region | Aspartate binding | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 133_153 | 68 | 565.0 | Transmembrane | Helical | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 262_285 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 295_322 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 344_365 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 411_437 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 498_519 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 99_119 | 68 | 565.0 | Transmembrane | Helical | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 133_153 | 68 | 565.0 | Transmembrane | Helical | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 262_285 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 295_322 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 344_365 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 411_437 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 498_519 | 68 | 565.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 99_119 | 68 | 565.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MED26 | chr19:16738682 | chr19:15083730 | ENST00000263390 | - | 1 | 3 | 228_325 | 24 | 601.0 | Compositional bias | Note=Pro-rich |
| Hgene | MED26 | chr19:16738683 | chr19:15083729 | ENST00000263390 | - | 1 | 3 | 228_325 | 24 | 601.0 | Compositional bias | Note=Pro-rich |
| Hgene | MED26 | chr19:16738682 | chr19:15083730 | ENST00000263390 | - | 1 | 3 | 10_87 | 24 | 601.0 | Domain | TFIIS N-terminal |
| Hgene | MED26 | chr19:16738683 | chr19:15083729 | ENST00000263390 | - | 1 | 3 | 10_87 | 24 | 601.0 | Domain | TFIIS N-terminal |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 1_55 | 68 | 565.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 1_55 | 68 | 565.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC1A6 | chr19:16738682 | chr19:15083730 | ENST00000221742 | -1 | 9 | 56_76 | 68 | 565.0 | Transmembrane | Helical | |
| Tgene | SLC1A6 | chr19:16738683 | chr19:15083729 | ENST00000221742 | -1 | 9 | 56_76 | 68 | 565.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for MED26-SLC1A6 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >52750_52750_1_MED26-SLC1A6_MED26_chr19_16738682_ENST00000263390_SLC1A6_chr19_15083730_ENST00000221742_length(transcript)=2054nt_BP=335nt TCACTGGAGTTTGGTTCTCGAGGCGGCGGCAGCGAGCGGCGGCGGCTCCGGCGGCTCCTCCTCCTCCTTTGGTGGCGGCGGCGGCCCGGG ACCGAGACCCGGCCCCGGCTCCCCTCTCCGCCACCCCGCGCGCCCCCAACCCCCCGGCGCCGCGGGGAACATGCGGCTCCGGCTCCCGGA GGCCGGCGAGTGAGTGACCCCCGTCCCCCGCCTCAGCCCGCCCGCCCGCTGGCCCGGCCGCAACCCCCCGCCTCGCCCAGGCAATGACAG CGGCTCCGGCGTCTCCGCAGCAGATCAGGGACCGGCTGCTGCAGGCCATCGACCCCCAGAGCAACGATAGACCATGAGCAGCCATGGCAA CAGCCTGTTCCTGCGGGAGAGCGGCCAGCGGCTGGGCCGGGTGGGCTGGCTGCAGCGGCTGCAGGAAAGCCTGCAGCAGAGAGCACTGCG CACGCGCCTGCGCCTGCAGACCATGACCCTCGAGCACGTGCTGCGCTTCCTGCGCCGAAACGCCTTCATTCTGCTGACGGTCAGCGCCGT GGTCATTGGGGTCAGCCTGGCCTTTGCCCTGCGCCCATATCAGCTCACCTACCGCCAGATCAAGTACTTCTCTTTTCCTGGAGAGCTTCT GATGAGGATGCTGCAGATGCTGGTGTTACCTCTCATTGTCTCCAGCCTGGTCACAGGTATGGCATCCCTGGACAACAAGGCCACGGGGCG GATGGGGATGCGGGCAGCTGTGTACTACATGGTGACCACCATCATCGCGGTCTTCATCGGCATCCTCATGGTCACCATCATCCATCCCGG GAAGGGCTCCAAGGAGGGGCTGCACCGGGAGGGCCGGATCGAGACCATCCCCACAGCTGATGCCTTCATGGACCTGATCAGAAATATGTT TCCACCAAACCTTGTGGAGGCCTGCTTCAAACAGTTCAAGACGCAGTACAGCACGAGGGTGGTAACCAGGACCATGGTGAGGACAGAGAA CGGGTCTGAGCCGGGTGCCTCCATGCCTCCTCCATTCTCAGTGGAGAACGGAACCAGCTTCCTGGAAAATGTCACTCGGGCCTTGGGTAC CCTGCAGGAGATGCTGAGCTTTGAGGAGACTGTACCCGTGCCTGGCTCCGCCAATGGCATCAACGCCCTGGGCCTCGTGGTCTTCTCTGT GGCCTTTGGGCTGGTCATTGGTGGCATGAAACACAAGGGCAGAGTCCTCAGGGACTTCTTCGACAGCCTCAATGAGGCTATTATGAGGCT GGTGGGCATCATTATCTGGTATGCACCTGTGGGCATCCTGTTCCTGATTGCTGGGAAGATTCTGGAGATGGAAGACATGGCCGTCCTGGG GGGTCAGCTGGGCATGTACACCCTGACCGTCATCGTGGGCCTGTTCCTCCATGCCGGCATTGTCCTTCCCCTCATCTACTTCCTCGTCAC TCACCGGAACCCCTTCCCCTTCATTGGGGGCATGCTACAAGCCCTCATCACCGCTATGGGCACGTCTTCCAGCTCGGCAACGCTGCCCAT CACCTTCCGCTGCCTGGAGGAGGGCCTGGGTGTGGACCGCCGCATCACCAGGTTCGTCCTGCCCGTGGGCGCCACGGTCAACATGGATGG CACTGCCCTCTACGAGGCCCTGGCTGCCATCTTCATTGCTCAAGTTAACAACTACGAGCTCAACCTGGGTCAGATCACAACCATCAGCAT CACGGCCACAGCAGCCAGTGTTGGGGCTGCTGGCATCCCCCAGGCGGGTCTGGTCACCATGGTCATTGTGCTTACGTCGGTCGGCTTGCC CACGGAAGACATCACGCTCATCATTGCCGTGGACTGGTTCCTTGACCGGCTTCGCACAATGACCAACGTACTGGGGGACTCAATTGGAGC GGCCGTCATCGAGCACTTGTCTCAGCGGGAGCTGGAGCTTCAGGAAGCTGAGCTTACCCTCCCCAGCCTGGGGAAACCCTACAAGTCCCT >52750_52750_1_MED26-SLC1A6_MED26_chr19_16738682_ENST00000263390_SLC1A6_chr19_15083730_ENST00000221742_length(amino acids)=564AA_BP= MSSHGNSLFLRESGQRLGRVGWLQRLQESLQQRALRTRLRLQTMTLEHVLRFLRRNAFILLTVSAVVIGVSLAFALRPYQLTYRQIKYFS FPGELLMRMLQMLVLPLIVSSLVTGMASLDNKATGRMGMRAAVYYMVTTIIAVFIGILMVTIIHPGKGSKEGLHREGRIETIPTADAFMD LIRNMFPPNLVEACFKQFKTQYSTRVVTRTMVRTENGSEPGASMPPPFSVENGTSFLENVTRALGTLQEMLSFEETVPVPGSANGINALG LVVFSVAFGLVIGGMKHKGRVLRDFFDSLNEAIMRLVGIIIWYAPVGILFLIAGKILEMEDMAVLGGQLGMYTLTVIVGLFLHAGIVLPL IYFLVTHRNPFPFIGGMLQALITAMGTSSSSATLPITFRCLEEGLGVDRRITRFVLPVGATVNMDGTALYEALAAIFIAQVNNYELNLGQ ITTISITATAASVGAAGIPQAGLVTMVIVLTSVGLPTEDITLIIAVDWFLDRLRTMTNVLGDSIGAAVIEHLSQRELELQEAELTLPSLG -------------------------------------------------------------- >52750_52750_2_MED26-SLC1A6_MED26_chr19_16738682_ENST00000263390_SLC1A6_chr19_15083730_ENST00000430939_length(transcript)=1993nt_BP=335nt TCACTGGAGTTTGGTTCTCGAGGCGGCGGCAGCGAGCGGCGGCGGCTCCGGCGGCTCCTCCTCCTCCTTTGGTGGCGGCGGCGGCCCGGG ACCGAGACCCGGCCCCGGCTCCCCTCTCCGCCACCCCGCGCGCCCCCAACCCCCCGGCGCCGCGGGGAACATGCGGCTCCGGCTCCCGGA GGCCGGCGAGTGAGTGACCCCCGTCCCCCGCCTCAGCCCGCCCGCCCGCTGGCCCGGCCGCAACCCCCCGCCTCGCCCAGGCAATGACAG CGGCTCCGGCGTCTCCGCAGCAGATCAGGGACCGGCTGCTGCAGGCCATCGACCCCCAGAGCAACATAGACCATGAGCAGCCATGGCAAC AGCCTGTTCCTGCGGGAGAGCGGCCAGCGGCTGGGCCGGGTGGGCTGGCTGCAGCGGCTGCAGGAAAGCCTGCAGCAGAGAGCACTGCGC ACGCGCCTGCGCCTGCAGACCATGACCCTCGAGCACGTGCTGCGCTTCCTGCGCCGAAACGCCTTCATTCTGCTGACGGTCAGCGCCGTG GTCATTGGGGTCAGCCTGGCCTTTGCCCTGCGCCCATATCAGCTCACCTACCGCCAGATCAAGTACTTCTCTTTTCCTGGAGAGCTTCTG ATGAGGATGCTGCAGATGCTGGTGTTACCTCTCATTGTCTCCAGCCTGGTCACAGAAATATGTTTCCACCAAACCTTGTGGAGGCCTGCT TCAAACAGTTCAAGACGCAGTACAGCACGAGGGTGGTAACCAGGACCATGGTGAGGACAGAGAACGGGTCTGAGCCGGGTGCCTCCATGC CTCCTCCATTCTCAGTGGAGAACGGAACCAGCTTCCTGGAAAATGTCACTCGGGCCTTGGGTACCCTGCAGGAGATGCTGAGCTTTGAGG AGACTGTACCCGTGCCTGGCTCCGCCAATGGCATCAACGCCCTGGGCCTCGTGGTCTTCTCTGTGGCCTTTGGGCTGGTCATTGGTGGCA TGAAACACAAGGGCAGAGTCCTCAGGGACTTCTTCGACAGCCTCAATGAGGCTATTATGAGGCTGGTGGGCATCATTATCTGGTATGCAC CTGTGGGCATCCTGTTCCTGATTGCTGGGAAGATTCTGGAGATGGAAGACATGGCCGTCCTGGGGGGTCAGCTGGGCATGTACACCCTGA CCGTCATCGTGGGCCTGTTCCTCCATGCCGGCATTGTCCTTCCCCTCATCTACTTCCTCGTCACTCACCGGAACCCCTTCCCCTTCATTG GGGGCATGCTACAAGCCCTCATCACCGCTATGGGCACGTCTTCCAGCTCGGCAACGCTGCCCATCACCTTCCGCTGCCTGGAGGAGGGCC TGGGTGTGGACCGCCGCATCACCAGGTTCGTCCTGCCCGTGGGCGCCACGGTCAACATGGATGGCACTGCCCTCTACGAGGCCCTGGCTG CCATCTTCATTGCTCAAGTTAACAACTACGAGCTCAACCTGGGTCAGATCACAACCATCAGCATCACGGCCACAGCAGCCAGTGTTGGGG CTGCTGGCATCCCCCAGGCGGGTCTGGTCACCATGGTCATTGTGCTTACGTCGGTCGGCTTGCCCACGGAAGACATCACGCTCATCATTG CCGTGGACTGGTTCCTTGACCGGCTTCGCACAATGACCAACGTACTGGGGGACTCAATTGGAGCGGCCGTCATCGAGCACTTGTCTCAGC GGGAGCTGGAGCTTCAGGAAGCTGAGCTTACCCTCCCCAGCCTGGGGAAACCCTACAAGTCCCTCATGGCACAGGAGAAGGGGGCATCCC GGGGACGGGGAGGCAACGAGAGTGCTATGTGAGGGGCCTCCAGCTCTGCCCCCCCAGAGAGGAGGGAAGGGGGCTGGGGAGGGGAGTCCT GGTGACACATCTGTTGCCCAACTGACCGTGGGCTGAACACACGTTCTGCTTGACTCATTTAGGGGGGAGGGAAAAGTGAAATAAAGGAGC >52750_52750_2_MED26-SLC1A6_MED26_chr19_16738682_ENST00000263390_SLC1A6_chr19_15083730_ENST00000430939_length(amino acids)=522AA_BP=24 MTAAPASPQQIRDRLLQAIDPQSNIDHEQPWQQPVPAGERPAAGPGGLAAAAAGKPAAESTAHAPAPADHDPRARAALPAPKRLHSADGQ RRGHWGQPGLCPAPISAHLPPDQVLLFSWRASDEDAADAGVTSHCLQPGHRNMFPPNLVEACFKQFKTQYSTRVVTRTMVRTENGSEPGA SMPPPFSVENGTSFLENVTRALGTLQEMLSFEETVPVPGSANGINALGLVVFSVAFGLVIGGMKHKGRVLRDFFDSLNEAIMRLVGIIIW YAPVGILFLIAGKILEMEDMAVLGGQLGMYTLTVIVGLFLHAGIVLPLIYFLVTHRNPFPFIGGMLQALITAMGTSSSSATLPITFRCLE EGLGVDRRITRFVLPVGATVNMDGTALYEALAAIFIAQVNNYELNLGQITTISITATAASVGAAGIPQAGLVTMVIVLTSVGLPTEDITL -------------------------------------------------------------- >52750_52750_3_MED26-SLC1A6_MED26_chr19_16738683_ENST00000263390_SLC1A6_chr19_15083729_ENST00000221742_length(transcript)=2054nt_BP=335nt TCACTGGAGTTTGGTTCTCGAGGCGGCGGCAGCGAGCGGCGGCGGCTCCGGCGGCTCCTCCTCCTCCTTTGGTGGCGGCGGCGGCCCGGG ACCGAGACCCGGCCCCGGCTCCCCTCTCCGCCACCCCGCGCGCCCCCAACCCCCCGGCGCCGCGGGGAACATGCGGCTCCGGCTCCCGGA GGCCGGCGAGTGAGTGACCCCCGTCCCCCGCCTCAGCCCGCCCGCCCGCTGGCCCGGCCGCAACCCCCCGCCTCGCCCAGGCAATGACAG CGGCTCCGGCGTCTCCGCAGCAGATCAGGGACCGGCTGCTGCAGGCCATCGACCCCCAGAGCAACGATAGACCATGAGCAGCCATGGCAA CAGCCTGTTCCTGCGGGAGAGCGGCCAGCGGCTGGGCCGGGTGGGCTGGCTGCAGCGGCTGCAGGAAAGCCTGCAGCAGAGAGCACTGCG CACGCGCCTGCGCCTGCAGACCATGACCCTCGAGCACGTGCTGCGCTTCCTGCGCCGAAACGCCTTCATTCTGCTGACGGTCAGCGCCGT GGTCATTGGGGTCAGCCTGGCCTTTGCCCTGCGCCCATATCAGCTCACCTACCGCCAGATCAAGTACTTCTCTTTTCCTGGAGAGCTTCT GATGAGGATGCTGCAGATGCTGGTGTTACCTCTCATTGTCTCCAGCCTGGTCACAGGTATGGCATCCCTGGACAACAAGGCCACGGGGCG GATGGGGATGCGGGCAGCTGTGTACTACATGGTGACCACCATCATCGCGGTCTTCATCGGCATCCTCATGGTCACCATCATCCATCCCGG GAAGGGCTCCAAGGAGGGGCTGCACCGGGAGGGCCGGATCGAGACCATCCCCACAGCTGATGCCTTCATGGACCTGATCAGAAATATGTT TCCACCAAACCTTGTGGAGGCCTGCTTCAAACAGTTCAAGACGCAGTACAGCACGAGGGTGGTAACCAGGACCATGGTGAGGACAGAGAA CGGGTCTGAGCCGGGTGCCTCCATGCCTCCTCCATTCTCAGTGGAGAACGGAACCAGCTTCCTGGAAAATGTCACTCGGGCCTTGGGTAC CCTGCAGGAGATGCTGAGCTTTGAGGAGACTGTACCCGTGCCTGGCTCCGCCAATGGCATCAACGCCCTGGGCCTCGTGGTCTTCTCTGT GGCCTTTGGGCTGGTCATTGGTGGCATGAAACACAAGGGCAGAGTCCTCAGGGACTTCTTCGACAGCCTCAATGAGGCTATTATGAGGCT GGTGGGCATCATTATCTGGTATGCACCTGTGGGCATCCTGTTCCTGATTGCTGGGAAGATTCTGGAGATGGAAGACATGGCCGTCCTGGG GGGTCAGCTGGGCATGTACACCCTGACCGTCATCGTGGGCCTGTTCCTCCATGCCGGCATTGTCCTTCCCCTCATCTACTTCCTCGTCAC TCACCGGAACCCCTTCCCCTTCATTGGGGGCATGCTACAAGCCCTCATCACCGCTATGGGCACGTCTTCCAGCTCGGCAACGCTGCCCAT CACCTTCCGCTGCCTGGAGGAGGGCCTGGGTGTGGACCGCCGCATCACCAGGTTCGTCCTGCCCGTGGGCGCCACGGTCAACATGGATGG CACTGCCCTCTACGAGGCCCTGGCTGCCATCTTCATTGCTCAAGTTAACAACTACGAGCTCAACCTGGGTCAGATCACAACCATCAGCAT CACGGCCACAGCAGCCAGTGTTGGGGCTGCTGGCATCCCCCAGGCGGGTCTGGTCACCATGGTCATTGTGCTTACGTCGGTCGGCTTGCC CACGGAAGACATCACGCTCATCATTGCCGTGGACTGGTTCCTTGACCGGCTTCGCACAATGACCAACGTACTGGGGGACTCAATTGGAGC GGCCGTCATCGAGCACTTGTCTCAGCGGGAGCTGGAGCTTCAGGAAGCTGAGCTTACCCTCCCCAGCCTGGGGAAACCCTACAAGTCCCT >52750_52750_3_MED26-SLC1A6_MED26_chr19_16738683_ENST00000263390_SLC1A6_chr19_15083729_ENST00000221742_length(amino acids)=564AA_BP=0 MSSHGNSLFLRESGQRLGRVGWLQRLQESLQQRALRTRLRLQTMTLEHVLRFLRRNAFILLTVSAVVIGVSLAFALRPYQLTYRQIKYFS FPGELLMRMLQMLVLPLIVSSLVTGMASLDNKATGRMGMRAAVYYMVTTIIAVFIGILMVTIIHPGKGSKEGLHREGRIETIPTADAFMD LIRNMFPPNLVEACFKQFKTQYSTRVVTRTMVRTENGSEPGASMPPPFSVENGTSFLENVTRALGTLQEMLSFEETVPVPGSANGINALG LVVFSVAFGLVIGGMKHKGRVLRDFFDSLNEAIMRLVGIIIWYAPVGILFLIAGKILEMEDMAVLGGQLGMYTLTVIVGLFLHAGIVLPL IYFLVTHRNPFPFIGGMLQALITAMGTSSSSATLPITFRCLEEGLGVDRRITRFVLPVGATVNMDGTALYEALAAIFIAQVNNYELNLGQ ITTISITATAASVGAAGIPQAGLVTMVIVLTSVGLPTEDITLIIAVDWFLDRLRTMTNVLGDSIGAAVIEHLSQRELELQEAELTLPSLG -------------------------------------------------------------- >52750_52750_4_MED26-SLC1A6_MED26_chr19_16738683_ENST00000263390_SLC1A6_chr19_15083729_ENST00000430939_length(transcript)=1993nt_BP=335nt TCACTGGAGTTTGGTTCTCGAGGCGGCGGCAGCGAGCGGCGGCGGCTCCGGCGGCTCCTCCTCCTCCTTTGGTGGCGGCGGCGGCCCGGG ACCGAGACCCGGCCCCGGCTCCCCTCTCCGCCACCCCGCGCGCCCCCAACCCCCCGGCGCCGCGGGGAACATGCGGCTCCGGCTCCCGGA GGCCGGCGAGTGAGTGACCCCCGTCCCCCGCCTCAGCCCGCCCGCCCGCTGGCCCGGCCGCAACCCCCCGCCTCGCCCAGGCAATGACAG CGGCTCCGGCGTCTCCGCAGCAGATCAGGGACCGGCTGCTGCAGGCCATCGACCCCCAGAGCAACATAGACCATGAGCAGCCATGGCAAC AGCCTGTTCCTGCGGGAGAGCGGCCAGCGGCTGGGCCGGGTGGGCTGGCTGCAGCGGCTGCAGGAAAGCCTGCAGCAGAGAGCACTGCGC ACGCGCCTGCGCCTGCAGACCATGACCCTCGAGCACGTGCTGCGCTTCCTGCGCCGAAACGCCTTCATTCTGCTGACGGTCAGCGCCGTG GTCATTGGGGTCAGCCTGGCCTTTGCCCTGCGCCCATATCAGCTCACCTACCGCCAGATCAAGTACTTCTCTTTTCCTGGAGAGCTTCTG ATGAGGATGCTGCAGATGCTGGTGTTACCTCTCATTGTCTCCAGCCTGGTCACAGAAATATGTTTCCACCAAACCTTGTGGAGGCCTGCT TCAAACAGTTCAAGACGCAGTACAGCACGAGGGTGGTAACCAGGACCATGGTGAGGACAGAGAACGGGTCTGAGCCGGGTGCCTCCATGC CTCCTCCATTCTCAGTGGAGAACGGAACCAGCTTCCTGGAAAATGTCACTCGGGCCTTGGGTACCCTGCAGGAGATGCTGAGCTTTGAGG AGACTGTACCCGTGCCTGGCTCCGCCAATGGCATCAACGCCCTGGGCCTCGTGGTCTTCTCTGTGGCCTTTGGGCTGGTCATTGGTGGCA TGAAACACAAGGGCAGAGTCCTCAGGGACTTCTTCGACAGCCTCAATGAGGCTATTATGAGGCTGGTGGGCATCATTATCTGGTATGCAC CTGTGGGCATCCTGTTCCTGATTGCTGGGAAGATTCTGGAGATGGAAGACATGGCCGTCCTGGGGGGTCAGCTGGGCATGTACACCCTGA CCGTCATCGTGGGCCTGTTCCTCCATGCCGGCATTGTCCTTCCCCTCATCTACTTCCTCGTCACTCACCGGAACCCCTTCCCCTTCATTG GGGGCATGCTACAAGCCCTCATCACCGCTATGGGCACGTCTTCCAGCTCGGCAACGCTGCCCATCACCTTCCGCTGCCTGGAGGAGGGCC TGGGTGTGGACCGCCGCATCACCAGGTTCGTCCTGCCCGTGGGCGCCACGGTCAACATGGATGGCACTGCCCTCTACGAGGCCCTGGCTG CCATCTTCATTGCTCAAGTTAACAACTACGAGCTCAACCTGGGTCAGATCACAACCATCAGCATCACGGCCACAGCAGCCAGTGTTGGGG CTGCTGGCATCCCCCAGGCGGGTCTGGTCACCATGGTCATTGTGCTTACGTCGGTCGGCTTGCCCACGGAAGACATCACGCTCATCATTG CCGTGGACTGGTTCCTTGACCGGCTTCGCACAATGACCAACGTACTGGGGGACTCAATTGGAGCGGCCGTCATCGAGCACTTGTCTCAGC GGGAGCTGGAGCTTCAGGAAGCTGAGCTTACCCTCCCCAGCCTGGGGAAACCCTACAAGTCCCTCATGGCACAGGAGAAGGGGGCATCCC GGGGACGGGGAGGCAACGAGAGTGCTATGTGAGGGGCCTCCAGCTCTGCCCCCCCAGAGAGGAGGGAAGGGGGCTGGGGAGGGGAGTCCT GGTGACACATCTGTTGCCCAACTGACCGTGGGCTGAACACACGTTCTGCTTGACTCATTTAGGGGGGAGGGAAAAGTGAAATAAAGGAGC >52750_52750_4_MED26-SLC1A6_MED26_chr19_16738683_ENST00000263390_SLC1A6_chr19_15083729_ENST00000430939_length(amino acids)=522AA_BP=24 MTAAPASPQQIRDRLLQAIDPQSNIDHEQPWQQPVPAGERPAAGPGGLAAAAAGKPAAESTAHAPAPADHDPRARAALPAPKRLHSADGQ RRGHWGQPGLCPAPISAHLPPDQVLLFSWRASDEDAADAGVTSHCLQPGHRNMFPPNLVEACFKQFKTQYSTRVVTRTMVRTENGSEPGA SMPPPFSVENGTSFLENVTRALGTLQEMLSFEETVPVPGSANGINALGLVVFSVAFGLVIGGMKHKGRVLRDFFDSLNEAIMRLVGIIIW YAPVGILFLIAGKILEMEDMAVLGGQLGMYTLTVIVGLFLHAGIVLPLIYFLVTHRNPFPFIGGMLQALITAMGTSSSSATLPITFRCLE EGLGVDRRITRFVLPVGATVNMDGTALYEALAAIFIAQVNNYELNLGQITTISITATAASVGAAGIPQAGLVTMVIVLTSVGLPTEDITL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MED26-SLC1A6 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MED26-SLC1A6 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MED26-SLC1A6 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
