
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MEMO1-MCU (FusionGDB2 ID:52979) |
Fusion Gene Summary for MEMO1-MCU |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MEMO1-MCU | Fusion gene ID: 52979 | Hgene | Tgene | Gene symbol | MEMO1 | MCU | Gene ID | 51072 | 90550 |
| Gene name | mediator of cell motility 1 | mitochondrial calcium uniporter | |
| Synonyms | C2orf4|CGI-27|MEMO|NS5ATP7 | C10orf42|CCDC109A|HsMCU | |
| Cytomap | 2p22.3 | 10q22.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein MEMO1C21orf19-like proteinHCV NS5A-transactivated protein 7hepatitis C virus NS5A-transactivated protein 7mediator of ErbB2-driven cell motility 1 | calcium uniporter protein, mitochondrialcoiled-coil domain-containing protein 109A | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | Q9Y316 | Q96AQ8 | |
| Ensembl transtripts involved in fusion gene | ENST00000295065, ENST00000379383, ENST00000404530, ENST00000426310, ENST00000490459, ENST00000407893, | ENST00000605416, ENST00000357157, ENST00000373053, ENST00000536019, | |
| Fusion gene scores | * DoF score | 6 X 6 X 4=144 | 7 X 5 X 5=175 |
| # samples | 6 | 7 | |
| ** MAII score | log2(6/144*10)=-1.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/175*10)=-1.32192809488736 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MEMO1 [Title/Abstract] AND MCU [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MEMO1(32142995)-MCU(74594117), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | MEMO1-MCU seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. MEMO1-MCU seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | MCU | GO:0006851 | mitochondrial calcium ion transmembrane transport | 21685888|24560927 |
| Tgene | MCU | GO:0019722 | calcium-mediated signaling | 21685886|21685888 |
| Tgene | MCU | GO:0036444 | calcium import into the mitochondrion | 22925203 |
| Tgene | MCU | GO:0051259 | protein complex oligomerization | 21685886 |
| Tgene | MCU | GO:0051561 | positive regulation of mitochondrial calcium ion concentration | 21685888|24560927 |
 Fusion gene breakpoints across MEMO1 (5'-gene) Fusion gene breakpoints across MEMO1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
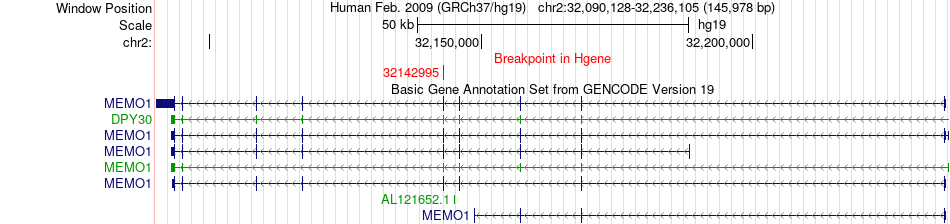 |
 Fusion gene breakpoints across MCU (3'-gene) Fusion gene breakpoints across MCU (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
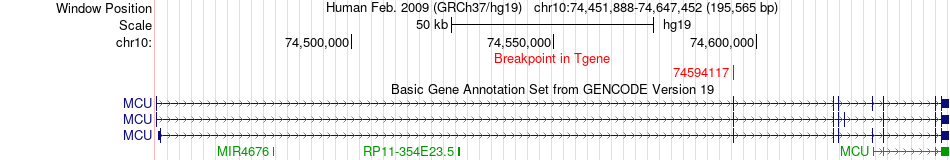 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | ACC | TCGA-OR-A5K5-01A | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
Top |
Fusion Gene ORF analysis for MEMO1-MCU |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000295065 | ENST00000605416 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5CDS-intron | ENST00000379383 | ENST00000605416 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5CDS-intron | ENST00000404530 | ENST00000605416 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5CDS-intron | ENST00000426310 | ENST00000605416 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5UTR-3CDS | ENST00000490459 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5UTR-3CDS | ENST00000490459 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5UTR-3CDS | ENST00000490459 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| 5UTR-intron | ENST00000490459 | ENST00000605416 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000295065 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000295065 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000295065 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000379383 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000379383 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000379383 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000426310 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000426310 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| Frame-shift | ENST00000426310 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| In-frame | ENST00000404530 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| In-frame | ENST00000404530 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| In-frame | ENST00000404530 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| intron-3CDS | ENST00000407893 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| intron-3CDS | ENST00000407893 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| intron-3CDS | ENST00000407893 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
| intron-intron | ENST00000407893 | ENST00000605416 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000404530 | MEMO1 | chr2 | 32142995 | - | ENST00000373053 | MCU | chr10 | 74594117 | + | 3265 | 487 | 517 | 1392 | 291 |
| ENST00000404530 | MEMO1 | chr2 | 32142995 | - | ENST00000357157 | MCU | chr10 | 74594117 | + | 3202 | 487 | 517 | 1329 | 270 |
| ENST00000404530 | MEMO1 | chr2 | 32142995 | - | ENST00000536019 | MCU | chr10 | 74594117 | + | 3263 | 487 | 517 | 1392 | 291 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000404530 | ENST00000373053 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + | 0.024543503 | 0.97545654 |
| ENST00000404530 | ENST00000357157 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + | 0.023811832 | 0.97618824 |
| ENST00000404530 | ENST00000536019 | MEMO1 | chr2 | 32142995 | - | MCU | chr10 | 74594117 | + | 0.021663027 | 0.97833705 |
Top |
Fusion Genomic Features for MEMO1-MCU |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
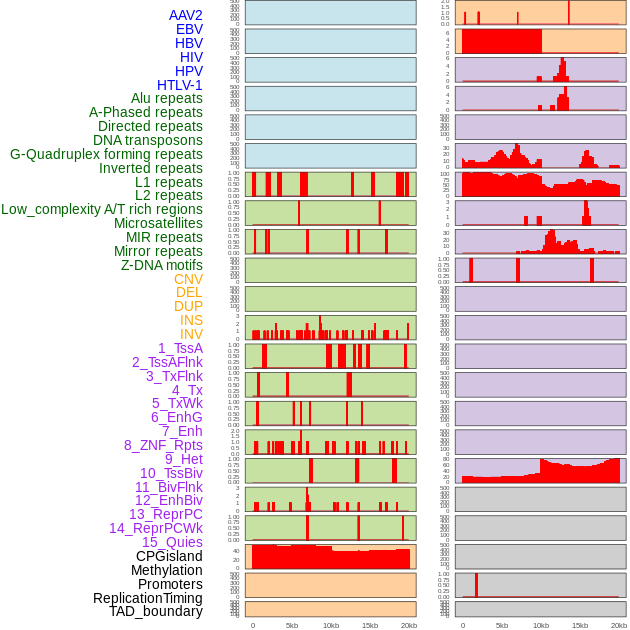
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
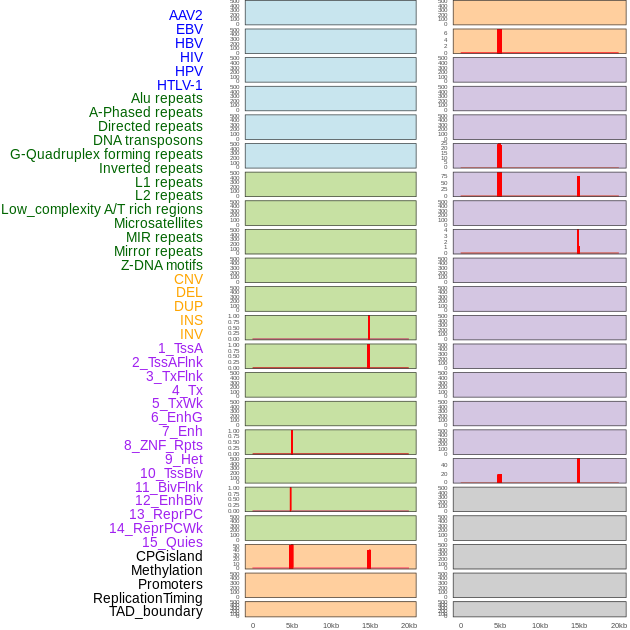
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for MEMO1-MCU |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:32142995/chr10:74594117) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MEMO1 | MCU |
| FUNCTION: May control cell migration by relaying extracellular chemotactic signals to the microtubule cytoskeleton. Mediator of ERBB2 signaling. The MEMO1-RHOA-DIAPH1 signaling pathway plays an important role in ERBB2-dependent stabilization of microtubules at the cell cortex. It controls the localization of APC and CLASP2 to the cell membrane, via the regulation of GSK3B activity. In turn, membrane-bound APC allows the localization of the MACF1 to the cell membrane, which is required for microtubule capture and stabilization. Is required for breast carcinoma cell migration. {ECO:0000269|PubMed:15156151, ECO:0000269|PubMed:20937854}. | FUNCTION: Key regulator of mitochondrial calcium uniporter (MCU) required for calcium entry into mitochondrion (PubMed:23178883, PubMed:26445506, PubMed:27184846, PubMed:26976564). Plays a direct role in uniporter-mediated calcium uptake via a direct interaction with MCU (PubMed:23178883). Probably involved in the assembly of the membrane components of the uniporter complex (uniplex) (PubMed:27184846). {ECO:0000269|PubMed:23178883, ECO:0000269|PubMed:26445506, ECO:0000269|PubMed:26976564, ECO:0000269|PubMed:27184846}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 192_223 | 50 | 331.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 311_339 | 50 | 331.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 192_223 | 50 | 352.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 311_339 | 50 | 352.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 192_223 | 1 | 303.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 311_339 | 1 | 303.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 216_234 | 50 | 331.0 | Region | Outer juxtamembrane helix (OJMH) | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 283_292 | 50 | 331.0 | Region | Inner juxtamembrane helix (IJMH) | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 75_165 | 50 | 331.0 | Region | N-terminal MCU domain | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 216_234 | 50 | 352.0 | Region | Outer juxtamembrane helix (OJMH) | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 283_292 | 50 | 352.0 | Region | Inner juxtamembrane helix (IJMH) | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 75_165 | 50 | 352.0 | Region | N-terminal MCU domain | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 216_234 | 1 | 303.0 | Region | Outer juxtamembrane helix (OJMH) | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 283_292 | 1 | 303.0 | Region | Inner juxtamembrane helix (IJMH) | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 75_165 | 1 | 303.0 | Region | N-terminal MCU domain | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 257_265 | 50 | 331.0 | Topological domain | Mitochondrial intermembrane | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 284_351 | 50 | 331.0 | Topological domain | Mitochondrial matrix | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 51_233 | 50 | 331.0 | Topological domain | Mitochondrial matrix | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 257_265 | 50 | 352.0 | Topological domain | Mitochondrial intermembrane | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 284_351 | 50 | 352.0 | Topological domain | Mitochondrial matrix | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 51_233 | 50 | 352.0 | Topological domain | Mitochondrial matrix | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 257_265 | 1 | 303.0 | Topological domain | Mitochondrial intermembrane | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 284_351 | 1 | 303.0 | Topological domain | Mitochondrial matrix | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 51_233 | 1 | 303.0 | Topological domain | Mitochondrial matrix | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 234_256 | 50 | 331.0 | Transmembrane | Helical | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000357157 | 0 | 8 | 266_283 | 50 | 331.0 | Transmembrane | Helical | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 234_256 | 50 | 352.0 | Transmembrane | Helical | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000373053 | 0 | 8 | 266_283 | 50 | 352.0 | Transmembrane | Helical | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 234_256 | 1 | 303.0 | Transmembrane | Helical | |
| Tgene | MCU | chr2:32142995 | chr10:74594117 | ENST00000536019 | 0 | 8 | 266_283 | 1 | 303.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for MEMO1-MCU |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >52979_52979_1_MEMO1-MCU_MEMO1_chr2_32142995_ENST00000404530_MCU_chr10_74594117_ENST00000357157_length(transcript)=3202nt_BP=487nt ATGCTCAGTGGGCAGGACCCGTTAGAGTGGCGAGCGGCACAGGCACCAAGATGTCCAACCGAGTGGTCTGCCGAGAAGCCAGTCACGCCG GGAGCTGGTACACAGCCTCAGGACCGCAGCTGAATGCACAGCTAGAAGGTTGGCTTTCACAAGTACAGTCTACAAAAAGACCTGCTAGAG CCATTATTGCCCCCCATGCAGGATATACGTACTGTGGGTCTTGTGCTGCCCATGCTTATAAACAAGTGGATCCGTCTATTACCCGGAGAA TTTTCATCCTTGGGCCTTCTCATCATGTGCCCCTCTCTCGATGTGCACTTTCCAGTGTGGATATATATAGGACACCTCTGTATGACCTTC GTATTGACCAAAAGATTTACGGAGAACTGTGGAAGACAGGAATGTTTGAACGCATGTCTCTGCAGACAGATGAAGATGAACACAGTATTG AAATGCATTTGCCTTATACAGCTAAAGCCATGGAAAGGTACACCAGAGGATCGCTTCCTGGCAGAATTTGGGAGCTGTTTATTGCAGCAC TGTTGTGCCCTCTGATGATGTTACAGTGGTTTATCAAAATGGGTTACCTGTGATATCTGTGAGGCTACCATCCCGGCGTGAACGCTGTCA GTTCACACTCAAGCCTATCTCTGACTCTGTTGGTGTATTTTTACGACAACTGCAAGAAGAGGATCGGGGAATTGACAGAGTTGCTATCTA TTCACCAGATGGTGTTCGCGTTGCTGCTTCAACAGGAATAGACCTCCTCCTCCTTGATGACTTTAAGCTGGTCATTAATGACTTAACATA CCACGTACGACCACCAAAAAGAGTAGAGATGGGGTTTTGCCATGTTGGCCAGAATGGTTTTGAACTCCTGACCTCAAGTTATCTACCTGC CTCAGCCTCCCAAAGTGCCGAGATTATAGCCGTACGAATTGAGATTAGCAGAAAAGCTGAGAAGAGGACCACTTTGGTGCTATGGGGTGG CCTTGCCTACATGGCCACACAGTTTGGCATTTTGGCCCGGCTTACCTGGTGGGAATATTCCTGGGACATCATGGAGCCAGTAACATACTT CATCACTTATGGAAGTGCCATGGCAATGTATGCATATTTTGTAATGACACGCCAGGAATATGTTTATCCAGAAGCCAGAGACAGACAATA CTTACTATTTTTCCATAAAGGAGCCAAAAAGTCACGTTTTGACCTAGAGAAATACAATCAACTCAAGGATGCAATTGCTCAGGCAGAAAT GGACCTTAAGAGACTGAGAGACCCATTACAAGTACATCTGCCTCTCCGACAAATTGGTGAAAAAGATTGATCTGCAAAAAGCCTCTGAAT CCTGGCAGAAGGAACACCTGTTTGCCTTTTTAATTAAAGCATTGCAGGTGGAAGCTGGGAGCCATGTGGGGGGTAGAGCGTTTTTACCTT TAATTATAAAACAAAAACAGAAAGGATCTGAGGGAAGAAGGGAATGTTAAAACCTGAGGATCAGGCATTGTGGAATATAAGCTCAAAGGG CTTAGTGAATATTGTCTTAACCAAGTATCTCAGTTTCTGGATGAAAATGATGCAGTTATATAGTTGAGAGATTCATAAAGAGAAAACAAT GCTGGGGGTGTTCGTTTCTTGCATCTTCTTTGCAGAGTCAGCAAAAGAGTAACACACCAGCACCCCACTCGACTCTATTTGTTTTTAATT TAACTGTCCCTATTTTTGACATAGGAGTAAATAAATATACTAGAAAAGCAAATTCTCATGATATGCTAAAATATCATTAGCATTTATTTT AAATTGGACCCAGTCTCTGCAGAGTTACCAGGAATCTTTCCTTCCAGCATCCCTTTACTGACCACCTACCTGTACCTCTTGGTTACACTC ATTTTTTCCATTTGATAATTGGAACCAACTTATAACTGTTTAATAATTGACACTTTAGATTATCTCTTAATACCTTCTTAAATGTCTATA TATCCCAGTGCTCTGGATCAGTGTCTAAAAATCACTGGCAACACTGCATGAGGTTGTTGGTTTTGTTTTGTTTTATTAATTAGTCTTTCA CAGGAGGAATAATTGCCCTCCTTTATATACTTATCTATTGATAATCCCCTCTCCCTCCAGAACACAAATCAGAGGGAAAGGGGGTGTTCA GCTGTACTACCAAATCAGGAAGATGTAAGGTTTACAAATTGGCTAAGAATCATGGCTCTGTAGCCATTTCAACCAGAATAATTTTATTGC TAATCTGCTTTGTGTGACAGCATTCCAGGCCAGCCAGATGGGACTGCCTTGTCTGGAGGCTTTGTTCATCTCGAAGGACACACACTTCCA CACTGTTTGTGAGCCCTCCCACCTCCACAACTTCAGTTGTAAATCAAGTGTGTGGATCTCAAAGGGTGCAATTTATCTTTATATAGGAAT ACATTTCTAGGGCTTCCTTCAAGCCCACTCTCTTCACCCTATTTTTTCTTATCTTAAATTGAGAGAAAGAGAATTAATCTTATACTTTGT CAAAACATTTTCTACCATATTTCCAGATGACATCTGCGCTTGAAGAGTCAAAGGAATCTGTGTCTAATATCCTGTTTTTAACTGCTGTAG GGGCAGGATGGAAAGGATGATGGGGGCTGCCACACCACTGATTGGCCTTTTCTTTCACGTGATTCATCCTTCCTCATTGTGGCAAGGAGT TTCTTTCTCTTTTTCTTCCTCCTTTGGGATCATTGTGTATGAAAAGAAAAACTTTAAATGACAAACCCAGACTCCAGGTGCCTTGCAAAG GTTGAAGGCCAGCCAGGATTGCTGCTGCTGCTGCTACTCCTGCCAACACCCCTTTCATTGGCATGACGGAATGAAAGGATGCATGTCTCC ACTTCCTGACCCTCCGCCCACTTCCTTCTCCCTCCACCACCCCCAGTCGTCAGCTCCTTCCCTCATTTATTTTTGTTAAGTTGTGTGAAT TATTTTTAACCCATTTATCCTGTTTGTGCATAGGGTTTTTAAGAAGAAACAGCACAGTGCAACGAGCAAATCTTTTTGGGGTGTGTGGGA AGCAAGGGAGGGAGGACATGGAGAAAAGTTCTTTAAACAAATAGCAAACTATTGAACATGTGTAAAATCCTGTATCATTTATGAAATATG >52979_52979_1_MEMO1-MCU_MEMO1_chr2_32142995_ENST00000404530_MCU_chr10_74594117_ENST00000357157_length(amino acids)=270AA_BP= MGAVYCSTVVPSDDVTVVYQNGLPVISVRLPSRRERCQFTLKPISDSVGVFLRQLQEEDRGIDRVAIYSPDGVRVAASTGIDLLLLDDFK LVINDLTYHVRPPKRVEMGFCHVGQNGFELLTSSYLPASASQSAEIIAVRIEISRKAEKRTTLVLWGGLAYMATQFGILARLTWWEYSWD IMEPVTYFITYGSAMAMYAYFVMTRQEYVYPEARDRQYLLFFHKGAKKSRFDLEKYNQLKDAIAQAEMDLKRLRDPLQVHLPLRQIGEKD -------------------------------------------------------------- >52979_52979_2_MEMO1-MCU_MEMO1_chr2_32142995_ENST00000404530_MCU_chr10_74594117_ENST00000373053_length(transcript)=3265nt_BP=487nt ATGCTCAGTGGGCAGGACCCGTTAGAGTGGCGAGCGGCACAGGCACCAAGATGTCCAACCGAGTGGTCTGCCGAGAAGCCAGTCACGCCG GGAGCTGGTACACAGCCTCAGGACCGCAGCTGAATGCACAGCTAGAAGGTTGGCTTTCACAAGTACAGTCTACAAAAAGACCTGCTAGAG CCATTATTGCCCCCCATGCAGGATATACGTACTGTGGGTCTTGTGCTGCCCATGCTTATAAACAAGTGGATCCGTCTATTACCCGGAGAA TTTTCATCCTTGGGCCTTCTCATCATGTGCCCCTCTCTCGATGTGCACTTTCCAGTGTGGATATATATAGGACACCTCTGTATGACCTTC GTATTGACCAAAAGATTTACGGAGAACTGTGGAAGACAGGAATGTTTGAACGCATGTCTCTGCAGACAGATGAAGATGAACACAGTATTG AAATGCATTTGCCTTATACAGCTAAAGCCATGGAAAGGTACACCAGAGGATCGCTTCCTGGCAGAATTTGGGAGCTGTTTATTGCAGCAC TGTTGTGCCCTCTGATGATGTTACAGTGGTTTATCAAAATGGGTTACCTGTGATATCTGTGAGGCTACCATCCCGGCGTGAACGCTGTCA GTTCACACTCAAGCCTATCTCTGACTCTGTTGGTGTATTTTTACGACAACTGCAAGAAGAGGATCGGGGAATTGACAGAGTTGCTATCTA TTCACCAGATGGTGTTCGCGTTGCTGCTTCAACAGGAATAGACCTCCTCCTCCTTGATGACTTTAAGCTGGTCATTAATGACTTAACATA CCACGTACGACCACCAAAAAGAGACCTCTTAAGTCATGAAAATGCAGCAACGCTGAATGATGTAAAGACATTGGTCCAGCAACTATACAC CACACTGTGCATTGAGCAGCACCAGTTAAACAAGGAAAGGGAGCTTATTGAAAGACTAGAGGATCTCAAAGAGCAGCTGGCTCCCCTGGA AAAGGTACGAATTGAGATTAGCAGAAAAGCTGAGAAGAGGACCACTTTGGTGCTATGGGGTGGCCTTGCCTACATGGCCACACAGTTTGG CATTTTGGCCCGGCTTACCTGGTGGGAATATTCCTGGGACATCATGGAGCCAGTAACATACTTCATCACTTATGGAAGTGCCATGGCAAT GTATGCATATTTTGTAATGACACGCCAGGAATATGTTTATCCAGAAGCCAGAGACAGACAATACTTACTATTTTTCCATAAAGGAGCCAA AAAGTCACGTTTTGACCTAGAGAAATACAATCAACTCAAGGATGCAATTGCTCAGGCAGAAATGGACCTTAAGAGACTGAGAGACCCATT ACAAGTACATCTGCCTCTCCGACAAATTGGTGAAAAAGATTGATCTGCAAAAAGCCTCTGAATCCTGGCAGAAGGAACACCTGTTTGCCT TTTTAATTAAAGCATTGCAGGTGGAAGCTGGGAGCCATGTGGGGGGTAGAGCGTTTTTACCTTTAATTATAAAACAAAAACAGAAAGGAT CTGAGGGAAGAAGGGAATGTTAAAACCTGAGGATCAGGCATTGTGGAATATAAGCTCAAAGGGCTTAGTGAATATTGTCTTAACCAAGTA TCTCAGTTTCTGGATGAAAATGATGCAGTTATATAGTTGAGAGATTCATAAAGAGAAAACAATGCTGGGGGTGTTCGTTTCTTGCATCTT CTTTGCAGAGTCAGCAAAAGAGTAACACACCAGCACCCCACTCGACTCTATTTGTTTTTAATTTAACTGTCCCTATTTTTGACATAGGAG TAAATAAATATACTAGAAAAGCAAATTCTCATGATATGCTAAAATATCATTAGCATTTATTTTAAATTGGACCCAGTCTCTGCAGAGTTA CCAGGAATCTTTCCTTCCAGCATCCCTTTACTGACCACCTACCTGTACCTCTTGGTTACACTCATTTTTTCCATTTGATAATTGGAACCA ACTTATAACTGTTTAATAATTGACACTTTAGATTATCTCTTAATACCTTCTTAAATGTCTATATATCCCAGTGCTCTGGATCAGTGTCTA AAAATCACTGGCAACACTGCATGAGGTTGTTGGTTTTGTTTTGTTTTATTAATTAGTCTTTCACAGGAGGAATAATTGCCCTCCTTTATA TACTTATCTATTGATAATCCCCTCTCCCTCCAGAACACAAATCAGAGGGAAAGGGGGTGTTCAGCTGTACTACCAAATCAGGAAGATGTA AGGTTTACAAATTGGCTAAGAATCATGGCTCTGTAGCCATTTCAACCAGAATAATTTTATTGCTAATCTGCTTTGTGTGACAGCATTCCA GGCCAGCCAGATGGGACTGCCTTGTCTGGAGGCTTTGTTCATCTCGAAGGACACACACTTCCACACTGTTTGTGAGCCCTCCCACCTCCA CAACTTCAGTTGTAAATCAAGTGTGTGGATCTCAAAGGGTGCAATTTATCTTTATATAGGAATACATTTCTAGGGCTTCCTTCAAGCCCA CTCTCTTCACCCTATTTTTTCTTATCTTAAATTGAGAGAAAGAGAATTAATCTTATACTTTGTCAAAACATTTTCTACCATATTTCCAGA TGACATCTGCGCTTGAAGAGTCAAAGGAATCTGTGTCTAATATCCTGTTTTTAACTGCTGTAGGGGCAGGATGGAAAGGATGATGGGGGC TGCCACACCACTGATTGGCCTTTTCTTTCACGTGATTCATCCTTCCTCATTGTGGCAAGGAGTTTCTTTCTCTTTTTCTTCCTCCTTTGG GATCATTGTGTATGAAAAGAAAAACTTTAAATGACAAACCCAGACTCCAGGTGCCTTGCAAAGGTTGAAGGCCAGCCAGGATTGCTGCTG CTGCTGCTACTCCTGCCAACACCCCTTTCATTGGCATGACGGAATGAAAGGATGCATGTCTCCACTTCCTGACCCTCCGCCCACTTCCTT CTCCCTCCACCACCCCCAGTCGTCAGCTCCTTCCCTCATTTATTTTTGTTAAGTTGTGTGAATTATTTTTAACCCATTTATCCTGTTTGT GCATAGGGTTTTTAAGAAGAAACAGCACAGTGCAACGAGCAAATCTTTTTGGGGTGTGTGGGAAGCAAGGGAGGGAGGACATGGAGAAAA GTTCTTTAAACAAATAGCAAACTATTGAACATGTGTAAAATCCTGTATCATTTATGAAATATGTATAAAAAGCAATGTACCTTCTGGAAC >52979_52979_2_MEMO1-MCU_MEMO1_chr2_32142995_ENST00000404530_MCU_chr10_74594117_ENST00000373053_length(amino acids)=291AA_BP= MGAVYCSTVVPSDDVTVVYQNGLPVISVRLPSRRERCQFTLKPISDSVGVFLRQLQEEDRGIDRVAIYSPDGVRVAASTGIDLLLLDDFK LVINDLTYHVRPPKRDLLSHENAATLNDVKTLVQQLYTTLCIEQHQLNKERELIERLEDLKEQLAPLEKVRIEISRKAEKRTTLVLWGGL AYMATQFGILARLTWWEYSWDIMEPVTYFITYGSAMAMYAYFVMTRQEYVYPEARDRQYLLFFHKGAKKSRFDLEKYNQLKDAIAQAEMD -------------------------------------------------------------- >52979_52979_3_MEMO1-MCU_MEMO1_chr2_32142995_ENST00000404530_MCU_chr10_74594117_ENST00000536019_length(transcript)=3263nt_BP=487nt ATGCTCAGTGGGCAGGACCCGTTAGAGTGGCGAGCGGCACAGGCACCAAGATGTCCAACCGAGTGGTCTGCCGAGAAGCCAGTCACGCCG GGAGCTGGTACACAGCCTCAGGACCGCAGCTGAATGCACAGCTAGAAGGTTGGCTTTCACAAGTACAGTCTACAAAAAGACCTGCTAGAG CCATTATTGCCCCCCATGCAGGATATACGTACTGTGGGTCTTGTGCTGCCCATGCTTATAAACAAGTGGATCCGTCTATTACCCGGAGAA TTTTCATCCTTGGGCCTTCTCATCATGTGCCCCTCTCTCGATGTGCACTTTCCAGTGTGGATATATATAGGACACCTCTGTATGACCTTC GTATTGACCAAAAGATTTACGGAGAACTGTGGAAGACAGGAATGTTTGAACGCATGTCTCTGCAGACAGATGAAGATGAACACAGTATTG AAATGCATTTGCCTTATACAGCTAAAGCCATGGAAAGGTACACCAGAGGATCGCTTCCTGGCAGAATTTGGGAGCTGTTTATTGCAGCAC TGTTGTGCCCTCTGATGATGTTACAGTGGTTTATCAAAATGGGTTACCTGTGATATCTGTGAGGCTACCATCCCGGCGTGAACGCTGTCA GTTCACACTCAAGCCTATCTCTGACTCTGTTGGTGTATTTTTACGACAACTGCAAGAAGAGGATCGGGGAATTGACAGAGTTGCTATCTA TTCACCAGATGGTGTTCGCGTTGCTGCTTCAACAGGAATAGACCTCCTCCTCCTTGATGACTTTAAGCTGGTCATTAATGACTTAACATA CCACGTACGACCACCAAAAAGAGACCTCTTAAGTCATGAAAATGCAGCAACGCTGAATGATGTAAAGACATTGGTCCAGCAACTATACAC CACACTGTGCATTGAGCAGCACCAGTTAAACAAGGAAAGGGAGCTTATTGAAAGACTAGAGGATCTCAAAGAGCAGCTGGCTCCCCTGGA AAAGGTACGAATTGAGATTAGCAGAAAAGCTGAGAAGAGGACCACTTTGGTGCTATGGGGTGGCCTTGCCTACATGGCCACACAGTTTGG CATTTTGGCCCGGCTTACCTGGTGGGAATATTCCTGGGACATCATGGAGCCAGTAACATACTTCATCACTTATGGAAGTGCCATGGCAAT GTATGCATATTTTGTAATGACACGCCAGGAATATGTTTATCCAGAAGCCAGAGACAGACAATACTTACTATTTTTCCATAAAGGAGCCAA AAAGTCACGTTTTGACCTAGAGAAATACAATCAACTCAAGGATGCAATTGCTCAGGCAGAAATGGACCTTAAGAGACTGAGAGACCCATT ACAAGTACATCTGCCTCTCCGACAAATTGGTGAAAAAGATTGATCTGCAAAAAGCCTCTGAATCCTGGCAGAAGGAACACCTGTTTGCCT TTTTAATTAAAGCATTGCAGGTGGAAGCTGGGAGCCATGTGGGGGGTAGAGCGTTTTTACCTTTAATTATAAAACAAAAACAGAAAGGAT CTGAGGGAAGAAGGGAATGTTAAAACCTGAGGATCAGGCATTGTGGAATATAAGCTCAAAGGGCTTAGTGAATATTGTCTTAACCAAGTA TCTCAGTTTCTGGATGAAAATGATGCAGTTATATAGTTGAGAGATTCATAAAGAGAAAACAATGCTGGGGGTGTTCGTTTCTTGCATCTT CTTTGCAGAGTCAGCAAAAGAGTAACACACCAGCACCCCACTCGACTCTATTTGTTTTTAATTTAACTGTCCCTATTTTTGACATAGGAG TAAATAAATATACTAGAAAAGCAAATTCTCATGATATGCTAAAATATCATTAGCATTTATTTTAAATTGGACCCAGTCTCTGCAGAGTTA CCAGGAATCTTTCCTTCCAGCATCCCTTTACTGACCACCTACCTGTACCTCTTGGTTACACTCATTTTTTCCATTTGATAATTGGAACCA ACTTATAACTGTTTAATAATTGACACTTTAGATTATCTCTTAATACCTTCTTAAATGTCTATATATCCCAGTGCTCTGGATCAGTGTCTA AAAATCACTGGCAACACTGCATGAGGTTGTTGGTTTTGTTTTGTTTTATTAATTAGTCTTTCACAGGAGGAATAATTGCCCTCCTTTATA TACTTATCTATTGATAATCCCCTCTCCCTCCAGAACACAAATCAGAGGGAAAGGGGGTGTTCAGCTGTACTACCAAATCAGGAAGATGTA AGGTTTACAAATTGGCTAAGAATCATGGCTCTGTAGCCATTTCAACCAGAATAATTTTATTGCTAATCTGCTTTGTGTGACAGCATTCCA GGCCAGCCAGATGGGACTGCCTTGTCTGGAGGCTTTGTTCATCTCGAAGGACACACACTTCCACACTGTTTGTGAGCCCTCCCACCTCCA CAACTTCAGTTGTAAATCAAGTGTGTGGATCTCAAAGGGTGCAATTTATCTTTATATAGGAATACATTTCTAGGGCTTCCTTCAAGCCCA CTCTCTTCACCCTATTTTTTCTTATCTTAAATTGAGAGAAAGAGAATTAATCTTATACTTTGTCAAAACATTTTCTACCATATTTCCAGA TGACATCTGCGCTTGAAGAGTCAAAGGAATCTGTGTCTAATATCCTGTTTTTAACTGCTGTAGGGGCAGGATGGAAAGGATGATGGGGGC TGCCACACCACTGATTGGCCTTTTCTTTCACGTGATTCATCCTTCCTCATTGTGGCAAGGAGTTTCTTTCTCTTTTTCTTCCTCCTTTGG GATCATTGTGTATGAAAAGAAAAACTTTAAATGACAAACCCAGACTCCAGGTGCCTTGCAAAGGTTGAAGGCCAGCCAGGATTGCTGCTG CTGCTGCTACTCCTGCCAACACCCCTTTCATTGGCATGACGGAATGAAAGGATGCATGTCTCCACTTCCTGACCCTCCGCCCACTTCCTT CTCCCTCCACCACCCCCAGTCGTCAGCTCCTTCCCTCATTTATTTTTGTTAAGTTGTGTGAATTATTTTTAACCCATTTATCCTGTTTGT GCATAGGGTTTTTAAGAAGAAACAGCACAGTGCAACGAGCAAATCTTTTTGGGGTGTGTGGGAAGCAAGGGAGGGAGGACATGGAGAAAA GTTCTTTAAACAAATAGCAAACTATTGAACATGTGTAAAATCCTGTATCATTTATGAAATATGTATAAAAAGCAATGTACCTTCTGGAAC >52979_52979_3_MEMO1-MCU_MEMO1_chr2_32142995_ENST00000404530_MCU_chr10_74594117_ENST00000536019_length(amino acids)=291AA_BP= MGAVYCSTVVPSDDVTVVYQNGLPVISVRLPSRRERCQFTLKPISDSVGVFLRQLQEEDRGIDRVAIYSPDGVRVAASTGIDLLLLDDFK LVINDLTYHVRPPKRDLLSHENAATLNDVKTLVQQLYTTLCIEQHQLNKERELIERLEDLKEQLAPLEKVRIEISRKAEKRTTLVLWGGL AYMATQFGILARLTWWEYSWDIMEPVTYFITYGSAMAMYAYFVMTRQEYVYPEARDRQYLLFFHKGAKKSRFDLEKYNQLKDAIAQAEMD -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MEMO1-MCU |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MEMO1-MCU |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MEMO1-MCU |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
