
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:METAP1-IQGAP1 (FusionGDB2 ID:53009) |
Fusion Gene Summary for METAP1-IQGAP1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: METAP1-IQGAP1 | Fusion gene ID: 53009 | Hgene | Tgene | Gene symbol | METAP1 | IQGAP1 | Gene ID | 23173 | 8826 |
| Gene name | methionyl aminopeptidase 1 | IQ motif containing GTPase activating protein 1 | |
| Synonyms | MAP1A|MetAP1A | HUMORFA01|SAR1|p195 | |
| Cytomap | 4q23 | 15q26.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | methionine aminopeptidase 1MAP 1metAP 1peptidase M 1 | ras GTPase-activating-like protein IQGAP1RasGAP-like with IQ motifs | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | P53582 | P46940 | |
| Ensembl transtripts involved in fusion gene | ENST00000296411, ENST00000544031, ENST00000506548, | ENST00000560020, ENST00000268182, ENST00000560738, | |
| Fusion gene scores | * DoF score | 3 X 3 X 3=27 | 17 X 15 X 7=1785 |
| # samples | 3 | 19 | |
| ** MAII score | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(19/1785*10)=-3.23185275058551 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: METAP1 [Title/Abstract] AND IQGAP1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | METAP1(99966461)-IQGAP1(91030186), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | IQGAP1 | GO:0071277 | cellular response to calcium ion | 18567582 |
 Fusion gene breakpoints across METAP1 (5'-gene) Fusion gene breakpoints across METAP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
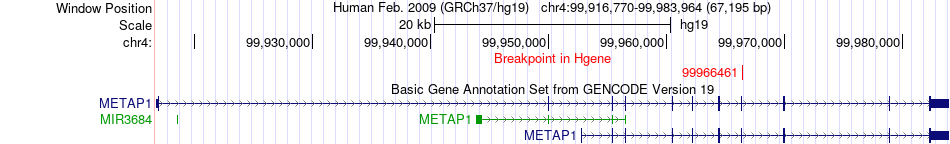 |
 Fusion gene breakpoints across IQGAP1 (3'-gene) Fusion gene breakpoints across IQGAP1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
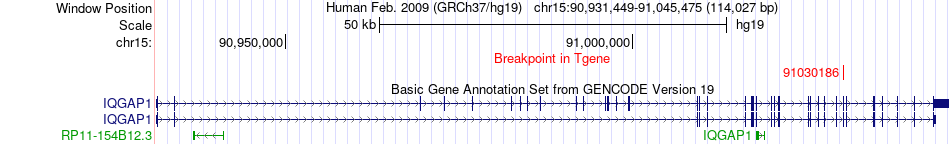 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-EQ-A4SO-01A | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
Top |
Fusion Gene ORF analysis for METAP1-IQGAP1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000296411 | ENST00000560020 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| 5CDS-intron | ENST00000544031 | ENST00000560020 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| In-frame | ENST00000296411 | ENST00000268182 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| In-frame | ENST00000296411 | ENST00000560738 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| In-frame | ENST00000544031 | ENST00000268182 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| In-frame | ENST00000544031 | ENST00000560738 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| intron-3CDS | ENST00000506548 | ENST00000268182 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| intron-3CDS | ENST00000506548 | ENST00000560738 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
| intron-intron | ENST00000506548 | ENST00000560020 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000296411 | METAP1 | chr4 | 99966461 | + | ENST00000268182 | IQGAP1 | chr15 | 91030186 | + | 4006 | 921 | 134 | 1870 | 578 |
| ENST00000296411 | METAP1 | chr4 | 99966461 | + | ENST00000560738 | IQGAP1 | chr15 | 91030186 | + | 2134 | 921 | 134 | 1870 | 578 |
| ENST00000544031 | METAP1 | chr4 | 99966461 | + | ENST00000268182 | IQGAP1 | chr15 | 91030186 | + | 3722 | 637 | 0 | 1586 | 528 |
| ENST00000544031 | METAP1 | chr4 | 99966461 | + | ENST00000560738 | IQGAP1 | chr15 | 91030186 | + | 1850 | 637 | 0 | 1586 | 528 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000296411 | ENST00000268182 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + | 0.000265983 | 0.999734 |
| ENST00000296411 | ENST00000560738 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + | 0.001622478 | 0.99837756 |
| ENST00000544031 | ENST00000268182 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + | 0.000365332 | 0.9996346 |
| ENST00000544031 | ENST00000560738 | METAP1 | chr4 | 99966461 | + | IQGAP1 | chr15 | 91030186 | + | 0.001886282 | 0.99811375 |
Top |
Fusion Genomic Features for METAP1-IQGAP1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
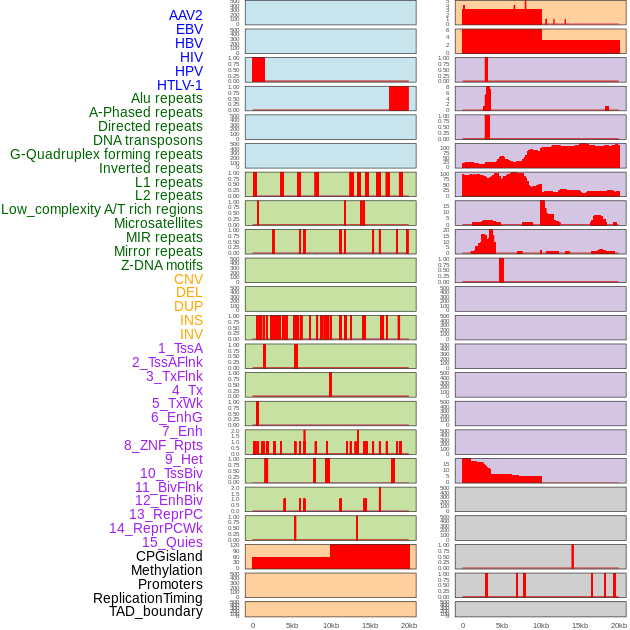 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
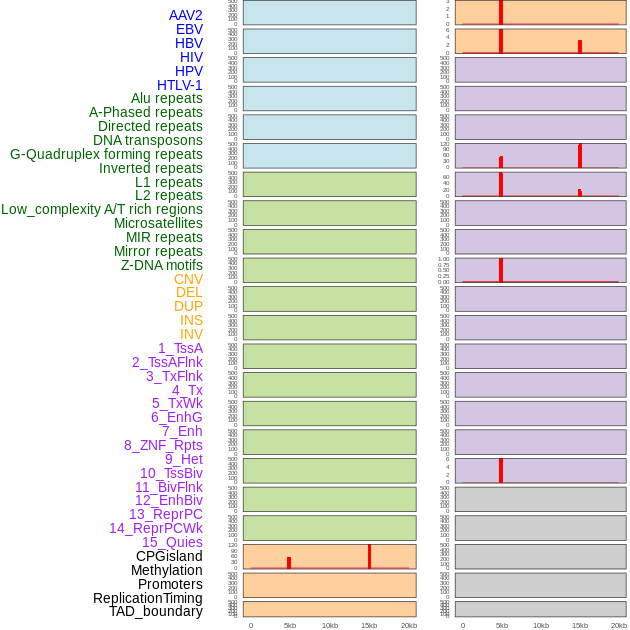 |
Top |
Fusion Protein Features for METAP1-IQGAP1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr4:99966461/chr15:91030186) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| METAP1 | IQGAP1 |
| FUNCTION: Cotranslationally removes the N-terminal methionine from nascent proteins. The N-terminal methionine is often cleaved when the second residue in the primary sequence is small and uncharged (Met-Ala-, Cys, Gly, Pro, Ser, Thr, or Val). Required for normal progression through the cell cycle. {ECO:0000255|HAMAP-Rule:MF_03174, ECO:0000269|PubMed:16274222, ECO:0000269|PubMed:17114291}. | FUNCTION: Plays a crucial role in regulating the dynamics and assembly of the actin cytoskeleton. Binds to activated CDC42 but does not stimulate its GTPase activity. It associates with calmodulin. Could serve as an assembly scaffold for the organization of a multimolecular complex that would interface incoming signals to the reorganization of the actin cytoskeleton at the plasma membrane. May promote neurite outgrowth (PubMed:15695813). May play a possible role in cell cycle regulation by contributing to cell cycle progression after DNA replication arrest (PubMed:20883816). {ECO:0000269|PubMed:15695813, ECO:0000269|PubMed:20883816}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | METAP1 | chr4:99966461 | chr15:91030186 | ENST00000296411 | + | 8 | 11 | 9_52 | 262 | 387.0 | Region | Zinc finger-like%3B important for proper ribosome association |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 1004_1237 | 1341 | 1658.0 | Domain | Ras-GAP | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 44_159 | 1341 | 1658.0 | Domain | Calponin-homology (CH) | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 679_712 | 1341 | 1658.0 | Domain | WW | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 745_774 | 1341 | 1658.0 | Domain | IQ 1 | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 775_804 | 1341 | 1658.0 | Domain | IQ 2 | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 805_834 | 1341 | 1658.0 | Domain | IQ 3 | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 835_864 | 1341 | 1658.0 | Domain | IQ 4 | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 1276_1657 | 1341 | 1658.0 | Region | Note=C2 | |
| Tgene | IQGAP1 | chr4:99966461 | chr15:91030186 | ENST00000268182 | 30 | 38 | 956_1274 | 1341 | 1658.0 | Region | Note=C1 |
Top |
Fusion Gene Sequence for METAP1-IQGAP1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >53009_53009_1_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000296411_IQGAP1_chr15_91030186_ENST00000268182_length(transcript)=4006nt_BP=921nt CTCCCGCGCCCAGGCCAGCGGCGGGCCCAGCTCCTCCCCCGACTCGGTCTCTCTCCCCTCCCCTCCGCCCGGCAGTTCCTCCCTCCCGCC GCCGCCTCTTCCTCGGTGAGGCGCTCTTCCAGCGGGCAGGCAGCATGGCGGCCGTGGAGACGCGGGTGTGCGAGACAGACGGCTGCAGCA GTGAGGCCAAGCTCCAGTGTCCCACTTGCATCAAGCTGGGCATCCAGGGCTCGTACTTCTGCTCGCAGGAATGTTTTAAAGGAAGTTGGG CTACTCACAAGTTACTACATAAGAAAGCAAAAGATGAAAAGGCGAAGCGAGAAGTGTCTTCCTGGACTGTGGAAGGTGATATTAATACTG ACCCATGGGCAGGTTATCGATATACTGGTAAACTCAGACCACATTATCCACTGATGCCAACAAGGCCAGTGCCAAGTTATATTCAAAGAC CAGATTATGCTGATCATCCCTTAGGAATGTCTGAATCTGAACAGGCTCTTAAAGGTACTTCTCAGATTAAATTACTCTCATCTGAAGATA TAGAAGGGATGCGACTTGTATGTAGGCTTGCTAGAGAAGTTTTGGATGTTGCTGCCGGCATGATTAAACCAGGTGTAACTACTGAAGAAA TAGATCACGCTGTACACTTAGCATGTATTGCAAGAAATTGCTACCCTTCTCCCCTGAATTATTATAATTTCCCAAAGTCTTGTTGTACCT CAGTGAATGAAGTCATTTGCCATGGAATACCAGACAGAAGGCCCTTACAAGAAGGTGACATTGTTAATGTGGATATCACTCTTTATCGCA ATGGTTATCATGGGGACCTGAATGAGACATTTTTTGTTGGAGAAGTGGATGATGGAGCACGGAAACTTGTTCAGACCACATATGAGTGCC TGATGCAAGCCATTGATGCAGGGGAAAGCTCTGGCAATTTAAATGACCCAAATAAGGAGGCACTGGCTAAGACGGAAGTGTCTCTCACCC TGACCAACAAGTTCGACGTGCCTGGAGATGAGAATGCAGAAATGGATGCTCGAACCATCTTACTGAATACAAAACGTTTAATTGTGGATG TCATCCGGTTCCAGCCAGGAGAGACCTTGACTGAAATCCTAGAAACACCAGCCACCAGTGAACAGGAAGCAGAACATCAGAGAGCCATGC AGAGACGTGCTATCCGTGATGCCAAAACACCTGACAAGATGAAAAAGTCAAAATCTGTAAAGGAAGACAGCAACCTCACTCTTCAAGAGA AGAAAGAGAAGATCCAGACAGGTTTAAAGAAGCTAACAGAGCTTGGAACCGTGGACCCAAAGAACAAATACCAGGAACTGATCAACGACA TTGCCAGGGATATTCGGAATCAGCGGAGGTACCGACAGAGGAGAAAGGCCGAACTAGTGAAACTGCAACAGACATACGCTGCTCTGAACT CTAAGGCCACCTTTTATGGGGAGCAGGTGGATTACTATAAAAGCTATATCAAAACCTGCTTGGATAACTTAGCCAGCAAGGGCAAAGTCT CCAAAAAGCCTAGGGAAATGAAAGGAAAGAAAAGCAAAAAGATTTCTCTGAAATATACAGCAGCAAGACTACATGAAAAAGGAGTTCTTC TGGAAATTGAGGACCTGCAAGTGAATCAGTTTAAAAATGTTATATTTGAAATCAGTCCAACAGAAGAAGTTGGAGACTTCGAAGTGAAAG CCAAATTCATGGGAGTTCAAATGGAGACTTTTATGTTACATTATCAGGACCTGCTGCAGCTACAGTATGAAGGAGTTGCAGTCATGAAAT TATTTGATAGAGCTAAAGTAAATGTCAACCTCCTGATCTTCCTTCTCAACAAAAAGTTCTACGGGAAGTAATTGATCGTTTGCTGCCAGC CCAGAAGGATGAAGGAAAGAAGCACCTCACAGCTCCTTTCTAGGTCCTTCTTTCCTCATTGGAAGCAAAGACCTAGCCAACAACAGCACC TCAATCTGATACACTCCCGATGCCACATTTTTAACTCCTCTCGCTCTGATGGGACATTTGTTACCCTTTTTTCATAGTGAAATTGTGTTT CAGGCTTAGTCTGACCTTTCTGGTTTCTTCATTTTCTTCCATTACTTAGGAAAGAGTGGAAACTCCACTAAAATTTCTCTGTGTTGTTAC AGTCTTAGAGGTTGCAGTACTATATTGTAAGCTTTGGTGTTTGTTTAATTAGCAATAGGGATGGTAGGATTCAAATGTGTGTCATTTAGA AGTGGAAGCTATTAGCACCAATGACATAAATACATACAAGACACACAACTAAAATGTCATGTTATTAACAGTTATTAGGTTGTCATTTAA AAATAAAGTTCCTTTATATTTCTGTCCCATCAGGAAAACTGAAGGATATGGGGAATCATTGGTTATCTTCCATTGTGTTTTTCTTTATGG ACAGGAGCTAATGGAAGTGACAGTCATGTTCAAAGGAAGCATTTCTAGAAAAAAGGAGATAATGTTTTTAAATTTCATTATCAAACTTGG GCAATTCTGTTTGTGTAACTCCCCGACTAGTGGATGGGAGAGTCCCATTGCTAAAATTCAGCTACTCAGATAAATTCAGAATGGGTCAAG GCACCTGCCTGTTTTTGTTGGTGCACAGAGATTGACTTGATTCAGAGAGACAATTCACTCCATCCCTATGGCAGAGGAATGGGTTAGCCC TAATGTAGAATGTCATTGTTTTTAAAACTGTTTTATATCTTAAGAGTGCCTTATTAAAGTATAGATGTATGTCTTAAAATGTGGGTGATA GGAATTTTAAAGATTTATATAATGCATCAAAAGCCTTAGAATAAGAAAAGCTTTTTTTAAATTGCTTTATCTGTATATCTGAACTCTTGA AACTTATAGCTAAAACACTAGGATTTATCTGCAGTGTTCAGGGAGATAATTCTGCCTTTAATTGTCTAAAACAAAAACAAAACCAGCCAA CCTATGTTACACGTGAGATTAAAACCAATTTTTTCCCCATTTTTTCTCCTTTTTTCTCTTGCTGCCCACATTGTGCCTTTATTTTATGAG CCCCAGTTTTCTGGGCTTAGTTTAAAAAAAAAATCAAGTCTAAACATTGCATTTAGAAAGCTTTTGTTCTTGGATAAAAAGTCATACACT TTAAAAAAAAAAAAAACTTTTTCCAGGAAAATATATTGAAATCATGCTGCTGAGCCTCTATTTTCTTTCTTTGATGTTTTGATTCAGTAT TCTTTTATCATAAATTTTTAGCATTTAAAAATTCACTGATGTACATTAAGCCAATAAACTGCTTTAATGAATAACAAACTATGTAGTGTG TCCCTATTATAAATGCATTGGAGAAGTATTTTTATGAGACTCTTTACTCAGGTGCATGGTTACAGCCCACAGGGAGGCATGGAGTGCCAT GGAAGGATTCGCCACTACCCAGACCTTGTTTTTTGTTGTATTTTGGAAGACAGGTTTTTTAAAGAAACATTTTCCTCAGATTAAAAGATG ATGCTATTACAACTAGCATTGCCTCAAAAACTGGGACCAACCAAAGTGTGTCAACCCTGTTTCCTTAAAAGAGGCTATGAATCCCAAAGG CCACATCCAAGACAGGCAATAATGAGCAGAGTTTACAGCTCCTTTAATAAAATGTGTCAGTAATTTTAAGGTTTATAGTTCCCTCAACAC AATTGCTAATGCAGAATAGTGTAAAATGCGCTTCAAGAATGTTGATGATGATGATATAGAATTGTGGCTTTAGTAGCACAGAGGATGCCC CAACAAACTCATGGCGTTGAAACCACACAGTTCTCATTACTGTTATTTATTAGCTGTAGCATTCTCTGTCTCCTCTCTCTCCTCCTTTGA CCTTCTCCTCGACCAGCCATCATGACATTTACCATGAATTTACTTCCTCCCAAGAGTTTGGACTGCCCGTCAGATTGTTGCTGCACATAG >53009_53009_1_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000296411_IQGAP1_chr15_91030186_ENST00000268182_length(amino acids)=578AA_BP=262 MAAVETRVCETDGCSSEAKLQCPTCIKLGIQGSYFCSQECFKGSWATHKLLHKKAKDEKAKREVSSWTVEGDINTDPWAGYRYTGKLRPH YPLMPTRPVPSYIQRPDYADHPLGMSESEQALKGTSQIKLLSSEDIEGMRLVCRLAREVLDVAAGMIKPGVTTEEIDHAVHLACIARNCY PSPLNYYNFPKSCCTSVNEVICHGIPDRRPLQEGDIVNVDITLYRNGYHGDLNETFFVGEVDDGARKLVQTTYECLMQAIDAGESSGNLN DPNKEALAKTEVSLTLTNKFDVPGDENAEMDARTILLNTKRLIVDVIRFQPGETLTEILETPATSEQEAEHQRAMQRRAIRDAKTPDKMK KSKSVKEDSNLTLQEKKEKIQTGLKKLTELGTVDPKNKYQELINDIARDIRNQRRYRQRRKAELVKLQQTYAALNSKATFYGEQVDYYKS YIKTCLDNLASKGKVSKKPREMKGKKSKKISLKYTAARLHEKGVLLEIEDLQVNQFKNVIFEISPTEEVGDFEVKAKFMGVQMETFMLHY -------------------------------------------------------------- >53009_53009_2_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000296411_IQGAP1_chr15_91030186_ENST00000560738_length(transcript)=2134nt_BP=921nt CTCCCGCGCCCAGGCCAGCGGCGGGCCCAGCTCCTCCCCCGACTCGGTCTCTCTCCCCTCCCCTCCGCCCGGCAGTTCCTCCCTCCCGCC GCCGCCTCTTCCTCGGTGAGGCGCTCTTCCAGCGGGCAGGCAGCATGGCGGCCGTGGAGACGCGGGTGTGCGAGACAGACGGCTGCAGCA GTGAGGCCAAGCTCCAGTGTCCCACTTGCATCAAGCTGGGCATCCAGGGCTCGTACTTCTGCTCGCAGGAATGTTTTAAAGGAAGTTGGG CTACTCACAAGTTACTACATAAGAAAGCAAAAGATGAAAAGGCGAAGCGAGAAGTGTCTTCCTGGACTGTGGAAGGTGATATTAATACTG ACCCATGGGCAGGTTATCGATATACTGGTAAACTCAGACCACATTATCCACTGATGCCAACAAGGCCAGTGCCAAGTTATATTCAAAGAC CAGATTATGCTGATCATCCCTTAGGAATGTCTGAATCTGAACAGGCTCTTAAAGGTACTTCTCAGATTAAATTACTCTCATCTGAAGATA TAGAAGGGATGCGACTTGTATGTAGGCTTGCTAGAGAAGTTTTGGATGTTGCTGCCGGCATGATTAAACCAGGTGTAACTACTGAAGAAA TAGATCACGCTGTACACTTAGCATGTATTGCAAGAAATTGCTACCCTTCTCCCCTGAATTATTATAATTTCCCAAAGTCTTGTTGTACCT CAGTGAATGAAGTCATTTGCCATGGAATACCAGACAGAAGGCCCTTACAAGAAGGTGACATTGTTAATGTGGATATCACTCTTTATCGCA ATGGTTATCATGGGGACCTGAATGAGACATTTTTTGTTGGAGAAGTGGATGATGGAGCACGGAAACTTGTTCAGACCACATATGAGTGCC TGATGCAAGCCATTGATGCAGGGGAAAGCTCTGGCAATTTAAATGACCCAAATAAGGAGGCACTGGCTAAGACGGAAGTGTCTCTCACCC TGACCAACAAGTTCGACGTGCCTGGAGATGAGAATGCAGAAATGGATGCTCGAACCATCTTACTGAATACAAAACGTTTAATTGTGGATG TCATCCGGTTCCAGCCAGGAGAGACCTTGACTGAAATCCTAGAAACACCAGCCACCAGTGAACAGGAAGCAGAACATCAGAGAGCCATGC AGAGACGTGCTATCCGTGATGCCAAAACACCTGACAAGATGAAAAAGTCAAAATCTGTAAAGGAAGACAGCAACCTCACTCTTCAAGAGA AGAAAGAGAAGATCCAGACAGGTTTAAAGAAGCTAACAGAGCTTGGAACCGTGGACCCAAAGAACAAATACCAGGAACTGATCAACGACA TTGCCAGGGATATTCGGAATCAGCGGAGGTACCGACAGAGGAGAAAGGCCGAACTAGTGAAACTGCAACAGACATACGCTGCTCTGAACT CTAAGGCCACCTTTTATGGGGAGCAGGTGGATTACTATAAAAGCTATATCAAAACCTGCTTGGATAACTTAGCCAGCAAGGGCAAAGTCT CCAAAAAGCCTAGGGAAATGAAAGGAAAGAAAAGCAAAAAGATTTCTCTGAAATATACAGCAGCAAGACTACATGAAAAAGGAGTTCTTC TGGAAATTGAGGACCTGCAAGTGAATCAGTTTAAAAATGTTATATTTGAAATCAGTCCAACAGAAGAAGTTGGAGACTTCGAAGTGAAAG CCAAATTCATGGGAGTTCAAATGGAGACTTTTATGTTACATTATCAGGACCTGCTGCAGCTACAGTATGAAGGAGTTGCAGTCATGAAAT TATTTGATAGAGCTAAAGTAAATGTCAACCTCCTGATCTTCCTTCTCAACAAAAAGTTCTACGGGAAGTAATTGATCGTTTGCTGCCAGC CCAGAAGGATGAAGGAAAGAAGCACCTCACAGCTCCTTTCTAGGTCCTTCTTTCCTCATTGGAAGCAAAGACCTAGCCAACAACAGCACC TCAATCTGATACACTCCCGATGCCACATTTTTAACTCCTCTCGCTCTGATGGGACATTTGTTACCCTTTTTTCATAGTGAAATTGTGTTT >53009_53009_2_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000296411_IQGAP1_chr15_91030186_ENST00000560738_length(amino acids)=578AA_BP=262 MAAVETRVCETDGCSSEAKLQCPTCIKLGIQGSYFCSQECFKGSWATHKLLHKKAKDEKAKREVSSWTVEGDINTDPWAGYRYTGKLRPH YPLMPTRPVPSYIQRPDYADHPLGMSESEQALKGTSQIKLLSSEDIEGMRLVCRLAREVLDVAAGMIKPGVTTEEIDHAVHLACIARNCY PSPLNYYNFPKSCCTSVNEVICHGIPDRRPLQEGDIVNVDITLYRNGYHGDLNETFFVGEVDDGARKLVQTTYECLMQAIDAGESSGNLN DPNKEALAKTEVSLTLTNKFDVPGDENAEMDARTILLNTKRLIVDVIRFQPGETLTEILETPATSEQEAEHQRAMQRRAIRDAKTPDKMK KSKSVKEDSNLTLQEKKEKIQTGLKKLTELGTVDPKNKYQELINDIARDIRNQRRYRQRRKAELVKLQQTYAALNSKATFYGEQVDYYKS YIKTCLDNLASKGKVSKKPREMKGKKSKKISLKYTAARLHEKGVLLEIEDLQVNQFKNVIFEISPTEEVGDFEVKAKFMGVQMETFMLHY -------------------------------------------------------------- >53009_53009_3_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000544031_IQGAP1_chr15_91030186_ENST00000268182_length(transcript)=3722nt_BP=637nt ATGTGTACCATGCATAAAGATGAAAAGGCGAAGCGAGAAGTGTCTTCCTGGACTGTGGAAGGTGATATTAATACTGACCCATGGGCAGGT TATCGATATACTGGTAAACTCAGACCACATTATCCACTGATGCCAACAAGGCCAGTGCCAAGTTATATTCAAAGACCAGATTATGCTGAT CATCCCTTAGGAATGTCTGAATCTGAACAGGCTCTTAAAGGTACTTCTCAGATTAAATTACTCTCATCTGAAGATATAGAAGGGATGCGA CTTGTATGTAGGCTTGCTAGAGAAGTTTTGGATGTTGCTGCCGGCATGATTAAACCAGGTGTAACTACTGAAGAAATAGATCACGCTGTA CACTTAGCATGTATTGCAAGAAATTGCTACCCTTCTCCCCTGAATTATTATAATTTCCCAAAGTCTTGTTGTACCTCAGTGAATGAAGTC ATTTGCCATGGAATACCAGACAGAAGGCCCTTACAAGAAGGTGACATTGTTAATGTGGATATCACTCTTTATCGCAATGGTTATCATGGG GACCTGAATGAGACATTTTTTGTTGGAGAAGTGGATGATGGAGCACGGAAACTTGTTCAGACCACATATGAGTGCCTGATGCAAGCCATT GATGCAGGGGAAAGCTCTGGCAATTTAAATGACCCAAATAAGGAGGCACTGGCTAAGACGGAAGTGTCTCTCACCCTGACCAACAAGTTC GACGTGCCTGGAGATGAGAATGCAGAAATGGATGCTCGAACCATCTTACTGAATACAAAACGTTTAATTGTGGATGTCATCCGGTTCCAG CCAGGAGAGACCTTGACTGAAATCCTAGAAACACCAGCCACCAGTGAACAGGAAGCAGAACATCAGAGAGCCATGCAGAGACGTGCTATC CGTGATGCCAAAACACCTGACAAGATGAAAAAGTCAAAATCTGTAAAGGAAGACAGCAACCTCACTCTTCAAGAGAAGAAAGAGAAGATC CAGACAGGTTTAAAGAAGCTAACAGAGCTTGGAACCGTGGACCCAAAGAACAAATACCAGGAACTGATCAACGACATTGCCAGGGATATT CGGAATCAGCGGAGGTACCGACAGAGGAGAAAGGCCGAACTAGTGAAACTGCAACAGACATACGCTGCTCTGAACTCTAAGGCCACCTTT TATGGGGAGCAGGTGGATTACTATAAAAGCTATATCAAAACCTGCTTGGATAACTTAGCCAGCAAGGGCAAAGTCTCCAAAAAGCCTAGG GAAATGAAAGGAAAGAAAAGCAAAAAGATTTCTCTGAAATATACAGCAGCAAGACTACATGAAAAAGGAGTTCTTCTGGAAATTGAGGAC CTGCAAGTGAATCAGTTTAAAAATGTTATATTTGAAATCAGTCCAACAGAAGAAGTTGGAGACTTCGAAGTGAAAGCCAAATTCATGGGA GTTCAAATGGAGACTTTTATGTTACATTATCAGGACCTGCTGCAGCTACAGTATGAAGGAGTTGCAGTCATGAAATTATTTGATAGAGCT AAAGTAAATGTCAACCTCCTGATCTTCCTTCTCAACAAAAAGTTCTACGGGAAGTAATTGATCGTTTGCTGCCAGCCCAGAAGGATGAAG GAAAGAAGCACCTCACAGCTCCTTTCTAGGTCCTTCTTTCCTCATTGGAAGCAAAGACCTAGCCAACAACAGCACCTCAATCTGATACAC TCCCGATGCCACATTTTTAACTCCTCTCGCTCTGATGGGACATTTGTTACCCTTTTTTCATAGTGAAATTGTGTTTCAGGCTTAGTCTGA CCTTTCTGGTTTCTTCATTTTCTTCCATTACTTAGGAAAGAGTGGAAACTCCACTAAAATTTCTCTGTGTTGTTACAGTCTTAGAGGTTG CAGTACTATATTGTAAGCTTTGGTGTTTGTTTAATTAGCAATAGGGATGGTAGGATTCAAATGTGTGTCATTTAGAAGTGGAAGCTATTA GCACCAATGACATAAATACATACAAGACACACAACTAAAATGTCATGTTATTAACAGTTATTAGGTTGTCATTTAAAAATAAAGTTCCTT TATATTTCTGTCCCATCAGGAAAACTGAAGGATATGGGGAATCATTGGTTATCTTCCATTGTGTTTTTCTTTATGGACAGGAGCTAATGG AAGTGACAGTCATGTTCAAAGGAAGCATTTCTAGAAAAAAGGAGATAATGTTTTTAAATTTCATTATCAAACTTGGGCAATTCTGTTTGT GTAACTCCCCGACTAGTGGATGGGAGAGTCCCATTGCTAAAATTCAGCTACTCAGATAAATTCAGAATGGGTCAAGGCACCTGCCTGTTT TTGTTGGTGCACAGAGATTGACTTGATTCAGAGAGACAATTCACTCCATCCCTATGGCAGAGGAATGGGTTAGCCCTAATGTAGAATGTC ATTGTTTTTAAAACTGTTTTATATCTTAAGAGTGCCTTATTAAAGTATAGATGTATGTCTTAAAATGTGGGTGATAGGAATTTTAAAGAT TTATATAATGCATCAAAAGCCTTAGAATAAGAAAAGCTTTTTTTAAATTGCTTTATCTGTATATCTGAACTCTTGAAACTTATAGCTAAA ACACTAGGATTTATCTGCAGTGTTCAGGGAGATAATTCTGCCTTTAATTGTCTAAAACAAAAACAAAACCAGCCAACCTATGTTACACGT GAGATTAAAACCAATTTTTTCCCCATTTTTTCTCCTTTTTTCTCTTGCTGCCCACATTGTGCCTTTATTTTATGAGCCCCAGTTTTCTGG GCTTAGTTTAAAAAAAAAATCAAGTCTAAACATTGCATTTAGAAAGCTTTTGTTCTTGGATAAAAAGTCATACACTTTAAAAAAAAAAAA AACTTTTTCCAGGAAAATATATTGAAATCATGCTGCTGAGCCTCTATTTTCTTTCTTTGATGTTTTGATTCAGTATTCTTTTATCATAAA TTTTTAGCATTTAAAAATTCACTGATGTACATTAAGCCAATAAACTGCTTTAATGAATAACAAACTATGTAGTGTGTCCCTATTATAAAT GCATTGGAGAAGTATTTTTATGAGACTCTTTACTCAGGTGCATGGTTACAGCCCACAGGGAGGCATGGAGTGCCATGGAAGGATTCGCCA CTACCCAGACCTTGTTTTTTGTTGTATTTTGGAAGACAGGTTTTTTAAAGAAACATTTTCCTCAGATTAAAAGATGATGCTATTACAACT AGCATTGCCTCAAAAACTGGGACCAACCAAAGTGTGTCAACCCTGTTTCCTTAAAAGAGGCTATGAATCCCAAAGGCCACATCCAAGACA GGCAATAATGAGCAGAGTTTACAGCTCCTTTAATAAAATGTGTCAGTAATTTTAAGGTTTATAGTTCCCTCAACACAATTGCTAATGCAG AATAGTGTAAAATGCGCTTCAAGAATGTTGATGATGATGATATAGAATTGTGGCTTTAGTAGCACAGAGGATGCCCCAACAAACTCATGG CGTTGAAACCACACAGTTCTCATTACTGTTATTTATTAGCTGTAGCATTCTCTGTCTCCTCTCTCTCCTCCTTTGACCTTCTCCTCGACC AGCCATCATGACATTTACCATGAATTTACTTCCTCCCAAGAGTTTGGACTGCCCGTCAGATTGTTGCTGCACATAGTTGCCTTTGTATCT >53009_53009_3_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000544031_IQGAP1_chr15_91030186_ENST00000268182_length(amino acids)=528AA_BP=212 MCTMHKDEKAKREVSSWTVEGDINTDPWAGYRYTGKLRPHYPLMPTRPVPSYIQRPDYADHPLGMSESEQALKGTSQIKLLSSEDIEGMR LVCRLAREVLDVAAGMIKPGVTTEEIDHAVHLACIARNCYPSPLNYYNFPKSCCTSVNEVICHGIPDRRPLQEGDIVNVDITLYRNGYHG DLNETFFVGEVDDGARKLVQTTYECLMQAIDAGESSGNLNDPNKEALAKTEVSLTLTNKFDVPGDENAEMDARTILLNTKRLIVDVIRFQ PGETLTEILETPATSEQEAEHQRAMQRRAIRDAKTPDKMKKSKSVKEDSNLTLQEKKEKIQTGLKKLTELGTVDPKNKYQELINDIARDI RNQRRYRQRRKAELVKLQQTYAALNSKATFYGEQVDYYKSYIKTCLDNLASKGKVSKKPREMKGKKSKKISLKYTAARLHEKGVLLEIED -------------------------------------------------------------- >53009_53009_4_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000544031_IQGAP1_chr15_91030186_ENST00000560738_length(transcript)=1850nt_BP=637nt ATGTGTACCATGCATAAAGATGAAAAGGCGAAGCGAGAAGTGTCTTCCTGGACTGTGGAAGGTGATATTAATACTGACCCATGGGCAGGT TATCGATATACTGGTAAACTCAGACCACATTATCCACTGATGCCAACAAGGCCAGTGCCAAGTTATATTCAAAGACCAGATTATGCTGAT CATCCCTTAGGAATGTCTGAATCTGAACAGGCTCTTAAAGGTACTTCTCAGATTAAATTACTCTCATCTGAAGATATAGAAGGGATGCGA CTTGTATGTAGGCTTGCTAGAGAAGTTTTGGATGTTGCTGCCGGCATGATTAAACCAGGTGTAACTACTGAAGAAATAGATCACGCTGTA CACTTAGCATGTATTGCAAGAAATTGCTACCCTTCTCCCCTGAATTATTATAATTTCCCAAAGTCTTGTTGTACCTCAGTGAATGAAGTC ATTTGCCATGGAATACCAGACAGAAGGCCCTTACAAGAAGGTGACATTGTTAATGTGGATATCACTCTTTATCGCAATGGTTATCATGGG GACCTGAATGAGACATTTTTTGTTGGAGAAGTGGATGATGGAGCACGGAAACTTGTTCAGACCACATATGAGTGCCTGATGCAAGCCATT GATGCAGGGGAAAGCTCTGGCAATTTAAATGACCCAAATAAGGAGGCACTGGCTAAGACGGAAGTGTCTCTCACCCTGACCAACAAGTTC GACGTGCCTGGAGATGAGAATGCAGAAATGGATGCTCGAACCATCTTACTGAATACAAAACGTTTAATTGTGGATGTCATCCGGTTCCAG CCAGGAGAGACCTTGACTGAAATCCTAGAAACACCAGCCACCAGTGAACAGGAAGCAGAACATCAGAGAGCCATGCAGAGACGTGCTATC CGTGATGCCAAAACACCTGACAAGATGAAAAAGTCAAAATCTGTAAAGGAAGACAGCAACCTCACTCTTCAAGAGAAGAAAGAGAAGATC CAGACAGGTTTAAAGAAGCTAACAGAGCTTGGAACCGTGGACCCAAAGAACAAATACCAGGAACTGATCAACGACATTGCCAGGGATATT CGGAATCAGCGGAGGTACCGACAGAGGAGAAAGGCCGAACTAGTGAAACTGCAACAGACATACGCTGCTCTGAACTCTAAGGCCACCTTT TATGGGGAGCAGGTGGATTACTATAAAAGCTATATCAAAACCTGCTTGGATAACTTAGCCAGCAAGGGCAAAGTCTCCAAAAAGCCTAGG GAAATGAAAGGAAAGAAAAGCAAAAAGATTTCTCTGAAATATACAGCAGCAAGACTACATGAAAAAGGAGTTCTTCTGGAAATTGAGGAC CTGCAAGTGAATCAGTTTAAAAATGTTATATTTGAAATCAGTCCAACAGAAGAAGTTGGAGACTTCGAAGTGAAAGCCAAATTCATGGGA GTTCAAATGGAGACTTTTATGTTACATTATCAGGACCTGCTGCAGCTACAGTATGAAGGAGTTGCAGTCATGAAATTATTTGATAGAGCT AAAGTAAATGTCAACCTCCTGATCTTCCTTCTCAACAAAAAGTTCTACGGGAAGTAATTGATCGTTTGCTGCCAGCCCAGAAGGATGAAG GAAAGAAGCACCTCACAGCTCCTTTCTAGGTCCTTCTTTCCTCATTGGAAGCAAAGACCTAGCCAACAACAGCACCTCAATCTGATACAC TCCCGATGCCACATTTTTAACTCCTCTCGCTCTGATGGGACATTTGTTACCCTTTTTTCATAGTGAAATTGTGTTTCAGGCTTAGTCTGA >53009_53009_4_METAP1-IQGAP1_METAP1_chr4_99966461_ENST00000544031_IQGAP1_chr15_91030186_ENST00000560738_length(amino acids)=528AA_BP=212 MCTMHKDEKAKREVSSWTVEGDINTDPWAGYRYTGKLRPHYPLMPTRPVPSYIQRPDYADHPLGMSESEQALKGTSQIKLLSSEDIEGMR LVCRLAREVLDVAAGMIKPGVTTEEIDHAVHLACIARNCYPSPLNYYNFPKSCCTSVNEVICHGIPDRRPLQEGDIVNVDITLYRNGYHG DLNETFFVGEVDDGARKLVQTTYECLMQAIDAGESSGNLNDPNKEALAKTEVSLTLTNKFDVPGDENAEMDARTILLNTKRLIVDVIRFQ PGETLTEILETPATSEQEAEHQRAMQRRAIRDAKTPDKMKKSKSVKEDSNLTLQEKKEKIQTGLKKLTELGTVDPKNKYQELINDIARDI RNQRRYRQRRKAELVKLQQTYAALNSKATFYGEQVDYYKSYIKTCLDNLASKGKVSKKPREMKGKKSKKISLKYTAARLHEKGVLLEIED -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for METAP1-IQGAP1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for METAP1-IQGAP1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for METAP1-IQGAP1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
