
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:METTL16-FAM101B (FusionGDB2 ID:53061) |
Fusion Gene Summary for METTL16-FAM101B |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: METTL16-FAM101B | Fusion gene ID: 53061 | Hgene | Tgene | Gene symbol | METTL16 | FAM101B | Gene ID | 79066 | 359845 |
| Gene name | methyltransferase like 16 | refilin B | |
| Synonyms | METT10D | CFM1|FAM101B | |
| Cytomap | 17p13.3 | 17p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | RNA N6-adenosine-methyltransferase METTL16N6-adenosine-methyltransferase METTL16U6 small nuclear RNA (adenine-(43)-N(6))-methyltransferaseU6 snRNA methyltransferasemethyltransferase 10 domain containingmethyltransferase 10 domain-containing proteinm | refilin-Bfamily with sequence similarity 101, member Bfilamin-interacting protein FAM101Bprotein FAM101BrefilinBregulator of filamin protein B | |
| Modification date | 20200320 | 20200313 | |
| UniProtAcc | Q86W50 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000571669, ENST00000263092, ENST00000538844, | ENST00000329099, | |
| Fusion gene scores | * DoF score | 10 X 12 X 7=840 | 2 X 2 X 2=8 |
| # samples | 12 | 2 | |
| ** MAII score | log2(12/840*10)=-2.8073549220576 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(2/8*10)=1.32192809488736 | |
| Context | PubMed: METTL16 [Title/Abstract] AND FAM101B [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | METTL16(2344784)-FAM101B(293285), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | METTL16 | GO:0006556 | S-adenosylmethionine biosynthetic process | 28525753|30197297 |
| Hgene | METTL16 | GO:0010608 | posttranscriptional regulation of gene expression | 28525753 |
| Hgene | METTL16 | GO:0048024 | regulation of mRNA splicing, via spliceosome | 28525753 |
| Hgene | METTL16 | GO:0080009 | mRNA methylation | 30197297|30197299 |
| Hgene | METTL16 | GO:0120049 | snRNA (adenine-N6)-methylation | 28525753|29051200 |
| Hgene | METTL16 | GO:1905869 | negative regulation of 3'-UTR-mediated mRNA stabilization | 28525753 |
 Fusion gene breakpoints across METTL16 (5'-gene) Fusion gene breakpoints across METTL16 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
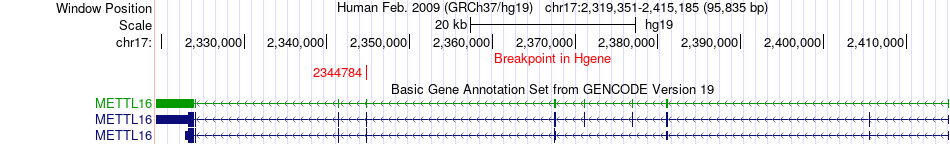 |
 Fusion gene breakpoints across FAM101B (3'-gene) Fusion gene breakpoints across FAM101B (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
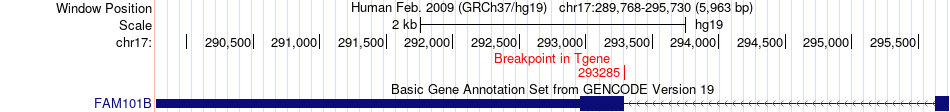 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUSC | TCGA-98-8020 | METTL16 | chr17 | 2344784 | - | FAM101B | chr17 | 293285 | - |
Top |
Fusion Gene ORF analysis for METTL16-FAM101B |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5UTR-3CDS | ENST00000571669 | ENST00000329099 | METTL16 | chr17 | 2344784 | - | FAM101B | chr17 | 293285 | - |
| In-frame | ENST00000263092 | ENST00000329099 | METTL16 | chr17 | 2344784 | - | FAM101B | chr17 | 293285 | - |
| In-frame | ENST00000538844 | ENST00000329099 | METTL16 | chr17 | 2344784 | - | FAM101B | chr17 | 293285 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000263092 | METTL16 | chr17 | 2344784 | - | ENST00000329099 | FAM101B | chr17 | 293285 | - | 4443 | 926 | 104 | 1246 | 380 |
| ENST00000538844 | METTL16 | chr17 | 2344784 | - | ENST00000329099 | FAM101B | chr17 | 293285 | - | 4188 | 671 | 527 | 991 | 154 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000263092 | ENST00000329099 | METTL16 | chr17 | 2344784 | - | FAM101B | chr17 | 293285 | - | 0.005163384 | 0.9948367 |
| ENST00000538844 | ENST00000329099 | METTL16 | chr17 | 2344784 | - | FAM101B | chr17 | 293285 | - | 0.6385494 | 0.36145064 |
Top |
Fusion Genomic Features for METTL16-FAM101B |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
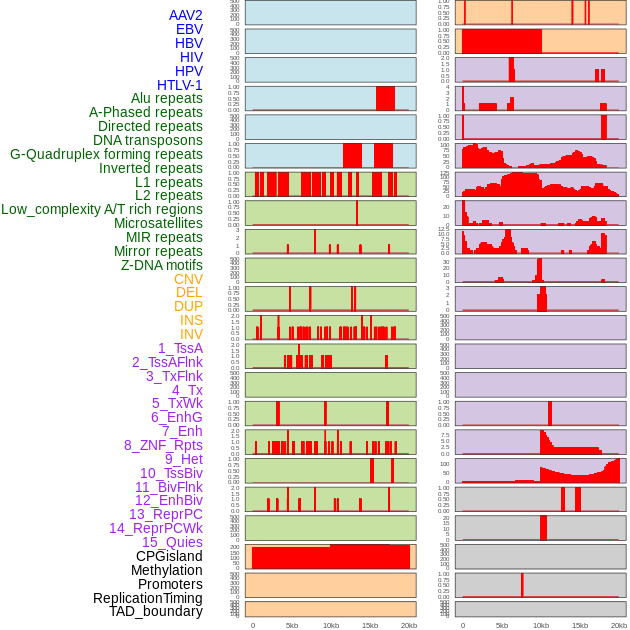 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for METTL16-FAM101B |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr17:2344784/chr17:293285) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| METTL16 | . |
| FUNCTION: RNA N6-methyltransferase that methylates adenosine residues at the N(6) position of a subset of RNAs and is involved in S-adenosyl-L-methionine homeostasis by regulating expression of MAT2A transcripts (PubMed:28525753, PubMed:30197299, PubMed:30197297). Able to N6-methylate a subset of mRNAs and U6 small nuclear RNAs (U6 snRNAs) (PubMed:28525753). In contrast to the METTL3-METTL14 heterodimer, only able to methylate a limited number of RNAs: requires both a 5'UACAGAGAA-3' nonamer sequence and a specific RNA structure (PubMed:28525753, PubMed:30197299, PubMed:30197297). Plays a key role in S-adenosyl-L-methionine homeostasis by mediating N6-methylation of MAT2A mRNAs, altering splicing and/or stability of MAT2A transcripts: in presence of S-adenosyl-L-methionine, binds the 3'-UTR region of MAT2A mRNA and specifically N6-methylates the first hairpin of MAT2A mRNA, impairing MAT2A expression (PubMed:28525753). In S-adenosyl-L-methionine-limiting conditions, binds the 3'-UTR region of MAT2A mRNA but stalls due to the lack of a methyl donor, preventing N6-methylation and promoting expression of MAT2A (PubMed:28525753). In addition to mRNAs, also able to mediate N6-methylation of U6 small nuclear RNA (U6 snRNA): specifically N6-methylates adenine in position 43 of U6 snRNAs (PubMed:28525753, PubMed:29051200, PubMed:32266935). Also able to bind various lncRNAs, such as 7SK snRNA (7SK RNA) or 7SL RNA (PubMed:29051200). Specifically binds the 3'-end of the MALAT1 long non-coding RNA (PubMed:27872311). {ECO:0000269|PubMed:27872311, ECO:0000269|PubMed:28525753, ECO:0000269|PubMed:29051200, ECO:0000269|PubMed:30197297, ECO:0000269|PubMed:30197299, ECO:0000269|PubMed:32266935}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 163_167 | 266 | 563.0 | Region | K-loop |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 17_20 | 266 | 563.0 | Region | RNA-binding |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 199_211 | 266 | 563.0 | Region | RNA-binding |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 250_254 | 266 | 563.0 | Region | RNA-binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 277_283 | 266 | 563.0 | Region | RNA-binding |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 289_400 | 266 | 563.0 | Region | VCR 1 |
| Hgene | METTL16 | chr17:2344784 | chr17:293285 | ENST00000263092 | - | 7 | 10 | 514_562 | 266 | 563.0 | Region | VCR 2 |
Top |
Fusion Gene Sequence for METTL16-FAM101B |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >53061_53061_1_METTL16-FAM101B_METTL16_chr17_2344784_ENST00000263092_FAM101B_chr17_293285_ENST00000329099_length(transcript)=4443nt_BP=926nt AAGATGGCGGCGCGCTGGGAGCGTATCATCTGCGTTTCTAGGAGCTTCGCTATGCGGCTGCTTTAAGATTCTAGGGTTGTACAGGCCCAC GCCAGACACGACGTCTGGCAGGAACCTCGGCCTCAGAGATGGCTCTGAGTAAATCAATGCATGCAAGAAATAGATACAAGGACAAACCTC CTGACTTTGCATATCTGGCATCCAAATATCCAGATTTTAAGCAGCATGTTCAGATAAATCTGAATGGAAGAGTGAGCCTTAATTTTAAAG ACCCCGAAGCAGTCAGAGCTCTGACGTGTACTCTCCTAAGGGAAGATTTTGGACTTTCTATTGATATTCCATTGGAGAGACTAATTCCCA CAGTTCCCTTGAGACTCAACTATATTCACTGGGTAGAAGATCTGATCGGTCACCAGGATTCTGACAAAAGTACTCTCCGAAGAGGAATTG ACATAGGCACGGGGGCATCTTGCATCTACCCCTTACTTGGAGCAACCTTGAATGGCTGGTATTTCCTCGCAACAGAAGTGGATGATATGT GTTTCAACTATGCAAAGAAAAATGTGGAACAGAATAACTTATCTGATCTCATAAAAGTGGTGAAAGTGCCACAGAAGACACTCCTGATGG ATGCTCTTAAAGAAGAATCTGAGATAATCTATGACTTTTGCATGTGCAACCCTCCCTTTTTTGCCAATCAATTGGAAGCCAAGGGAGTAA ACTCACGAAATCCTCGAAGACCTCCGCCTAGTTCTGTTAATACAGGAGGCATCACAGAGATCATGGCAGAAGGAGGTGAATTAGAGTTTG TTAAAAGGATCATCCATGACAGTCTACAACTTAAAAAAAGATTAAGATGGTATAGCTGCATGCTGGGAAAGAAATGCAGCCTGGCGCCTC TGAAGGAGGAGCTTCGCATACAAGGGGTACACCTCCTTGGTCAAGTACGACTCCGAGAGGCACTTCATCGACGACGTGCAGCTGCCCCTG GGCCTGGCGGTGGCCTCCTGCAGCCAGACGGTCACCTGTGTCCCCAATGGCACGTGGCGCAACTACAAGGCCGAGGTGCGCTTCGAGCCA CGCCACAGGCCCACCCGCTTCCTCAGTACCACCATCGTGTACCCCAAGTACCCCAAGGCCGTCTACACCACCACCCTGGATTACAACTGC CGCAAGACGCTGAGGAGGTTTCTGTCCAGCGTGGAGCTCGAAGCCGCGGAGCTCCCGGGCAGCGACGACCTCTCTGATGAATGCTGACCC AGCAGGCTGGGCTGGGGTCGGACCGGCTCTCCCGCTGTCCTGCCCCCGACTGCCCTGGCCAAGCTGGGCGACCTTACCCTTCACGTCATT CCAGCCCCAGAGGAGTCTAACCTAGAGATACCTCCAAGCGCGGCCGGGGGAAGAGAAAAAAAAGAAAAAATCACTGCATCTCATAATTAT TGAGATCTTTGTGTTGTAATTTTCAAGCTGTTTTTAGAGGGAATATATGTCCTGGTTTGCTGCTGTGTTGTTTCCCCAAAGTCTCAATCA GATGAGGCAACAAAAAGACCACCAGAATTGCAGGAAAGCAGCAGCAGTCTGGGATAGGGAGGGGGGAGGAGAGCTCTCCCTCCGTGGTCA GTTTGTCAGAAGAAAAGCATGGAAAAAGTGAGATTTAAGAAATCTCAGCCACGGGTGTCTTCACTGCACAAAAGTGGTGCTAATTCAGTA AACAGTGACACCTGTGTGGGTTCAATCTGTGGAGAGTTGAGTTCCATTCCTTTGTTTTTAAATTTCCACCTCCATTTGTGTTTCCTATTA ACCTAAACTCTTGTAATTCTCTAAATCTTTTTTCATGATATAAAAAAAAAAGAAAATCCCAAACCACAATTTAAGATGCCTTTTTTTTTT AAAAAAAAAGGTTCTCCGTGTTGCCACCTGCTCCGGGAAGGCCAGGCCGATGGGCGGCCAGGTTTGGGAGCAGCGGCTTCGGCTCCGATT GCTTCCTGACGGTCAGTTCCAAGCACAGGCTCTGAGAAACGCCTCGAACCCTTGGTATCTGGTGACCGTCTCCGCAGGCTTCCGTGAGTC TCCGATAGGTGTTTTGGGCTTTTGGAATACTTTCAGAATACTTTCCCATTTTTTCATAAGGCAAAACGGAGTGGACGGCCTTTTTTTGAG AGAAACAGCATTCTAAGAGTGTGCTTGGAAAACACACTTTTTGGGAGTCCACGTGGGGAGTGGACACTCAGGACAGGAAGCGGTGGCCCT AAGACCTGCGGCCACCCGGAGGAGGCAGACGCAGGCGATGACCTGGTTCATCATGAGAGGAGCTAGAGTTGACTCAGTCTTAGGATCTCG ATGGCGTCAGAATTGCAGAGGGTTTCTGCCAGTGCCTTAGGTCTAATTCAGAAAGCAAGGAAGGCTGGGCACGGTGGCTGGCGCCTGTAG TCTCAGCTACTTGGAAACTGAGGCAGGAGGATCTCTTGAGCCCAAGAATTCGAGACCGGTCTGGCAACATAGTGAGCCTCCATCTCTAAA AAAAAGTGGCAAAGCTGAGGAAATCTGTGCCCCCATCCCCCGACACCAGACAAGCTAAGGTGCATTCAGAGTAGGGAACGATGTCAAAAC CTGGCCGCCGTGGTGGCGCGCATTCCTCTCACACCACCGTACGGAGGCAGGTGACAGTCAACTGAAATGTCCCCAAGGACCGTGTGAATT CGGCTGGCGTGGCCCAGCAGCGGGGTGGACCAGGGTGCGGTGGCTTCAGACCTGCCTCCCGCCACCCCGTCCTCCCCTAACCCCTTCTTG GGACCTCCTGCCAATCCCTGTGGTGTTTGAGTAAAGAAAGTGGCTGATGGTTTTACTTTTTTCTTCCAATATTGGAAAGAACAAACTAGA TATTCCCCTTGAACACATCCAAATTATTTCCAGTGCCTGAGGGTTACCGAGCACTGTAAGCTCAGCTCGGCTGTAGCTCCGGGTTTCTGC CGATGGCCTCCATCTGCGCCGAAAGCTGAACTGAGCCCCGCGGTGCCCCGGAGCCACAGCAGCAGGTGCAGCGACAATCGCCGTCGTGCG CAGTCGCTGGAGTCCTGGAGAGCCACGCTGTCCTCTGCAGACAACAGCATCAGTCCGCTGTGCCCCAATGCATCGACGCCCTGTGGACGG GGCCTGCAGTCCACCCGTGGTCTGAGCTTGCCGGGCTGAAGCTCGGCTGGGACCCAGGGAGGGGCCACCCTGGGACCCACAGTCTCCTCA GTCACGTGCAGAGCAAAAAATGCTTCCTGTGTTTGGAAGAGCAAGTGACGCAGCATTTATTGATAAAAATGGATTTTTCTGTCTGATTTC ATGCTGTGGTCGTTTAAGTGGTCAATGTCTTTTTTTTTTTTTTTTTTGACATGGAGTCTCGCTCTGTTGCCCAGGCTGAGTGGAATGGCG CCAACTCAGCTCACTGCAACCTTCACCTCCCAGGCTCAAGCGACTCTCCTGCCTCAGCCTCCTGAGCAGCTGGGATTACAGGTGCCCACC ACCACGCCCAGCTAATTTTTGTATTTTTAGTAGAGACGGGGTTTCACCATGTTGGCCAGGCTGGTCTCGAACACCTGACCTTGTGATCCA CCCGCCTCGGCCTCCCAATAATTTGTAAGTTATGTTAGCGGGATCCTCAAGGCCTTGCTTTGCCCCGTGGAGACGCTTGCTCGGATGAGC TCAGGAAACAGTACCGGCTGCGTGGCAGGTCTGGGTGTTGTGTGCGAGGACGTGGCCTTTGAACACCGCTGTGTTCTCAGAGGTCCTTAG GAGATATTTTTTTTTGTCTTAGGGGGACTGTGTTAAGTTCAGACAAATCATGCTGGGTGTGTAGAGAGTGTGAAATACGTCAGTGAAGTA AGTAGCAGTGAGCGATTGTGAATGTGTAATGTAAATGGAAAACCGGGTTTTACCGTGTTAAGTTATTCACTAGGGAGCCAGTCGTAGTTC TTTGTAATCCTCTTTCTTCCAAACCTGCTTTGCTGAAAGTTGCAGAAAAGGAAGTGTGTGGAGAGAAACAGAACCCTTCAGGGTGGGTCA GAGGACGCCATCCACAGTGGATTCGTGTTCGTTTGCAGGTGGAAGCAGTGATTTTTAGGACCCACTGATTAAAAACAAACATTCCCAAGT GTCTCTGAGAGATGCTGTTTATTTGTTAATTAAAAAGCTTTTTTCTCTGTCTTTTAAATTATGGCTTTCATGTAATAAGGATATTTTTAG TGAAAAATTGTTTTCCTTTCAAATTACAGACCTTTTAAAAAAACTTAATTTGAGCGAGTACCTTTTCATTTGACACTTTTCCTGTTTCTA ACCTTAGGAAACCAGAATAGCGTTTGGCAGACACGACGTTTTCAGTTTACCTTTGACACCTGCCCCACTCCATTTTGCTTTGTGATGTCT >53061_53061_1_METTL16-FAM101B_METTL16_chr17_2344784_ENST00000263092_FAM101B_chr17_293285_ENST00000329099_length(amino acids)=380AA_BP=1 MAGTSASEMALSKSMHARNRYKDKPPDFAYLASKYPDFKQHVQINLNGRVSLNFKDPEAVRALTCTLLREDFGLSIDIPLERLIPTVPLR LNYIHWVEDLIGHQDSDKSTLRRGIDIGTGASCIYPLLGATLNGWYFLATEVDDMCFNYAKKNVEQNNLSDLIKVVKVPQKTLLMDALKE ESEIIYDFCMCNPPFFANQLEAKGVNSRNPRRPPPSSVNTGGITEIMAEGGELEFVKRIIHDSLQLKKRLRWYSCMLGKKCSLAPLKEEL RIQGVHLLGQVRLREALHRRRAAAPGPGGGLLQPDGHLCPQWHVAQLQGRGALRATPQAHPLPQYHHRVPQVPQGRLHHHPGLQLPQDAE -------------------------------------------------------------- >53061_53061_2_METTL16-FAM101B_METTL16_chr17_2344784_ENST00000538844_FAM101B_chr17_293285_ENST00000329099_length(transcript)=4188nt_BP=671nt ACAAGATGGCGGCGCGCTGGGAGCGTATCATCTGCGTTTCTAGGAGCTTCGCTATGCGGCTGCTTTAAGATTCTAGGGTTGTACAGGCCC ACGCCAGACACGACGTCTGGCAGGAACCTCGGCCTCAGAGATGGCTCTGAGTAAATCAATGCATGCAAGAAATAGATACAAGGACAAACC TCCTGACTTTGCATATCTGGCATCCAAATATCCAGATTTTAAGCAGCATGTTCAGATAAATCTGAATGGAAGAGTGAGCCTTAATTTTAA AGACCCCGAAGCAGTCAGAGCTCTGACGTGTACTCTCCTAAGGGAAGATTTTGGACTTTCTATTGATATTCCATTGGAGAGACTAATTCC CACAGTTCCCTTGAGACTCAACTATATTCACTGGGTAGAAGATCTGATCGGTCACCAGGATTCTGACAAAAGTACTCTCCGAAGAGGAAT TGACATAGGGAGTAAACTCACGAAATCCTCGAAGACCTCCGCCTAGTTCTGTTAATACAGGAGGCATCACAGAGATCATGGCAGAAGGAG GTGAATTAGAGTTTGTTAAAAGGATCATCCATGACAGTCTACAACTTAAAAAAAGATTAAGATGGTATAGCTGCATGCTGGGAAAGAAAT GCAGCCTGGCGCCTCTGAAGGAGGAGCTTCGCATACAAGGGGTACACCTCCTTGGTCAAGTACGACTCCGAGAGGCACTTCATCGACGAC GTGCAGCTGCCCCTGGGCCTGGCGGTGGCCTCCTGCAGCCAGACGGTCACCTGTGTCCCCAATGGCACGTGGCGCAACTACAAGGCCGAG GTGCGCTTCGAGCCACGCCACAGGCCCACCCGCTTCCTCAGTACCACCATCGTGTACCCCAAGTACCCCAAGGCCGTCTACACCACCACC CTGGATTACAACTGCCGCAAGACGCTGAGGAGGTTTCTGTCCAGCGTGGAGCTCGAAGCCGCGGAGCTCCCGGGCAGCGACGACCTCTCT GATGAATGCTGACCCAGCAGGCTGGGCTGGGGTCGGACCGGCTCTCCCGCTGTCCTGCCCCCGACTGCCCTGGCCAAGCTGGGCGACCTT ACCCTTCACGTCATTCCAGCCCCAGAGGAGTCTAACCTAGAGATACCTCCAAGCGCGGCCGGGGGAAGAGAAAAAAAAGAAAAAATCACT GCATCTCATAATTATTGAGATCTTTGTGTTGTAATTTTCAAGCTGTTTTTAGAGGGAATATATGTCCTGGTTTGCTGCTGTGTTGTTTCC CCAAAGTCTCAATCAGATGAGGCAACAAAAAGACCACCAGAATTGCAGGAAAGCAGCAGCAGTCTGGGATAGGGAGGGGGGAGGAGAGCT CTCCCTCCGTGGTCAGTTTGTCAGAAGAAAAGCATGGAAAAAGTGAGATTTAAGAAATCTCAGCCACGGGTGTCTTCACTGCACAAAAGT GGTGCTAATTCAGTAAACAGTGACACCTGTGTGGGTTCAATCTGTGGAGAGTTGAGTTCCATTCCTTTGTTTTTAAATTTCCACCTCCAT TTGTGTTTCCTATTAACCTAAACTCTTGTAATTCTCTAAATCTTTTTTCATGATATAAAAAAAAAAGAAAATCCCAAACCACAATTTAAG ATGCCTTTTTTTTTTAAAAAAAAAGGTTCTCCGTGTTGCCACCTGCTCCGGGAAGGCCAGGCCGATGGGCGGCCAGGTTTGGGAGCAGCG GCTTCGGCTCCGATTGCTTCCTGACGGTCAGTTCCAAGCACAGGCTCTGAGAAACGCCTCGAACCCTTGGTATCTGGTGACCGTCTCCGC AGGCTTCCGTGAGTCTCCGATAGGTGTTTTGGGCTTTTGGAATACTTTCAGAATACTTTCCCATTTTTTCATAAGGCAAAACGGAGTGGA CGGCCTTTTTTTGAGAGAAACAGCATTCTAAGAGTGTGCTTGGAAAACACACTTTTTGGGAGTCCACGTGGGGAGTGGACACTCAGGACA GGAAGCGGTGGCCCTAAGACCTGCGGCCACCCGGAGGAGGCAGACGCAGGCGATGACCTGGTTCATCATGAGAGGAGCTAGAGTTGACTC AGTCTTAGGATCTCGATGGCGTCAGAATTGCAGAGGGTTTCTGCCAGTGCCTTAGGTCTAATTCAGAAAGCAAGGAAGGCTGGGCACGGT GGCTGGCGCCTGTAGTCTCAGCTACTTGGAAACTGAGGCAGGAGGATCTCTTGAGCCCAAGAATTCGAGACCGGTCTGGCAACATAGTGA GCCTCCATCTCTAAAAAAAAGTGGCAAAGCTGAGGAAATCTGTGCCCCCATCCCCCGACACCAGACAAGCTAAGGTGCATTCAGAGTAGG GAACGATGTCAAAACCTGGCCGCCGTGGTGGCGCGCATTCCTCTCACACCACCGTACGGAGGCAGGTGACAGTCAACTGAAATGTCCCCA AGGACCGTGTGAATTCGGCTGGCGTGGCCCAGCAGCGGGGTGGACCAGGGTGCGGTGGCTTCAGACCTGCCTCCCGCCACCCCGTCCTCC CCTAACCCCTTCTTGGGACCTCCTGCCAATCCCTGTGGTGTTTGAGTAAAGAAAGTGGCTGATGGTTTTACTTTTTTCTTCCAATATTGG AAAGAACAAACTAGATATTCCCCTTGAACACATCCAAATTATTTCCAGTGCCTGAGGGTTACCGAGCACTGTAAGCTCAGCTCGGCTGTA GCTCCGGGTTTCTGCCGATGGCCTCCATCTGCGCCGAAAGCTGAACTGAGCCCCGCGGTGCCCCGGAGCCACAGCAGCAGGTGCAGCGAC AATCGCCGTCGTGCGCAGTCGCTGGAGTCCTGGAGAGCCACGCTGTCCTCTGCAGACAACAGCATCAGTCCGCTGTGCCCCAATGCATCG ACGCCCTGTGGACGGGGCCTGCAGTCCACCCGTGGTCTGAGCTTGCCGGGCTGAAGCTCGGCTGGGACCCAGGGAGGGGCCACCCTGGGA CCCACAGTCTCCTCAGTCACGTGCAGAGCAAAAAATGCTTCCTGTGTTTGGAAGAGCAAGTGACGCAGCATTTATTGATAAAAATGGATT TTTCTGTCTGATTTCATGCTGTGGTCGTTTAAGTGGTCAATGTCTTTTTTTTTTTTTTTTTTGACATGGAGTCTCGCTCTGTTGCCCAGG CTGAGTGGAATGGCGCCAACTCAGCTCACTGCAACCTTCACCTCCCAGGCTCAAGCGACTCTCCTGCCTCAGCCTCCTGAGCAGCTGGGA TTACAGGTGCCCACCACCACGCCCAGCTAATTTTTGTATTTTTAGTAGAGACGGGGTTTCACCATGTTGGCCAGGCTGGTCTCGAACACC TGACCTTGTGATCCACCCGCCTCGGCCTCCCAATAATTTGTAAGTTATGTTAGCGGGATCCTCAAGGCCTTGCTTTGCCCCGTGGAGACG CTTGCTCGGATGAGCTCAGGAAACAGTACCGGCTGCGTGGCAGGTCTGGGTGTTGTGTGCGAGGACGTGGCCTTTGAACACCGCTGTGTT CTCAGAGGTCCTTAGGAGATATTTTTTTTTGTCTTAGGGGGACTGTGTTAAGTTCAGACAAATCATGCTGGGTGTGTAGAGAGTGTGAAA TACGTCAGTGAAGTAAGTAGCAGTGAGCGATTGTGAATGTGTAATGTAAATGGAAAACCGGGTTTTACCGTGTTAAGTTATTCACTAGGG AGCCAGTCGTAGTTCTTTGTAATCCTCTTTCTTCCAAACCTGCTTTGCTGAAAGTTGCAGAAAAGGAAGTGTGTGGAGAGAAACAGAACC CTTCAGGGTGGGTCAGAGGACGCCATCCACAGTGGATTCGTGTTCGTTTGCAGGTGGAAGCAGTGATTTTTAGGACCCACTGATTAAAAA CAAACATTCCCAAGTGTCTCTGAGAGATGCTGTTTATTTGTTAATTAAAAAGCTTTTTTCTCTGTCTTTTAAATTATGGCTTTCATGTAA TAAGGATATTTTTAGTGAAAAATTGTTTTCCTTTCAAATTACAGACCTTTTAAAAAAACTTAATTTGAGCGAGTACCTTTTCATTTGACA CTTTTCCTGTTTCTAACCTTAGGAAACCAGAATAGCGTTTGGCAGACACGACGTTTTCAGTTTACCTTTGACACCTGCCCCACTCCATTT >53061_53061_2_METTL16-FAM101B_METTL16_chr17_2344784_ENST00000538844_FAM101B_chr17_293285_ENST00000329099_length(amino acids)=154AA_BP=48 MAEGGELEFVKRIIHDSLQLKKRLRWYSCMLGKKCSLAPLKEELRIQGVHLLGQVRLREALHRRRAAAPGPGGGLLQPDGHLCPQWHVAQ -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for METTL16-FAM101B |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for METTL16-FAM101B |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for METTL16-FAM101B |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
