
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MICALL2-CA3 (FusionGDB2 ID:53544) |
Fusion Gene Summary for MICALL2-CA3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MICALL2-CA3 | Fusion gene ID: 53544 | Hgene | Tgene | Gene symbol | MICALL2 | CA3 | Gene ID | 79778 | 761 |
| Gene name | MICAL like 2 | carbonic anhydrase 3 | |
| Synonyms | JRAB|MICAL-L2 | CAIII|Car3 | |
| Cytomap | 7p22.3 | 8q21.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | MICAL-like protein 2junctional Rab13-binding proteinmolecule interacting with CasL-like 2 | carbonic anhydrase 3CA-IIIHEL-S-167mPcarbonate dehydratase IIIcarbonic anhydrase III, muscle specificcarbonic anhydrase IIIIepididymis secretory sperm binding protein Li 167mP | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q8IY33 | P07451 | |
| Ensembl transtripts involved in fusion gene | ENST00000297508, ENST00000405088, ENST00000471899, | ENST00000285381, | |
| Fusion gene scores | * DoF score | 12 X 8 X 8=768 | 7 X 7 X 8=392 |
| # samples | 16 | 13 | |
| ** MAII score | log2(16/768*10)=-2.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(13/392*10)=-1.59234203108675 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MICALL2 [Title/Abstract] AND CA3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MICALL2(1481828)-CA3(86354302), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across MICALL2 (5'-gene) Fusion gene breakpoints across MICALL2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
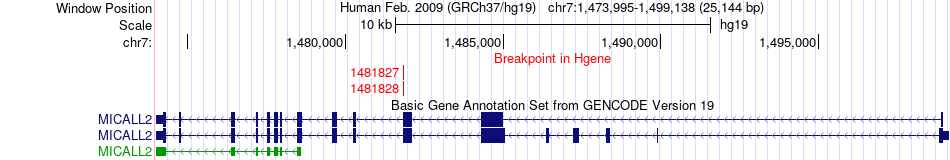 |
 Fusion gene breakpoints across CA3 (3'-gene) Fusion gene breakpoints across CA3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
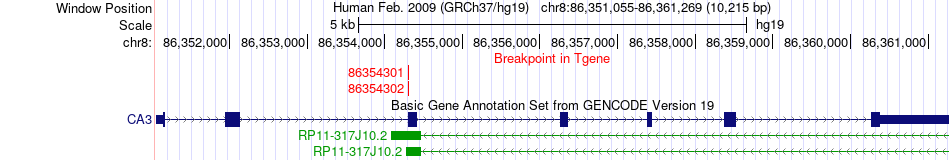 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | ESCA | TCGA-V5-A7RB-01A | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + |
| ChimerDB4 | ESCA | TCGA-V5-A7RB | MICALL2 | chr7 | 1481827 | - | CA3 | chr8 | 86354301 | + |
| ChimerDB4 | ESCA | TCGA-V5-A7RB | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + |
Top |
Fusion Gene ORF analysis for MICALL2-CA3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000297508 | ENST00000285381 | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + |
| In-frame | ENST00000297508 | ENST00000285381 | MICALL2 | chr7 | 1481827 | - | CA3 | chr8 | 86354301 | + |
| In-frame | ENST00000405088 | ENST00000285381 | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + |
| In-frame | ENST00000405088 | ENST00000285381 | MICALL2 | chr7 | 1481827 | - | CA3 | chr8 | 86354301 | + |
| intron-3CDS | ENST00000471899 | ENST00000285381 | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + |
| intron-3CDS | ENST00000471899 | ENST00000285381 | MICALL2 | chr7 | 1481827 | - | CA3 | chr8 | 86354301 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000405088 | MICALL2 | chr7 | 1481828 | - | ENST00000285381 | CA3 | chr8 | 86354302 | + | 2513 | 1075 | 0 | 1625 | 541 |
| ENST00000297508 | MICALL2 | chr7 | 1481828 | - | ENST00000285381 | CA3 | chr8 | 86354302 | + | 3325 | 1887 | 92 | 2437 | 781 |
| ENST00000405088 | MICALL2 | chr7 | 1481827 | - | ENST00000285381 | CA3 | chr8 | 86354301 | + | 2513 | 1075 | 0 | 1625 | 541 |
| ENST00000297508 | MICALL2 | chr7 | 1481827 | - | ENST00000285381 | CA3 | chr8 | 86354301 | + | 3325 | 1887 | 92 | 2437 | 781 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000405088 | ENST00000285381 | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + | 0.07791868 | 0.9220813 |
| ENST00000297508 | ENST00000285381 | MICALL2 | chr7 | 1481828 | - | CA3 | chr8 | 86354302 | + | 0.017403414 | 0.9825966 |
| ENST00000405088 | ENST00000285381 | MICALL2 | chr7 | 1481827 | - | CA3 | chr8 | 86354301 | + | 0.07791868 | 0.9220813 |
| ENST00000297508 | ENST00000285381 | MICALL2 | chr7 | 1481827 | - | CA3 | chr8 | 86354301 | + | 0.017403414 | 0.9825966 |
Top |
Fusion Genomic Features for MICALL2-CA3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
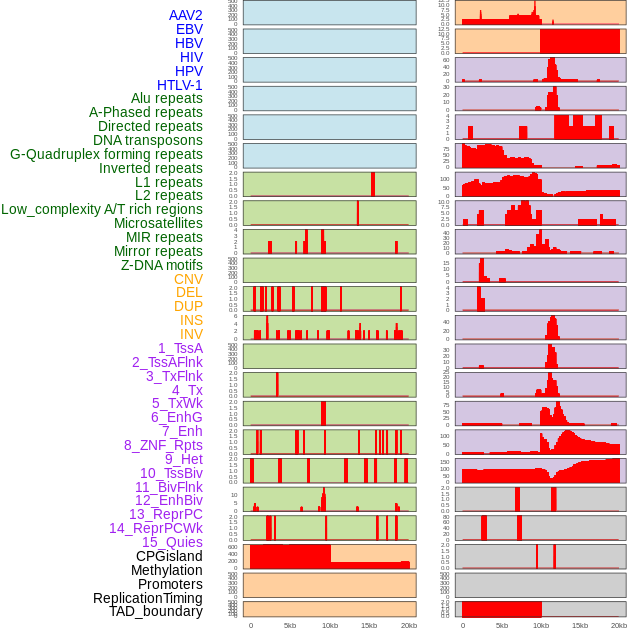 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
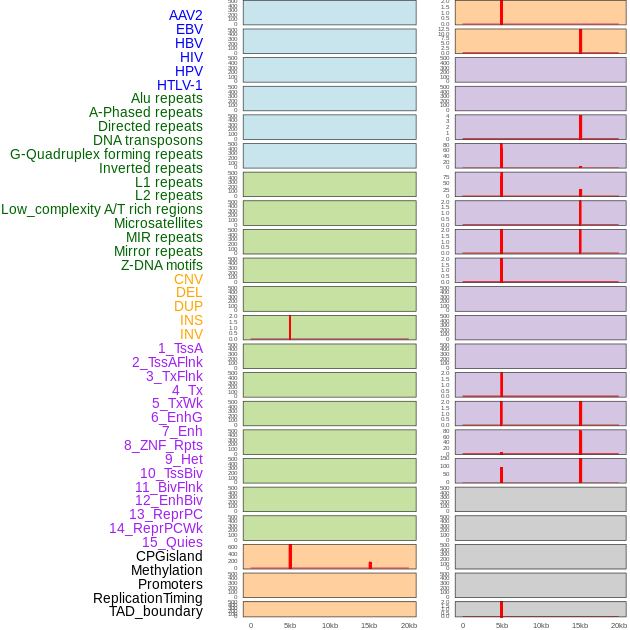 |
Top |
Fusion Protein Features for MICALL2-CA3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:1481828/chr8:86354302) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MICALL2 | CA3 |
| FUNCTION: Effector of small Rab GTPases which is involved in junctional complexes assembly through the regulation of cell adhesion molecules transport to the plasma membrane and actin cytoskeleton reorganization. Regulates the endocytic recycling of occludins, claudins and E-cadherin to the plasma membrane and may thereby regulate the establishment of tight junctions and adherens junctions. In parallel, may regulate actin cytoskeleton reorganization directly through interaction with F-actin or indirectly through actinins and filamins. Most probably involved in the processes of epithelial cell differentiation, cell spreading and neurite outgrowth (By similarity). {ECO:0000250}. | FUNCTION: Reversible hydration of carbon dioxide. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 356_359 | 570 | 905.0 | Compositional bias | Note=Poly-Ala |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 392_397 | 570 | 905.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 356_359 | 358 | 693.0 | Compositional bias | Note=Poly-Ala |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 356_359 | 570 | 905.0 | Compositional bias | Note=Poly-Ala |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 392_397 | 570 | 905.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 356_359 | 358 | 693.0 | Compositional bias | Note=Poly-Ala |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 186_248 | 570 | 905.0 | Domain | LIM zinc-binding |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 1_107 | 570 | 905.0 | Domain | Calponin-homology (CH) |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 186_248 | 358 | 693.0 | Domain | LIM zinc-binding |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 1_107 | 358 | 693.0 | Domain | Calponin-homology (CH) |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 186_248 | 570 | 905.0 | Domain | LIM zinc-binding |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 1_107 | 570 | 905.0 | Domain | Calponin-homology (CH) |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 186_248 | 358 | 693.0 | Domain | LIM zinc-binding |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 1_107 | 358 | 693.0 | Domain | Calponin-homology (CH) |
| Tgene | CA3 | chr7:1481827 | chr8:86354301 | ENST00000285381 | 1 | 7 | 198_199 | 77 | 261.0 | Region | Substrate binding | |
| Tgene | CA3 | chr7:1481828 | chr8:86354302 | ENST00000285381 | 1 | 7 | 198_199 | 77 | 261.0 | Region | Substrate binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 735_771 | 570 | 905.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 735_771 | 358 | 693.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 735_771 | 570 | 905.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 735_771 | 358 | 693.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 663_666 | 570 | 905.0 | Compositional bias | Note=Poly-Arg |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 392_397 | 358 | 693.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 663_666 | 358 | 693.0 | Compositional bias | Note=Poly-Arg |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 663_666 | 570 | 905.0 | Compositional bias | Note=Poly-Arg |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 392_397 | 358 | 693.0 | Compositional bias | Note=Poly-Ser |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 663_666 | 358 | 693.0 | Compositional bias | Note=Poly-Arg |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 723_874 | 570 | 905.0 | Domain | bMERB |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 723_874 | 358 | 693.0 | Domain | bMERB |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 723_874 | 570 | 905.0 | Domain | bMERB |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 723_874 | 358 | 693.0 | Domain | bMERB |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000297508 | - | 7 | 17 | 261_697 | 570 | 905.0 | Region | Mediates targeting to the cell plasma membrane |
| Hgene | MICALL2 | chr7:1481827 | chr8:86354301 | ENST00000405088 | - | 3 | 13 | 261_697 | 358 | 693.0 | Region | Mediates targeting to the cell plasma membrane |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000297508 | - | 7 | 17 | 261_697 | 570 | 905.0 | Region | Mediates targeting to the cell plasma membrane |
| Hgene | MICALL2 | chr7:1481828 | chr8:86354302 | ENST00000405088 | - | 3 | 13 | 261_697 | 358 | 693.0 | Region | Mediates targeting to the cell plasma membrane |
| Tgene | CA3 | chr7:1481827 | chr8:86354301 | ENST00000285381 | 1 | 7 | 3_259 | 77 | 261.0 | Domain | Alpha-carbonic anhydrase | |
| Tgene | CA3 | chr7:1481828 | chr8:86354302 | ENST00000285381 | 1 | 7 | 3_259 | 77 | 261.0 | Domain | Alpha-carbonic anhydrase | |
| Tgene | CA3 | chr7:1481827 | chr8:86354301 | ENST00000285381 | 1 | 7 | 64_67 | 77 | 261.0 | Region | Note=Involved in proton transfer | |
| Tgene | CA3 | chr7:1481828 | chr8:86354302 | ENST00000285381 | 1 | 7 | 64_67 | 77 | 261.0 | Region | Note=Involved in proton transfer |
Top |
Fusion Gene Sequence for MICALL2-CA3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >53544_53544_1_MICALL2-CA3_MICALL2_chr7_1481827_ENST00000297508_CA3_chr8_86354301_ENST00000285381_length(transcript)=3325nt_BP=1887nt CTTCTCTTAGGGACCCGGCACTCGTCGCCCTCAGTCCGGCTCAGCTGGGGCTGGGCGCCGTGGGTCTGGCGGTTCCGTAGCGGTCCCAGC GTCTGTCCCGCCGGCCGGGCGGTCGCGGCACAGCCGCGGGAAGGTGTCGGAGGGCGGTTCCGCCGCGCGGCGGGCGCGCCGCCCACATGG CGGCCATCAGGGCGCTGCAACAGTGGTGCCGGCAGCAGTGCGAGGGCTACCGCGACGTGAATATCTGCAACATGACCACGTCGTTCCGCG ACGGCCTGGCTTTCTGCGCCATCCTGCACCGCCACCGGCCCGACCTCATAAACTTCAGTGCTCTCAAGAAGGAAAATATTTATGAAAACA ATAAACTGGCCTTCCGCGTGGCCGAGGAGCACTTGGGCATCCCAGCCTTGCTGGATGCCGAGGACATGGTGGCCTTGAAGGTGCCTGACC GGCTGAGCATCTTGACCTACGTGTCCCAGTATTACAACTACTTCCACGGCCGCTCCCCCATTGGGGGCATGGCAGGCGTGAAGAGGGCCT CGGAGGACTCTGAGGAGGAGCCGTCAGGGAAGAAGGCTCCAGTCCAGGCGGCCAAGCTGCCCTCGCCCGCCCCAGCCCGGAAGCCTCCAC TATCTCCAGCCCAGACAAACCCTGTGGTCCAGAGGAGGAATGAGGGTGCAGGGGGCCCGCCCCCCAAGACTGACCAGGCATTGGCGGGCA GCTTGGTCAGCAGCACCTGCGGGGTCTGCGGCAAGCACGTGCACCTGGTACAGCGGCACCTGGCCGACGGGAGGCTTTACCACCGGAGCT GCTTCAGGTGTAAGCAGTGCTCCTGCACGCTGCACTCGGGGGCCTACAAGGCCACAGGAGAGCCGGGCACCTTCGTCTGCACCAGCCACC TCCCCGCAGCCGCCTCTGCAAGCCCCAAGTTGACGGGTCTGGTCCCCCGACAGCCAGGGGCCATGGGTGTGGATTCCAGGACCTCCTGTT CCCCACAGAAGGCCCAGGAGGCAAACAAGGCCAGACCGTCGGCCTGGGAGCCTGCTGCGGGCAACTCGCCTGCCAGGGCTTCCGTTCCAG CTGCACCCAACCCTGCAGCCACCAGCGCCACGTCCGTCCACGTGAGGAGCCCAGCCAGGCCCTCTGAGAGCCGCCTGGCCCCCACTCCCA CGGAGGGGAAAGTCCGCCCTCGTGTGACCAATAGCTCCCCGATGGGCTGGTCGTCAGCTGCCCCGTGCACAGCAGCGGCTGCCTCCCATC CCGCCGTGCCCCCGAGTGCCCCAGACCCTCGCCCGGCCACACCCCAGGGTGGGGGAGCCCCCCGAGTGGCAGCTCCTCAAACCACACTCA GTTCAAGCTCCACATCTGCAGCCACGGTGGACCCCCCAGCCTGGACCCCGTCCGCCTCCAGGACCCAGCAGGCCCGGAATAAGTTTTTCC AAACATCAGCAGTGCCCCCCGGCACCAGCCTTTCTGGCAGAGGTCCCACCCCGTCACTTGTTCTATCCAAGGACAGCAGCAAGGAGCAGG CGCGGAACTTCCTCAAGCAGGCCCTCTCAGCGCTGGAAGAGGCTGGCGCTCCGGCGCCTGGCAGGCCCTCCCCAGCCACTGCCGCTGTTC CCAGTTCTCAGCCCAAAACTGAAGCACCACAAGCAAGTCCCTTAGCCAAGCCGTTACAGTCCTCGTCTCCCCGGGTGCTTGGCCTCCCTT CGAGGATGGAACCGCCAGCCCCGCTGAGCACGAGCAGTACCTCTCAGGCATCCGCGTTGCCCCCGGCAGGCAGGAGGAACTTGGCGGAAT CCTCAGGGGTCGGCAGGGTGGGTGCTGGCTCCAGGCCGAAGCCAGAGGCCCCGATGGCAAAGGGTAAAAGCACCACCTTAACGCAGGTGC TGAGAGGGGGTCCTCTCCCTGGACCCTACCGACTTCGCCAGTTTCATCTTCACTGGGGCTCTTCGGATGATCATGGCTCTGAGCACACCG TGGATGGAGTCAAGTATGCAGCGGAGCTTCATTTGGTTCACTGGAACCCGAAGTATAACACTTTTAAAGAAGCCCTGAAGCAGCGCGATG GGATCGCTGTGATTGGCATTTTTCTGAAGATAGGACATGAGAATGGCGAGTTCCAGATTTTCCTTGATGCATTGGACAAGATTAAGACAA AGGGCAAGGAGGCGCCCTTCACAAAGTTTGACCCATCCTGCCTGTTCCCGGCATGCCGGGACTACTGGACCTACCAGGGCTCATTCACCA CGCCGCCCTGCGAGGAATGCATTGTGTGGCTGCTGCTGAAGGAGCCCATGACCGTGAGCTCTGACCAGATGGCCAAGCTGCGGAGCCTCC TCTCCAGTGCTGAGAACGAGCCCCCAGTGCCTCTTGTGAGCAACTGGCGACCTCCACAGCCTATCAATAACAGGGTGGTGAGAGCTTCCT TCAAATGAGGCTGCTGGATCTTGCCCTCTTCAGGAAAGGAAACCTACCATTGGAGAGCTTGGTTCCTTGCCTCCTTCTGGTGCTCCTTAC TCCAAGTCTATTTCATTTTTCCACACTGAGCAATGAATGTGAGAGATGTGGTCACCAAGATCTAAGTTACTTGTTGAAAGAAAGTTACTT TCGACAAGATCTAATATGAAAGCATAGATTTCACATTTGATCTCTGTAATAATCATCTTTCCTATAAAAGTAGCATTTTTGGTAAAGTTT CAAAGAAGAAGAAACAGAGATGGAAGAGTAAAGATATTTTTAAAATGGCTAGCTATTGGGCACCAGTTTTTCTGTTATCTAAAATTTCAC ACAACTTCATTGTTTTTATTTTTATATTATGAGTTGTCCATCTTAAAGAAATATGAGTAATTCTACATGTAGTAGAGGTGTATGAAGATC ATATAACAATTAAACATAAGCCAGAAATTAAAATGACTATAGACAGCAAGAATTGAGCTAATAATATGTTTTAACTCTTAACACCAGCAA GAAGTCAGTCATTTATTGAAGTTTTAGCTACTAAGATTACTTGGTTTTGATTACCAGTGAAAAGAAAACACAATACAATCAGGAGTTTTC AAATTTTTGATTCAGTATTTGAATTTCTTCTTCATAAATGTAGTTGAATTTATCCTAGTATTTTTCTTTACCTGAAGGAGGGCCATTTAT TTTTAATTTCACTACATTTTTCTTTGCATGATTATTAAAATAAAAACTGCCTCTGTTGTGTTTCTCACTGGAGGCTGGAATGAATGATCA >53544_53544_1_MICALL2-CA3_MICALL2_chr7_1481827_ENST00000297508_CA3_chr8_86354301_ENST00000285381_length(amino acids)=781AA_BP=598 MSRRPGGRGTAAGRCRRAVPPRGGRAAHMAAIRALQQWCRQQCEGYRDVNICNMTTSFRDGLAFCAILHRHRPDLINFSALKKENIYENN KLAFRVAEEHLGIPALLDAEDMVALKVPDRLSILTYVSQYYNYFHGRSPIGGMAGVKRASEDSEEEPSGKKAPVQAAKLPSPAPARKPPL SPAQTNPVVQRRNEGAGGPPPKTDQALAGSLVSSTCGVCGKHVHLVQRHLADGRLYHRSCFRCKQCSCTLHSGAYKATGEPGTFVCTSHL PAAASASPKLTGLVPRQPGAMGVDSRTSCSPQKAQEANKARPSAWEPAAGNSPARASVPAAPNPAATSATSVHVRSPARPSESRLAPTPT EGKVRPRVTNSSPMGWSSAAPCTAAAASHPAVPPSAPDPRPATPQGGGAPRVAAPQTTLSSSSTSAATVDPPAWTPSASRTQQARNKFFQ TSAVPPGTSLSGRGPTPSLVLSKDSSKEQARNFLKQALSALEEAGAPAPGRPSPATAAVPSSQPKTEAPQASPLAKPLQSSSPRVLGLPS RMEPPAPLSTSSTSQASALPPAGRRNLAESSGVGRVGAGSRPKPEAPMAKGKSTTLTQVLRGGPLPGPYRLRQFHLHWGSSDDHGSEHTV DGVKYAAELHLVHWNPKYNTFKEALKQRDGIAVIGIFLKIGHENGEFQIFLDALDKIKTKGKEAPFTKFDPSCLFPACRDYWTYQGSFTT -------------------------------------------------------------- >53544_53544_2_MICALL2-CA3_MICALL2_chr7_1481827_ENST00000405088_CA3_chr8_86354301_ENST00000285381_length(transcript)=2513nt_BP=1075nt ATGGCGGCCATCAGGGCGCTGCAACAGTGGTGCCGGCAGCAGTGCGAGGGCTACCGCGACGTGAATATCTGCAACATGACCACCCACCTC CCCGCAGCCGCCTCTGCAAGCCCCAAGTTGACGGGTCTGGTCCCCCGACAGCCAGGGGCCATGGGTGTGGATTCCAGGACCTCCTGTTCC CCACAGAAGGCCCAGGAGGCAAACAAGGCCAGACCGTCGGCCTGGGAGCCTGCTGCGGGCAACTCGCCTGCCAGGGCTTCCGTTCCAGCT GCACCCAACCCTGCAGCCACCAGCGCCACGTCCGTCCACGTGAGGAGCCCAGCCAGGCCCTCTGAGAGCCGCCTGGCCCCCACTCCCACG GAGGGGAAAGTCCGCCCTCGTGTGACCAATAGCTCCCCGATGGGCTGGTCGTCAGCTGCCCCGTGCACAGCAGCGGCTGCCTCCCATCCC GCCGTGCCCCCGAGTGCCCCAGACCCTCGCCCGGCCACACCCCAGGGTGGGGGAGCCCCCCGAGTGGCAGCTCCTCAAACCACACTCAGT TCAAGCTCCACATCTGCAGCCACGGTGGACCCCCCAGCCTGGACCCCGTCCGCCTCCAGGACCCAGCAGGCCCGGAATAAGTTTTTCCAA ACATCAGCAGTGCCCCCCGGCACCAGCCTTTCTGGCAGAGGTCCCACCCCGTCACTTGTTCTATCCAAGGACAGCAGCAAGGAGCAGGCG CGGAACTTCCTCAAGCAGGCCCTCTCAGCGCTGGAAGAGGCTGGCGCTCCGGCGCCTGGCAGGCCCTCCCCAGCCACTGCCGCTGTTCCC AGTTCTCAGCCCAAAACTGAAGCACCACAAGCAAGTCCCTTAGCCAAGCCGTTACAGTCCTCGTCTCCCCGGGTGCTTGGCCTCCCTTCG AGGATGGAACCGCCAGCCCCGCTGAGCACGAGCAGTACCTCTCAGGCATCCGCGTTGCCCCCGGCAGGCAGGAGGAACTTGGCGGAATCC TCAGGGGTCGGCAGGGTGGGTGCTGGCTCCAGGCCGAAGCCAGAGGCCCCGATGGCAAAGGGTAAAAGCACCACCTTAACGCAGGTGCTG AGAGGGGGTCCTCTCCCTGGACCCTACCGACTTCGCCAGTTTCATCTTCACTGGGGCTCTTCGGATGATCATGGCTCTGAGCACACCGTG GATGGAGTCAAGTATGCAGCGGAGCTTCATTTGGTTCACTGGAACCCGAAGTATAACACTTTTAAAGAAGCCCTGAAGCAGCGCGATGGG ATCGCTGTGATTGGCATTTTTCTGAAGATAGGACATGAGAATGGCGAGTTCCAGATTTTCCTTGATGCATTGGACAAGATTAAGACAAAG GGCAAGGAGGCGCCCTTCACAAAGTTTGACCCATCCTGCCTGTTCCCGGCATGCCGGGACTACTGGACCTACCAGGGCTCATTCACCACG CCGCCCTGCGAGGAATGCATTGTGTGGCTGCTGCTGAAGGAGCCCATGACCGTGAGCTCTGACCAGATGGCCAAGCTGCGGAGCCTCCTC TCCAGTGCTGAGAACGAGCCCCCAGTGCCTCTTGTGAGCAACTGGCGACCTCCACAGCCTATCAATAACAGGGTGGTGAGAGCTTCCTTC AAATGAGGCTGCTGGATCTTGCCCTCTTCAGGAAAGGAAACCTACCATTGGAGAGCTTGGTTCCTTGCCTCCTTCTGGTGCTCCTTACTC CAAGTCTATTTCATTTTTCCACACTGAGCAATGAATGTGAGAGATGTGGTCACCAAGATCTAAGTTACTTGTTGAAAGAAAGTTACTTTC GACAAGATCTAATATGAAAGCATAGATTTCACATTTGATCTCTGTAATAATCATCTTTCCTATAAAAGTAGCATTTTTGGTAAAGTTTCA AAGAAGAAGAAACAGAGATGGAAGAGTAAAGATATTTTTAAAATGGCTAGCTATTGGGCACCAGTTTTTCTGTTATCTAAAATTTCACAC AACTTCATTGTTTTTATTTTTATATTATGAGTTGTCCATCTTAAAGAAATATGAGTAATTCTACATGTAGTAGAGGTGTATGAAGATCAT ATAACAATTAAACATAAGCCAGAAATTAAAATGACTATAGACAGCAAGAATTGAGCTAATAATATGTTTTAACTCTTAACACCAGCAAGA AGTCAGTCATTTATTGAAGTTTTAGCTACTAAGATTACTTGGTTTTGATTACCAGTGAAAAGAAAACACAATACAATCAGGAGTTTTCAA ATTTTTGATTCAGTATTTGAATTTCTTCTTCATAAATGTAGTTGAATTTATCCTAGTATTTTTCTTTACCTGAAGGAGGGCCATTTATTT TTAATTTCACTACATTTTTCTTTGCATGATTATTAAAATAAAAACTGCCTCTGTTGTGTTTCTCACTGGAGGCTGGAATGAATGATCACT >53544_53544_2_MICALL2-CA3_MICALL2_chr7_1481827_ENST00000405088_CA3_chr8_86354301_ENST00000285381_length(amino acids)=541AA_BP=358 MAAIRALQQWCRQQCEGYRDVNICNMTTHLPAAASASPKLTGLVPRQPGAMGVDSRTSCSPQKAQEANKARPSAWEPAAGNSPARASVPA APNPAATSATSVHVRSPARPSESRLAPTPTEGKVRPRVTNSSPMGWSSAAPCTAAAASHPAVPPSAPDPRPATPQGGGAPRVAAPQTTLS SSSTSAATVDPPAWTPSASRTQQARNKFFQTSAVPPGTSLSGRGPTPSLVLSKDSSKEQARNFLKQALSALEEAGAPAPGRPSPATAAVP SSQPKTEAPQASPLAKPLQSSSPRVLGLPSRMEPPAPLSTSSTSQASALPPAGRRNLAESSGVGRVGAGSRPKPEAPMAKGKSTTLTQVL RGGPLPGPYRLRQFHLHWGSSDDHGSEHTVDGVKYAAELHLVHWNPKYNTFKEALKQRDGIAVIGIFLKIGHENGEFQIFLDALDKIKTK GKEAPFTKFDPSCLFPACRDYWTYQGSFTTPPCEECIVWLLLKEPMTVSSDQMAKLRSLLSSAENEPPVPLVSNWRPPQPINNRVVRASF -------------------------------------------------------------- >53544_53544_3_MICALL2-CA3_MICALL2_chr7_1481828_ENST00000297508_CA3_chr8_86354302_ENST00000285381_length(transcript)=3325nt_BP=1887nt CTTCTCTTAGGGACCCGGCACTCGTCGCCCTCAGTCCGGCTCAGCTGGGGCTGGGCGCCGTGGGTCTGGCGGTTCCGTAGCGGTCCCAGC GTCTGTCCCGCCGGCCGGGCGGTCGCGGCACAGCCGCGGGAAGGTGTCGGAGGGCGGTTCCGCCGCGCGGCGGGCGCGCCGCCCACATGG CGGCCATCAGGGCGCTGCAACAGTGGTGCCGGCAGCAGTGCGAGGGCTACCGCGACGTGAATATCTGCAACATGACCACGTCGTTCCGCG ACGGCCTGGCTTTCTGCGCCATCCTGCACCGCCACCGGCCCGACCTCATAAACTTCAGTGCTCTCAAGAAGGAAAATATTTATGAAAACA ATAAACTGGCCTTCCGCGTGGCCGAGGAGCACTTGGGCATCCCAGCCTTGCTGGATGCCGAGGACATGGTGGCCTTGAAGGTGCCTGACC GGCTGAGCATCTTGACCTACGTGTCCCAGTATTACAACTACTTCCACGGCCGCTCCCCCATTGGGGGCATGGCAGGCGTGAAGAGGGCCT CGGAGGACTCTGAGGAGGAGCCGTCAGGGAAGAAGGCTCCAGTCCAGGCGGCCAAGCTGCCCTCGCCCGCCCCAGCCCGGAAGCCTCCAC TATCTCCAGCCCAGACAAACCCTGTGGTCCAGAGGAGGAATGAGGGTGCAGGGGGCCCGCCCCCCAAGACTGACCAGGCATTGGCGGGCA GCTTGGTCAGCAGCACCTGCGGGGTCTGCGGCAAGCACGTGCACCTGGTACAGCGGCACCTGGCCGACGGGAGGCTTTACCACCGGAGCT GCTTCAGGTGTAAGCAGTGCTCCTGCACGCTGCACTCGGGGGCCTACAAGGCCACAGGAGAGCCGGGCACCTTCGTCTGCACCAGCCACC TCCCCGCAGCCGCCTCTGCAAGCCCCAAGTTGACGGGTCTGGTCCCCCGACAGCCAGGGGCCATGGGTGTGGATTCCAGGACCTCCTGTT CCCCACAGAAGGCCCAGGAGGCAAACAAGGCCAGACCGTCGGCCTGGGAGCCTGCTGCGGGCAACTCGCCTGCCAGGGCTTCCGTTCCAG CTGCACCCAACCCTGCAGCCACCAGCGCCACGTCCGTCCACGTGAGGAGCCCAGCCAGGCCCTCTGAGAGCCGCCTGGCCCCCACTCCCA CGGAGGGGAAAGTCCGCCCTCGTGTGACCAATAGCTCCCCGATGGGCTGGTCGTCAGCTGCCCCGTGCACAGCAGCGGCTGCCTCCCATC CCGCCGTGCCCCCGAGTGCCCCAGACCCTCGCCCGGCCACACCCCAGGGTGGGGGAGCCCCCCGAGTGGCAGCTCCTCAAACCACACTCA GTTCAAGCTCCACATCTGCAGCCACGGTGGACCCCCCAGCCTGGACCCCGTCCGCCTCCAGGACCCAGCAGGCCCGGAATAAGTTTTTCC AAACATCAGCAGTGCCCCCCGGCACCAGCCTTTCTGGCAGAGGTCCCACCCCGTCACTTGTTCTATCCAAGGACAGCAGCAAGGAGCAGG CGCGGAACTTCCTCAAGCAGGCCCTCTCAGCGCTGGAAGAGGCTGGCGCTCCGGCGCCTGGCAGGCCCTCCCCAGCCACTGCCGCTGTTC CCAGTTCTCAGCCCAAAACTGAAGCACCACAAGCAAGTCCCTTAGCCAAGCCGTTACAGTCCTCGTCTCCCCGGGTGCTTGGCCTCCCTT CGAGGATGGAACCGCCAGCCCCGCTGAGCACGAGCAGTACCTCTCAGGCATCCGCGTTGCCCCCGGCAGGCAGGAGGAACTTGGCGGAAT CCTCAGGGGTCGGCAGGGTGGGTGCTGGCTCCAGGCCGAAGCCAGAGGCCCCGATGGCAAAGGGTAAAAGCACCACCTTAACGCAGGTGC TGAGAGGGGGTCCTCTCCCTGGACCCTACCGACTTCGCCAGTTTCATCTTCACTGGGGCTCTTCGGATGATCATGGCTCTGAGCACACCG TGGATGGAGTCAAGTATGCAGCGGAGCTTCATTTGGTTCACTGGAACCCGAAGTATAACACTTTTAAAGAAGCCCTGAAGCAGCGCGATG GGATCGCTGTGATTGGCATTTTTCTGAAGATAGGACATGAGAATGGCGAGTTCCAGATTTTCCTTGATGCATTGGACAAGATTAAGACAA AGGGCAAGGAGGCGCCCTTCACAAAGTTTGACCCATCCTGCCTGTTCCCGGCATGCCGGGACTACTGGACCTACCAGGGCTCATTCACCA CGCCGCCCTGCGAGGAATGCATTGTGTGGCTGCTGCTGAAGGAGCCCATGACCGTGAGCTCTGACCAGATGGCCAAGCTGCGGAGCCTCC TCTCCAGTGCTGAGAACGAGCCCCCAGTGCCTCTTGTGAGCAACTGGCGACCTCCACAGCCTATCAATAACAGGGTGGTGAGAGCTTCCT TCAAATGAGGCTGCTGGATCTTGCCCTCTTCAGGAAAGGAAACCTACCATTGGAGAGCTTGGTTCCTTGCCTCCTTCTGGTGCTCCTTAC TCCAAGTCTATTTCATTTTTCCACACTGAGCAATGAATGTGAGAGATGTGGTCACCAAGATCTAAGTTACTTGTTGAAAGAAAGTTACTT TCGACAAGATCTAATATGAAAGCATAGATTTCACATTTGATCTCTGTAATAATCATCTTTCCTATAAAAGTAGCATTTTTGGTAAAGTTT CAAAGAAGAAGAAACAGAGATGGAAGAGTAAAGATATTTTTAAAATGGCTAGCTATTGGGCACCAGTTTTTCTGTTATCTAAAATTTCAC ACAACTTCATTGTTTTTATTTTTATATTATGAGTTGTCCATCTTAAAGAAATATGAGTAATTCTACATGTAGTAGAGGTGTATGAAGATC ATATAACAATTAAACATAAGCCAGAAATTAAAATGACTATAGACAGCAAGAATTGAGCTAATAATATGTTTTAACTCTTAACACCAGCAA GAAGTCAGTCATTTATTGAAGTTTTAGCTACTAAGATTACTTGGTTTTGATTACCAGTGAAAAGAAAACACAATACAATCAGGAGTTTTC AAATTTTTGATTCAGTATTTGAATTTCTTCTTCATAAATGTAGTTGAATTTATCCTAGTATTTTTCTTTACCTGAAGGAGGGCCATTTAT TTTTAATTTCACTACATTTTTCTTTGCATGATTATTAAAATAAAAACTGCCTCTGTTGTGTTTCTCACTGGAGGCTGGAATGAATGATCA >53544_53544_3_MICALL2-CA3_MICALL2_chr7_1481828_ENST00000297508_CA3_chr8_86354302_ENST00000285381_length(amino acids)=781AA_BP=598 MSRRPGGRGTAAGRCRRAVPPRGGRAAHMAAIRALQQWCRQQCEGYRDVNICNMTTSFRDGLAFCAILHRHRPDLINFSALKKENIYENN KLAFRVAEEHLGIPALLDAEDMVALKVPDRLSILTYVSQYYNYFHGRSPIGGMAGVKRASEDSEEEPSGKKAPVQAAKLPSPAPARKPPL SPAQTNPVVQRRNEGAGGPPPKTDQALAGSLVSSTCGVCGKHVHLVQRHLADGRLYHRSCFRCKQCSCTLHSGAYKATGEPGTFVCTSHL PAAASASPKLTGLVPRQPGAMGVDSRTSCSPQKAQEANKARPSAWEPAAGNSPARASVPAAPNPAATSATSVHVRSPARPSESRLAPTPT EGKVRPRVTNSSPMGWSSAAPCTAAAASHPAVPPSAPDPRPATPQGGGAPRVAAPQTTLSSSSTSAATVDPPAWTPSASRTQQARNKFFQ TSAVPPGTSLSGRGPTPSLVLSKDSSKEQARNFLKQALSALEEAGAPAPGRPSPATAAVPSSQPKTEAPQASPLAKPLQSSSPRVLGLPS RMEPPAPLSTSSTSQASALPPAGRRNLAESSGVGRVGAGSRPKPEAPMAKGKSTTLTQVLRGGPLPGPYRLRQFHLHWGSSDDHGSEHTV DGVKYAAELHLVHWNPKYNTFKEALKQRDGIAVIGIFLKIGHENGEFQIFLDALDKIKTKGKEAPFTKFDPSCLFPACRDYWTYQGSFTT -------------------------------------------------------------- >53544_53544_4_MICALL2-CA3_MICALL2_chr7_1481828_ENST00000405088_CA3_chr8_86354302_ENST00000285381_length(transcript)=2513nt_BP=1075nt ATGGCGGCCATCAGGGCGCTGCAACAGTGGTGCCGGCAGCAGTGCGAGGGCTACCGCGACGTGAATATCTGCAACATGACCACCCACCTC CCCGCAGCCGCCTCTGCAAGCCCCAAGTTGACGGGTCTGGTCCCCCGACAGCCAGGGGCCATGGGTGTGGATTCCAGGACCTCCTGTTCC CCACAGAAGGCCCAGGAGGCAAACAAGGCCAGACCGTCGGCCTGGGAGCCTGCTGCGGGCAACTCGCCTGCCAGGGCTTCCGTTCCAGCT GCACCCAACCCTGCAGCCACCAGCGCCACGTCCGTCCACGTGAGGAGCCCAGCCAGGCCCTCTGAGAGCCGCCTGGCCCCCACTCCCACG GAGGGGAAAGTCCGCCCTCGTGTGACCAATAGCTCCCCGATGGGCTGGTCGTCAGCTGCCCCGTGCACAGCAGCGGCTGCCTCCCATCCC GCCGTGCCCCCGAGTGCCCCAGACCCTCGCCCGGCCACACCCCAGGGTGGGGGAGCCCCCCGAGTGGCAGCTCCTCAAACCACACTCAGT TCAAGCTCCACATCTGCAGCCACGGTGGACCCCCCAGCCTGGACCCCGTCCGCCTCCAGGACCCAGCAGGCCCGGAATAAGTTTTTCCAA ACATCAGCAGTGCCCCCCGGCACCAGCCTTTCTGGCAGAGGTCCCACCCCGTCACTTGTTCTATCCAAGGACAGCAGCAAGGAGCAGGCG CGGAACTTCCTCAAGCAGGCCCTCTCAGCGCTGGAAGAGGCTGGCGCTCCGGCGCCTGGCAGGCCCTCCCCAGCCACTGCCGCTGTTCCC AGTTCTCAGCCCAAAACTGAAGCACCACAAGCAAGTCCCTTAGCCAAGCCGTTACAGTCCTCGTCTCCCCGGGTGCTTGGCCTCCCTTCG AGGATGGAACCGCCAGCCCCGCTGAGCACGAGCAGTACCTCTCAGGCATCCGCGTTGCCCCCGGCAGGCAGGAGGAACTTGGCGGAATCC TCAGGGGTCGGCAGGGTGGGTGCTGGCTCCAGGCCGAAGCCAGAGGCCCCGATGGCAAAGGGTAAAAGCACCACCTTAACGCAGGTGCTG AGAGGGGGTCCTCTCCCTGGACCCTACCGACTTCGCCAGTTTCATCTTCACTGGGGCTCTTCGGATGATCATGGCTCTGAGCACACCGTG GATGGAGTCAAGTATGCAGCGGAGCTTCATTTGGTTCACTGGAACCCGAAGTATAACACTTTTAAAGAAGCCCTGAAGCAGCGCGATGGG ATCGCTGTGATTGGCATTTTTCTGAAGATAGGACATGAGAATGGCGAGTTCCAGATTTTCCTTGATGCATTGGACAAGATTAAGACAAAG GGCAAGGAGGCGCCCTTCACAAAGTTTGACCCATCCTGCCTGTTCCCGGCATGCCGGGACTACTGGACCTACCAGGGCTCATTCACCACG CCGCCCTGCGAGGAATGCATTGTGTGGCTGCTGCTGAAGGAGCCCATGACCGTGAGCTCTGACCAGATGGCCAAGCTGCGGAGCCTCCTC TCCAGTGCTGAGAACGAGCCCCCAGTGCCTCTTGTGAGCAACTGGCGACCTCCACAGCCTATCAATAACAGGGTGGTGAGAGCTTCCTTC AAATGAGGCTGCTGGATCTTGCCCTCTTCAGGAAAGGAAACCTACCATTGGAGAGCTTGGTTCCTTGCCTCCTTCTGGTGCTCCTTACTC CAAGTCTATTTCATTTTTCCACACTGAGCAATGAATGTGAGAGATGTGGTCACCAAGATCTAAGTTACTTGTTGAAAGAAAGTTACTTTC GACAAGATCTAATATGAAAGCATAGATTTCACATTTGATCTCTGTAATAATCATCTTTCCTATAAAAGTAGCATTTTTGGTAAAGTTTCA AAGAAGAAGAAACAGAGATGGAAGAGTAAAGATATTTTTAAAATGGCTAGCTATTGGGCACCAGTTTTTCTGTTATCTAAAATTTCACAC AACTTCATTGTTTTTATTTTTATATTATGAGTTGTCCATCTTAAAGAAATATGAGTAATTCTACATGTAGTAGAGGTGTATGAAGATCAT ATAACAATTAAACATAAGCCAGAAATTAAAATGACTATAGACAGCAAGAATTGAGCTAATAATATGTTTTAACTCTTAACACCAGCAAGA AGTCAGTCATTTATTGAAGTTTTAGCTACTAAGATTACTTGGTTTTGATTACCAGTGAAAAGAAAACACAATACAATCAGGAGTTTTCAA ATTTTTGATTCAGTATTTGAATTTCTTCTTCATAAATGTAGTTGAATTTATCCTAGTATTTTTCTTTACCTGAAGGAGGGCCATTTATTT TTAATTTCACTACATTTTTCTTTGCATGATTATTAAAATAAAAACTGCCTCTGTTGTGTTTCTCACTGGAGGCTGGAATGAATGATCACT >53544_53544_4_MICALL2-CA3_MICALL2_chr7_1481828_ENST00000405088_CA3_chr8_86354302_ENST00000285381_length(amino acids)=541AA_BP=358 MAAIRALQQWCRQQCEGYRDVNICNMTTHLPAAASASPKLTGLVPRQPGAMGVDSRTSCSPQKAQEANKARPSAWEPAAGNSPARASVPA APNPAATSATSVHVRSPARPSESRLAPTPTEGKVRPRVTNSSPMGWSSAAPCTAAAASHPAVPPSAPDPRPATPQGGGAPRVAAPQTTLS SSSTSAATVDPPAWTPSASRTQQARNKFFQTSAVPPGTSLSGRGPTPSLVLSKDSSKEQARNFLKQALSALEEAGAPAPGRPSPATAAVP SSQPKTEAPQASPLAKPLQSSSPRVLGLPSRMEPPAPLSTSSTSQASALPPAGRRNLAESSGVGRVGAGSRPKPEAPMAKGKSTTLTQVL RGGPLPGPYRLRQFHLHWGSSDDHGSEHTVDGVKYAAELHLVHWNPKYNTFKEALKQRDGIAVIGIFLKIGHENGEFQIFLDALDKIKTK GKEAPFTKFDPSCLFPACRDYWTYQGSFTTPPCEECIVWLLLKEPMTVSSDQMAKLRSLLSSAENEPPVPLVSNWRPPQPINNRVVRASF -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MICALL2-CA3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MICALL2-CA3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MICALL2-CA3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
