
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MKNK2-RNF126 (FusionGDB2 ID:54107) |
Fusion Gene Summary for MKNK2-RNF126 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MKNK2-RNF126 | Fusion gene ID: 54107 | Hgene | Tgene | Gene symbol | MKNK2 | RNF126 | Gene ID | 2872 | 55658 |
| Gene name | MAPK interacting serine/threonine kinase 2 | ring finger protein 126 | |
| Synonyms | GPRK7|MNK2 | - | |
| Cytomap | 19p13.3 | 19p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | MAP kinase-interacting serine/threonine-protein kinase 2G protein-coupled receptor kinase 7MAP kinase interacting serine/threonine kinase 2MAP kinase signal-integrating kinase 2MAPK signal-integrating kinase 2 | E3 ubiquitin-protein ligase RNF126 | |
| Modification date | 20200313 | 20200320 | |
| UniProtAcc | Q9HBH9 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000250896, ENST00000309340, ENST00000541165, ENST00000588014, ENST00000591142, ENST00000591588, ENST00000591601, | ENST00000292363, | |
| Fusion gene scores | * DoF score | 12 X 12 X 6=864 | 8 X 3 X 6=144 |
| # samples | 13 | 9 | |
| ** MAII score | log2(13/864*10)=-2.73251968913501 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/144*10)=-0.678071905112638 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MKNK2 [Title/Abstract] AND RNF126 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MKNK2(2041839)-RNF126(652884), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | MKNK2-RNF126 seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. MKNK2-RNF126 seems lost the major protein functional domain in Hgene partner, which is a kinase due to the frame-shifted ORF. MKNK2-RNF126 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MKNK2 | GO:0006468 | protein phosphorylation | 11463832 |
| Hgene | MKNK2 | GO:0030097 | hemopoiesis | 21149447 |
| Hgene | MKNK2 | GO:0035556 | intracellular signal transduction | 11463832 |
| Hgene | MKNK2 | GO:0071243 | cellular response to arsenic-containing substance | 18299328 |
| Tgene | RNF126 | GO:0006511 | ubiquitin-dependent protein catabolic process | 24981174 |
| Tgene | RNF126 | GO:0006513 | protein monoubiquitination | 23277564 |
| Tgene | RNF126 | GO:0043161 | proteasome-mediated ubiquitin-dependent protein catabolic process | 24981174 |
 Fusion gene breakpoints across MKNK2 (5'-gene) Fusion gene breakpoints across MKNK2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
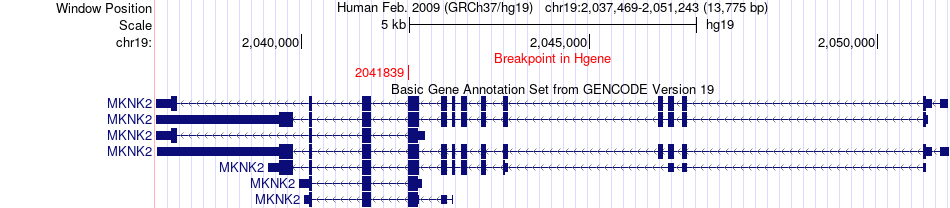 |
 Fusion gene breakpoints across RNF126 (3'-gene) Fusion gene breakpoints across RNF126 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
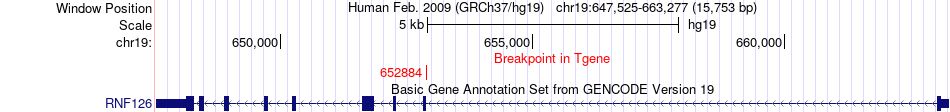 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LAML | TCGA-AB-2897-03A | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| ChimerDB4 | LAML | TCGA-AB-2897_61GABAAXX_5 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
Top |
Fusion Gene ORF analysis for MKNK2-RNF126 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000250896 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| Frame-shift | ENST00000309340 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| Frame-shift | ENST00000541165 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| Frame-shift | ENST00000588014 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| In-frame | ENST00000591142 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| In-frame | ENST00000591588 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
| In-frame | ENST00000591601 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000591142 | MKNK2 | chr19 | 2041839 | - | ENST00000292363 | RNF126 | chr19 | 652884 | - | 1764 | 301 | 124 | 1161 | 345 |
| ENST00000591601 | MKNK2 | chr19 | 2041839 | - | ENST00000292363 | RNF126 | chr19 | 652884 | - | 2444 | 981 | 15 | 1841 | 608 |
| ENST00000591588 | MKNK2 | chr19 | 2041839 | - | ENST00000292363 | RNF126 | chr19 | 652884 | - | 1706 | 243 | 66 | 1103 | 345 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000591142 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - | 0.015574513 | 0.98442554 |
| ENST00000591601 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - | 0.02815003 | 0.97185004 |
| ENST00000591588 | ENST00000292363 | MKNK2 | chr19 | 2041839 | - | RNF126 | chr19 | 652884 | - | 0.016567195 | 0.98343277 |
Top |
Fusion Genomic Features for MKNK2-RNF126 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
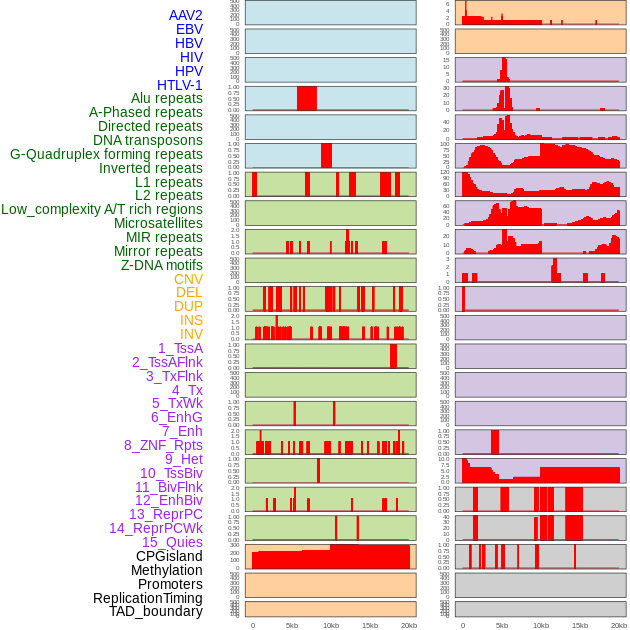 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for MKNK2-RNF126 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:2041839/chr19:652884) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MKNK2 | . |
| FUNCTION: Serine/threonine-protein kinase that phosphorylates SFPQ/PSF, HNRNPA1 and EIF4E. May play a role in the response to environmental stress and cytokines. Appears to regulate translation by phosphorylating EIF4E, thus increasing the affinity of this protein for the 7-methylguanosine-containing mRNA cap. Required for mediating PP2A-inhibition-induced EIF4E phosphorylation. Triggers EIF4E shuttling from cytoplasm to nucleus. Isoform 1 displays a high basal kinase activity, but isoform 2 exhibits a very low kinase activity. Acts as a mediator of the suppressive effects of IFNgamma on hematopoiesis. Negative regulator for signals that control generation of arsenic trioxide As(2)O(3)-dependent apoptosis and anti-leukemic responses. Involved in anti-apoptotic signaling in response to serum withdrawal. {ECO:0000269|PubMed:11154262, ECO:0000269|PubMed:11463832, ECO:0000269|PubMed:12897141, ECO:0000269|PubMed:16111636, ECO:0000269|PubMed:17965020, ECO:0000269|PubMed:18299328, ECO:0000269|PubMed:20823271, ECO:0000269|PubMed:20927323, ECO:0000269|PubMed:21149447}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000250896 | - | 11 | 14 | 60_66 | 315 | 466.0 | Motif | Note=Nuclear localization signal |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000309340 | - | 11 | 14 | 60_66 | 315 | 415.0 | Motif | Note=Nuclear localization signal |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000591601 | - | 10 | 13 | 60_66 | 315 | 466.0 | Motif | Note=Nuclear localization signal |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000250896 | - | 11 | 14 | 90_98 | 315 | 466.0 | Nucleotide binding | ATP |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000309340 | - | 11 | 14 | 90_98 | 315 | 415.0 | Nucleotide binding | ATP |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000591601 | - | 10 | 13 | 90_98 | 315 | 466.0 | Nucleotide binding | ATP |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000250896 | - | 11 | 14 | 160_162 | 315 | 466.0 | Region | Note=Staurosporine binding |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000309340 | - | 11 | 14 | 160_162 | 315 | 415.0 | Region | Note=Staurosporine binding |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000591601 | - | 10 | 13 | 160_162 | 315 | 466.0 | Region | Note=Staurosporine binding |
| Tgene | RNF126 | chr19:2041839 | chr19:652884 | ENST00000292363 | 0 | 9 | 289_303 | 25 | 312.0 | Compositional bias | Note=Ser-rich | |
| Tgene | RNF126 | chr19:2041839 | chr19:652884 | ENST00000292363 | 0 | 9 | 229_270 | 25 | 312.0 | Zinc finger | RING-type |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000250896 | - | 11 | 14 | 84_388 | 315 | 466.0 | Domain | Protein kinase |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000309340 | - | 11 | 14 | 84_388 | 315 | 415.0 | Domain | Protein kinase |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000591601 | - | 10 | 13 | 84_388 | 315 | 466.0 | Domain | Protein kinase |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000250896 | - | 11 | 14 | 444_448 | 315 | 466.0 | Motif | MAP kinase binding |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000309340 | - | 11 | 14 | 444_448 | 315 | 415.0 | Motif | MAP kinase binding |
| Hgene | MKNK2 | chr19:2041839 | chr19:652884 | ENST00000591601 | - | 10 | 13 | 444_448 | 315 | 466.0 | Motif | MAP kinase binding |
| Tgene | RNF126 | chr19:2041839 | chr19:652884 | ENST00000292363 | 0 | 9 | 13_32 | 25 | 312.0 | Zinc finger | C4-type |
Top |
Fusion Gene Sequence for MKNK2-RNF126 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >54107_54107_1_MKNK2-RNF126_MKNK2_chr19_2041839_ENST00000591142_RNF126_chr19_652884_ENST00000292363_length(transcript)=1764nt_BP=301nt GGGGCGGGGTCCGAGCCGGCTGGCGCAGGTGACCCGCAGTTACCCGGGCTCGGTTCCCACGTAATGCTGGGAAACCCCGGCTGACGCGCC CCTCCCCCGCCCGCAGTGCGGCTCGGCGGAGTACATGGCCCCGGAGGTAGTGGAGGCCTTCAGCGAGGAGGCTAGCATCTACGACAAGCG CTGCGACCTGTGGAGCCTGGGCGTCATCTTGTATATCCTACTCAGCGGCTACCCGCCCTTCGTGGGCCGCTGTGGCAGCGACTGCGGCTG GGACCGCGGCGAGGCCTGCCCTGCCTGCCAGGATTATATCTGTCCAAGATGCGAGTCTGGTTTTATCGAGGAGCTTCCGGAAGAGACCAG GAGCACAGAAAATGGTTCTGCCCCCTCCACAGCTCCCACAGACCAGAGCCGGCCACCGTTGGAGCACGTGGACCAGCACCTGTTCACGCT GCCGCAGGGCTACGGACAGTTTGCTTTCGGCATCTTCGATGACAGCTTCGAGATCCCCACGTTCCCTCCTGGGGCGCAGGCTGACGACGG CAGGGACCCTGAGAGCCGGCGGGAGAGAGACCATCCGTCCCGGCACCGGTACGGCGCCCGACAGCCCCGCGCCCGCCTCACCACGCGGCG GGCCACCGGCCGGCACGAAGGCGTCCCCACGCTGGAAGGGATCATCCAGCAGCTCGTCAACGGCATCATCACGCCCGCCACCATCCCCAG CCTGGGCCCCTGGGGAGTCCTGCACTCAAACCCTATGGACTACGCCTGGGGGGCCAACGGCCTGGATGCCATCATCACACAGCTCCTCAA TCAGTTTGAAAACACAGGCCCCCCACCGGCAGATAAAGAGAAAATCCAGGCCCTCCCCACCGTCCCCGTCACTGAGGAGCACGTAGGCTC CGGGCTCGAGTGCCCTGTGTGCAAGGACGACTACGCGCTGGGTGAGCGTGTGCGGCAGCTGCCCTGCAACCACCTGTTCCACGACGGCTG CATCGTGCCCTGGCTGGAGCAGCACGACAGCTGCCCCGTCTGCCGAAAAAGCCTCACGGGACAGAACACGGCCACGAACCCCCCTGGCCT CACTGGGGTGAGCTTCTCCTCCTCGTCGTCATCGTCCTCCTCCAGCTCGCCCAGCAACGAGAACGCCACAAGCAACTCGTGAGCCCACGT CGGCCGTCGGGAAAGCACGGGGCCTTTCCCACCCACCCTCAGCCAGCGCCACACGGCACCCACAGACTGGGTGCCCCGGCGGCGCCACGC TTGGCTGGTCAGCGCTGCAGGTCCCGCCTGTTCCAGGGCAGGACCCGGGCCCGGCCCACCGGCCCCCTGGCTTGGGAAGGCGTGGGCCAC ATGGTCCCCTCTGGGGCGTCCCCAGCCTCCCCGTCCTCTGTCTAACCTCACCCTCTAAACGTTCAGCGGTGGAAAGATTTTTATAATTTT AAATTATTACTGCTTTGAAATAAACGGACGTTTGAGCTCACGTGGCGGCCATGCATGGTTTGTGGCGGAAGACGGGGCCCAGCGGCTCCC CCAGCGCTGAAGCACGGGGTTCAGCGGCTCCCCCAGTGCTGCAGGTGCCGCCAGGACGGGCCACGTGGGCCCCCAGCCTCCCCCTCGGCC GCCTCCACGACCCGGAGTAGGGGGGCTCGGGACCATGAGGATGACCAGCAAAATTCAAGAACAAAACTGCTCCAACAGACTTTTTTAAAG >54107_54107_1_MKNK2-RNF126_MKNK2_chr19_2041839_ENST00000591142_RNF126_chr19_652884_ENST00000292363_length(amino acids)=345AA_BP=58 MAPEVVEAFSEEASIYDKRCDLWSLGVILYILLSGYPPFVGRCGSDCGWDRGEACPACQDYICPRCESGFIEELPEETRSTENGSAPSTA PTDQSRPPLEHVDQHLFTLPQGYGQFAFGIFDDSFEIPTFPPGAQADDGRDPESRRERDHPSRHRYGARQPRARLTTRRATGRHEGVPTL EGIIQQLVNGIITPATIPSLGPWGVLHSNPMDYAWGANGLDAIITQLLNQFENTGPPPADKEKIQALPTVPVTEEHVGSGLECPVCKDDY -------------------------------------------------------------- >54107_54107_2_MKNK2-RNF126_MKNK2_chr19_2041839_ENST00000591588_RNF126_chr19_652884_ENST00000292363_length(transcript)=1706nt_BP=243nt ACGTAATGCTGGGAAACCCCGGCTGACGCGCCCCTCCCCCGCCCGCAGTGCGGCTCGGCGGAGTACATGGCCCCGGAGGTAGTGGAGGCC TTCAGCGAGGAGGCTAGCATCTACGACAAGCGCTGCGACCTGTGGAGCCTGGGCGTCATCTTGTATATCCTACTCAGCGGCTACCCGCCC TTCGTGGGCCGCTGTGGCAGCGACTGCGGCTGGGACCGCGGCGAGGCCTGCCCTGCCTGCCAGGATTATATCTGTCCAAGATGCGAGTCT GGTTTTATCGAGGAGCTTCCGGAAGAGACCAGGAGCACAGAAAATGGTTCTGCCCCCTCCACAGCTCCCACAGACCAGAGCCGGCCACCG TTGGAGCACGTGGACCAGCACCTGTTCACGCTGCCGCAGGGCTACGGACAGTTTGCTTTCGGCATCTTCGATGACAGCTTCGAGATCCCC ACGTTCCCTCCTGGGGCGCAGGCTGACGACGGCAGGGACCCTGAGAGCCGGCGGGAGAGAGACCATCCGTCCCGGCACCGGTACGGCGCC CGACAGCCCCGCGCCCGCCTCACCACGCGGCGGGCCACCGGCCGGCACGAAGGCGTCCCCACGCTGGAAGGGATCATCCAGCAGCTCGTC AACGGCATCATCACGCCCGCCACCATCCCCAGCCTGGGCCCCTGGGGAGTCCTGCACTCAAACCCTATGGACTACGCCTGGGGGGCCAAC GGCCTGGATGCCATCATCACACAGCTCCTCAATCAGTTTGAAAACACAGGCCCCCCACCGGCAGATAAAGAGAAAATCCAGGCCCTCCCC ACCGTCCCCGTCACTGAGGAGCACGTAGGCTCCGGGCTCGAGTGCCCTGTGTGCAAGGACGACTACGCGCTGGGTGAGCGTGTGCGGCAG CTGCCCTGCAACCACCTGTTCCACGACGGCTGCATCGTGCCCTGGCTGGAGCAGCACGACAGCTGCCCCGTCTGCCGAAAAAGCCTCACG GGACAGAACACGGCCACGAACCCCCCTGGCCTCACTGGGGTGAGCTTCTCCTCCTCGTCGTCATCGTCCTCCTCCAGCTCGCCCAGCAAC GAGAACGCCACAAGCAACTCGTGAGCCCACGTCGGCCGTCGGGAAAGCACGGGGCCTTTCCCACCCACCCTCAGCCAGCGCCACACGGCA CCCACAGACTGGGTGCCCCGGCGGCGCCACGCTTGGCTGGTCAGCGCTGCAGGTCCCGCCTGTTCCAGGGCAGGACCCGGGCCCGGCCCA CCGGCCCCCTGGCTTGGGAAGGCGTGGGCCACATGGTCCCCTCTGGGGCGTCCCCAGCCTCCCCGTCCTCTGTCTAACCTCACCCTCTAA ACGTTCAGCGGTGGAAAGATTTTTATAATTTTAAATTATTACTGCTTTGAAATAAACGGACGTTTGAGCTCACGTGGCGGCCATGCATGG TTTGTGGCGGAAGACGGGGCCCAGCGGCTCCCCCAGCGCTGAAGCACGGGGTTCAGCGGCTCCCCCAGTGCTGCAGGTGCCGCCAGGACG GGCCACGTGGGCCCCCAGCCTCCCCCTCGGCCGCCTCCACGACCCGGAGTAGGGGGGCTCGGGACCATGAGGATGACCAGCAAAATTCAA >54107_54107_2_MKNK2-RNF126_MKNK2_chr19_2041839_ENST00000591588_RNF126_chr19_652884_ENST00000292363_length(amino acids)=345AA_BP=58 MAPEVVEAFSEEASIYDKRCDLWSLGVILYILLSGYPPFVGRCGSDCGWDRGEACPACQDYICPRCESGFIEELPEETRSTENGSAPSTA PTDQSRPPLEHVDQHLFTLPQGYGQFAFGIFDDSFEIPTFPPGAQADDGRDPESRRERDHPSRHRYGARQPRARLTTRRATGRHEGVPTL EGIIQQLVNGIITPATIPSLGPWGVLHSNPMDYAWGANGLDAIITQLLNQFENTGPPPADKEKIQALPTVPVTEEHVGSGLECPVCKDDY -------------------------------------------------------------- >54107_54107_3_MKNK2-RNF126_MKNK2_chr19_2041839_ENST00000591601_RNF126_chr19_652884_ENST00000292363_length(transcript)=2444nt_BP=981nt CGGTCCCCTCCCCCGCTGGCGGGGCCCGGACAGAAGATGGTGCAGAAGAAACCAGCCGAACTTCAGGGTTTCCACCGTTCGTTCAAGGGG CAGAACCCCTTCGAGCTGGCCTTCTCCCTAGACCAGCCCGACCACGGAGACTCTGACTTTGGCCTGCAGTGCTCAGCCCGCCCTGACATG CCCGCCAGCCAGCCCATTGACATCCCGGACGCCAAGAAGAGGGGCAAGAAGAAGAAGCGCGGCCGGGCCACCGACAGCTTCTCGGGCAGG TTTGAAGACGTCTACCAGCTGCAGGAAGATGTGCTGGGGGAGGGCGCTCATGCCCGAGTGCAGACCTGCATCAACCTGATCACCAGCCAG GAGTACGCCGTCAAGATCATTGAGAAGCAGCCAGGCCACATTCGGAGCAGGGTTTTCAGGGAGGTGGAGATGCTGTACCAGTGCCAGGGA CACAGGAACGTCCTAGAGCTGATTGAGTTCTTCGAGGAGGAGGACCGCTTCTACCTGGTGTTTGAGAAGATGCGGGGAGGCTCCATCCTG AGCCACATCCACAAGCGCCGGCACTTCAACGAGCTGGAGGCCAGCGTGGTGGTGCAGGACGTGGCCAGCGCCTTGGACTTTCTGCATAAC AAAGGCATCGCCCACAGGGACCTAAAGCCGGAAAACATCCTCTGTGAGCACCCCAACCAGGTCTCCCCCGTGAAGATCTGTGACTTCGAC CTGGGCAGCGGCATCAAACTCAACGGGGACTGCTCCCCTATCTCCACCCCGGAGCTGCTCACTCCGTGCGGCTCGGCGGAGTACATGGCC CCGGAGGTAGTGGAGGCCTTCAGCGAGGAGGCTAGCATCTACGACAAGCGCTGCGACCTGTGGAGCCTGGGCGTCATCTTGTATATCCTA CTCAGCGGCTACCCGCCCTTCGTGGGCCGCTGTGGCAGCGACTGCGGCTGGGACCGCGGCGAGGCCTGCCCTGCCTGCCAGGATTATATC TGTCCAAGATGCGAGTCTGGTTTTATCGAGGAGCTTCCGGAAGAGACCAGGAGCACAGAAAATGGTTCTGCCCCCTCCACAGCTCCCACA GACCAGAGCCGGCCACCGTTGGAGCACGTGGACCAGCACCTGTTCACGCTGCCGCAGGGCTACGGACAGTTTGCTTTCGGCATCTTCGAT GACAGCTTCGAGATCCCCACGTTCCCTCCTGGGGCGCAGGCTGACGACGGCAGGGACCCTGAGAGCCGGCGGGAGAGAGACCATCCGTCC CGGCACCGGTACGGCGCCCGACAGCCCCGCGCCCGCCTCACCACGCGGCGGGCCACCGGCCGGCACGAAGGCGTCCCCACGCTGGAAGGG ATCATCCAGCAGCTCGTCAACGGCATCATCACGCCCGCCACCATCCCCAGCCTGGGCCCCTGGGGAGTCCTGCACTCAAACCCTATGGAC TACGCCTGGGGGGCCAACGGCCTGGATGCCATCATCACACAGCTCCTCAATCAGTTTGAAAACACAGGCCCCCCACCGGCAGATAAAGAG AAAATCCAGGCCCTCCCCACCGTCCCCGTCACTGAGGAGCACGTAGGCTCCGGGCTCGAGTGCCCTGTGTGCAAGGACGACTACGCGCTG GGTGAGCGTGTGCGGCAGCTGCCCTGCAACCACCTGTTCCACGACGGCTGCATCGTGCCCTGGCTGGAGCAGCACGACAGCTGCCCCGTC TGCCGAAAAAGCCTCACGGGACAGAACACGGCCACGAACCCCCCTGGCCTCACTGGGGTGAGCTTCTCCTCCTCGTCGTCATCGTCCTCC TCCAGCTCGCCCAGCAACGAGAACGCCACAAGCAACTCGTGAGCCCACGTCGGCCGTCGGGAAAGCACGGGGCCTTTCCCACCCACCCTC AGCCAGCGCCACACGGCACCCACAGACTGGGTGCCCCGGCGGCGCCACGCTTGGCTGGTCAGCGCTGCAGGTCCCGCCTGTTCCAGGGCA GGACCCGGGCCCGGCCCACCGGCCCCCTGGCTTGGGAAGGCGTGGGCCACATGGTCCCCTCTGGGGCGTCCCCAGCCTCCCCGTCCTCTG TCTAACCTCACCCTCTAAACGTTCAGCGGTGGAAAGATTTTTATAATTTTAAATTATTACTGCTTTGAAATAAACGGACGTTTGAGCTCA CGTGGCGGCCATGCATGGTTTGTGGCGGAAGACGGGGCCCAGCGGCTCCCCCAGCGCTGAAGCACGGGGTTCAGCGGCTCCCCCAGTGCT GCAGGTGCCGCCAGGACGGGCCACGTGGGCCCCCAGCCTCCCCCTCGGCCGCCTCCACGACCCGGAGTAGGGGGGCTCGGGACCATGAGG ATGACCAGCAAAATTCAAGAACAAAACTGCTCCAACAGACTTTTTTAAAGGAAAAAATATGTGTATCTTGAAAGCTATTTAAAATACACT >54107_54107_3_MKNK2-RNF126_MKNK2_chr19_2041839_ENST00000591601_RNF126_chr19_652884_ENST00000292363_length(amino acids)=608AA_BP=321 MAGPGQKMVQKKPAELQGFHRSFKGQNPFELAFSLDQPDHGDSDFGLQCSARPDMPASQPIDIPDAKKRGKKKKRGRATDSFSGRFEDVY QLQEDVLGEGAHARVQTCINLITSQEYAVKIIEKQPGHIRSRVFREVEMLYQCQGHRNVLELIEFFEEEDRFYLVFEKMRGGSILSHIHK RRHFNELEASVVVQDVASALDFLHNKGIAHRDLKPENILCEHPNQVSPVKICDFDLGSGIKLNGDCSPISTPELLTPCGSAEYMAPEVVE AFSEEASIYDKRCDLWSLGVILYILLSGYPPFVGRCGSDCGWDRGEACPACQDYICPRCESGFIEELPEETRSTENGSAPSTAPTDQSRP PLEHVDQHLFTLPQGYGQFAFGIFDDSFEIPTFPPGAQADDGRDPESRRERDHPSRHRYGARQPRARLTTRRATGRHEGVPTLEGIIQQL VNGIITPATIPSLGPWGVLHSNPMDYAWGANGLDAIITQLLNQFENTGPPPADKEKIQALPTVPVTEEHVGSGLECPVCKDDYALGERVR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MKNK2-RNF126 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MKNK2-RNF126 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MKNK2-RNF126 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
