
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MNAT1-AP5M1 (FusionGDB2 ID:54455) |
Fusion Gene Summary for MNAT1-AP5M1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MNAT1-AP5M1 | Fusion gene ID: 54455 | Hgene | Tgene | Gene symbol | MNAT1 | AP5M1 | Gene ID | 4331 | 55745 |
| Gene name | MNAT1 component of CDK activating kinase | adaptor related protein complex 5 subunit mu 1 | |
| Synonyms | CAP35|MAT1|RNF66|TFB3 | C14orf108|MUDENG|Mu5|MuD | |
| Cytomap | 14q23.1 | 14q22.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | CDK-activating kinase assembly factor MAT1CDK7/cyclin-H assembly factorMNAT CDK-activating kinase assembly factor 1MNAT1, CDK activating kinase assembly factorRING finger protein 66RING finger protein MAT1cyclin G1 interacting proteinmenage a trois | AP-5 complex subunit mu-1AP-5 complex subunit muMHD domain-containing death-inducing proteinMU-2/AP1M2 domain containing, death-inducingMu-2 related death-inducingadapter-related protein complex 5 mu subunitadapter-related protein complex 5 subunit | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | P51948 | Q9H0R1 | |
| Ensembl transtripts involved in fusion gene | ENST00000261245, ENST00000539616, ENST00000555545, | ENST00000556723, ENST00000261558, ENST00000431972, | |
| Fusion gene scores | * DoF score | 10 X 6 X 7=420 | 3 X 4 X 2=24 |
| # samples | 10 | 4 | |
| ** MAII score | log2(10/420*10)=-2.0703893278914 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/24*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: MNAT1 [Title/Abstract] AND AP5M1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MNAT1(61346553)-AP5M1(57740962), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MNAT1 | GO:0006357 | regulation of transcription by RNA polymerase II | 10801852 |
| Hgene | MNAT1 | GO:0006366 | transcription by RNA polymerase II | 9852112 |
| Hgene | MNAT1 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 8692841 |
| Hgene | MNAT1 | GO:1905775 | negative regulation of DNA helicase activity | 11445587 |
 Fusion gene breakpoints across MNAT1 (5'-gene) Fusion gene breakpoints across MNAT1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
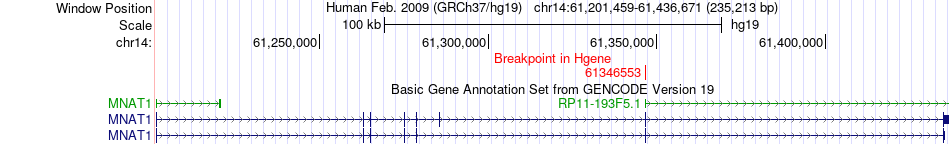 |
 Fusion gene breakpoints across AP5M1 (3'-gene) Fusion gene breakpoints across AP5M1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
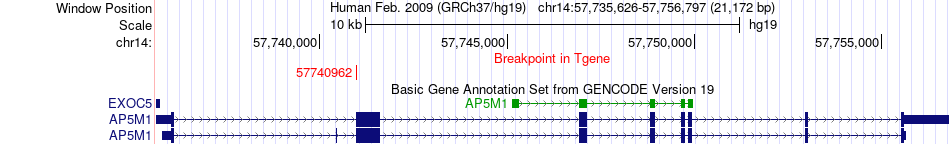 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUSC | TCGA-LA-A7SW-01A | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| ChimerDB4 | LUSC | TCGA-LA-A7SW | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
Top |
Fusion Gene ORF analysis for MNAT1-AP5M1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000261245 | ENST00000556723 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| 5CDS-intron | ENST00000539616 | ENST00000556723 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| In-frame | ENST00000261245 | ENST00000261558 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| In-frame | ENST00000261245 | ENST00000431972 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| In-frame | ENST00000539616 | ENST00000261558 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| In-frame | ENST00000539616 | ENST00000431972 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| intron-3CDS | ENST00000555545 | ENST00000261558 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| intron-3CDS | ENST00000555545 | ENST00000431972 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
| intron-intron | ENST00000555545 | ENST00000556723 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000261245 | MNAT1 | chr14 | 61346553 | + | ENST00000261558 | AP5M1 | chr14 | 57740962 | + | 3504 | 910 | 5 | 2308 | 767 |
| ENST00000261245 | MNAT1 | chr14 | 61346553 | + | ENST00000431972 | AP5M1 | chr14 | 57740962 | + | 2350 | 910 | 5 | 2308 | 767 |
| ENST00000539616 | MNAT1 | chr14 | 61346553 | + | ENST00000261558 | AP5M1 | chr14 | 57740962 | + | 3364 | 770 | 9 | 2168 | 719 |
| ENST00000539616 | MNAT1 | chr14 | 61346553 | + | ENST00000431972 | AP5M1 | chr14 | 57740962 | + | 2210 | 770 | 9 | 2168 | 719 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000261245 | ENST00000261558 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + | 0.000264493 | 0.99973553 |
| ENST00000261245 | ENST00000431972 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + | 0.00058159 | 0.9994184 |
| ENST00000539616 | ENST00000261558 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + | 0.000329578 | 0.9996704 |
| ENST00000539616 | ENST00000431972 | MNAT1 | chr14 | 61346553 | + | AP5M1 | chr14 | 57740962 | + | 0.000722382 | 0.99927765 |
Top |
Fusion Genomic Features for MNAT1-AP5M1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
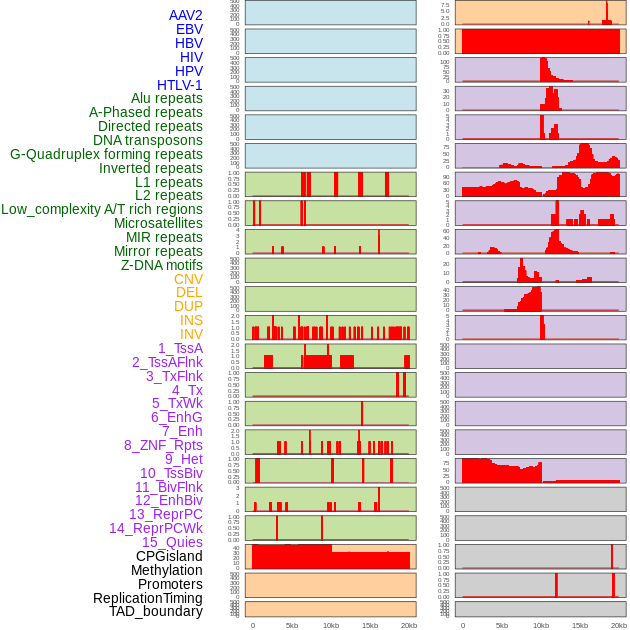 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
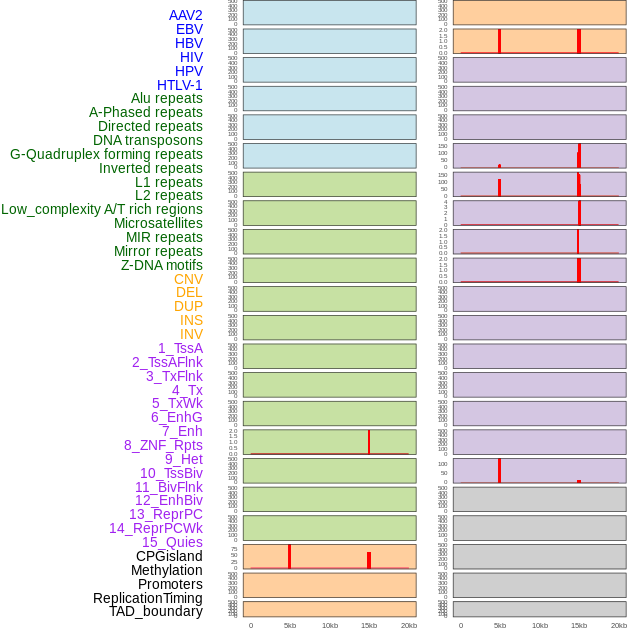 |
Top |
Fusion Protein Features for MNAT1-AP5M1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr14:61346553/chr14:57740962) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MNAT1 | AP5M1 |
| FUNCTION: Stabilizes the cyclin H-CDK7 complex to form a functional CDK-activating kinase (CAK) enzymatic complex. CAK activates the cyclin-associated kinases CDK1, CDK2, CDK4 and CDK6 by threonine phosphorylation. CAK complexed to the core-TFIIH basal transcription factor activates RNA polymerase II by serine phosphorylation of the repetitive C-terminal domain (CTD) of its large subunit (POLR2A), allowing its escape from the promoter and elongation of the transcripts. Involved in cell cycle control and in RNA transcription by RNA polymerase II. {ECO:0000269|PubMed:10024882}. | FUNCTION: As part of AP-5, a probable fifth adaptor protein complex it may be involved in endosomal transport. According to PubMed:18395520, it may play a role in cell death. {ECO:0000269|PubMed:18395520, ECO:0000269|PubMed:22022230}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MNAT1 | chr14:61346553 | chr14:57740962 | ENST00000261245 | + | 7 | 8 | 142_161 | 269 | 310.0 | Domain | UIM |
| Hgene | MNAT1 | chr14:61346553 | chr14:57740962 | ENST00000539616 | + | 6 | 7 | 142_161 | 227 | 268.0 | Domain | UIM |
| Hgene | MNAT1 | chr14:61346553 | chr14:57740962 | ENST00000261245 | + | 7 | 8 | 6_50 | 269 | 310.0 | Zinc finger | RING-type |
| Hgene | MNAT1 | chr14:61346553 | chr14:57740962 | ENST00000539616 | + | 6 | 7 | 6_50 | 227 | 268.0 | Zinc finger | RING-type |
| Tgene | AP5M1 | chr14:61346553 | chr14:57740962 | ENST00000261558 | 0 | 8 | 206_476 | 24 | 491.0 | Domain | MHD |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for MNAT1-AP5M1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >54455_54455_1_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000261245_AP5M1_chr14_57740962_ENST00000261558_length(transcript)=3504nt_BP=910nt AGCGCCTGTTGGTAGGAACCTGCTTGGTCGCGTCTGAGGGGGCTTGTAGGTGGCTCTGGCTGAAACAGGCGCCTGCGAGAGTCTGTAGGA GGGAAACCGCCATGGACGATCAGGGTTGCCCTCGGTGTAAGACCACCAAATATCGGAACCCCTCCTTGAAGCTGATGGTGAATGTGTGCG GACACACTCTCTGTGAAAGTTGTGTAGATTTACTGTTTGTGAGAGGAGCTGGAAACTGCCCTGAGTGTGGTACTCCACTCAGAAAGAGCA ACTTCAGGGTACAACTCTTTGAAGATCCCACTGTTGACAAGGAGGTTGAGATCAGGAAAAAAGTGCTAAAGATATACAATAAAAGGGAAG AAGATTTTCCTAGTCTAAGAGAATACAATGATTTCTTGGAAGAAGTGGAAGAAATTGTTTTCAACTTGACCAACAATGTGGATTTGGACA ACACCAAAAAGAAAATGGAGATATACCAAAAGGAAAACAAAGATGTTATTCAGAAAAATAAATTAAAGCTGACTCGAGAACAGGAAGAAC TGGAAGAAGCTTTAGAAGTGGAACGACAGGAAAATGAACAAAGAAGATTATTTATACAAAAAGAAGAACAACTGCAGCAGATTCTAAAAA GGAAGAATAAGCAGGCTTTTTTAGATGAGCTGGAGAGTTCTGATCTCCCTGTTGCTCTGCTTTTGGCTCAGCATAAAGATAGATCTACCC AATTAGAAATGCAACTTGAGAAACCCAAACCTGTAAAACCAGTGACGTTTTCCACAGGCATCAAAATGGGTCAACATATTTCACTGGCAC CTATTCACAAGCTTGAAGAAGCTCTGTATGAATACCAGCCACTGCAGATAGAGACATATGGACCACATGTTCCTGAGCTTGAGATGCTAG GAAGACTTGGACGGTATCCAACTGTTGAAAAACGAGCCAGAGTCTTCAATGGAGCAAGTTATGTGCCTGTTCCTGAAGATGGTCCCTTTC TTAAAGCACTGCTCTTTGAACTTAGATTATTGGATGATGATAAAGACTTCGTTGAGAGTCGTGATAGCTGTTCACGCATCAATAAAACAT CCATTTATGGACTCCTGATAGGAGGTGAAGAACTCTGGCCAGTTGTTGCTTTTCTGAAGAATGACATGATATATGCTTGTGTTCCACTAG TTGAACAAACTCTGTCCCCTCGTCCGCCACTAATTAGTGTCAGTGGAGTTTCACAAGGCTTTGAATTTCTTTTTGGGATACAGGATTTTC TTTATTCAGGTCAAAAAAATGACTCTGAGCTGAATACAAAATTGAGCCAGTTGCCTGACTTGCTTCTGCAGGCTTGTCCATTTGGTACTT TATTAGATGCCAACTTACAGAATTCATTAGATAATACCAATTTTGCATCTGTGACTCAGCCACAGAAACAGCCAGCTTGGAAAACTGGGA CGTACAAAGGAAAACCACAAGTTTCTATTTCTATCACTGAAAAGGTAAAATCCATGCAATATGATAAACAGGGTATAGCAGATACATGGC AAGTTGTTGGAACAGTGACTTGCAAGTGTGATTTGGAAGGAATCATGCCAAATGTTACCATCAGCTTGAGTCTCCCCACCAATGGATCTC CACTTCAGGATATTCTAGTTCACCCTTGTGTAACTTCTCTTGACTCTGCAATTCTGACTTCTAGTAGTATTGATGCAATGGATGACTCTG CATTTAGTGGGCCTTACAAATTTCCATTCACTCCACCTTTAGAGTCATTCAACTTATGCTTCTACACTTCCCAGGTCCCTGTCCCACCAA TTTTGGGTTTTTATCAAATGAAGGAGGAAGAAGTACAACTAAGAATAACCATTAATTTAAAACTTCATGAAAGTGTGAAAAATAATTTTG AATTCTGTGAAGCCCATATACCTTTTTACAATAGAGGTCCAATTACACATTTGGAATACAAAACTAGTTTTGGCCAGCTTGAAGTATTTC GAGAGAAAAGCTTATTGATCTGGATTATTGGCCAGAAGTTCCCAAAATCAATGGAAATTAGTCTTTCTGGAACTGTAACTTTTGGAGCCA AGAGCCATGAGAAGCAGCCATTTGACCCAATTTGTACTGGAGAAACAGCATATTTAAAGCTTCATTTTAGGATCTTAGATTACACACTTA CTGGATGTTATGCAGATCAGCATTCAGTTCAAGTTTTTGCATCAGGAAAACCAAAAATAAGTGCACACCGGAAACTAATTTCTTCTGATT ATTACATCTGGAATTCTAAAGCCCCTGCTCCAGTAACATATGGATCATTATTATTGTAATAGTCTCATGTTTAAATGGGATTATATAATG ATAACAGTTTAAAGAAAATCATAATCTTATATTTTTAATGTGGATGCATATAACCTGTGAGTGAAAAATCACTGAATGATTTAATTGTAA AAGTAGTCTTATGTGGTGTTTGTAGTCTGATAGAGCTTGAAAGGACATTTTAAAAGCTAATGTCTCCAATTTTGTTAACCTTCGATTTTA TGCCAGTATAATTCAGAACATAGAAAAGTAATGATTCACTTGGGCTCATTTTAGACTGGTCCTGGGTCACCCTGCCACACTTGTTTCCTA GTGTTTCTGTGGCAGACATTGCTAATCAATTACAGCCCTTTTCTGTACTGAGCCTTGGATAAAGGGTCAGGCTCCTTTTTAGTTCAGAGA TTCAGGCAGCCACTCCCAGTGGGTTGTAGATAATGTGCAAGATAAAAACTATTTTCTCTTCCAAATCTAAGTACTAAGCTCCTAGTATAA GGTGTTGTTACAGAATACCAGAGACCATGTTAGAGACAACTACATCTCTTCAAAAAACAGCCAACAGAGACAAAGGAAAAGTGTTTAAAT AGTAAGCTGTTCTTCTTAATCAGAACTATCCTATTGACTAATAAATAATCTGCATAATTCTACTTAAGGTGTGTAATCTCTGTTCTAGAG TTAGTTTTTAAGTAAGCTTGTTAATCTGCCACTTTGACATTTTGCTTAGGATGTCAGTAGCCATATTAAGATGTGTAGAATACCTTCAGA AGATGATCATAGTGTTTTGTAATCATTTAATGTCTGCAGCCAAATTTTTAAAGGTAATTTAGACCTAATACTGCTCTTGCTGTGTCTTAT TAAGTTAAAATTAATGAATGAATTCTGGTAAAAATTCAAAAGGCACTCTGTGAGTAGAGAGTATCATTTAAGCTTATTTTAGTCACATGT AGTATATATCTCCTTAAAGCTGTCACTCTCACTTTCTTACCATTCTCTTGATTTCTTCAGAAACCATCTAGTCATCATCTTTATACTCTA CCTGCTTCTGCAATTATATATCATATTATGTTTTCAGAGCAGTTCATTGTCAAGTTGGACTTTAAGTGACCATTCAAGAAAAGATGAAAT >54455_54455_1_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000261245_AP5M1_chr14_57740962_ENST00000261558_length(amino acids)=767AA_BP=302 MLVGTCLVASEGACRWLWLKQAPARVCRRETAMDDQGCPRCKTTKYRNPSLKLMVNVCGHTLCESCVDLLFVRGAGNCPECGTPLRKSNF RVQLFEDPTVDKEVEIRKKVLKIYNKREEDFPSLREYNDFLEEVEEIVFNLTNNVDLDNTKKKMEIYQKENKDVIQKNKLKLTREQEELE EALEVERQENEQRRLFIQKEEQLQQILKRKNKQAFLDELESSDLPVALLLAQHKDRSTQLEMQLEKPKPVKPVTFSTGIKMGQHISLAPI HKLEEALYEYQPLQIETYGPHVPELEMLGRLGRYPTVEKRARVFNGASYVPVPEDGPFLKALLFELRLLDDDKDFVESRDSCSRINKTSI YGLLIGGEELWPVVAFLKNDMIYACVPLVEQTLSPRPPLISVSGVSQGFEFLFGIQDFLYSGQKNDSELNTKLSQLPDLLLQACPFGTLL DANLQNSLDNTNFASVTQPQKQPAWKTGTYKGKPQVSISITEKVKSMQYDKQGIADTWQVVGTVTCKCDLEGIMPNVTISLSLPTNGSPL QDILVHPCVTSLDSAILTSSSIDAMDDSAFSGPYKFPFTPPLESFNLCFYTSQVPVPPILGFYQMKEEEVQLRITINLKLHESVKNNFEF CEAHIPFYNRGPITHLEYKTSFGQLEVFREKSLLIWIIGQKFPKSMEISLSGTVTFGAKSHEKQPFDPICTGETAYLKLHFRILDYTLTG -------------------------------------------------------------- >54455_54455_2_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000261245_AP5M1_chr14_57740962_ENST00000431972_length(transcript)=2350nt_BP=910nt AGCGCCTGTTGGTAGGAACCTGCTTGGTCGCGTCTGAGGGGGCTTGTAGGTGGCTCTGGCTGAAACAGGCGCCTGCGAGAGTCTGTAGGA GGGAAACCGCCATGGACGATCAGGGTTGCCCTCGGTGTAAGACCACCAAATATCGGAACCCCTCCTTGAAGCTGATGGTGAATGTGTGCG GACACACTCTCTGTGAAAGTTGTGTAGATTTACTGTTTGTGAGAGGAGCTGGAAACTGCCCTGAGTGTGGTACTCCACTCAGAAAGAGCA ACTTCAGGGTACAACTCTTTGAAGATCCCACTGTTGACAAGGAGGTTGAGATCAGGAAAAAAGTGCTAAAGATATACAATAAAAGGGAAG AAGATTTTCCTAGTCTAAGAGAATACAATGATTTCTTGGAAGAAGTGGAAGAAATTGTTTTCAACTTGACCAACAATGTGGATTTGGACA ACACCAAAAAGAAAATGGAGATATACCAAAAGGAAAACAAAGATGTTATTCAGAAAAATAAATTAAAGCTGACTCGAGAACAGGAAGAAC TGGAAGAAGCTTTAGAAGTGGAACGACAGGAAAATGAACAAAGAAGATTATTTATACAAAAAGAAGAACAACTGCAGCAGATTCTAAAAA GGAAGAATAAGCAGGCTTTTTTAGATGAGCTGGAGAGTTCTGATCTCCCTGTTGCTCTGCTTTTGGCTCAGCATAAAGATAGATCTACCC AATTAGAAATGCAACTTGAGAAACCCAAACCTGTAAAACCAGTGACGTTTTCCACAGGCATCAAAATGGGTCAACATATTTCACTGGCAC CTATTCACAAGCTTGAAGAAGCTCTGTATGAATACCAGCCACTGCAGATAGAGACATATGGACCACATGTTCCTGAGCTTGAGATGCTAG GAAGACTTGGACGGTATCCAACTGTTGAAAAACGAGCCAGAGTCTTCAATGGAGCAAGTTATGTGCCTGTTCCTGAAGATGGTCCCTTTC TTAAAGCACTGCTCTTTGAACTTAGATTATTGGATGATGATAAAGACTTCGTTGAGAGTCGTGATAGCTGTTCACGCATCAATAAAACAT CCATTTATGGACTCCTGATAGGAGGTGAAGAACTCTGGCCAGTTGTTGCTTTTCTGAAGAATGACATGATATATGCTTGTGTTCCACTAG TTGAACAAACTCTGTCCCCTCGTCCGCCACTAATTAGTGTCAGTGGAGTTTCACAAGGCTTTGAATTTCTTTTTGGGATACAGGATTTTC TTTATTCAGGTCAAAAAAATGACTCTGAGCTGAATACAAAATTGAGCCAGTTGCCTGACTTGCTTCTGCAGGCTTGTCCATTTGGTACTT TATTAGATGCCAACTTACAGAATTCATTAGATAATACCAATTTTGCATCTGTGACTCAGCCACAGAAACAGCCAGCTTGGAAAACTGGGA CGTACAAAGGAAAACCACAAGTTTCTATTTCTATCACTGAAAAGGTAAAATCCATGCAATATGATAAACAGGGTATAGCAGATACATGGC AAGTTGTTGGAACAGTGACTTGCAAGTGTGATTTGGAAGGAATCATGCCAAATGTTACCATCAGCTTGAGTCTCCCCACCAATGGATCTC CACTTCAGGATATTCTAGTTCACCCTTGTGTAACTTCTCTTGACTCTGCAATTCTGACTTCTAGTAGTATTGATGCAATGGATGACTCTG CATTTAGTGGGCCTTACAAATTTCCATTCACTCCACCTTTAGAGTCATTCAACTTATGCTTCTACACTTCCCAGGTCCCTGTCCCACCAA TTTTGGGTTTTTATCAAATGAAGGAGGAAGAAGTACAACTAAGAATAACCATTAATTTAAAACTTCATGAAAGTGTGAAAAATAATTTTG AATTCTGTGAAGCCCATATACCTTTTTACAATAGAGGTCCAATTACACATTTGGAATACAAAACTAGTTTTGGCCAGCTTGAAGTATTTC GAGAGAAAAGCTTATTGATCTGGATTATTGGCCAGAAGTTCCCAAAATCAATGGAAATTAGTCTTTCTGGAACTGTAACTTTTGGAGCCA AGAGCCATGAGAAGCAGCCATTTGACCCAATTTGTACTGGAGAAACAGCATATTTAAAGCTTCATTTTAGGATCTTAGATTACACACTTA CTGGATGTTATGCAGATCAGCATTCAGTTCAAGTTTTTGCATCAGGAAAACCAAAAATAAGTGCACACCGGAAACTAATTTCTTCTGATT ATTACATCTGGAATTCTAAAGCCCCTGCTCCAGTAACATATGGATCATTATTATTGTAATAGTCTCATGTTTAAATGGGATTATATAATG >54455_54455_2_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000261245_AP5M1_chr14_57740962_ENST00000431972_length(amino acids)=767AA_BP=302 MLVGTCLVASEGACRWLWLKQAPARVCRRETAMDDQGCPRCKTTKYRNPSLKLMVNVCGHTLCESCVDLLFVRGAGNCPECGTPLRKSNF RVQLFEDPTVDKEVEIRKKVLKIYNKREEDFPSLREYNDFLEEVEEIVFNLTNNVDLDNTKKKMEIYQKENKDVIQKNKLKLTREQEELE EALEVERQENEQRRLFIQKEEQLQQILKRKNKQAFLDELESSDLPVALLLAQHKDRSTQLEMQLEKPKPVKPVTFSTGIKMGQHISLAPI HKLEEALYEYQPLQIETYGPHVPELEMLGRLGRYPTVEKRARVFNGASYVPVPEDGPFLKALLFELRLLDDDKDFVESRDSCSRINKTSI YGLLIGGEELWPVVAFLKNDMIYACVPLVEQTLSPRPPLISVSGVSQGFEFLFGIQDFLYSGQKNDSELNTKLSQLPDLLLQACPFGTLL DANLQNSLDNTNFASVTQPQKQPAWKTGTYKGKPQVSISITEKVKSMQYDKQGIADTWQVVGTVTCKCDLEGIMPNVTISLSLPTNGSPL QDILVHPCVTSLDSAILTSSSIDAMDDSAFSGPYKFPFTPPLESFNLCFYTSQVPVPPILGFYQMKEEEVQLRITINLKLHESVKNNFEF CEAHIPFYNRGPITHLEYKTSFGQLEVFREKSLLIWIIGQKFPKSMEISLSGTVTFGAKSHEKQPFDPICTGETAYLKLHFRILDYTLTG -------------------------------------------------------------- >54455_54455_3_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000539616_AP5M1_chr14_57740962_ENST00000261558_length(transcript)=3364nt_BP=770nt GGAACCTGCTTGGTCGCGTCTGAGGGGGCTTGTAGGTGGCTCTGGCTGAAACAGGCGCCTGCGAGAGTCTGTAGGAGGGAAACCGCCATG GACGATCAGGGTTGCCCTCGGTGTAAGACCACCAAATATCGGAACCCCTCCTTGAAGCTGATGGTGAATGTGTGCGGACACACTCTCTGT GAAAGTTGTGTAGATTTACTGTTTGTGAGAGGAGCTGGAAACTGCCCTGAGTGTGGTACTCCACTCAGAAAGAGCAACTTCAGGGTACAA CTCTTTGAAGATCCCACTGTTGACAAGGAGGTTGAGATCAGGAAAAAAGTGCTAAAGATATACAATAAAAGGGAAGAAGATTTTCCTAGT CTAAGAGAATACAATGATTTCTTGGAAGAAGTGGAAGAAATTGTTTTCAACTTGACCAACAATGTGGATTTGGACAACACCAAAAAGAAA ATGGAGATATACCAAAAGGAAAACAAAGATGTTATTCAGAAAAATAAATTAAAGCTGACTCGAGAACAGGAAGAACTGGAAGAAGCTTTA GAAGTGGAACGACAGGAAAATGAACAAAGAAGATTATTTATACAAAAAGAAGAACAACTGCAGCAGATTCTAAAAAGGAAGAATAAGCAG GCTTTTTTAGATGAGCTGGGTCAACATATTTCACTGGCACCTATTCACAAGCTTGAAGAAGCTCTGTATGAATACCAGCCACTGCAGATA GAGACATATGGACCACATGTTCCTGAGCTTGAGATGCTAGGAAGACTTGGACGGTATCCAACTGTTGAAAAACGAGCCAGAGTCTTCAAT GGAGCAAGTTATGTGCCTGTTCCTGAAGATGGTCCCTTTCTTAAAGCACTGCTCTTTGAACTTAGATTATTGGATGATGATAAAGACTTC GTTGAGAGTCGTGATAGCTGTTCACGCATCAATAAAACATCCATTTATGGACTCCTGATAGGAGGTGAAGAACTCTGGCCAGTTGTTGCT TTTCTGAAGAATGACATGATATATGCTTGTGTTCCACTAGTTGAACAAACTCTGTCCCCTCGTCCGCCACTAATTAGTGTCAGTGGAGTT TCACAAGGCTTTGAATTTCTTTTTGGGATACAGGATTTTCTTTATTCAGGTCAAAAAAATGACTCTGAGCTGAATACAAAATTGAGCCAG TTGCCTGACTTGCTTCTGCAGGCTTGTCCATTTGGTACTTTATTAGATGCCAACTTACAGAATTCATTAGATAATACCAATTTTGCATCT GTGACTCAGCCACAGAAACAGCCAGCTTGGAAAACTGGGACGTACAAAGGAAAACCACAAGTTTCTATTTCTATCACTGAAAAGGTAAAA TCCATGCAATATGATAAACAGGGTATAGCAGATACATGGCAAGTTGTTGGAACAGTGACTTGCAAGTGTGATTTGGAAGGAATCATGCCA AATGTTACCATCAGCTTGAGTCTCCCCACCAATGGATCTCCACTTCAGGATATTCTAGTTCACCCTTGTGTAACTTCTCTTGACTCTGCA ATTCTGACTTCTAGTAGTATTGATGCAATGGATGACTCTGCATTTAGTGGGCCTTACAAATTTCCATTCACTCCACCTTTAGAGTCATTC AACTTATGCTTCTACACTTCCCAGGTCCCTGTCCCACCAATTTTGGGTTTTTATCAAATGAAGGAGGAAGAAGTACAACTAAGAATAACC ATTAATTTAAAACTTCATGAAAGTGTGAAAAATAATTTTGAATTCTGTGAAGCCCATATACCTTTTTACAATAGAGGTCCAATTACACAT TTGGAATACAAAACTAGTTTTGGCCAGCTTGAAGTATTTCGAGAGAAAAGCTTATTGATCTGGATTATTGGCCAGAAGTTCCCAAAATCA ATGGAAATTAGTCTTTCTGGAACTGTAACTTTTGGAGCCAAGAGCCATGAGAAGCAGCCATTTGACCCAATTTGTACTGGAGAAACAGCA TATTTAAAGCTTCATTTTAGGATCTTAGATTACACACTTACTGGATGTTATGCAGATCAGCATTCAGTTCAAGTTTTTGCATCAGGAAAA CCAAAAATAAGTGCACACCGGAAACTAATTTCTTCTGATTATTACATCTGGAATTCTAAAGCCCCTGCTCCAGTAACATATGGATCATTA TTATTGTAATAGTCTCATGTTTAAATGGGATTATATAATGATAACAGTTTAAAGAAAATCATAATCTTATATTTTTAATGTGGATGCATA TAACCTGTGAGTGAAAAATCACTGAATGATTTAATTGTAAAAGTAGTCTTATGTGGTGTTTGTAGTCTGATAGAGCTTGAAAGGACATTT TAAAAGCTAATGTCTCCAATTTTGTTAACCTTCGATTTTATGCCAGTATAATTCAGAACATAGAAAAGTAATGATTCACTTGGGCTCATT TTAGACTGGTCCTGGGTCACCCTGCCACACTTGTTTCCTAGTGTTTCTGTGGCAGACATTGCTAATCAATTACAGCCCTTTTCTGTACTG AGCCTTGGATAAAGGGTCAGGCTCCTTTTTAGTTCAGAGATTCAGGCAGCCACTCCCAGTGGGTTGTAGATAATGTGCAAGATAAAAACT ATTTTCTCTTCCAAATCTAAGTACTAAGCTCCTAGTATAAGGTGTTGTTACAGAATACCAGAGACCATGTTAGAGACAACTACATCTCTT CAAAAAACAGCCAACAGAGACAAAGGAAAAGTGTTTAAATAGTAAGCTGTTCTTCTTAATCAGAACTATCCTATTGACTAATAAATAATC TGCATAATTCTACTTAAGGTGTGTAATCTCTGTTCTAGAGTTAGTTTTTAAGTAAGCTTGTTAATCTGCCACTTTGACATTTTGCTTAGG ATGTCAGTAGCCATATTAAGATGTGTAGAATACCTTCAGAAGATGATCATAGTGTTTTGTAATCATTTAATGTCTGCAGCCAAATTTTTA AAGGTAATTTAGACCTAATACTGCTCTTGCTGTGTCTTATTAAGTTAAAATTAATGAATGAATTCTGGTAAAAATTCAAAAGGCACTCTG TGAGTAGAGAGTATCATTTAAGCTTATTTTAGTCACATGTAGTATATATCTCCTTAAAGCTGTCACTCTCACTTTCTTACCATTCTCTTG ATTTCTTCAGAAACCATCTAGTCATCATCTTTATACTCTACCTGCTTCTGCAATTATATATCATATTATGTTTTCAGAGCAGTTCATTGT CAAGTTGGACTTTAAGTGACCATTCAAGAAAAGATGAAATCTCACGAACCTCAAAACTTCATTCATGTCTTTTTACAAATGAGAAAAAAA >54455_54455_3_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000539616_AP5M1_chr14_57740962_ENST00000261558_length(amino acids)=719AA_BP=254 MVASEGACRWLWLKQAPARVCRRETAMDDQGCPRCKTTKYRNPSLKLMVNVCGHTLCESCVDLLFVRGAGNCPECGTPLRKSNFRVQLFE DPTVDKEVEIRKKVLKIYNKREEDFPSLREYNDFLEEVEEIVFNLTNNVDLDNTKKKMEIYQKENKDVIQKNKLKLTREQEELEEALEVE RQENEQRRLFIQKEEQLQQILKRKNKQAFLDELGQHISLAPIHKLEEALYEYQPLQIETYGPHVPELEMLGRLGRYPTVEKRARVFNGAS YVPVPEDGPFLKALLFELRLLDDDKDFVESRDSCSRINKTSIYGLLIGGEELWPVVAFLKNDMIYACVPLVEQTLSPRPPLISVSGVSQG FEFLFGIQDFLYSGQKNDSELNTKLSQLPDLLLQACPFGTLLDANLQNSLDNTNFASVTQPQKQPAWKTGTYKGKPQVSISITEKVKSMQ YDKQGIADTWQVVGTVTCKCDLEGIMPNVTISLSLPTNGSPLQDILVHPCVTSLDSAILTSSSIDAMDDSAFSGPYKFPFTPPLESFNLC FYTSQVPVPPILGFYQMKEEEVQLRITINLKLHESVKNNFEFCEAHIPFYNRGPITHLEYKTSFGQLEVFREKSLLIWIIGQKFPKSMEI -------------------------------------------------------------- >54455_54455_4_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000539616_AP5M1_chr14_57740962_ENST00000431972_length(transcript)=2210nt_BP=770nt GGAACCTGCTTGGTCGCGTCTGAGGGGGCTTGTAGGTGGCTCTGGCTGAAACAGGCGCCTGCGAGAGTCTGTAGGAGGGAAACCGCCATG GACGATCAGGGTTGCCCTCGGTGTAAGACCACCAAATATCGGAACCCCTCCTTGAAGCTGATGGTGAATGTGTGCGGACACACTCTCTGT GAAAGTTGTGTAGATTTACTGTTTGTGAGAGGAGCTGGAAACTGCCCTGAGTGTGGTACTCCACTCAGAAAGAGCAACTTCAGGGTACAA CTCTTTGAAGATCCCACTGTTGACAAGGAGGTTGAGATCAGGAAAAAAGTGCTAAAGATATACAATAAAAGGGAAGAAGATTTTCCTAGT CTAAGAGAATACAATGATTTCTTGGAAGAAGTGGAAGAAATTGTTTTCAACTTGACCAACAATGTGGATTTGGACAACACCAAAAAGAAA ATGGAGATATACCAAAAGGAAAACAAAGATGTTATTCAGAAAAATAAATTAAAGCTGACTCGAGAACAGGAAGAACTGGAAGAAGCTTTA GAAGTGGAACGACAGGAAAATGAACAAAGAAGATTATTTATACAAAAAGAAGAACAACTGCAGCAGATTCTAAAAAGGAAGAATAAGCAG GCTTTTTTAGATGAGCTGGGTCAACATATTTCACTGGCACCTATTCACAAGCTTGAAGAAGCTCTGTATGAATACCAGCCACTGCAGATA GAGACATATGGACCACATGTTCCTGAGCTTGAGATGCTAGGAAGACTTGGACGGTATCCAACTGTTGAAAAACGAGCCAGAGTCTTCAAT GGAGCAAGTTATGTGCCTGTTCCTGAAGATGGTCCCTTTCTTAAAGCACTGCTCTTTGAACTTAGATTATTGGATGATGATAAAGACTTC GTTGAGAGTCGTGATAGCTGTTCACGCATCAATAAAACATCCATTTATGGACTCCTGATAGGAGGTGAAGAACTCTGGCCAGTTGTTGCT TTTCTGAAGAATGACATGATATATGCTTGTGTTCCACTAGTTGAACAAACTCTGTCCCCTCGTCCGCCACTAATTAGTGTCAGTGGAGTT TCACAAGGCTTTGAATTTCTTTTTGGGATACAGGATTTTCTTTATTCAGGTCAAAAAAATGACTCTGAGCTGAATACAAAATTGAGCCAG TTGCCTGACTTGCTTCTGCAGGCTTGTCCATTTGGTACTTTATTAGATGCCAACTTACAGAATTCATTAGATAATACCAATTTTGCATCT GTGACTCAGCCACAGAAACAGCCAGCTTGGAAAACTGGGACGTACAAAGGAAAACCACAAGTTTCTATTTCTATCACTGAAAAGGTAAAA TCCATGCAATATGATAAACAGGGTATAGCAGATACATGGCAAGTTGTTGGAACAGTGACTTGCAAGTGTGATTTGGAAGGAATCATGCCA AATGTTACCATCAGCTTGAGTCTCCCCACCAATGGATCTCCACTTCAGGATATTCTAGTTCACCCTTGTGTAACTTCTCTTGACTCTGCA ATTCTGACTTCTAGTAGTATTGATGCAATGGATGACTCTGCATTTAGTGGGCCTTACAAATTTCCATTCACTCCACCTTTAGAGTCATTC AACTTATGCTTCTACACTTCCCAGGTCCCTGTCCCACCAATTTTGGGTTTTTATCAAATGAAGGAGGAAGAAGTACAACTAAGAATAACC ATTAATTTAAAACTTCATGAAAGTGTGAAAAATAATTTTGAATTCTGTGAAGCCCATATACCTTTTTACAATAGAGGTCCAATTACACAT TTGGAATACAAAACTAGTTTTGGCCAGCTTGAAGTATTTCGAGAGAAAAGCTTATTGATCTGGATTATTGGCCAGAAGTTCCCAAAATCA ATGGAAATTAGTCTTTCTGGAACTGTAACTTTTGGAGCCAAGAGCCATGAGAAGCAGCCATTTGACCCAATTTGTACTGGAGAAACAGCA TATTTAAAGCTTCATTTTAGGATCTTAGATTACACACTTACTGGATGTTATGCAGATCAGCATTCAGTTCAAGTTTTTGCATCAGGAAAA CCAAAAATAAGTGCACACCGGAAACTAATTTCTTCTGATTATTACATCTGGAATTCTAAAGCCCCTGCTCCAGTAACATATGGATCATTA >54455_54455_4_MNAT1-AP5M1_MNAT1_chr14_61346553_ENST00000539616_AP5M1_chr14_57740962_ENST00000431972_length(amino acids)=719AA_BP=254 MVASEGACRWLWLKQAPARVCRRETAMDDQGCPRCKTTKYRNPSLKLMVNVCGHTLCESCVDLLFVRGAGNCPECGTPLRKSNFRVQLFE DPTVDKEVEIRKKVLKIYNKREEDFPSLREYNDFLEEVEEIVFNLTNNVDLDNTKKKMEIYQKENKDVIQKNKLKLTREQEELEEALEVE RQENEQRRLFIQKEEQLQQILKRKNKQAFLDELGQHISLAPIHKLEEALYEYQPLQIETYGPHVPELEMLGRLGRYPTVEKRARVFNGAS YVPVPEDGPFLKALLFELRLLDDDKDFVESRDSCSRINKTSIYGLLIGGEELWPVVAFLKNDMIYACVPLVEQTLSPRPPLISVSGVSQG FEFLFGIQDFLYSGQKNDSELNTKLSQLPDLLLQACPFGTLLDANLQNSLDNTNFASVTQPQKQPAWKTGTYKGKPQVSISITEKVKSMQ YDKQGIADTWQVVGTVTCKCDLEGIMPNVTISLSLPTNGSPLQDILVHPCVTSLDSAILTSSSIDAMDDSAFSGPYKFPFTPPLESFNLC FYTSQVPVPPILGFYQMKEEEVQLRITINLKLHESVKNNFEFCEAHIPFYNRGPITHLEYKTSFGQLEVFREKSLLIWIIGQKFPKSMEI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MNAT1-AP5M1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MNAT1-AP5M1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MNAT1-AP5M1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
