
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MUC2-KIRREL3 (FusionGDB2 ID:56022) |
Fusion Gene Summary for MUC2-KIRREL3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MUC2-KIRREL3 | Fusion gene ID: 56022 | Hgene | Tgene | Gene symbol | MUC2 | KIRREL3 | Gene ID | 4583 | 84623 |
| Gene name | mucin 2, oligomeric mucus/gel-forming | kirre like nephrin family adhesion molecule 3 | |
| Synonyms | MLP|MUC-2|SMUC | KIRRE|MRD4|NEPH2|PRO4502 | |
| Cytomap | 11p15.5 | 11q24.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | mucin-2mucin 2, intestinal/tracheal | kin of IRRE-like protein 3kin of IRRE like 3kin of irregular chiasm-like protein 3nephrin-like 2nephrin-like protein 2 | |
| Modification date | 20200320 | 20200313 | |
| UniProtAcc | Q02817 | Q8IZU9 | |
| Ensembl transtripts involved in fusion gene | ENST00000359061, ENST00000441003, ENST00000333592, ENST00000361558, | ENST00000416561, ENST00000525704, ENST00000533026, ENST00000525144, ENST00000529097, | |
| Fusion gene scores | * DoF score | 9 X 9 X 4=324 | 11 X 8 X 4=352 |
| # samples | 9 | 10 | |
| ** MAII score | log2(9/324*10)=-1.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(10/352*10)=-1.81557542886257 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MUC2 [Title/Abstract] AND KIRREL3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MUC2(1092801)-KIRREL3(126294560), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | MUC2-KIRREL3 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. MUC2-KIRREL3 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across MUC2 (5'-gene) Fusion gene breakpoints across MUC2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
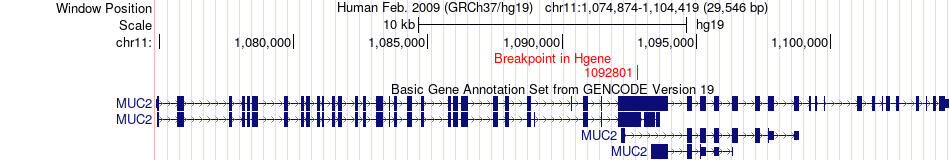 |
 Fusion gene breakpoints across KIRREL3 (3'-gene) Fusion gene breakpoints across KIRREL3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
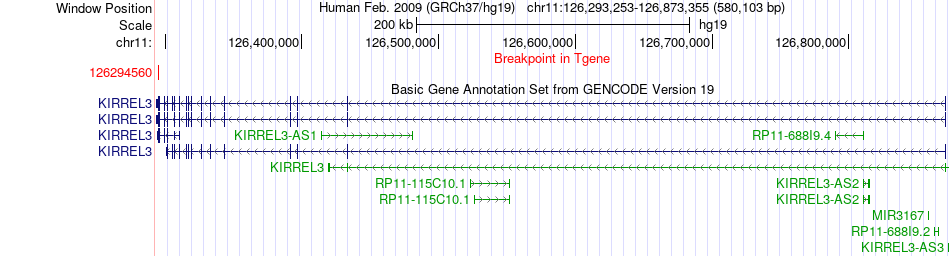 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | COAD | TCGA-AA-3518-01A | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
Top |
Fusion Gene ORF analysis for MUC2-KIRREL3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000359061 | ENST00000416561 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| 5CDS-intron | ENST00000359061 | ENST00000525704 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| 5CDS-intron | ENST00000359061 | ENST00000533026 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| 5CDS-intron | ENST00000441003 | ENST00000416561 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| 5CDS-intron | ENST00000441003 | ENST00000525704 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| 5CDS-intron | ENST00000441003 | ENST00000533026 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| Frame-shift | ENST00000359061 | ENST00000525144 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| Frame-shift | ENST00000359061 | ENST00000529097 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| Frame-shift | ENST00000441003 | ENST00000525144 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| In-frame | ENST00000441003 | ENST00000529097 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-3CDS | ENST00000333592 | ENST00000525144 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-3CDS | ENST00000333592 | ENST00000529097 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-3CDS | ENST00000361558 | ENST00000525144 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-3CDS | ENST00000361558 | ENST00000529097 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-intron | ENST00000333592 | ENST00000416561 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-intron | ENST00000333592 | ENST00000525704 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-intron | ENST00000333592 | ENST00000533026 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-intron | ENST00000361558 | ENST00000416561 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-intron | ENST00000361558 | ENST00000525704 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
| intron-intron | ENST00000361558 | ENST00000533026 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000441003 | MUC2 | chr11 | 1092801 | + | ENST00000529097 | KIRREL3 | chr11 | 126294560 | - | 6228 | 5057 | 15 | 4841 | 1608 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000441003 | ENST00000529097 | MUC2 | chr11 | 1092801 | + | KIRREL3 | chr11 | 126294560 | - | 0.00130668 | 0.9986933 |
Top |
Fusion Genomic Features for MUC2-KIRREL3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
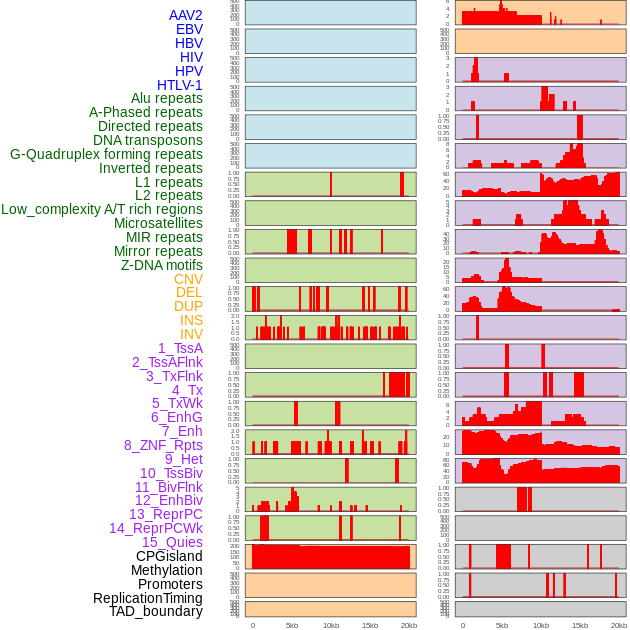 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for MUC2-KIRREL3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:1092801/chr11:126294560) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MUC2 | KIRREL3 |
| FUNCTION: Coats the epithelia of the intestines, airways, and other mucus membrane-containing organs. Thought to provide a protective, lubricating barrier against particles and infectious agents at mucosal surfaces. Major constituent of both the inner and outer mucus layers of the colon and may play a role in excluding bacteria from the inner mucus layer. {ECO:0000269|PubMed:19432394}. | FUNCTION: Synaptic adhesion molecule required for the formation of target-specific synapses. Required for formation of target-specific synapses at hippocampal mossy fiber synapses. Required for formation of mossy fiber filopodia, the synaptic structures connecting dentate granule and GABA neurons. Probably acts as a homophilic adhesion molecule that promotes trans-cellular interactions and stabilize mossy fiber filipodia contact and subsequent synapse formation. Required for the coalescence of vomeronasal sensory neuron axons. May be involved in the hematopoietic supportive capacity of stroma cells; the secreted extracellular domain is directly responsible for supporting hematopoietic stem cells. {ECO:0000250|UniProtKB:Q8BR86}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 701_767 | 0 | 779.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 701_767 | 0 | 601.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 147_243 | 0 | 779.0 | Domain | Note=Ig-like C2-type 2 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 249_330 | 0 | 779.0 | Domain | Note=Ig-like C2-type 3 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 335_415 | 0 | 779.0 | Domain | Note=Ig-like C2-type 4 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 419_515 | 0 | 779.0 | Domain | Note=Ig-like C2-type 5 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 48_142 | 0 | 779.0 | Domain | Note=Ig-like C2-type 1 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 147_243 | 0 | 601.0 | Domain | Note=Ig-like C2-type 2 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 249_330 | 0 | 601.0 | Domain | Note=Ig-like C2-type 3 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 335_415 | 0 | 601.0 | Domain | Note=Ig-like C2-type 4 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 419_515 | 0 | 601.0 | Domain | Note=Ig-like C2-type 5 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 48_142 | 0 | 601.0 | Domain | Note=Ig-like C2-type 1 | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 22_535 | 0 | 779.0 | Topological domain | Extracellular | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 557_778 | 0 | 779.0 | Topological domain | Cytoplasmic | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 22_535 | 0 | 601.0 | Topological domain | Extracellular | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 557_778 | 0 | 601.0 | Topological domain | Cytoplasmic | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525144 | 0 | 17 | 536_556 | 0 | 779.0 | Transmembrane | Helical | |
| Tgene | KIRREL3 | chr11:1092801 | chr11:126294560 | ENST00000525704 | 0 | 14 | 536_556 | 0 | 601.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 295_351 | 0 | 1780.0 | Domain | Note=TIL |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 36_240 | 0 | 1780.0 | Domain | VWFD 1 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 390_604 | 0 | 1780.0 | Domain | VWFD 2 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 4480_4690 | 0 | 1780.0 | Domain | VWFD 4 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 4815_4886 | 0 | 1780.0 | Domain | VWFC 1 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 4924_4991 | 0 | 1780.0 | Domain | VWFC 2 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 5075_5160 | 0 | 1780.0 | Domain | CTCK |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 859_1065 | 0 | 1780.0 | Domain | VWFD 3 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1401_1747 | 0 | 1780.0 | Region | Note=Approximate repeats |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1401_1416 | 0 | 1780.0 | Repeat | Note=1 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1417_1432 | 0 | 1780.0 | Repeat | Note=2 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1433_1448 | 0 | 1780.0 | Repeat | Note=3 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1449_1464 | 0 | 1780.0 | Repeat | Note=4 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1465_1471 | 0 | 1780.0 | Repeat | Note=5 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1472_1478 | 0 | 1780.0 | Repeat | Note=6 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1479_1494 | 0 | 1780.0 | Repeat | Note=7A |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1495_1517 | 0 | 1780.0 | Repeat | Note=7B |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1518_1533 | 0 | 1780.0 | Repeat | Note=8A |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1534_1556 | 0 | 1780.0 | Repeat | Note=8B |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1557_1572 | 0 | 1780.0 | Repeat | Note=9A |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1573_1596 | 0 | 1780.0 | Repeat | Note=9B |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1597_1612 | 0 | 1780.0 | Repeat | Note=10A |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1613_1635 | 0 | 1780.0 | Repeat | Note=10B |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1636_1651 | 0 | 1780.0 | Repeat | Note=11A |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1652_1675 | 0 | 1780.0 | Repeat | Note=11B |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1676_1683 | 0 | 1780.0 | Repeat | Note=12 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1684_1699 | 0 | 1780.0 | Repeat | Note=13 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1700_1715 | 0 | 1780.0 | Repeat | Note=14 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1716_1731 | 0 | 1780.0 | Repeat | Note=15 |
| Hgene | MUC2 | chr11:1092801 | chr11:126294560 | ENST00000359061 | + | 1 | 32 | 1732_1747 | 0 | 1780.0 | Repeat | Note=16 |
Top |
Fusion Gene Sequence for MUC2-KIRREL3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >56022_56022_1_MUC2-KIRREL3_MUC2_chr11_1092801_ENST00000441003_KIRREL3_chr11_126294560_ENST00000529097_length(transcript)=6228nt_BP=5057nt CAACCCACACCGCCCCTGCCAGCCACCATGGGGCTGCCACTAGCCCGCCTGGCGGCTGTGTGCCTGGCCCTGTCTTTGGCAGGGGGCTCG GAGCTCCAGACAGAGGGCAGAACCCGAAACCACGGCCACAACGTCTGCAGCACCTGGGGCAACTTCCACTACAAGACCTTCGACGGGGAC GTCTTCCGCTTCCCCGGCCTCTGCGACTACAACTTCGCCTCCGACTGCCGAGGCTCCTACAAGGAATTTGCTGTGCACCTGAAGCGGGGT CCGGGCCAGGCTGAGGCCCCCGCCGGGGTGGAGTCCATCCTGCTGACCATCAAGGATGACACCATCTACCTCACCCGCCACCTGGCTGTG CTTAACGGGGCCGTGGTCAGCACCCCGCACTACAGCCCCGGGCTGCTCATTGAGAAGAGCGATGCCTACACCAAAGTCTACTCCCGCGCC GGCCTCACCCTCATGTGGAACCGGGAGGATGCACTCATGCTGGAGCTGGACACTAAGTTCCGGAACCACACCTGTGGCCTCTGCGGGGAC TACAACGGCCTGCAGAGCTATTCAGAATTCCTCTCTGACGGCGTGCTCTTCAGTCCCCTGGAGTTTGGGAACATGCAGAAGATCAACCAG CCCGATGTGGTGTGTGAGGATCCCGAGGAGGAGGTGGCCCCCGCATCCTGCTCCGAGCACCGCGCCGAGTGTGAGAGGCTGCTGACCGCC GAGGCCTTCGCGGACTGTCAGGACCTGGTGCCGCTGGAGCCGTATCTGCGCGCCTGCCAGCAGGACCGCTGCCGGTGCCCGGGCGGTGAC ACCTGCGTCTGCAGCACCGTGGCCGAGTTCTCCCGCCAGTGCTCCCACGCCGGCGGCCGGCCCGGGAACTGGAGGACCGCCACGCTCTGC CCCAAGACCTGCCCCGGGAACCTGGTGTACCTGGAGAGCGGCTCGCCCTGCATGGACACCTGCTCACACCTGGAGGTGAGCAGCCTGTGC GAGGAGCACCGCATGGACGGCTGTTTCTGCCCAGAAGGCACCGTATATGACGACATCGGGGACAGTGGCTGCGTTCCTGTGAGCCAGTGC CACTGCAGGCTGCACGGACACCTGTACACACCGGGCCAGGAGATCACCAATGACTGCGAGCAGTGTGTCTGTAACGCTGGCCGCTGGGTG TGCAAAGACCTGCCCTGCCCCGGCACCTGTGCCCTGGAAGGCGGCTCCCACATCACCACCTTCGATGGGAAGACGTACACCTTCCACGGG GACTGCTACTATGTCCTGGCCAAGGGTGACCACAACGATTCCTACGCTCTCCTGGGCGAGCTGGCCCCCTGTGGCTCCACAGACAAGCAG ACCTGCCTGAAGACGGTGGTGCTGCTGGCTGACAAGAAGAAGAATGTGGTGGTCTTCAAGTCCGATGGCAGTGTACTGCTCAACGAGCTG CAGGTGAACCTGCCCCACGTGACCGCGAGCTTCTCTGTCTTCCGCCCGTCTTCCTACCACATCATGGTGAGCATGGCCATTGGCGTCCGG CTGCAGGTGCAGCTGGCCCCAGTCATGCAACTCTTTGTGACACTGGACCAGGCCTCCCAGGGGCAGGTGCAGGGCCTCTGCGGGAACTTC AACGGCCTGGAAGGTGACGACTTCAAGACGGCCAGCGGGCTGGTGGAGGCCACGGGGGCCGGCTTTGCCAACACCTGGAAGGCACAGTCA AGCTGCCATGACAAGCTGGACTGGTTGGACGATCCCTGCTCCCTGAACATCGAGAGCGCCAACTACGCCGAGCACTGGTGCTCCCTCCTG AAGAAGACAGAGACCCCCTTTGGCAGGTGCCACTCGGCTGTGGACCCTGCTGAGTATTACAAGAGGTGCAAATATGACACGTGTAACTGT CAGAACAATGAGGACTGCCTGTGCGCCGCCCTGTCCTCCTACGCGCGCGCCTGCACCGCCAAGGGCGTCATGCTGTGGGGCTGGCGGGAG CATGTCTGCAACAAGGATGTGGGCTCCTGCCCCAACTCGCAGGTCTTCCTGTACAACCTGACCACCTGCCAGCAGACCTGCCGCTCCCTC TCCGAGGCCGACAGCCACTGTCTCGAGGGCTTTGCGCCTGTGGACGGCTGCGGCTGCCCTGACCACACCTTCCTGGACGAGAAGGGCCGC TGCGTACCCCTGGCCAAGTGCTCCTGTTACCACCGCGGTCTCTACCTGGAGGCGGGGGATGTGGTCGTCAGGCAGGAAGAACGATGTGTG TGCCGGGATGGGCGGCTGCACTGTAGGCAGATCCGGCTGATCGGCCAGAGCTGCACGGCCCCAAAGATCCACATGGACTGCAGCAACCTG ACTGCACTGGCCACCTCGAAGCCCCGAGCCCTCAGCTGCCAGACGCTGGCCGCCGGCTATTACCACACAGAGTGTGTCAGTGGCTGTGTG TGCCCCGACGGGCTGATGGATGACGGCCGGGGTGGCTGCGTGGTGGAGAAGGAATGCCCTTGCGTCCATAACAACGACCTGTATTCTTCC GGCGCCAAGATCAAGGTGGACTGCAATACCTGCACCTGCAAGAGAGGACGCTGGGTGTGCACCCAGGCTGTGTGCCATGGCACCTGCTCC ATTTACGGGAGTGGCCACTACATCACCTTTGATGGGAAGTACTACGACTTTGACGGACACTGCTCCTACGTGGCTGTTCAGGACTACTGC GGCCAGAACTCCTCACTGGGCTCATTCAGCATCATCACCGAGAACGTCCCCTGTGGCACTACGGGCGTCACCTGCTCCAAGGCCATCAAG ATCTTCATGGGGAGGACGGAGCTGAAGTTGGAAGACAAGCACCGTGTGGTGATCCAGCGTGATGAGGGTCACCACGTGGCCTACACCACG CGGGAGGTGGGCCAGTACCTGGTGGTGGAGTCCAGCACGGGCATCATCGTCATCTGGGACAAGAGGACCACCGTGTTCATCAAGCTGGCT CCCTCCTACAAGGGCACCGTGTGTGGCCTGTGTGGGAACTTTGACCACCGCTCCAACAACGACTTCACCACGCGGGACCACATGGTGGTG AGCAGCGAGCTGGACTTCGGGAACAGCTGGAAGGAGGCCCCCACCTGCCCAGATGTGAGCACCAACCCCGAGCCCTGCAGCCTGAACCCG CACCGCCGCTCCTGGGCCGAGAAGCAGTGCAGCATCCTCAAAAGCAGCGTGTTCAGCATCTGCCACAGCAAGGTGGACCCCAAGCCCTTC TACGAGGCCTGTGTGCACGACTCGTGCTCCTGTGACACGGGTGGGGACTGTGAGTGCTTCTGCTCTGCCGTGGCCTCCTACGCCCAGGAG TGTACCAAAGAGGGGGCCTGCGTGTTCTGGAGGACGCCGGACCTGTGCCCCATATTCTGCGACTACTACAACCCTCCGCATGAGTGTGAG TGGCACTATGAGCCATGTGGGAACCGGAGCTTCGAGACCTGCAGGACCATCAACGGCATCCACTCCAACATCTCCGTGTCCTACCTGGAG GGCTGCTACCCCCGGTGCCCCAAGGACAGGCCCATCTATGAGGAGGATCTGAAGAAGTGTGTCACTGCAGACAAGTGTGGCTGCTATGTC GAGGACACCCACTACCCACCTGGAGCATCGGTTCCCACCGAGGAGACCTGCAAGTCCTGCGTGTGTACCAACTCCTCCCAAGTCGTCTGC AGGCCGGAGGAAGGAAAGATTCTTAACCAGACCCAGGATGGCGCCTTCTGCTACTGGGAGATCTGTGGCCCCAACGGGACGGTGGAGAAG CACTTCAACATCTGTTCCATTACGACACGCCCGTCCACCCTGACCACCTTCACCACCATCACCCTCCCCACCACCCCCACCACCTTCACC ACTACCACCACCACCACCACCCCGACCTCCAGCACAGTTTTATCAACAACTCCGAACCACCACCACCACGGTGACCCCAACCCCAACACC CACCGGCACACAGACCCCAACAACGACACCCATCACCACCACCACCACGGTGACCCCAACCCCAACACCCACCGGCACACAGACCCCAAC ATCGACACCCATCACCACCACCACTACGGTGACCCCAACCCCAACACCCACCGGCACACAGACCCCAACCACGACACCCATCACCACCAC CACTACGGTGACCCCAACCCCAACACCCACTGGCACACAGACCCCAACCCCAACAGCCATCACCACCACCACTACGGTGACCCCAACCCC AACACCCACCGGCACACAGACCCCAACCACGACACCCATCACCACCACCACTACGGTGACACCAACCCCAACACCCACCGGCACACAGTC CCCAACCCCAACAGCCATCACCACCACCACTACGGTGACCCCAACCCCAACACCCACCGGCACACAGACCCCAACATCGACACCCATCAC CACCACCACTACGGTGACCCCAACCCCAACACCCACCGGCACACAGACCCCAACCCCGACACCCATCTCCACCACCACTACGGTGACCCC AACCCCAACACCCACCGGCACACAGACCCCAACCACGACACCCATCACCACCACCACCACGGTGACCCCAACCCCGACACCCACCGGCAC ACAGACCCCAACCACGGTACTCATCACCACCACCACTACGATGACCCCAACCCCAACACCCACCAGCACAAAGAGTACAACCGTGACACC CATCACCACCACAACTACGGTGACCGCAACCCCAACACCCACCGGCACACAGACCCCAACCATGATACCCATCAGCACCACCACTACGGT GACCCCAACCCCAACACCCACCACTGGAAGCACGGGGCCCCCCACCCACACAAGCACAGCACCCATTGCTGAGTTGACCACATCCAATCC TCCGCCTGAGTCCTCAACCCCTCAGACCTCTCGGTCCACCTCTTCCCCTCTCACGGAGTCAACCACCCTTCTGAGTACCCTACCACCTGC CATTGAGATGACCAGCACGGCCCCACCCTCCACACCCACGGCACCCACGACCACGAGCGGAGGCCACACACTGTCTCCACCGCCCAGCAC CACCACGTCCCCTCCAGCTTCCTCCTCCCACCACTCCCAGTCCTCGTCCCAGAACTCTGACCCCAGTCGACCCCTGCAGCGGCGGATGCA GACTCACGTCTAAGGATCACACACCGCGGGTGGGGACGGGCCAGGGAAGAGGTCAGGGCACGTTCTGGTTGTCCAGGGACGAGGGGTACT TTGCAGAGGACACCAGAATTGGCCACTTCCAGGACAGCCTCCCAGCGCCTCTGCCACTGCCTTCCTTCGAAGCTCTGATCAAGCACAAAT CTGGGTCCCCAGGTGCTGTGTGCCAGAGGTGGGCGGGTGGGGAGACAGACAGAGGCTGCGGCTGAGTGCGCTGTGCTTAGTGCTGGACAC CCGTGTCCCCGGCCCTTTCCTGGAGGCCCCTCTACCACCTGCTCTGCCCACAGGCACAAGTGGCAGCTATAACTCTGCTTTCATGAAACT GCGGTCCACTCTCTGGTCTCTCTGTGGGCTCTACCCCTCGCTGACCAGAAGCTCTACCTACCCCTGTGCCTGTGCTCCCATACAGCCCTG GGGAGAAGGGGATGACGTCTTCCCAGCACTGAGCTGCCCCAGAAACCCCGGCTCCCCACTGCTGCTCATAGCCCATACCCTGGAGGCTGA CAAGCCAGAAATGGCCTTGGCTAAAGGAGCCTCTCTCTCACCAGGCTGGCCGGGAGCCCACCCCCAATTTGTTTGGTGTTTTGTGTCCAT ACTCTTGCAGTTCTGTCCTTGGACTTGATGCCGCTGAACTCTGCGGTGGGACCGGTCCGGTCAGAGCCTGGTGTACTGGGGGGAGGGAGG GAGGAGGGAGCCTGTGCTGACGGAGCACCTCGCCGGGTGTGCCCCTCCTGGGCTGTGTGACCCCAGCCTCCCCACCCACCTCCTGCTTTG TGTACTCCTCCCCTCCCCCTCAGCACAATCGGAGTTCATATAAGAAGTGCGGGAGCTTCTCTGGTCAGGGTTCTCTGAACACTTATGGAG AGAGTGCTTCCTGGGAAGTGTGGCGTTTGAAGGGGCTGGAGGGCAGGTCTTTAAGATGGCGAGACTGCCCTTCTCAGCTGATAAACACAA GAACGGCGATCCTGTCTTCAGTAAGGCTCCACGAGAAGAGAGGAAGTATATCTACACCTCAACCCTCCTAGTCACCACCTGAAATAAATG >56022_56022_1_MUC2-KIRREL3_MUC2_chr11_1092801_ENST00000441003_KIRREL3_chr11_126294560_ENST00000529097_length(amino acids)=1608AA_BP= MPATMGLPLARLAAVCLALSLAGGSELQTEGRTRNHGHNVCSTWGNFHYKTFDGDVFRFPGLCDYNFASDCRGSYKEFAVHLKRGPGQAE APAGVESILLTIKDDTIYLTRHLAVLNGAVVSTPHYSPGLLIEKSDAYTKVYSRAGLTLMWNREDALMLELDTKFRNHTCGLCGDYNGLQ SYSEFLSDGVLFSPLEFGNMQKINQPDVVCEDPEEEVAPASCSEHRAECERLLTAEAFADCQDLVPLEPYLRACQQDRCRCPGGDTCVCS TVAEFSRQCSHAGGRPGNWRTATLCPKTCPGNLVYLESGSPCMDTCSHLEVSSLCEEHRMDGCFCPEGTVYDDIGDSGCVPVSQCHCRLH GHLYTPGQEITNDCEQCVCNAGRWVCKDLPCPGTCALEGGSHITTFDGKTYTFHGDCYYVLAKGDHNDSYALLGELAPCGSTDKQTCLKT VVLLADKKKNVVVFKSDGSVLLNELQVNLPHVTASFSVFRPSSYHIMVSMAIGVRLQVQLAPVMQLFVTLDQASQGQVQGLCGNFNGLEG DDFKTASGLVEATGAGFANTWKAQSSCHDKLDWLDDPCSLNIESANYAEHWCSLLKKTETPFGRCHSAVDPAEYYKRCKYDTCNCQNNED CLCAALSSYARACTAKGVMLWGWREHVCNKDVGSCPNSQVFLYNLTTCQQTCRSLSEADSHCLEGFAPVDGCGCPDHTFLDEKGRCVPLA KCSCYHRGLYLEAGDVVVRQEERCVCRDGRLHCRQIRLIGQSCTAPKIHMDCSNLTALATSKPRALSCQTLAAGYYHTECVSGCVCPDGL MDDGRGGCVVEKECPCVHNNDLYSSGAKIKVDCNTCTCKRGRWVCTQAVCHGTCSIYGSGHYITFDGKYYDFDGHCSYVAVQDYCGQNSS LGSFSIITENVPCGTTGVTCSKAIKIFMGRTELKLEDKHRVVIQRDEGHHVAYTTREVGQYLVVESSTGIIVIWDKRTTVFIKLAPSYKG TVCGLCGNFDHRSNNDFTTRDHMVVSSELDFGNSWKEAPTCPDVSTNPEPCSLNPHRRSWAEKQCSILKSSVFSICHSKVDPKPFYEACV HDSCSCDTGGDCECFCSAVASYAQECTKEGACVFWRTPDLCPIFCDYYNPPHECEWHYEPCGNRSFETCRTINGIHSNISVSYLEGCYPR CPKDRPIYEEDLKKCVTADKCGCYVEDTHYPPGASVPTEETCKSCVCTNSSQVVCRPEEGKILNQTQDGAFCYWEICGPNGTVEKHFNIC SITTRPSTLTTFTTITLPTTPTTFTTTTTTTTPTSSTVLSTTPNHHHHGDPNPNTHRHTDPNNDTHHHHHHGDPNPNTHRHTDPNIDTHH HHHYGDPNPNTHRHTDPNHDTHHHHHYGDPNPNTHWHTDPNPNSHHHHHYGDPNPNTHRHTDPNHDTHHHHHYGDTNPNTHRHTVPNPNS HHHHHYGDPNPNTHRHTDPNIDTHHHHHYGDPNPNTHRHTDPNPDTHLHHHYGDPNPNTHRHTDPNHDTHHHHHHGDPNPDTHRHTDPNH -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MUC2-KIRREL3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MUC2-KIRREL3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MUC2-KIRREL3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
