
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MYRFL-KCNMB4 (FusionGDB2 ID:56841) |
Fusion Gene Summary for MYRFL-KCNMB4 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MYRFL-KCNMB4 | Fusion gene ID: 56841 | Hgene | Tgene | Gene symbol | MYRFL | KCNMB4 | Gene ID | 196446 | 27345 |
| Gene name | myelin regulatory factor like | potassium calcium-activated channel subfamily M regulatory beta subunit 4 | |
| Synonyms | C12orf15|C12orf28|bcm1377 | - | |
| Cytomap | 12q15 | 12q15 | |
| Type of gene | protein-coding | protein-coding | |
| Description | myelin regulatory factor-like protein | calcium-activated potassium channel subunit beta-4BK channel beta subunit 4BK channel subunit beta-4BKbeta4MaxiK channel beta-subunit 4big potassium channel beta subunit 4calcium-activated potassium channel, subfamily M subunit beta-4charybdotoxin | |
| Modification date | 20200322 | 20200313 | |
| UniProtAcc | Q96LU7 | Q86W47 | |
| Ensembl transtripts involved in fusion gene | ENST00000299350, ENST00000547771, ENST00000552032, ENST00000535034, | ENST00000258111, | |
| Fusion gene scores | * DoF score | 3 X 3 X 2=18 | 15 X 8 X 8=960 |
| # samples | 3 | 16 | |
| ** MAII score | log2(3/18*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(16/960*10)=-2.58496250072116 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MYRFL [Title/Abstract] AND KCNMB4 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MYRFL(70321528)-KCNMB4(70824265), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | KCNMB4 | GO:0001508 | action potential | 10692449 |
| Tgene | KCNMB4 | GO:0005513 | detection of calcium ion | 10692449 |
| Tgene | KCNMB4 | GO:0006813 | potassium ion transport | 10692449 |
| Tgene | KCNMB4 | GO:0019228 | neuronal action potential | 10692449 |
 Fusion gene breakpoints across MYRFL (5'-gene) Fusion gene breakpoints across MYRFL (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
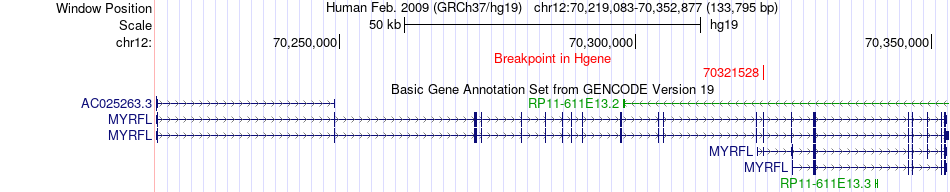 |
 Fusion gene breakpoints across KCNMB4 (3'-gene) Fusion gene breakpoints across KCNMB4 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
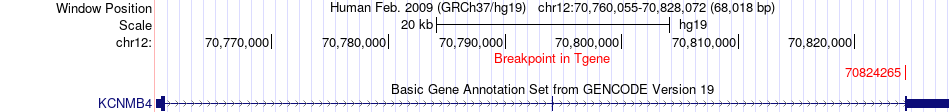 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SARC | TCGA-DX-AB2H-01A | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + |
Top |
Fusion Gene ORF analysis for MYRFL-KCNMB4 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5UTR-3CDS | ENST00000299350 | ENST00000258111 | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + |
| In-frame | ENST00000547771 | ENST00000258111 | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + |
| In-frame | ENST00000552032 | ENST00000258111 | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + |
| intron-3CDS | ENST00000535034 | ENST00000258111 | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000552032 | MYRFL | chr12 | 70321528 | + | ENST00000258111 | KCNMB4 | chr12 | 70824265 | + | 5852 | 2044 | 214 | 2052 | 612 |
| ENST00000547771 | MYRFL | chr12 | 70321528 | + | ENST00000258111 | KCNMB4 | chr12 | 70824265 | + | 5840 | 2032 | 202 | 2040 | 612 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000552032 | ENST00000258111 | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + | 0.001302915 | 0.9986971 |
| ENST00000547771 | ENST00000258111 | MYRFL | chr12 | 70321528 | + | KCNMB4 | chr12 | 70824265 | + | 0.001294578 | 0.9987054 |
Top |
Fusion Genomic Features for MYRFL-KCNMB4 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
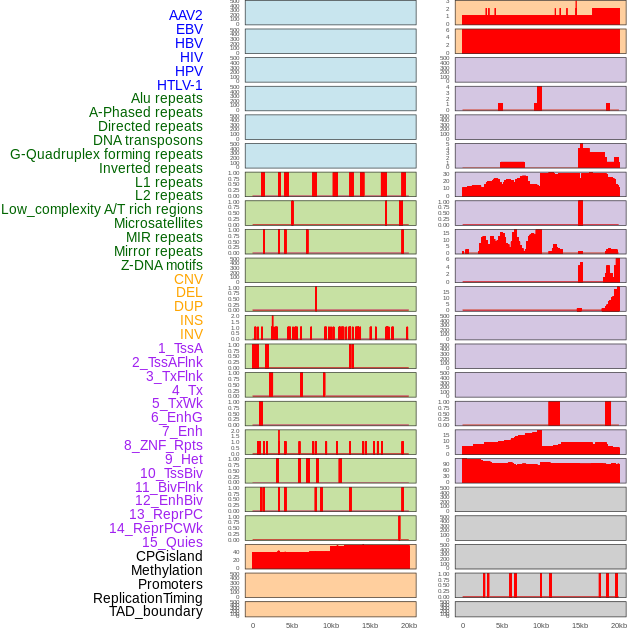 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
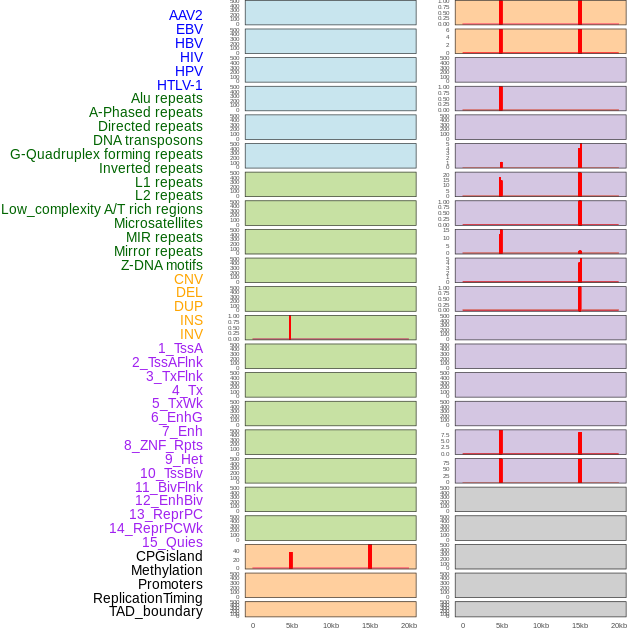 |
Top |
Fusion Protein Features for MYRFL-KCNMB4 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:70321528/chr12:70824265) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MYRFL | KCNMB4 |
| FUNCTION: Regulatory subunit of the calcium activated potassium KCNMA1 (maxiK) channel. Modulates the calcium sensitivity and gating kinetics of KCNMA1, thereby contributing to KCNMA1 channel diversity. Decreases the gating kinetics and calcium sensitivity of the KCNMA1 channel, but with fast deactivation kinetics. May decrease KCNMA1 channel openings at low calcium concentrations but increases channel openings at high calcium concentrations. Makes KCNMA1 channel resistant to 100 nM charybdotoxin (CTX) toxin concentrations. {ECO:0000269|PubMed:10692449, ECO:0000269|PubMed:10792058, ECO:0000269|PubMed:10828459}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MYRFL | chr12:70321528 | chr12:70824265 | ENST00000552032 | + | 15 | 25 | 543_575 | 610 | 911.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | MYRFL | chr12:70321528 | chr12:70824265 | ENST00000552032 | + | 15 | 25 | 142_405 | 610 | 911.0 | DNA binding | NDT80 |
| Hgene | MYRFL | chr12:70321528 | chr12:70824265 | ENST00000552032 | + | 15 | 25 | 451_559 | 610 | 911.0 | Domain | Peptidase S74 |
| Tgene | KCNMB4 | chr12:70321528 | chr12:70824265 | ENST00000258111 | 1 | 3 | 189_210 | 154 | 211.0 | Topological domain | Cytoplasmic | |
| Tgene | KCNMB4 | chr12:70321528 | chr12:70824265 | ENST00000258111 | 1 | 3 | 168_188 | 154 | 211.0 | Transmembrane | Helical%3B Name%3D2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MYRFL | chr12:70321528 | chr12:70824265 | ENST00000552032 | + | 15 | 25 | 628_648 | 610 | 911.0 | Transmembrane | Helical |
| Tgene | KCNMB4 | chr12:70321528 | chr12:70824265 | ENST00000258111 | 1 | 3 | 1_19 | 154 | 211.0 | Topological domain | Cytoplasmic | |
| Tgene | KCNMB4 | chr12:70321528 | chr12:70824265 | ENST00000258111 | 1 | 3 | 41_167 | 154 | 211.0 | Topological domain | Extracellular | |
| Tgene | KCNMB4 | chr12:70321528 | chr12:70824265 | ENST00000258111 | 1 | 3 | 20_40 | 154 | 211.0 | Transmembrane | Helical%3B Name%3D1 |
Top |
Fusion Gene Sequence for MYRFL-KCNMB4 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >56841_56841_1_MYRFL-KCNMB4_MYRFL_chr12_70321528_ENST00000547771_KCNMB4_chr12_70824265_ENST00000258111_length(transcript)=5840nt_BP=2032nt CTTCCAGTTTTATAATCACATTTGAAAACTCAACCATCTTTCTGGGCACAATTTGATGGCTGAAGATATGCTTCACTCCAATAAAAAGAA AATGAAGATTTTTCAAGAGCATTCGTAGGCTTCGAATCAAAAGGACAGTACTTATTTCCTGAGGTCTGATAACCGTTGTTTTATGAGGAA ATAGTACTAGGATCACAGTACCATGGATGTGGTAGGCGAAAATGAGGCCCTGCAGCAGTTCTTTGAAGCTCAGGGAGCCAATGGTACTCT GGAGAACCCAGCCCTGGACACAAGTCTGTTGGAGGAATTTCTGGGCAATGACTTTGATTTGGGGGCCTTGCAACGCCAGCTCCCAGACAC CCCGCCCTATTCTGCATCCGACTCATGCTCTCCTCCGCAGGTCAAAGGTGCATGCTACCCAACCCTGAGGCCCACAGCTGGGAGGACTCC AGCTCCTTTTCTCCACCCCACAGCTGCTCCTGCGATGCCGCCCATGCACCCGCTGCAGAGCACCTCTGGAATGGGCGACTCCTGCCAAAT CCACGGAGGCTTTCACAGCTGCCACTCAAACGCCAGTCATCTTGCCACCCCCCTGGACCAATCCGTGTCCTCCCATCTGGGGATAGGTTG TTCTTACCCTCAGCAGCCTCTGTGTCACAGCCCTGGAGCCTCATTGCCTCCAACCAAGAAGAGAAAGTGCACGCAGGCACTGGAGGACTC CGGGGAATGCCGAGTGTGGGCCTGCCACTGCAGACCGATGACAAGTAGGAGTCGCAGCAGTGAAGTCCAGGACCCTGACAGTGAGGGACA GAACAGAATGCCTACAGACCAGTGCTCTCCTGCTCTGAAGTGGCAACCATGCCATAGTGTTCCTTGGCACAGCTTATTAAACAGTCATTA TGAAAAACTACCTGATGTGGGCTATCGAGTTGTAACAGACAAAGGATTTAATTTTTCACCAGCAGATGAAGCTTTTGTTTGCCAAAAGAA GAATCATTTCCAGATAACCATTCACATCCAAGTTTGGGGAAGTCCAAAATTTGTTGAAACCGAGATGGGCCTAAAGCCAATAGAAATGTT TTACTTGAAAGTTTTTGGCACTAAGGTGGAAGCTACCAATCAAATAATTGCTATTGAACAGTCCCAAGCAGATAGGAGCAAAAAGATTTT CAATCCTGTTAAAATTGACCTACTGGCTGACCAGGTCACCAAAGTAACACTGGGACGATTACACTTCAGCGAAACCACAGCAAATAATAT GAGAAAAAAGGGAAAACCAAATCCAGACCAGAGATACTTCATGTTGGTGGTTGGACTGTATGCTGCTAACCAAGACCAGTTCTATCTGTT GTCTGCCCACATCTCTGAAAGGATCATTGTAAGGGCCTCTAACCCTGGGCAGTTTGAAAATGACAGTGATGCATTGTGGCAGCGAGGACA AGTTCCAGAATCTATTGTCTGTCACGGTCGAGTAGGAATCAACACAGATGCGCCGGACGAAGCCCTGGTTGTCTGTGGCAACATGAAAGT GATGGGGACCATCATGCATCCCTCTGACAGCCGGGCAAAGCAGAATATCCAGGAGGTTGACACGAATGAACAGCTGAAAAGAATAGCCCA AATGAGAATTGTTGAATATGACTACAAACCTGAATTTGCATCTGCAATGGGAATAAACACTGCCCATCAAACAGGAATGATTGCCCAGGA GGTGCAAGAAATCCTGCCCAGAGCAGTAAGAGAGGTTGGTGATGTCACCTGCGGAAACGGAGAGACCTTGGAGAACTTCCTCATGGTGGA TAAGGACCAGATCTTTATGGAAAATGTAGGTGCAGTGAAGCAACTGTGCAAACTAACTAACAACCTTGAGGAAAGAATAGAAGAGTTAGA AATATGGAACAGAAAGCTGGCCCGGCTAAAGCGGCTCAGTAGTTGGAAGTCATCAGCCAGTGAAGCAAGCACAATCAGCAAATCTAGCAG AGCCGTTAGTGCATCTTCTCCAAGAAGGGCCGTTCATAAAAAAAACAACAAGACCAGATGATGTGCTTCTGCATCGCACTCATGATGAGA TTGTCCTCCTGCATTGCTTCCTCTGGCCCCTGGTGACATTTGTGGTGGGCGTTCTCATTGTGGTCCTGACCATCTGTGCCAAGAGCTTGG CGGTCAAGGCGGAAGCCATGAAGAAGCGCAAGTTCTCTTAAAGGGGAAGGAGGCTTGTAGAAAGCAAAGTACAGAAGCTGTACTCATCGG CACGCGTCCACCTGCGGAACCTGTGTTTCCTGGCGCAGGAGATGGACAGGGCCACGACAGGGCTCTGAGAGGCTCATCCCTCAGTGGCAA CAGAAACAGGCACAACTGGAAGACTTGGAACCTCAAAGCTTGTATTCCATCTGCTGTAGCAATGGCTAAAGGGTCAAGATCTTAGCTGTA TGGAGTAACTATTTCAGAAAACCCTATAAGAAGTTCATTTTCTTTCAAAAGTAACAGTATATTATTTGTACAGTGTAGTATACAAACCAT TATGATTTATGCTACTTAAAAATATTAAAATAGAGTGGTCTGTGTTATTTTCTATTTCCTTTTTTATGCTTAGAACACCAGGGTTTAAAA AAAAAAAAAAGGTGAGGACATCTGGGTCTCATTTGCTTCTGCTAGGTTAAACTTTTACTTGACAACAAGGATTCCTGCTGAAGTCTGAAC CTTACTGTGTAACCCTCAGTTTCCACTATTAAAGAGTATCTTTTGACGTCTGCTTGGAAAATGAATAGTATACTGGTAACTCAGTCTCCA GTCACCTCTGTGTCTCTTAAGCAAGAGATTCTAAAAGATTGGGAAAACATATCCTCCAACACCTGCCTTTGCCTAACCATTATTTTTCAC CAGATTACTTCTTAAGAGAGGGAGGTGATTCTGAAGAAGGCTTCTATCTCAAAAAGCACTGGGCTTCCTTATTCATCTGTTCTTGTTGTT TTTGACGGAGTTAAAAAAGTTTGTGTGCAATACAATATACATGATGTGAAGGACACTCTTCAGCTTAGTGAAACGCTGTTTTCATTTTTT TTTTTTTTTGTAGGTCAGAAAAAAACAACAAAATCAGTTCAAGCATTTTTTTTTCTTTGTCCTTGCCTTGATGTTATGAGTATTAAAACC AGGAGGATTGCTGCCATTGTGCAGTTTGCTTAGACAAACCTGGAGATGCAACCCAGCTCACATCATTGCTACTGATGAGCTTTCTGTGCC TTTATCAAAAGTTGATTGAGAAGACCATATTTCTTTGTATCTTTTTATAAACTCAAATTCCAAGTATCAAATCGCAGGTCTCAGTGAACA TCAAACCTATTTACTACATAGAATCAAACCTTTGTTTAGGTGAGATGTACATCGTTAGTGGAGGAAAAACTGACAACCTAATTTCATTTG TTTTCTTCTGATACTCTTCAGACATGCCTCTATTAGAATAAAGGTAAACTGGAATTTAAAGACAAGTTCCCCTCAGTTATTTCCATGGAG CTGTAATATGTATATATGGAGTGATGGTTTCCTGACCTTTAGTCCACATACCAATGTTTTCTTTTTTCTTTTTTTTTTTTTTTTTTGAGA TGGTGTCTCACTCTGTTGCCAGGCTGGAGTGCAGTGGCACGATCTCGGCTCACTACAGTCTCCACCTCCTGGGTTCAAGTCATTCCTCTG CCTCAGCCTCCCGAGTAGCTGGGACTACAGGCACGCACCACCACGCCTGGCTAATTTTTTTGTATTTTTAGTAGAGACGGGGTTTCACCG TGTTAGCCAGGATGGTCTCAATCTCCTGACCTTGTGATCTGCCCACTTCACCTCCCAAAGTGCTGGGATTACAGGCCAATGTTTTCTTAA TCTTAGAATGTGAATAACTGAAAATCATAGTCTGTGGAAAGGTGTTGAATTGAGTATAATCTTCTTCTGTTTATTTTTGTGTTTTGTTTT TTAACAGATGGGTATCTTGCTATGTTGCCCAGGATGGAGTGCAGTAGCTATTCACAGGTATGATCATAGCACACTGCAGCCTCAAGCTCC TGGGCTCAAGCGATCCCCCTCCCTCAGCCTCCCAAGTATCTGGGGTTACTGGTGTGCACCACCGTGCTTGGCTCCAATAATTTTTTTTCT AATTCAAAAGTTACAGTTTCACTGTGAAAAAGGCCTTGAACACACTATTTATGACATCTTTTGAGGCAGCTCCAGTGCCTTGACTTCAAT CCCAGTTTCCGGTTGCAGCATCCTTGTTGTCTTAGCAACACAGTGAACTATTCTGAAGCATAGAGTAACACGAAACTGGGAGTCCGAGAA ATAATCATCTCTGCATCACATTATGGGAGACGAAGTCTGCTTTATCCATTTTATCTTTATTCAGTTGTCTATGATTAATTGATTACAGAG TAGTAGATTAGAATAGTGCATGGATATACATTTGTGTTGAAAAAAGGGGAAGTTGATATATATCAATCTTAGTTTTCATTTATCAGTTTG ATATTCATGCATTTACACTAAACGCTTCCATTTATCCCGAAAAAGTATATGCAACTGTATTCTGTAGGTTGATTTTTGGAAAAGGGGAGA AGCACACTGAATTCATAAGGTCACATGTAGTCTTAAGGTCTTACTTGCTTACAGCCAATTAAATTTGAAGCACCTTATTTATACTTGTTA AAGGTAAAACCCAAAAGAACAAGCAGAGGACATTTTAAGGTCATAAAAGGTAAATAAGCTTACCTTCTTAATGTTTTCATTCTCTTTTTG TATAAATCAGAAAATGATCTAAACTGCTGTAACAAAGAGACCCCAAAATATGATGGCTCATGTAAGATAATTTATTTTTTTCTCACATAG CAATCCAGAAGTGGCTTCATTTCACAAGGTATTCAAGGGATATAGGAGTCATCTACCTTGTTAGTTCTCTTAATACCCAAGGGTATTGTT CTTTCCATGGTCAAAGCTGGCTCAAGACTTCCTAGCCTGTGAAAAAAGAAGAAGGTGGAGCAAGCCATTTCCTTTTTAGGAAATTACAGC CATCACTTCTGCCCACCGTCCATTCATGAATACTTACTATATAGCTATACCTAGCTTCAAGAAAGCCTGGGACGTGTCTCTAACTAGATG GACATGTGCCCTACTAAAACTCCAGGGAAAGGGTTCTATTACTAAAGCTAAAAAGAGGGGAATGAATACTAGAGTTAAAGACAAAAATGA TAGCAGCCAATGGCCCATGCCGTGATAATCTGCTGAGCAGGCATGATGGAGATCCCTTGCCCAGCAGAAAGTGTTCCTTGGTGAAATCAT GAATCTGCTATCTAGGAGAAACTCCCTTGTCCATTGTCTTCTGTGGCCACTAGTTTGACCTCTAGGAAAGTCTTGCTCGTCAGCTTCTGT GGCCCCGTCTGAAACTTTTGAGGGACATCGCAGCTTTTGCAGCCCCTGCTTGCTGGTGCAGACTTTTAGACCTAGATTGCCTTAGAGACT GAAAAATATACGCTTTTATAGGCCGGGGTTTTAGTTCATTTGACTGTAATAAAGACGTCAATGCCGTTTTTAATGTTTGACTGCTGACAT CTTTCAAGACTCACCTTTCCCTTCTCCCTTATGCTGCACATCTGGGCAAGCTGATGGAAGCATGGGTGCCTCCTCCTTTGGCCCCAGCAG GAAGTTCAAATCACGCAAGCCCTGGCATGCATGCAGGAAGCTTCACCCCAGCCTCACACTCTAAGACGGATAAAAGCCAAACCAATTAAG >56841_56841_1_MYRFL-KCNMB4_MYRFL_chr12_70321528_ENST00000547771_KCNMB4_chr12_70824265_ENST00000258111_length(amino acids)=612AA_BP= MDVVGENEALQQFFEAQGANGTLENPALDTSLLEEFLGNDFDLGALQRQLPDTPPYSASDSCSPPQVKGACYPTLRPTAGRTPAPFLHPT AAPAMPPMHPLQSTSGMGDSCQIHGGFHSCHSNASHLATPLDQSVSSHLGIGCSYPQQPLCHSPGASLPPTKKRKCTQALEDSGECRVWA CHCRPMTSRSRSSEVQDPDSEGQNRMPTDQCSPALKWQPCHSVPWHSLLNSHYEKLPDVGYRVVTDKGFNFSPADEAFVCQKKNHFQITI HIQVWGSPKFVETEMGLKPIEMFYLKVFGTKVEATNQIIAIEQSQADRSKKIFNPVKIDLLADQVTKVTLGRLHFSETTANNMRKKGKPN PDQRYFMLVVGLYAANQDQFYLLSAHISERIIVRASNPGQFENDSDALWQRGQVPESIVCHGRVGINTDAPDEALVVCGNMKVMGTIMHP SDSRAKQNIQEVDTNEQLKRIAQMRIVEYDYKPEFASAMGINTAHQTGMIAQEVQEILPRAVREVGDVTCGNGETLENFLMVDKDQIFME -------------------------------------------------------------- >56841_56841_2_MYRFL-KCNMB4_MYRFL_chr12_70321528_ENST00000552032_KCNMB4_chr12_70824265_ENST00000258111_length(transcript)=5852nt_BP=2044nt TGGTTGTATATCCTTCCAGTTTTATAATCACATTTGAAAACTCAACCATCTTTCTGGGCACAATTTGATGGCTGAAGATATGCTTCACTC CAATAAAAAGAAAATGAAGATTTTTCAAGAGCATTCGTAGGCTTCGAATCAAAAGGACAGTACTTATTTCCTGAGGTCTGATAACCGTTG TTTTATGAGGAAATAGTACTAGGATCACAGTACCATGGATGTGGTAGGCGAAAATGAGGCCCTGCAGCAGTTCTTTGAAGCTCAGGGAGC CAATGGTACTCTGGAGAACCCAGCCCTGGACACAAGTCTGTTGGAGGAATTTCTGGGCAATGACTTTGATTTGGGGGCCTTGCAACGCCA GCTCCCAGACACCCCGCCCTATTCTGCATCCGACTCATGCTCTCCTCCGCAGGTCAAAGGTGCATGCTACCCAACCCTGAGGCCCACAGC TGGGAGGACTCCAGCTCCTTTTCTCCACCCCACAGCTGCTCCTGCGATGCCGCCCATGCACCCGCTGCAGAGCACCTCTGGAATGGGCGA CTCCTGCCAAATCCACGGAGGCTTTCACAGCTGCCACTCAAACGCCAGTCATCTTGCCACCCCCCTGGACCAATCCGTGTCCTCCCATCT GGGGATAGGTTGTTCTTACCCTCAGCAGCCTCTGTGTCACAGCCCTGGAGCCTCATTGCCTCCAACCAAGAAGAGAAAGTGCACGCAGGC ACTGGAGGACTCCGGGGAATGCCGAGTGTGGGCCTGCCACTGCAGACCGATGACAAGTAGGAGTCGCAGCAGTGAAGTCCAGGACCCTGA CAGTGAGGGACAGAACAGAATGCCTACAGACCAGTGCTCTCCTGCTCTGAAGTGGCAACCATGCCATAGTGTTCCTTGGCACAGCTTATT AAACAGTCATTATGAAAAACTACCTGATGTGGGCTATCGAGTTGTAACAGACAAAGGATTTAATTTTTCACCAGCAGATGAAGCTTTTGT TTGCCAAAAGAAGAATCATTTCCAGATAACCATTCACATCCAAGTTTGGGGAAGTCCAAAATTTGTTGAAACCGAGATGGGCCTAAAGCC AATAGAAATGTTTTACTTGAAAGTTTTTGGCACTAAGGTGGAAGCTACCAATCAAATAATTGCTATTGAACAGTCCCAAGCAGATAGGAG CAAAAAGATTTTCAATCCTGTTAAAATTGACCTACTGGCTGACCAGGTCACCAAAGTAACACTGGGACGATTACACTTCAGCGAAACCAC AGCAAATAATATGAGAAAAAAGGGAAAACCAAATCCAGACCAGAGATACTTCATGTTGGTGGTTGGACTGTATGCTGCTAACCAAGACCA GTTCTATCTGTTGTCTGCCCACATCTCTGAAAGGATCATTGTAAGGGCCTCTAACCCTGGGCAGTTTGAAAATGACAGTGATGCATTGTG GCAGCGAGGACAAGTTCCAGAATCTATTGTCTGTCACGGTCGAGTAGGAATCAACACAGATGCGCCGGACGAAGCCCTGGTTGTCTGTGG CAACATGAAAGTGATGGGGACCATCATGCATCCCTCTGACAGCCGGGCAAAGCAGAATATCCAGGAGGTTGACACGAATGAACAGCTGAA AAGAATAGCCCAAATGAGAATTGTTGAATATGACTACAAACCTGAATTTGCATCTGCAATGGGAATAAACACTGCCCATCAAACAGGAAT GATTGCCCAGGAGGTGCAAGAAATCCTGCCCAGAGCAGTAAGAGAGGTTGGTGATGTCACCTGCGGAAACGGAGAGACCTTGGAGAACTT CCTCATGGTGGATAAGGACCAGATCTTTATGGAAAATGTAGGTGCAGTGAAGCAACTGTGCAAACTAACTAACAACCTTGAGGAAAGAAT AGAAGAGTTAGAAATATGGAACAGAAAGCTGGCCCGGCTAAAGCGGCTCAGTAGTTGGAAGTCATCAGCCAGTGAAGCAAGCACAATCAG CAAATCTAGCAGAGCCGTTAGTGCATCTTCTCCAAGAAGGGCCGTTCATAAAAAAAACAACAAGACCAGATGATGTGCTTCTGCATCGCA CTCATGATGAGATTGTCCTCCTGCATTGCTTCCTCTGGCCCCTGGTGACATTTGTGGTGGGCGTTCTCATTGTGGTCCTGACCATCTGTG CCAAGAGCTTGGCGGTCAAGGCGGAAGCCATGAAGAAGCGCAAGTTCTCTTAAAGGGGAAGGAGGCTTGTAGAAAGCAAAGTACAGAAGC TGTACTCATCGGCACGCGTCCACCTGCGGAACCTGTGTTTCCTGGCGCAGGAGATGGACAGGGCCACGACAGGGCTCTGAGAGGCTCATC CCTCAGTGGCAACAGAAACAGGCACAACTGGAAGACTTGGAACCTCAAAGCTTGTATTCCATCTGCTGTAGCAATGGCTAAAGGGTCAAG ATCTTAGCTGTATGGAGTAACTATTTCAGAAAACCCTATAAGAAGTTCATTTTCTTTCAAAAGTAACAGTATATTATTTGTACAGTGTAG TATACAAACCATTATGATTTATGCTACTTAAAAATATTAAAATAGAGTGGTCTGTGTTATTTTCTATTTCCTTTTTTATGCTTAGAACAC CAGGGTTTAAAAAAAAAAAAAAGGTGAGGACATCTGGGTCTCATTTGCTTCTGCTAGGTTAAACTTTTACTTGACAACAAGGATTCCTGC TGAAGTCTGAACCTTACTGTGTAACCCTCAGTTTCCACTATTAAAGAGTATCTTTTGACGTCTGCTTGGAAAATGAATAGTATACTGGTA ACTCAGTCTCCAGTCACCTCTGTGTCTCTTAAGCAAGAGATTCTAAAAGATTGGGAAAACATATCCTCCAACACCTGCCTTTGCCTAACC ATTATTTTTCACCAGATTACTTCTTAAGAGAGGGAGGTGATTCTGAAGAAGGCTTCTATCTCAAAAAGCACTGGGCTTCCTTATTCATCT GTTCTTGTTGTTTTTGACGGAGTTAAAAAAGTTTGTGTGCAATACAATATACATGATGTGAAGGACACTCTTCAGCTTAGTGAAACGCTG TTTTCATTTTTTTTTTTTTTTGTAGGTCAGAAAAAAACAACAAAATCAGTTCAAGCATTTTTTTTTCTTTGTCCTTGCCTTGATGTTATG AGTATTAAAACCAGGAGGATTGCTGCCATTGTGCAGTTTGCTTAGACAAACCTGGAGATGCAACCCAGCTCACATCATTGCTACTGATGA GCTTTCTGTGCCTTTATCAAAAGTTGATTGAGAAGACCATATTTCTTTGTATCTTTTTATAAACTCAAATTCCAAGTATCAAATCGCAGG TCTCAGTGAACATCAAACCTATTTACTACATAGAATCAAACCTTTGTTTAGGTGAGATGTACATCGTTAGTGGAGGAAAAACTGACAACC TAATTTCATTTGTTTTCTTCTGATACTCTTCAGACATGCCTCTATTAGAATAAAGGTAAACTGGAATTTAAAGACAAGTTCCCCTCAGTT ATTTCCATGGAGCTGTAATATGTATATATGGAGTGATGGTTTCCTGACCTTTAGTCCACATACCAATGTTTTCTTTTTTCTTTTTTTTTT TTTTTTTTGAGATGGTGTCTCACTCTGTTGCCAGGCTGGAGTGCAGTGGCACGATCTCGGCTCACTACAGTCTCCACCTCCTGGGTTCAA GTCATTCCTCTGCCTCAGCCTCCCGAGTAGCTGGGACTACAGGCACGCACCACCACGCCTGGCTAATTTTTTTGTATTTTTAGTAGAGAC GGGGTTTCACCGTGTTAGCCAGGATGGTCTCAATCTCCTGACCTTGTGATCTGCCCACTTCACCTCCCAAAGTGCTGGGATTACAGGCCA ATGTTTTCTTAATCTTAGAATGTGAATAACTGAAAATCATAGTCTGTGGAAAGGTGTTGAATTGAGTATAATCTTCTTCTGTTTATTTTT GTGTTTTGTTTTTTAACAGATGGGTATCTTGCTATGTTGCCCAGGATGGAGTGCAGTAGCTATTCACAGGTATGATCATAGCACACTGCA GCCTCAAGCTCCTGGGCTCAAGCGATCCCCCTCCCTCAGCCTCCCAAGTATCTGGGGTTACTGGTGTGCACCACCGTGCTTGGCTCCAAT AATTTTTTTTCTAATTCAAAAGTTACAGTTTCACTGTGAAAAAGGCCTTGAACACACTATTTATGACATCTTTTGAGGCAGCTCCAGTGC CTTGACTTCAATCCCAGTTTCCGGTTGCAGCATCCTTGTTGTCTTAGCAACACAGTGAACTATTCTGAAGCATAGAGTAACACGAAACTG GGAGTCCGAGAAATAATCATCTCTGCATCACATTATGGGAGACGAAGTCTGCTTTATCCATTTTATCTTTATTCAGTTGTCTATGATTAA TTGATTACAGAGTAGTAGATTAGAATAGTGCATGGATATACATTTGTGTTGAAAAAAGGGGAAGTTGATATATATCAATCTTAGTTTTCA TTTATCAGTTTGATATTCATGCATTTACACTAAACGCTTCCATTTATCCCGAAAAAGTATATGCAACTGTATTCTGTAGGTTGATTTTTG GAAAAGGGGAGAAGCACACTGAATTCATAAGGTCACATGTAGTCTTAAGGTCTTACTTGCTTACAGCCAATTAAATTTGAAGCACCTTAT TTATACTTGTTAAAGGTAAAACCCAAAAGAACAAGCAGAGGACATTTTAAGGTCATAAAAGGTAAATAAGCTTACCTTCTTAATGTTTTC ATTCTCTTTTTGTATAAATCAGAAAATGATCTAAACTGCTGTAACAAAGAGACCCCAAAATATGATGGCTCATGTAAGATAATTTATTTT TTTCTCACATAGCAATCCAGAAGTGGCTTCATTTCACAAGGTATTCAAGGGATATAGGAGTCATCTACCTTGTTAGTTCTCTTAATACCC AAGGGTATTGTTCTTTCCATGGTCAAAGCTGGCTCAAGACTTCCTAGCCTGTGAAAAAAGAAGAAGGTGGAGCAAGCCATTTCCTTTTTA GGAAATTACAGCCATCACTTCTGCCCACCGTCCATTCATGAATACTTACTATATAGCTATACCTAGCTTCAAGAAAGCCTGGGACGTGTC TCTAACTAGATGGACATGTGCCCTACTAAAACTCCAGGGAAAGGGTTCTATTACTAAAGCTAAAAAGAGGGGAATGAATACTAGAGTTAA AGACAAAAATGATAGCAGCCAATGGCCCATGCCGTGATAATCTGCTGAGCAGGCATGATGGAGATCCCTTGCCCAGCAGAAAGTGTTCCT TGGTGAAATCATGAATCTGCTATCTAGGAGAAACTCCCTTGTCCATTGTCTTCTGTGGCCACTAGTTTGACCTCTAGGAAAGTCTTGCTC GTCAGCTTCTGTGGCCCCGTCTGAAACTTTTGAGGGACATCGCAGCTTTTGCAGCCCCTGCTTGCTGGTGCAGACTTTTAGACCTAGATT GCCTTAGAGACTGAAAAATATACGCTTTTATAGGCCGGGGTTTTAGTTCATTTGACTGTAATAAAGACGTCAATGCCGTTTTTAATGTTT GACTGCTGACATCTTTCAAGACTCACCTTTCCCTTCTCCCTTATGCTGCACATCTGGGCAAGCTGATGGAAGCATGGGTGCCTCCTCCTT TGGCCCCAGCAGGAAGTTCAAATCACGCAAGCCCTGGCATGCATGCAGGAAGCTTCACCCCAGCCTCACACTCTAAGACGGATAAAAGCC AAACCAATTAAGCCGTTTCTCGACCCTCCTGGGAGCCTGCCCTATCTCCCTGGAAAGTCTCAGTATGTGAGTAATAAACCTTTTTATACC >56841_56841_2_MYRFL-KCNMB4_MYRFL_chr12_70321528_ENST00000552032_KCNMB4_chr12_70824265_ENST00000258111_length(amino acids)=612AA_BP= MDVVGENEALQQFFEAQGANGTLENPALDTSLLEEFLGNDFDLGALQRQLPDTPPYSASDSCSPPQVKGACYPTLRPTAGRTPAPFLHPT AAPAMPPMHPLQSTSGMGDSCQIHGGFHSCHSNASHLATPLDQSVSSHLGIGCSYPQQPLCHSPGASLPPTKKRKCTQALEDSGECRVWA CHCRPMTSRSRSSEVQDPDSEGQNRMPTDQCSPALKWQPCHSVPWHSLLNSHYEKLPDVGYRVVTDKGFNFSPADEAFVCQKKNHFQITI HIQVWGSPKFVETEMGLKPIEMFYLKVFGTKVEATNQIIAIEQSQADRSKKIFNPVKIDLLADQVTKVTLGRLHFSETTANNMRKKGKPN PDQRYFMLVVGLYAANQDQFYLLSAHISERIIVRASNPGQFENDSDALWQRGQVPESIVCHGRVGINTDAPDEALVVCGNMKVMGTIMHP SDSRAKQNIQEVDTNEQLKRIAQMRIVEYDYKPEFASAMGINTAHQTGMIAQEVQEILPRAVREVGDVTCGNGETLENFLMVDKDQIFME -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MYRFL-KCNMB4 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MYRFL-KCNMB4 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MYRFL-KCNMB4 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
