
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NAA15-LPAR1 (FusionGDB2 ID:56943) |
Fusion Gene Summary for NAA15-LPAR1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NAA15-LPAR1 | Fusion gene ID: 56943 | Hgene | Tgene | Gene symbol | NAA15 | LPAR1 | Gene ID | 80155 | 1902 |
| Gene name | N-alpha-acetyltransferase 15, NatA auxiliary subunit | lysophosphatidic acid receptor 1 | |
| Synonyms | Ga19|MRD50|NARG1|NAT1P|NATH|TBDN|TBDN100 | EDG2|GPR26|Gpcr26|LPA1|Mrec1.3|VZG1|edg-2|rec.1.3|vzg-1 | |
| Cytomap | 4q31.1 | 9q31.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | N-alpha-acetyltransferase 15, NatA auxiliary subunitN-terminal acetyltransferaseNMDA receptor regulated 1NMDA receptor-regulated protein 1gastric cancer antigen Ga19protein tubedown-1transcriptional coactivator tubedown-100tubedown-1 | lysophosphatidic acid receptor 1LPA receptor 1LPA-1endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2lysophosphatidic acid receptor Edg-2ventricular zone gene 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q9BXJ9 | Q92633 | |
| Ensembl transtripts involved in fusion gene | ENST00000296543, ENST00000398947, ENST00000480277, ENST00000515576, | ENST00000358883, ENST00000374430, ENST00000374431, ENST00000538760, ENST00000541779, | |
| Fusion gene scores | * DoF score | 18 X 10 X 7=1260 | 10 X 6 X 6=360 |
| # samples | 21 | 11 | |
| ** MAII score | log2(21/1260*10)=-2.58496250072116 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/360*10)=-1.71049338280502 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NAA15 [Title/Abstract] AND LPAR1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NAA15(140222985)-LPAR1(113638002), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | NAA15-LPAR1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. NAA15-LPAR1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. NAA15-LPAR1 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | NAA15 | GO:0006474 | N-terminal protein amino acid acetylation | 15496142 |
| Hgene | NAA15 | GO:0045893 | positive regulation of transcription, DNA-templated | 12145306 |
| Tgene | LPAR1 | GO:0007202 | activation of phospholipase C activity | 19306925 |
 Fusion gene breakpoints across NAA15 (5'-gene) Fusion gene breakpoints across NAA15 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
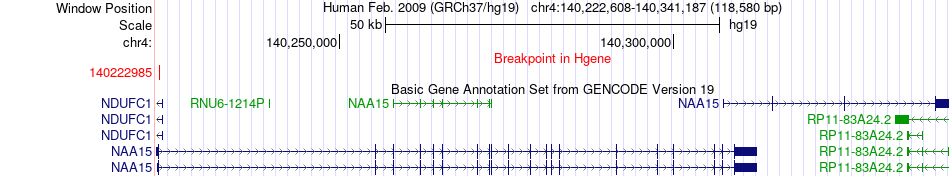 |
 Fusion gene breakpoints across LPAR1 (3'-gene) Fusion gene breakpoints across LPAR1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
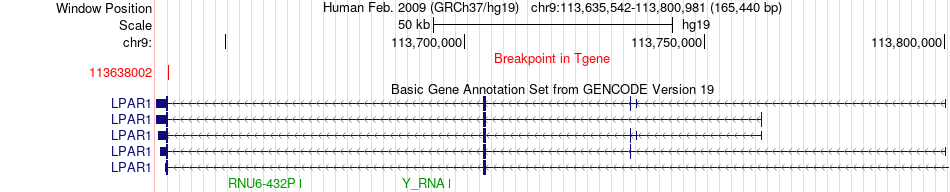 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | GBM | TCGA-06-0129-01A | NAA15 | chr4 | 140222985 | - | LPAR1 | chr9 | 113638002 | - |
| ChimerDB4 | GBM | TCGA-06-0129-01A | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| ChimerDB4 | GBM | TCGA-06-0129 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
Top |
Fusion Gene ORF analysis for NAA15-LPAR1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000296543 | ENST00000358883 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| Frame-shift | ENST00000296543 | ENST00000374430 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| Frame-shift | ENST00000296543 | ENST00000374431 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| Frame-shift | ENST00000398947 | ENST00000358883 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| Frame-shift | ENST00000398947 | ENST00000374430 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| Frame-shift | ENST00000398947 | ENST00000374431 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| In-frame | ENST00000296543 | ENST00000538760 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| In-frame | ENST00000296543 | ENST00000541779 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| In-frame | ENST00000398947 | ENST00000538760 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| In-frame | ENST00000398947 | ENST00000541779 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000480277 | ENST00000358883 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000480277 | ENST00000374430 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000480277 | ENST00000374431 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000480277 | ENST00000538760 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000480277 | ENST00000541779 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000515576 | ENST00000358883 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000515576 | ENST00000374430 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000515576 | ENST00000374431 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000515576 | ENST00000538760 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
| intron-3CDS | ENST00000515576 | ENST00000541779 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000296543 | NAA15 | chr4 | 140222985 | + | ENST00000541779 | LPAR1 | chr9 | 113638002 | - | 2836 | 377 | 277 | 678 | 133 |
| ENST00000296543 | NAA15 | chr4 | 140222985 | + | ENST00000538760 | LPAR1 | chr9 | 113638002 | - | 924 | 377 | 277 | 678 | 133 |
| ENST00000398947 | NAA15 | chr4 | 140222985 | + | ENST00000541779 | LPAR1 | chr9 | 113638002 | - | 2703 | 244 | 144 | 545 | 133 |
| ENST00000398947 | NAA15 | chr4 | 140222985 | + | ENST00000538760 | LPAR1 | chr9 | 113638002 | - | 791 | 244 | 144 | 545 | 133 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000296543 | ENST00000541779 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - | 0.64831036 | 0.35168967 |
| ENST00000296543 | ENST00000538760 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - | 0.37304795 | 0.626952 |
| ENST00000398947 | ENST00000541779 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - | 0.48540062 | 0.5145994 |
| ENST00000398947 | ENST00000538760 | NAA15 | chr4 | 140222985 | + | LPAR1 | chr9 | 113638002 | - | 0.24640578 | 0.75359416 |
Top |
Fusion Genomic Features for NAA15-LPAR1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for NAA15-LPAR1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr4:140222985/chr9:113638002) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| NAA15 | LPAR1 |
| FUNCTION: Auxillary subunit of the N-terminal acetyltransferase A (NatA) complex which displays alpha (N-terminal) acetyltransferase activity. The NAT activity may be important for vascular, hematopoietic and neuronal growth and development. Required to control retinal neovascularization in adult ocular endothelial cells. In complex with XRCC6 and XRCC5 (Ku80), up-regulates transcription from the osteocalcin promoter. {ECO:0000269|PubMed:11687548, ECO:0000269|PubMed:12145306, ECO:0000269|PubMed:15496142}. | FUNCTION: Receptor for lysophosphatidic acid (LPA) (PubMed:9070858, PubMed:19306925, PubMed:25025571, PubMed:26091040). Plays a role in the reorganization of the actin cytoskeleton, cell migration, differentiation and proliferation, and thereby contributes to the responses to tissue damage and infectious agents. Activates downstream signaling cascades via the G(i)/G(o), G(12)/G(13), and G(q) families of heteromeric G proteins. Signaling inhibits adenylyl cyclase activity and decreases cellular cAMP levels (PubMed:26091040). Signaling triggers an increase of cytoplasmic Ca(2+) levels (PubMed:19656035, PubMed:19733258, PubMed:26091040). Activates RALA; this leads to the activation of phospholipase C (PLC) and the formation of inositol 1,4,5-trisphosphate (PubMed:19306925). Signaling mediates activation of down-stream MAP kinases (By similarity). Contributes to the regulation of cell shape. Promotes Rho-dependent reorganization of the actin cytoskeleton in neuronal cells and neurite retraction (PubMed:26091040). Promotes the activation of Rho and the formation of actin stress fibers (PubMed:26091040). Promotes formation of lamellipodia at the leading edge of migrating cells via activation of RAC1 (By similarity). Through its function as lysophosphatidic acid receptor, plays a role in chemotaxis and cell migration, including responses to injury and wounding (PubMed:18066075, PubMed:19656035, PubMed:19733258). Plays a role in triggering inflammation in response to bacterial lipopolysaccharide (LPS) via its interaction with CD14. Promotes cell proliferation in response to lysophosphatidic acid. Required for normal skeleton development. May play a role in osteoblast differentiation. Required for normal brain development. Required for normal proliferation, survival and maturation of newly formed neurons in the adult dentate gyrus. Plays a role in pain perception and in the initiation of neuropathic pain (By similarity). {ECO:0000250|UniProtKB:P61793, ECO:0000269|PubMed:18066075, ECO:0000269|PubMed:19306925, ECO:0000269|PubMed:19656035, ECO:0000269|PubMed:19733258, ECO:0000269|PubMed:25025571, ECO:0000269|PubMed:26091040, ECO:0000269|PubMed:9070858, ECO:0000305|PubMed:11093753, ECO:0000305|PubMed:9069262}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 281_294 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 316_364 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 281_294 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 316_364 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 281_294 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 316_364 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 295_315 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D7 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 295_315 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D7 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 295_315 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D7 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 629_632 | 18 | 867.0 | Compositional bias | Note=Poly-Asp |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 612_629 | 18 | 867.0 | Motif | Bipartite nuclear localization signal |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 148_184 | 18 | 867.0 | Repeat | Note=TPR 3 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 224_257 | 18 | 867.0 | Repeat | Note=TPR 4 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 374_407 | 18 | 867.0 | Repeat | Note=TPR 5 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 409_441 | 18 | 867.0 | Repeat | Note=TPR 6 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 46_79 | 18 | 867.0 | Repeat | Note=TPR 1 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 485_518 | 18 | 867.0 | Repeat | Note=TPR 7 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 672_705 | 18 | 867.0 | Repeat | Note=TPR 8 |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 80_113 | 18 | 867.0 | Repeat | Note=TPR 2 |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 124_129 | 264 | 365.0 | Region | Lysophosphatidic acid binding | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 124_129 | 264 | 365.0 | Region | Lysophosphatidic acid binding | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 124_129 | 264 | 365.0 | Region | Lysophosphatidic acid binding | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 108_121 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 145_163 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 185_204 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 1_50 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 226_255 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 76_83 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 108_121 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 145_163 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 185_204 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 1_50 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 226_255 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 76_83 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 108_121 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 145_163 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 185_204 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 1_50 | 264 | 365.0 | Topological domain | Note=Extracellular | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 226_255 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 76_83 | 264 | 365.0 | Topological domain | Note=Cytoplasmic | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 122_144 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D3 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 164_184 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D4 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 205_225 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D5 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 256_280 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D6 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 51_75 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D1 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000358883 | 2 | 4 | 84_107 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D2 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 122_144 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D3 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 164_184 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D4 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 205_225 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D5 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 256_280 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D6 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 51_75 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D1 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374430 | 3 | 5 | 84_107 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D2 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 122_144 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D3 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 164_184 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D4 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 205_225 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D5 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 256_280 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D6 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 51_75 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D1 | |
| Tgene | LPAR1 | chr4:140222985 | chr9:113638002 | ENST00000374431 | 3 | 5 | 84_107 | 264 | 365.0 | Transmembrane | Note=Helical%3B Name%3D2 |
Top |
Fusion Gene Sequence for NAA15-LPAR1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >56943_56943_1_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000296543_LPAR1_chr9_113638002_ENST00000538760_length(transcript)=924nt_BP=377nt GTGTTAGGAAGCGTGACCCGGGTGGGAAAACCCTCCGCGTCCGCCATTTTGGCTGCCTCTGTCGGTCTGTTCAGTTACCACGTGAACCGC CGACGGAGACCCGTAGTGGGGGAGGCGGCGGCAGCGTTAAGTGAGAAAGGAAAAAAGACAACGAGGAAAAAGGAGGTGTCCGGGTAGGGC AACGCGGCGACACCCGAGGCCTGGTGGTGGCGGCGGATCGAGATATTCAAGGCTGAAGCAGCTACGGAACGGCAGCGGCGGCGGTCGGAC AAACTGACTGACCGAGCCGGGTGGTGGCGGGAGCAGCGGGAGCAGCCGGAACGATGCCGGCCGTGAGCCTCCCGCCCAAGGAGAATGCGC TCTTCAAGCGGATCTTGGGGCCTTTATCATCTGCTGGACTCCTGGATTGGTTTTGTTACTTCTAGACGTGTGCTGTCCACAGTGCGACGT GCTGGCCTATGAGAAATTCTTCCTTCTCCTTGCTGAATTCAACTCTGCCATGAACCCCATCATTTACTCCTACCGCGACAAAGAAATGAG CGCCACCTTTAGGCAGATCCTCTGCTGCCAGCGCAGTGAGAACCCCACCGGCCCCACAGAAGGCTCAGACCGCTCGGCTTCCTCCCTCAA CCACACCATCTTGGCTGGAGTTCACAGCAATGACCACTCTGTGGTTTAGAACGGAAACTGAGATGAGGAACCAGCCGTCCTCTCTTGGAG GATAAACAGCCTCCCCCTACCCAATTGCCAGGGCAAGGTGGGGTGTGAGAGAGGAGAAAAGTCAACTCATGTACTTAAACACTAACCAAT GACAGTATTTGTTCCTGGACCCCACAAGACTTGATATATATTGAAAATTAGCTTATGTGACAACCCTCATCTTGATCCCCATCCCTTCTG >56943_56943_1_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000296543_LPAR1_chr9_113638002_ENST00000538760_length(amino acids)=133AA_BP=33 MTEPGGGGSSGSSRNDAGREPPAQGECALQADLGAFIICWTPGLVLLLLDVCCPQCDVLAYEKFFLLLAEFNSAMNPIIYSYRDKEMSAT -------------------------------------------------------------- >56943_56943_2_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000296543_LPAR1_chr9_113638002_ENST00000541779_length(transcript)=2836nt_BP=377nt GTGTTAGGAAGCGTGACCCGGGTGGGAAAACCCTCCGCGTCCGCCATTTTGGCTGCCTCTGTCGGTCTGTTCAGTTACCACGTGAACCGC CGACGGAGACCCGTAGTGGGGGAGGCGGCGGCAGCGTTAAGTGAGAAAGGAAAAAAGACAACGAGGAAAAAGGAGGTGTCCGGGTAGGGC AACGCGGCGACACCCGAGGCCTGGTGGTGGCGGCGGATCGAGATATTCAAGGCTGAAGCAGCTACGGAACGGCAGCGGCGGCGGTCGGAC AAACTGACTGACCGAGCCGGGTGGTGGCGGGAGCAGCGGGAGCAGCCGGAACGATGCCGGCCGTGAGCCTCCCGCCCAAGGAGAATGCGC TCTTCAAGCGGATCTTGGGGCCTTTATCATCTGCTGGACTCCTGGATTGGTTTTGTTACTTCTAGACGTGTGCTGTCCACAGTGCGACGT GCTGGCCTATGAGAAATTCTTCCTTCTCCTTGCTGAATTCAACTCTGCCATGAACCCCATCATTTACTCCTACCGCGACAAAGAAATGAG CGCCACCTTTAGGCAGATCCTCTGCTGCCAGCGCAGTGAGAACCCCACCGGCCCCACAGAAGGCTCAGACCGCTCGGCTTCCTCCCTCAA CCACACCATCTTGGCTGGAGTTCACAGCAATGACCACTCTGTGGTTTAGAACGGAAACTGAGATGAGGAACCAGCCGTCCTCTCTTGGAG GATAAACAGCCTCCCCCTACCCAATTGCCAGGGCAAGGTGGGGTGTGAGAGAGGAGAAAAGTCAACTCATGTACTTAAACACTAACCAAT GACAGTATTTGTTCCTGGACCCCACAAGACTTGATATATATTGAAAATTAGCTTATGTGACAACCCTCATCTTGATCCCCATCCCTTCTG AAAGTAGGAAGTTGGAGCTCTTGCAATGGAATTCAAGAACAGACTCTGGAGTGTCCATTTAGACTACACTAACTAGACTTTTAAAAGATT TTGTGTGGTTTGGTGCAAGTCAGAATAAATTCTGGCTAGTTGAATCCACAACTTCATTTATATACAGGCTTCCCTTTTTTATTTTTAAAG GATACGTTTCACTTAATAAACACGTTTATGCCTATCAGCATGTTTGTGATGGATGAGACTATGGACTGCTTTTAAACTACCATAATTCCA TTTTTTCCCTTACATAGGAAAACTGTAAGTTGGAATTATCTTTTGTTTAGAAAGCATGCATGTAATGTATGTATGCAGTATGCCTTACTT AAAAAGATTAAAAGGATACTAATGTTAAATCTTCTAGGAAATAGAACCTAGACTTCAAAGCCAGTATTTGTTTAGGTCATGAAGCAAACA ATGCTCTAATCACAATATTAACTGTTTAATTAAAATGTTGTAACAAGTATAAAACAGGGAATGTAAGTTTATTACCAAAGTGATATGTAT TCCAAAAAAGTCATAGAAGATGAAGCACTATAATATTGTTCCCATATATTTAAAATACCCAAGTACATTCTAATTACCAGTATATCAGAG GAAAATTTTCGTAGTCTTTGTAAAATAATATACTCATCATAGAAAACTTGAAAAATGCAGAAATGTATAAAAAAGCAAAAATGATTACTG ATAATATCACAACCCAGAAGTAACCACCTTTAAAAAGCAACCCCCATGTATGCCTATATGTGTATTGTATACTTTTTTTACATAATTGGA GTCATACTGTAAACAGTTTTATAAGTAGATCTTTTTCATTGCAAAATTGCCACATTTTCTTATGGCATTAAAAATTTTACAAAAACATAA TTTTAATGGCTATATTATATTCCATTTAATGGATGCAACTCAGTTTATTTAACCATTCCCATGTTGTTAACTATTTAGGTTGTTTCTAAT TTTCATTATTATAAAGTTGCAGAAATTTGGTGTACATAAAACTGTCTCCATATAATTGATTATTAGGATATATTCCCATGAAGGATTCTT TTTTTAAAAAAATGTGAAATGTCATCTTGTACTTACACCTTTCATGAAAAGGGATTTCCTGCTTTTGTACTGCATGGGTGGCAGTTGTGA GGAAAAGCCAGTCAAATGACCTTTTTACAAAAGAAATGCAGTGGTCACTTCAGTTGAGAGTGACTTTTTAATACAACAAGATCAACTAGA AGAATTCAACTGTCTCAAGAATCAAGGTACCCCAATATATCTCGCAATTCCAAACTTTGTTTGAGGGACTCGTTATCCAGCTCTTGGTAG CCACACCTGCAATGTAAAATGGAAGAAAATGCAAAGAAACCAAATGTGCCGAGTGAATAAAGGATTGTCATATCAATTAGTGTGGGTCCA TCTGTAGCAAATGGGTTGAGTGTGTCAGTATGTGGTTGGTTACTGTGTATTCGCCAGGAATCACCCCGATAGGCTGCCACCCTATTAGGT GATACCTGTTTAATATGTTGTCCAGGTAGACTAGTAGTTGCATCAGTTTGCTGTAACAAGTAACCAGTGAGGTAACACAGTGGTGAAGCA GGTCAGGGGAGGTCAGGAGGATGTCTGAGAGAAAGAAGTCCGGGAGATGAATGGCTGTCTAGGAAGGAGGATGTCAGTGCACGGTTAGTG TTTGAGCAGAGGGCAGACTTGTAAAGTACCTGTAGTGAAAAGAATGTGGGGACCCGATTAGCAGAAAGGTGTTTGCACGTACTTTATACA AAATACAGAATACTTTATATTGGAAGTGAAAGAAATGAACGTGGACTTTTACACATGTGCATATTTTTCTGGAGGCTATATGGATTATTT >56943_56943_2_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000296543_LPAR1_chr9_113638002_ENST00000541779_length(amino acids)=133AA_BP=33 MTEPGGGGSSGSSRNDAGREPPAQGECALQADLGAFIICWTPGLVLLLLDVCCPQCDVLAYEKFFLLLAEFNSAMNPIIYSYRDKEMSAT -------------------------------------------------------------- >56943_56943_3_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000398947_LPAR1_chr9_113638002_ENST00000538760_length(transcript)=791nt_BP=244nt AGAAAGGAAAAAAGACAACGAGGAAAAAGGAGGTGTCCGGGTAGGGCAACGCGGCGACACCCGAGGCCTGGTGGTGGCGGCGGATCGAGA TATTCAAGGCTGAAGCAGCTACGGAACGGCAGCGGCGGCGGTCGGACAAACTGACTGACCGAGCCGGGTGGTGGCGGGAGCAGCGGGAGC AGCCGGAACGATGCCGGCCGTGAGCCTCCCGCCCAAGGAGAATGCGCTCTTCAAGCGGATCTTGGGGCCTTTATCATCTGCTGGACTCCT GGATTGGTTTTGTTACTTCTAGACGTGTGCTGTCCACAGTGCGACGTGCTGGCCTATGAGAAATTCTTCCTTCTCCTTGCTGAATTCAAC TCTGCCATGAACCCCATCATTTACTCCTACCGCGACAAAGAAATGAGCGCCACCTTTAGGCAGATCCTCTGCTGCCAGCGCAGTGAGAAC CCCACCGGCCCCACAGAAGGCTCAGACCGCTCGGCTTCCTCCCTCAACCACACCATCTTGGCTGGAGTTCACAGCAATGACCACTCTGTG GTTTAGAACGGAAACTGAGATGAGGAACCAGCCGTCCTCTCTTGGAGGATAAACAGCCTCCCCCTACCCAATTGCCAGGGCAAGGTGGGG TGTGAGAGAGGAGAAAAGTCAACTCATGTACTTAAACACTAACCAATGACAGTATTTGTTCCTGGACCCCACAAGACTTGATATATATTG >56943_56943_3_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000398947_LPAR1_chr9_113638002_ENST00000538760_length(amino acids)=133AA_BP=33 MTEPGGGGSSGSSRNDAGREPPAQGECALQADLGAFIICWTPGLVLLLLDVCCPQCDVLAYEKFFLLLAEFNSAMNPIIYSYRDKEMSAT -------------------------------------------------------------- >56943_56943_4_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000398947_LPAR1_chr9_113638002_ENST00000541779_length(transcript)=2703nt_BP=244nt AGAAAGGAAAAAAGACAACGAGGAAAAAGGAGGTGTCCGGGTAGGGCAACGCGGCGACACCCGAGGCCTGGTGGTGGCGGCGGATCGAGA TATTCAAGGCTGAAGCAGCTACGGAACGGCAGCGGCGGCGGTCGGACAAACTGACTGACCGAGCCGGGTGGTGGCGGGAGCAGCGGGAGC AGCCGGAACGATGCCGGCCGTGAGCCTCCCGCCCAAGGAGAATGCGCTCTTCAAGCGGATCTTGGGGCCTTTATCATCTGCTGGACTCCT GGATTGGTTTTGTTACTTCTAGACGTGTGCTGTCCACAGTGCGACGTGCTGGCCTATGAGAAATTCTTCCTTCTCCTTGCTGAATTCAAC TCTGCCATGAACCCCATCATTTACTCCTACCGCGACAAAGAAATGAGCGCCACCTTTAGGCAGATCCTCTGCTGCCAGCGCAGTGAGAAC CCCACCGGCCCCACAGAAGGCTCAGACCGCTCGGCTTCCTCCCTCAACCACACCATCTTGGCTGGAGTTCACAGCAATGACCACTCTGTG GTTTAGAACGGAAACTGAGATGAGGAACCAGCCGTCCTCTCTTGGAGGATAAACAGCCTCCCCCTACCCAATTGCCAGGGCAAGGTGGGG TGTGAGAGAGGAGAAAAGTCAACTCATGTACTTAAACACTAACCAATGACAGTATTTGTTCCTGGACCCCACAAGACTTGATATATATTG AAAATTAGCTTATGTGACAACCCTCATCTTGATCCCCATCCCTTCTGAAAGTAGGAAGTTGGAGCTCTTGCAATGGAATTCAAGAACAGA CTCTGGAGTGTCCATTTAGACTACACTAACTAGACTTTTAAAAGATTTTGTGTGGTTTGGTGCAAGTCAGAATAAATTCTGGCTAGTTGA ATCCACAACTTCATTTATATACAGGCTTCCCTTTTTTATTTTTAAAGGATACGTTTCACTTAATAAACACGTTTATGCCTATCAGCATGT TTGTGATGGATGAGACTATGGACTGCTTTTAAACTACCATAATTCCATTTTTTCCCTTACATAGGAAAACTGTAAGTTGGAATTATCTTT TGTTTAGAAAGCATGCATGTAATGTATGTATGCAGTATGCCTTACTTAAAAAGATTAAAAGGATACTAATGTTAAATCTTCTAGGAAATA GAACCTAGACTTCAAAGCCAGTATTTGTTTAGGTCATGAAGCAAACAATGCTCTAATCACAATATTAACTGTTTAATTAAAATGTTGTAA CAAGTATAAAACAGGGAATGTAAGTTTATTACCAAAGTGATATGTATTCCAAAAAAGTCATAGAAGATGAAGCACTATAATATTGTTCCC ATATATTTAAAATACCCAAGTACATTCTAATTACCAGTATATCAGAGGAAAATTTTCGTAGTCTTTGTAAAATAATATACTCATCATAGA AAACTTGAAAAATGCAGAAATGTATAAAAAAGCAAAAATGATTACTGATAATATCACAACCCAGAAGTAACCACCTTTAAAAAGCAACCC CCATGTATGCCTATATGTGTATTGTATACTTTTTTTACATAATTGGAGTCATACTGTAAACAGTTTTATAAGTAGATCTTTTTCATTGCA AAATTGCCACATTTTCTTATGGCATTAAAAATTTTACAAAAACATAATTTTAATGGCTATATTATATTCCATTTAATGGATGCAACTCAG TTTATTTAACCATTCCCATGTTGTTAACTATTTAGGTTGTTTCTAATTTTCATTATTATAAAGTTGCAGAAATTTGGTGTACATAAAACT GTCTCCATATAATTGATTATTAGGATATATTCCCATGAAGGATTCTTTTTTTAAAAAAATGTGAAATGTCATCTTGTACTTACACCTTTC ATGAAAAGGGATTTCCTGCTTTTGTACTGCATGGGTGGCAGTTGTGAGGAAAAGCCAGTCAAATGACCTTTTTACAAAAGAAATGCAGTG GTCACTTCAGTTGAGAGTGACTTTTTAATACAACAAGATCAACTAGAAGAATTCAACTGTCTCAAGAATCAAGGTACCCCAATATATCTC GCAATTCCAAACTTTGTTTGAGGGACTCGTTATCCAGCTCTTGGTAGCCACACCTGCAATGTAAAATGGAAGAAAATGCAAAGAAACCAA ATGTGCCGAGTGAATAAAGGATTGTCATATCAATTAGTGTGGGTCCATCTGTAGCAAATGGGTTGAGTGTGTCAGTATGTGGTTGGTTAC TGTGTATTCGCCAGGAATCACCCCGATAGGCTGCCACCCTATTAGGTGATACCTGTTTAATATGTTGTCCAGGTAGACTAGTAGTTGCAT CAGTTTGCTGTAACAAGTAACCAGTGAGGTAACACAGTGGTGAAGCAGGTCAGGGGAGGTCAGGAGGATGTCTGAGAGAAAGAAGTCCGG GAGATGAATGGCTGTCTAGGAAGGAGGATGTCAGTGCACGGTTAGTGTTTGAGCAGAGGGCAGACTTGTAAAGTACCTGTAGTGAAAAGA ATGTGGGGACCCGATTAGCAGAAAGGTGTTTGCACGTACTTTATACAAAATACAGAATACTTTATATTGGAAGTGAAAGAAATGAACGTG GACTTTTACACATGTGCATATTTTTCTGGAGGCTATATGGATTATTTGAAATGAAAAGCATACTAATTGGACTTTAATAAATCATTTTCA >56943_56943_4_NAA15-LPAR1_NAA15_chr4_140222985_ENST00000398947_LPAR1_chr9_113638002_ENST00000541779_length(amino acids)=133AA_BP=33 MTEPGGGGSSGSSRNDAGREPPAQGECALQADLGAFIICWTPGLVLLLLDVCCPQCDVLAYEKFFLLLAEFNSAMNPIIYSYRDKEMSAT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NAA15-LPAR1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | NAA15 | chr4:140222985 | chr9:113638002 | ENST00000296543 | + | 1 | 20 | 500_866 | 18.0 | 867.0 | HYPK |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NAA15-LPAR1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NAA15-LPAR1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
