
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NCAPH2-GEMIN2 (FusionGDB2 ID:57553) |
Fusion Gene Summary for NCAPH2-GEMIN2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NCAPH2-GEMIN2 | Fusion gene ID: 57553 | Hgene | Tgene | Gene symbol | NCAPH2 | GEMIN2 | Gene ID | 29781 | 8487 |
| Gene name | non-SMC condensin II complex subunit H2 | gem nuclear organelle associated protein 2 | |
| Synonyms | CAPH2 | SIP1|SIP1-delta | |
| Cytomap | 22q13.33 | 14q21.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | condensin-2 complex subunit H2CAP-H2 subunit of the condensin II complexCTA-384D8.36chromosome-associated protein H2kleisin beta | gem-associated protein 2SMN interacting protein 1-deltacomponent of gems 2gemin-2survival of motor neuron protein interacting protein 1 | |
| Modification date | 20200313 | 20200322 | |
| UniProtAcc | Q6IBW4 | O14893 | |
| Ensembl transtripts involved in fusion gene | ENST00000299821, ENST00000395698, ENST00000395701, ENST00000420993, ENST00000520297, | ENST00000250379, ENST00000308317, ENST00000396249, | |
| Fusion gene scores | * DoF score | 5 X 6 X 3=90 | 6 X 7 X 4=168 |
| # samples | 5 | 7 | |
| ** MAII score | log2(5/90*10)=-0.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/168*10)=-1.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NCAPH2 [Title/Abstract] AND GEMIN2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NCAPH2(50954977)-GEMIN2(39601161), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | GEMIN2 | GO:0000387 | spliceosomal snRNP assembly | 18984161 |
 Fusion gene breakpoints across NCAPH2 (5'-gene) Fusion gene breakpoints across NCAPH2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
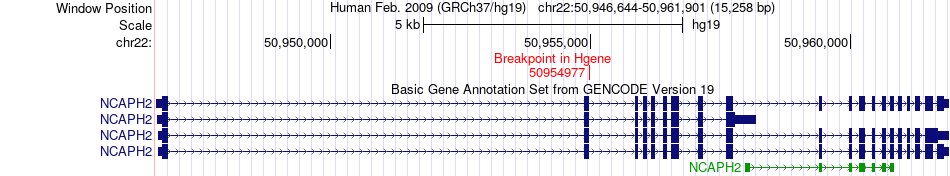 |
 Fusion gene breakpoints across GEMIN2 (3'-gene) Fusion gene breakpoints across GEMIN2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
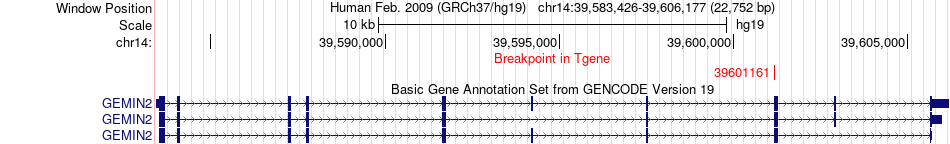 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-HU-A4GY | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
Top |
Fusion Gene ORF analysis for NCAPH2-GEMIN2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000299821 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000299821 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000299821 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000395698 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000395698 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000395698 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000395701 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000395701 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000395701 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000420993 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000420993 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| In-frame | ENST00000420993 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| intron-3CDS | ENST00000520297 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| intron-3CDS | ENST00000520297 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
| intron-3CDS | ENST00000520297 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000420993 | NCAPH2 | chr22 | 50954977 | + | ENST00000308317 | GEMIN2 | chr14 | 39601161 | + | 1036 | 332 | 122 | 541 | 139 |
| ENST00000420993 | NCAPH2 | chr22 | 50954977 | + | ENST00000396249 | GEMIN2 | chr14 | 39601161 | + | 500 | 332 | 122 | 451 | 109 |
| ENST00000420993 | NCAPH2 | chr22 | 50954977 | + | ENST00000250379 | GEMIN2 | chr14 | 39601161 | + | 838 | 332 | 122 | 541 | 139 |
| ENST00000395698 | NCAPH2 | chr22 | 50954977 | + | ENST00000308317 | GEMIN2 | chr14 | 39601161 | + | 1009 | 305 | 95 | 514 | 139 |
| ENST00000395698 | NCAPH2 | chr22 | 50954977 | + | ENST00000396249 | GEMIN2 | chr14 | 39601161 | + | 473 | 305 | 95 | 424 | 109 |
| ENST00000395698 | NCAPH2 | chr22 | 50954977 | + | ENST00000250379 | GEMIN2 | chr14 | 39601161 | + | 811 | 305 | 95 | 514 | 139 |
| ENST00000395701 | NCAPH2 | chr22 | 50954977 | + | ENST00000308317 | GEMIN2 | chr14 | 39601161 | + | 1008 | 304 | 94 | 513 | 139 |
| ENST00000395701 | NCAPH2 | chr22 | 50954977 | + | ENST00000396249 | GEMIN2 | chr14 | 39601161 | + | 472 | 304 | 94 | 423 | 109 |
| ENST00000395701 | NCAPH2 | chr22 | 50954977 | + | ENST00000250379 | GEMIN2 | chr14 | 39601161 | + | 810 | 304 | 94 | 513 | 139 |
| ENST00000299821 | NCAPH2 | chr22 | 50954977 | + | ENST00000308317 | GEMIN2 | chr14 | 39601161 | + | 992 | 288 | 78 | 497 | 139 |
| ENST00000299821 | NCAPH2 | chr22 | 50954977 | + | ENST00000396249 | GEMIN2 | chr14 | 39601161 | + | 456 | 288 | 78 | 407 | 109 |
| ENST00000299821 | NCAPH2 | chr22 | 50954977 | + | ENST00000250379 | GEMIN2 | chr14 | 39601161 | + | 794 | 288 | 78 | 497 | 139 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000420993 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.003700795 | 0.9962992 |
| ENST00000420993 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.016677784 | 0.98332226 |
| ENST00000420993 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.004102974 | 0.99589705 |
| ENST00000395698 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.002962964 | 0.99703705 |
| ENST00000395698 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.012328742 | 0.98767126 |
| ENST00000395698 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.002767575 | 0.9972324 |
| ENST00000395701 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.00267691 | 0.9973231 |
| ENST00000395701 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.011639975 | 0.98836005 |
| ENST00000395701 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.002481654 | 0.9975184 |
| ENST00000299821 | ENST00000308317 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.002914734 | 0.9970853 |
| ENST00000299821 | ENST00000396249 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.013400488 | 0.98659956 |
| ENST00000299821 | ENST00000250379 | NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.003071116 | 0.9969289 |
Top |
Fusion Genomic Features for NCAPH2-GEMIN2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.008866393 | 0.99113363 |
| NCAPH2 | chr22 | 50954977 | + | GEMIN2 | chr14 | 39601161 | + | 0.008866393 | 0.99113363 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for NCAPH2-GEMIN2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr22:50954977/chr14:39601161) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| NCAPH2 | GEMIN2 |
| FUNCTION: Regulatory subunit of the condensin-2 complex, a complex that seems to provide chromosomes with an additional level of organization and rigidity and in establishing mitotic chromosome architecture (PubMed:14532007). May promote the resolution of double-strand DNA catenanes (intertwines) between sister chromatids. Condensin-mediated compaction likely increases tension in catenated sister chromatids, providing directionality for type II topoisomerase-mediated strand exchanges toward chromatid decatenation. Required for decatenation of chromatin bridges at anaphase. Early in neurogenesis, may play an essential role to ensure accurate mitotic chromosome condensation in neuron stem cells, ultimately affecting neuron pool and cortex size (By similarity). Seems to have lineage-specific role in T-cell development (PubMed:14532007). {ECO:0000250|UniProtKB:Q8BSP2, ECO:0000269|PubMed:14532007}. | FUNCTION: The SMN complex plays a catalyst role in the assembly of small nuclear ribonucleoproteins (snRNPs), the building blocks of the spliceosome. Thereby, plays an important role in the splicing of cellular pre-mRNAs. Most spliceosomal snRNPs contain a common set of Sm proteins SNRPB, SNRPD1, SNRPD2, SNRPD3, SNRPE, SNRPF and SNRPG that assemble in a heptameric protein ring on the Sm site of the small nuclear RNA to form the core snRNP. In the cytosol, the Sm proteins SNRPD1, SNRPD2, SNRPE, SNRPF and SNRPG are trapped in an inactive 6S pICln-Sm complex by the chaperone CLNS1A that controls the assembly of the core snRNP. Dissociation by the SMN complex of CLNS1A from the trapped Sm proteins and their transfer to an SMN-Sm complex triggers the assembly of core snRNPs and their transport to the nucleus. {ECO:0000269|PubMed:18984161, ECO:0000269|PubMed:9323129}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | GEMIN2 | chr22:50954977 | chr14:39601161 | ENST00000250379 | 5 | 9 | 101_106 | 196 | 266.0 | Compositional bias | Note=Poly-Gln | |
| Tgene | GEMIN2 | chr22:50954977 | chr14:39601161 | ENST00000308317 | 6 | 10 | 101_106 | 211 | 281.0 | Compositional bias | Note=Poly-Gln | |
| Tgene | GEMIN2 | chr22:50954977 | chr14:39601161 | ENST00000396249 | 6 | 9 | 101_106 | 211 | 251.0 | Compositional bias | Note=Poly-Gln |
Top |
Fusion Gene Sequence for NCAPH2-GEMIN2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >57553_57553_1_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000299821_GEMIN2_chr14_39601161_ENST00000250379_length(transcript)=794nt_BP=288nt AACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCCCGGACATGGAGGACGTG GAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCCAGCTGGGCGAGTATCTG GAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGTTGATCCAGGGCTCTGCC TGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGGCTCATTCACTGATTCGG CAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCAAAGATGATGAGAGGGTTCCTGCTTTGAATTTATTAATCTGCTTG GTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATCTCTCAGGGATAGAAGATATTTCTCATGAAGG CAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCAACACATCTTCAACACTATGTGAAGGGTTCACATCTTAACCTGTG CAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAAATATCAATGGAAAATCCCACTCAGTTTTTGATGAACAGTTTGAA >57553_57553_1_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000299821_GEMIN2_chr14_39601161_ENST00000250379_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_2_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000299821_GEMIN2_chr14_39601161_ENST00000308317_length(transcript)=992nt_BP=288nt AACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCCCGGACATGGAGGACGTG GAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCCAGCTGGGCGAGTATCTG GAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGTTGATCCAGGGCTCTGCC TGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGGCTCATTCACTGATTCGG CAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCAAAGATGATGAGAGGGTTCCTGCTTTGAATTTATTAATCTGCTTG GTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATCTCTCAGGGATAGAAGATATTTCTCATGAAGG CAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCAACACATCTTCAACACTATGTGAAGGGTTCACATCTTAACCTGTG CAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAAATATCAATGGAAAATCCCACTCAGTTTTTGATGAACAGTTTGAA CAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAATTTTACATAGGTTCTTGGTGCTGTTTTGTTCTTTTTTTGTTTTT TGTTGTTTTGTTATTTACTTATATACATATAAAATTTTATTGAAAATATGTTTTGGTTACTAAAATTTTGTTTGACTCCTAACAAAAGAC AATGGATGGCCTTAGCATCAGAATTAAAATAATCTGGATTAAATGGCAATGTGTTCATAGTCAGCAATAAAATTAAACATTTTTCCCTTT >57553_57553_2_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000299821_GEMIN2_chr14_39601161_ENST00000308317_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_3_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000299821_GEMIN2_chr14_39601161_ENST00000396249_length(transcript)=456nt_BP=288nt AACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCCCGGACATGGAGGACGTG GAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCCAGCTGGGCGAGTATCTG GAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGTTGATCCAGGGCTCTGCC TGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGGCTCATTCACTGATTCGG CAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATC >57553_57553_3_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000299821_GEMIN2_chr14_39601161_ENST00000396249_length(amino acids)=109AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_4_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395698_GEMIN2_chr14_39601161_ENST00000250379_length(transcript)=811nt_BP=305nt CAGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCC CGGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCC AGCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGT TGATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGG CTCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCAAAGATGATGAGAGGGTTCCTGCTTTGA ATTTATTAATCTGCTTGGTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATCTCTCAGGGATAGAA GATATTTCTCATGAAGGCAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCAACACATCTTCAACACTATGTGAAGGGT TCACATCTTAACCTGTGCAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAAATATCAATGGAAAATCCCACTCAGTTT TTGATGAACAGTTTGAACAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAATTTTACATAGGTTCTTGGTGCTGTTTT >57553_57553_4_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395698_GEMIN2_chr14_39601161_ENST00000250379_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_5_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395698_GEMIN2_chr14_39601161_ENST00000308317_length(transcript)=1009nt_BP=305nt CAGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCC CGGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCC AGCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGT TGATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGG CTCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCAAAGATGATGAGAGGGTTCCTGCTTTGA ATTTATTAATCTGCTTGGTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATCTCTCAGGGATAGAA GATATTTCTCATGAAGGCAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCAACACATCTTCAACACTATGTGAAGGGT TCACATCTTAACCTGTGCAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAAATATCAATGGAAAATCCCACTCAGTTT TTGATGAACAGTTTGAACAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAATTTTACATAGGTTCTTGGTGCTGTTTT GTTCTTTTTTTGTTTTTTGTTGTTTTGTTATTTACTTATATACATATAAAATTTTATTGAAAATATGTTTTGGTTACTAAAATTTTGTTT GACTCCTAACAAAAGACAATGGATGGCCTTAGCATCAGAATTAAAATAATCTGGATTAAATGGCAATGTGTTCATAGTCAGCAATAAAAT >57553_57553_5_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395698_GEMIN2_chr14_39601161_ENST00000308317_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_6_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395698_GEMIN2_chr14_39601161_ENST00000396249_length(transcript)=473nt_BP=305nt CAGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCC CGGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCC AGCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGT TGATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGG CTCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGTATTTTGACCAACGTGATTTAGCTGATGAGCCA >57553_57553_6_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395698_GEMIN2_chr14_39601161_ENST00000396249_length(amino acids)=109AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_7_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395701_GEMIN2_chr14_39601161_ENST00000250379_length(transcript)=810nt_BP=304nt AGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCCC GGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCCA GCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGTT GATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGGC TCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCAAAGATGATGAGAGGGTTCCTGCTTTGAA TTTATTAATCTGCTTGGTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATCTCTCAGGGATAGAAG ATATTTCTCATGAAGGCAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCAACACATCTTCAACACTATGTGAAGGGTT CACATCTTAACCTGTGCAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAAATATCAATGGAAAATCCCACTCAGTTTT TGATGAACAGTTTGAACAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAATTTTACATAGGTTCTTGGTGCTGTTTTG >57553_57553_7_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395701_GEMIN2_chr14_39601161_ENST00000250379_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_8_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395701_GEMIN2_chr14_39601161_ENST00000308317_length(transcript)=1008nt_BP=304nt AGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCCC GGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCCA GCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGTT GATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGGC TCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCAAAGATGATGAGAGGGTTCCTGCTTTGAA TTTATTAATCTGCTTGGTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTTGATGTAGCTGATCTCTCAGGGATAGAAG ATATTTCTCATGAAGGCAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCAACACATCTTCAACACTATGTGAAGGGTT CACATCTTAACCTGTGCAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAAATATCAATGGAAAATCCCACTCAGTTTT TGATGAACAGTTTGAACAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAATTTTACATAGGTTCTTGGTGCTGTTTTG TTCTTTTTTTGTTTTTTGTTGTTTTGTTATTTACTTATATACATATAAAATTTTATTGAAAATATGTTTTGGTTACTAAAATTTTGTTTG ACTCCTAACAAAAGACAATGGATGGCCTTAGCATCAGAATTAAAATAATCTGGATTAAATGGCAATGTGTTCATAGTCAGCAATAAAATT >57553_57553_8_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395701_GEMIN2_chr14_39601161_ENST00000308317_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_9_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395701_GEMIN2_chr14_39601161_ENST00000396249_length(transcript)=472nt_BP=304nt AGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCTGCGGTCCCTTTGCCGCCCGTTCCCTCCC GGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCAAGAACTGGGAGGTGGACGTGGCGGCCCA GCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCACAATGAACTTCATTGAGGCAGCGTTGTT GATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTTGTCTTGAAAAGCCTTTGTTACCTGAGGC TCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGTATTTTGACCAACGTGATTTAGCTGATGAGCCAT >57553_57553_9_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000395701_GEMIN2_chr14_39601161_ENST00000396249_length(amino acids)=109AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_10_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000420993_GEMIN2_chr14_39601161_ENST00000250379_length(transcript)=838nt_BP=332nt GCGCCTACGCATTTTCCTGGGCGGGAACAGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCT GCGGTCCCTTTGCCGCCCGTTCCCTCCCGGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCA AGAACTGGGAGGTGGACGTGGCGGCCCAGCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCA CAATGAACTTCATTGAGGCAGCGTTGTTGATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTT GTCTTGAAAAGCCTTTGTTACCTGAGGCTCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCA AAGATGATGAGAGGGTTCCTGCTTTGAATTTATTAATCTGCTTGGTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTT GATGTAGCTGATCTCTCAGGGATAGAAGATATTTCTCATGAAGGCAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCA ACACATCTTCAACACTATGTGAAGGGTTCACATCTTAACCTGTGCAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAA ATATCAATGGAAAATCCCACTCAGTTTTTGATGAACAGTTTGAACAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAA >57553_57553_10_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000420993_GEMIN2_chr14_39601161_ENST00000250379_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_11_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000420993_GEMIN2_chr14_39601161_ENST00000308317_length(transcript)=1036nt_BP=332nt GCGCCTACGCATTTTCCTGGGCGGGAACAGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCT GCGGTCCCTTTGCCGCCCGTTCCCTCCCGGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCA AGAACTGGGAGGTGGACGTGGCGGCCCAGCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCA CAATGAACTTCATTGAGGCAGCGTTGTTGATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTT GTCTTGAAAAGCCTTTGTTACCTGAGGCTCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGATAGCA AAGATGATGAGAGGGTTCCTGCTTTGAATTTATTAATCTGCTTGGTTAGCAGGTATTTTGACCAACGTGATTTAGCTGATGAGCCATCTT GATGTAGCTGATCTCTCAGGGATAGAAGATATTTCTCATGAAGGCAGCCTAACTCTGAGGAAAACAATGCCAATTCAAGTACAGATTTCA ACACATCTTCAACACTATGTGAAGGGTTCACATCTTAACCTGTGCAATTCAGATTGATACTCAGAATATGGGTTGATTTGAATATCTGAA ATATCAATGGAAAATCCCACTCAGTTTTTGATGAACAGTTTGAACAGTTTTCTGTAATCAAGCAGCTTGCATAGAAATTGTATGATGAAA TTTTACATAGGTTCTTGGTGCTGTTTTGTTCTTTTTTTGTTTTTTGTTGTTTTGTTATTTACTTATATACATATAAAATTTTATTGAAAA TATGTTTTGGTTACTAAAATTTTGTTTGACTCCTAACAAAAGACAATGGATGGCCTTAGCATCAGAATTAAAATAATCTGGATTAAATGG >57553_57553_11_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000420993_GEMIN2_chr14_39601161_ENST00000308317_length(amino acids)=139AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- >57553_57553_12_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000420993_GEMIN2_chr14_39601161_ENST00000396249_length(transcript)=500nt_BP=332nt GCGCCTACGCATTTTCCTGGGCGGGAACAGCAAAATGGCGCCAGAACTAGTGGCGGGCTGAGGACGCCGTACCCCTCGGAAGGCAGCCCT GCGGTCCCTTTGCCGCCCGTTCCCTCCCGGACATGGAGGACGTGGAGGCGCGCTTCGCCCACCTCTTGCAGCCCATCCGCGACCTCACCA AGAACTGGGAGGTGGACGTGGCGGCCCAGCTGGGCGAGTATCTGGAGGAGCTGGATCAGATCTGCATTTCTTTTGACGAAGGCAAGACCA CAATGAACTTCATTGAGGCAGCGTTGTTGATCCAGGGCTCTGCCTGCGTCTACAGTAAGAAGGGAAGATGGCTTTATGCTTTATTGGCTT GTCTTGAAAAGCCTTTGTTACCTGAGGCTCATTCACTGATTCGGCAGCTTGCAAGAAGGTGCTCTGAAGTGAGGCTCTTAGTGGTATTTT >57553_57553_12_NCAPH2-GEMIN2_NCAPH2_chr22_50954977_ENST00000420993_GEMIN2_chr14_39601161_ENST00000396249_length(amino acids)=109AA_BP=70 MEDVEARFAHLLQPIRDLTKNWEVDVAAQLGEYLEELDQICISFDEGKTTMNFIEAALLIQGSACVYSKKGRWLYALLACLEKPLLPEAH -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NCAPH2-GEMIN2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NCAPH2-GEMIN2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NCAPH2-GEMIN2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
