
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NCOR2-DHX37 (FusionGDB2 ID:57865) |
Fusion Gene Summary for NCOR2-DHX37 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NCOR2-DHX37 | Fusion gene ID: 57865 | Hgene | Tgene | Gene symbol | NCOR2 | DHX37 | Gene ID | 9612 | 57647 |
| Gene name | nuclear receptor corepressor 2 | DEAH-box helicase 37 | |
| Synonyms | CTG26|N-CoR2|SMAP270|SMRT|SMRTE|SMRTE-tau|TNRC14|TRAC|TRAC-1|TRAC1 | DDX37|Dhr1|NEDBAVC | |
| Cytomap | 12q24.31 | 12q24.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | nuclear receptor corepressor 2CTG repeat protein 26T3 receptor-associating factorsilencing mediator for retinoid and thyroid hormone receptorsthyroid-, retinoic-acid-receptor-associated corepressor | probable ATP-dependent RNA helicase DHX37DEAD/DEAH box helicase DDX37DEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 37DEAH (Asp-Glu-Ala-His) box polypeptide 37DEAH box protein 37 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q9Y618 | Q8IY37 | |
| Ensembl transtripts involved in fusion gene | ENST00000356219, ENST00000397355, ENST00000404621, ENST00000405201, ENST00000429285, ENST00000404121, | ENST00000544745, ENST00000308736, | |
| Fusion gene scores | * DoF score | 35 X 38 X 19=25270 | 4 X 6 X 6=144 |
| # samples | 51 | 8 | |
| ** MAII score | log2(51/25270*10)=-5.63078460697328 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(8/144*10)=-0.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NCOR2 [Title/Abstract] AND DHX37 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | DHX37(125467056)-NCOR2(124922542), # samples:2 NCOR2(124950719)-DHX37(125450358), # samples:2 NCOR2(124970987)-DHX37(125470811), # samples:2 NCOR2(124846672)-DHX37(125435350), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | NCOR2-DHX37 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. NCOR2-DHX37 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. NCOR2-DHX37 seems lost the major protein functional domain in Hgene partner, which is a epigenetic factor due to the frame-shifted ORF. NCOR2-DHX37 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. NCOR2-DHX37 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across NCOR2 (5'-gene) Fusion gene breakpoints across NCOR2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
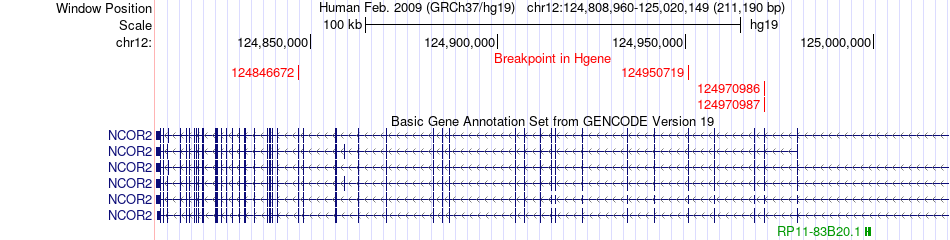 |
 Fusion gene breakpoints across DHX37 (3'-gene) Fusion gene breakpoints across DHX37 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
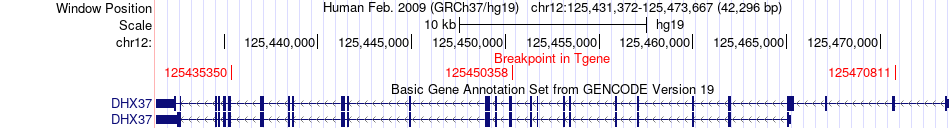 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | GBM | TCGA-06-0878-01A | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| ChimerDB4 | OV | TCGA-13-A5FT-01A | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| ChimerDB4 | OV | TCGA-13-A5FT | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| ChimerDB4 | STAD | TCGA-CG-4449-01A | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
Top |
Fusion Gene ORF analysis for NCOR2-DHX37 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000356219 | ENST00000544745 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000356219 | ENST00000544745 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000397355 | ENST00000544745 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000397355 | ENST00000544745 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000404621 | ENST00000544745 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000404621 | ENST00000544745 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000405201 | ENST00000544745 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000405201 | ENST00000544745 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000429285 | ENST00000544745 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5CDS-intron | ENST00000429285 | ENST00000544745 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| 5UTR-3CDS | ENST00000404121 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| 5UTR-3CDS | ENST00000404121 | ENST00000308736 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5UTR-3CDS | ENST00000404121 | ENST00000308736 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| 5UTR-3CDS | ENST00000404121 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| 5UTR-intron | ENST00000404121 | ENST00000544745 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| 5UTR-intron | ENST00000404121 | ENST00000544745 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000356219 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000356219 | ENST00000308736 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000356219 | ENST00000308736 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000356219 | ENST00000308736 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000356219 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000356219 | ENST00000544745 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000397355 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000397355 | ENST00000308736 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000397355 | ENST00000308736 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000397355 | ENST00000308736 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000397355 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000397355 | ENST00000544745 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000404121 | ENST00000308736 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000404121 | ENST00000544745 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000404621 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000404621 | ENST00000308736 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000404621 | ENST00000308736 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000404621 | ENST00000308736 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000404621 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000404621 | ENST00000544745 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000405201 | ENST00000308736 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000405201 | ENST00000308736 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000405201 | ENST00000308736 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000405201 | ENST00000544745 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000429285 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000429285 | ENST00000308736 | NCOR2 | chr12 | 124970987 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000429285 | ENST00000308736 | NCOR2 | chr12 | 124970986 | - | DHX37 | chr12 | 125470811 | - |
| Frame-shift | ENST00000429285 | ENST00000308736 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| Frame-shift | ENST00000429285 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| Frame-shift | ENST00000429285 | ENST00000544745 | NCOR2 | chr12 | 124846672 | - | DHX37 | chr12 | 125435350 | - |
| In-frame | ENST00000405201 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
| In-frame | ENST00000405201 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000405201 | NCOR2 | chr12 | 124950719 | - | ENST00000308736 | DHX37 | chr12 | 125450358 | - | 3565 | 706 | 1 | 2589 | 862 |
| ENST00000405201 | NCOR2 | chr12 | 124950719 | - | ENST00000544745 | DHX37 | chr12 | 125450358 | - | 3757 | 706 | 1 | 2643 | 880 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000405201 | ENST00000308736 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - | 0.003987408 | 0.99601257 |
| ENST00000405201 | ENST00000544745 | NCOR2 | chr12 | 124950719 | - | DHX37 | chr12 | 125450358 | - | 0.003411051 | 0.99658895 |
Top |
Fusion Genomic Features for NCOR2-DHX37 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
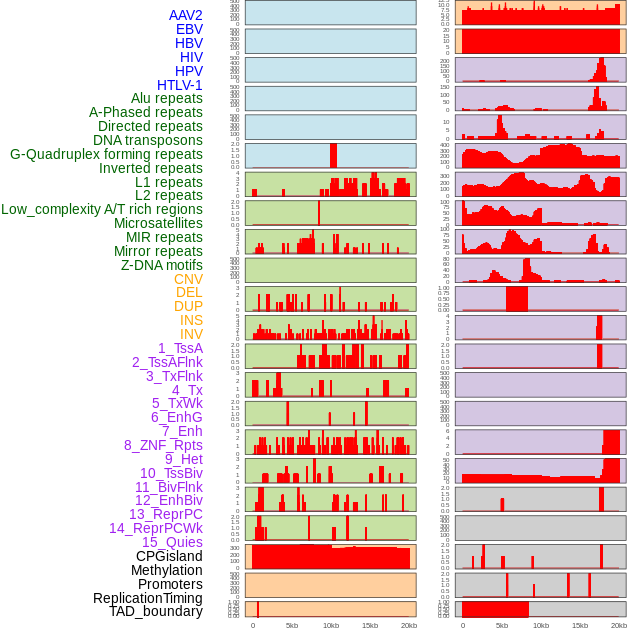 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for NCOR2-DHX37 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:125467056/chr12:124922542) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| NCOR2 | DHX37 |
| FUNCTION: Transcriptional corepressor (PubMed:20812024). Mediates the transcriptional repression activity of some nuclear receptors by promoting chromatin condensation, thus preventing access of the basal transcription. Isoform 1 and isoform 4 have different affinities for different nuclear receptors. Involved in the regulation BCL6-dependent of the germinal center (GC) reactions, mainly through the control of the GC B-cells proliferation and survival. Recruited by ZBTB7A to the androgen response elements/ARE on target genes, negatively regulates androgen receptor signaling and androgen-induced cell proliferation (PubMed:20812024). {ECO:0000269|PubMed:18212045, ECO:0000269|PubMed:20812024, ECO:0000269|PubMed:23911289}. | FUNCTION: ATP-binding RNA helicase that plays a role in maturation of the small ribosomal subunit in ribosome biogenesis (PubMed:30582406). Required for the release of the U3 snoRNP from pre-ribosomal particles (PubMed:30582406). Plays a role in early testis development (PubMed:31287541, PubMed:31337883). Probably plays also a role in brain development (PubMed:31256877). {ECO:0000269|PubMed:30582406, ECO:0000269|PubMed:31256877, ECO:0000269|PubMed:31287541, ECO:0000269|PubMed:31337883}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 174_215 | 235 | 2522.0 | Coiled coil | Ontology_term=ECO:0000255 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 522_561 | 235 | 2522.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 1384_1389 | 235 | 2522.0 | Compositional bias | Note=Poly-Pro |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 1839_1843 | 235 | 2522.0 | Compositional bias | Note=Poly-Gly |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 2476_2479 | 235 | 2522.0 | Compositional bias | Note=Poly-Pro |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 494_510 | 235 | 2522.0 | Compositional bias | Note=Poly-Gln |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 682_685 | 235 | 2522.0 | Compositional bias | Note=Poly-Lys |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 778_820 | 235 | 2522.0 | Compositional bias | Note=Pro-rich |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 995_1003 | 235 | 2522.0 | Compositional bias | Note=Poly-Pro |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 427_478 | 235 | 2522.0 | Domain | SANT 1 |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 610_661 | 235 | 2522.0 | Domain | SANT 2 |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 2136_2140 | 235 | 2522.0 | Motif | Note=CORNR box of ID1 |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 2339_2343 | 235 | 2522.0 | Motif | Note=CORNR box of ID2 |
| Tgene | DHX37 | chr12:124950719 | chr12:125450358 | ENST00000308736 | 11 | 27 | 262_429 | 530 | 1158.0 | Domain | Helicase ATP-binding | |
| Tgene | DHX37 | chr12:124950719 | chr12:125450358 | ENST00000308736 | 11 | 27 | 459_716 | 530 | 1158.0 | Domain | Helicase C-terminal | |
| Tgene | DHX37 | chr12:124950719 | chr12:125450358 | ENST00000308736 | 11 | 27 | 372_375 | 530 | 1158.0 | Motif | Note=DEAH box | |
| Tgene | DHX37 | chr12:124950719 | chr12:125450358 | ENST00000308736 | 11 | 27 | 275_282 | 530 | 1158.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for NCOR2-DHX37 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >57865_57865_1_NCOR2-DHX37_NCOR2_chr12_124950719_ENST00000405201_DHX37_chr12_125450358_ENST00000308736_length(transcript)=3565nt_BP=706nt CATGTCGGGATCCACACAGCCTGTGGCACAGACGTGGAGGGCCACTGAGCCCCGCTACCCGCCCCACAGCCTTTCCTACCCAGTGCAGAT CGCCCGGACGCACACGGACGTCGGGCTCCTGGAGTACCAGCACCACTCCCGCGACTATGCCTCCCACCTGTCGCCCGGCTCCATCATCCA GCCCCAGCGGCGGAGGCCCTCCCTGCTGTCTGAGTTCCAGCCCGGGAATGAACGGTCCCAGGAGCTCCACCTGCGGCCAGAGTCCCACTC ATACCTGCCCGAGCTGGGGAAGTCAGAGATGGAGTTCATTGAAAGCAAGCGCCCTCGGCTAGAGCTGCTGCCTGACCCCCTGCTGCGACC GTCACCCCTGCTGGCCACGGGCCAGCCTGCGGGATCTGAAGACCTCACCAAGGACCGTAGCCTGACGGGCAAGCTGGAACCGGTGTCTCC CCCCAGCCCCCCGCACACTGACCCTGAGCTGGAGCTGGTGCCGCCACGGCTGTCCAAGGAGGAGCTGATCCAGAACATGGACCGCGTGGA CCGAGAGATCACCATGGTAGAGCAGCAGATCTCTAAGCTGAAGAAGAAGCAGCAACAGCTGGAGGAGGAGGCTGCCAAGCCGCCCGAGCC TGAGAAGCCCGTGTCACCGCCGCCCATCGAGTCGAAGCACCGCAGCCTGGTGCAGATCATCTACGACGAGAACCGGGTGCTGCCCCAGAT CAACTTGGATCATTACTCGGTGTTACCGGCAGGCGAAGGCGATGAGGACAGGGAGGCAGAAGTGGATGAGGAAGAGGGGGCCCTGGACTC CGACCTCGATCTGGACCTGGGGGATGGCGGGCAAGATGGAGGTGAGCAGCCGGATGCCTCCCTCCCGCTCCACGTGCTCCCGCTGTACTC TCTGCTGGCCCCAGAGAAGCAAGCACAGGTCTTTAAGCCTCCACCGGAGGGGACTCGGTTGTGTGTTGTGGCCACCAATGTGGCCGAGAC GTCGCTTACCATCCCTGGCATCAAGTACGTGGTGGACTGTGGGAAGGTCAAGAAACGCTACTACGACCGCGTCACTGGCGTATCCTCCTT CCGTGTCACCTGGGTCTCCCAGGCATCAGCTGACCAGCGAGCGGGCAGAGCAGGACGGACGGAGCCCGGCCACTGCTACAGGCTGTATTC ATCTGCGGTTTTTGGTGACTTCGAGCAGTTTCCTCCTCCAGAAATCACCCGGAGGCCTGTTGAAGACTTAATCCTTCAAATGAAGGCGCT CAACGTTGAAAAGGTCATCAACTTCCCCTTCCCGACGCCCCCCTCCGTGGAAGCCCTTCTTGCCGCCGAGGAGCTGTTGATCGCACTGGG TGCCCTGCAACCGCCCCAGAAAGCAGAAAGGGTGAAGCAACTGCAGGAGAACCGGCTGAGCTGCCCCATCACTGCGCTGGGCCGGACAAT GGCCACATTCCCCGTGGCACCCCGCTACGCTAAGATGCTGGCACTGAGCCGACAACACGGCTGCCTGCCCTATGCCATCACCATCGTGGC CAGCATGACGGTGCGGGAGCTGTTTGAGGAGCTGGACAGACCAGCGGCCAGTGACGAGGAGCTCACCAGGCTGAAGAGCAAGCGGGCCCG GGTGGCCCAGATGAAGAGGACCTGGGCAGGGCAGGGGGCTTCTCTGAAGCTCGGCGACCTCATGGTGCTGCTGGGCGCCGTGGGAGCCTG TGAGTATGCCAGCTGCACACCCCAGTTTTGCGAAGCCAACGGGCTGCGGTACAAAGCCATGATGGAGATCCGGCGCCTGCGGGGCCAGCT GACCACCGCAGTCAATGCCGTGTGCCCCGAGGCTGAGCTCTTCGTGGATCCCAAGATGCAGCCGCCCACCGAGAGCCAGGTGACCTACCT GCGACAGATCGTGACGGCAGGCCTGGGGGACCACTTGGCCCGCAGGGTCCAGAGCGAGGAGATGCTGGAGGACAAGTGGAGGAACGCCTA CAAGACCCCTCTCCTCGACGACCCTGTCTTCATCCACCCCAGCTCCGTCCTTTTCAAAGAGCTCCCCGAGTTTGTGGTCTACCAGGAAAT CGTGGAGACCACTAAGATGTACATGAAAGGCGTCTCTAGCGTGGAGGTCCAGTGGATCCCGGCCCTGCTGCCCTCTTACTGCCAGTTTGA CAAGCCCCTGGAGGAACCAGCCCCTACATACTGCCCCGAGCGGGGGCGGGTGCTGTGTCACCGGGCCAGCGTGTTCTATCGCGTGGGCTG GCCGCTCCCCGCCATCGAGGTGGATTTTCCAGAGGGGATTGACCGCTACAAGCACTTTGCCCGGTTCCTGCTGGAAGGGCAGGTCTTCCG CAAGCTGGCCTCATACCGGAGCTGTCTGCTGTCCAGCCCCGGCACCATGCTGAAGACGTGGGCCAGGCTGCAGCCCCGTACGGAGAGCCT TCTGCGAGCCCTGGTTGCAGAGAAGGCTGACTGCCATGAAGCCTTGCTGGCTGCTTGGAAGAAAAACCCCAAATACCTGCTGGCTGAGTA CTGTGAGTGGCTTCCACAGGCCATGCACCCCGATATCGAGAAAGCCTGGCCCCCCACCACTGTCCACTGACCAGAAACCTGGCTGCAGGG CCGAGGACTGGTTTGGGGACTGGAGGGCTGGCAGCAGCCTGTCACCGTGCGACCGTGACCACCTGGCATGGGCTTCGTGGCCTGCTCTCA GGAAGTGGGTCAAGCCCTGGGAACCCTCATCCATGAGAGCTCGATCCCGTATGAAGGGTGCTGCCGCCCGTGCCATCTGGCCCGGGGGTG ACTTTTTGAACTGTTTATTATATGGTGGATGATGATTTCATCTCACGTGCTGGACGCTGTTCTGTTCAGTGTGCTCTTTGGACTACATTA GTCCCCTGTGGAGCAGCAGGGCTGGAGATCTCTGCAGTCCCTTCCCCGCCCGCCCTGCCAGAAGGCCGAGGAGGCACGTGGAGGGCCTCC TTCCTGCAATTCTTCCCTCTCCAGAGTCAGGGAGGGCTGCCCAGCCCTGGCCTCACAGCCGTCCCAGATGTTAGGTGAGCCACTGAGCTC TGTGTTGACCTTGAGGGGCCTGGCTGGGGGCCCCCAGGCTCCATGCCTTCTTGGGAGGGTGGCCGCCAACGCCTTTCCTGTGTTATGGCA ACAGGGAGTGGGCATCTCATCTGCCTGTGGTCAGCTCTCAGACGGCAGGGAGCGGAGCTGACGTTGGCTGTGCTTGGTCACCGCTGCCAT GCCGCAGAGGATGCGCCTAGCTGGGCTGGGGCCACACGACTATTATGTTGGCCTTGAACGGGGACTGCAGAGCCCTCAGTTTGTCTCCCT TGTTCCTCTGTGGCTGAGGTGGGAGGGGGAGGGTGGGGTAGGTCCCCCAGCAAGAAAGAGGGACAGGAGCACCCCAGGCAGGACCAAGGA GTCGGGAGGCCCCTGCCTTCTGTCCTCCATGGTGAGGGCACAGATGTCTCCCCAGAGCCCAGCGCTGGCAGAATGGATTCTGCTCCTGGC >57865_57865_1_NCOR2-DHX37_NCOR2_chr12_124950719_ENST00000405201_DHX37_chr12_125450358_ENST00000308736_length(amino acids)=862AA_BP=235 MSGSTQPVAQTWRATEPRYPPHSLSYPVQIARTHTDVGLLEYQHHSRDYASHLSPGSIIQPQRRRPSLLSEFQPGNERSQELHLRPESHS YLPELGKSEMEFIESKRPRLELLPDPLLRPSPLLATGQPAGSEDLTKDRSLTGKLEPVSPPSPPHTDPELELVPPRLSKEELIQNMDRVD REITMVEQQISKLKKKQQQLEEEAAKPPEPEKPVSPPPIESKHRSLVQIIYDENRVLPQINLDHYSVLPAGEGDEDREAEVDEEEGALDS DLDLDLGDGGQDGGEQPDASLPLHVLPLYSLLAPEKQAQVFKPPPEGTRLCVVATNVAETSLTIPGIKYVVDCGKVKKRYYDRVTGVSSF RVTWVSQASADQRAGRAGRTEPGHCYRLYSSAVFGDFEQFPPPEITRRPVEDLILQMKALNVEKVINFPFPTPPSVEALLAAEELLIALG ALQPPQKAERVKQLQENRLSCPITALGRTMATFPVAPRYAKMLALSRQHGCLPYAITIVASMTVRELFEELDRPAASDEELTRLKSKRAR VAQMKRTWAGQGASLKLGDLMVLLGAVGACEYASCTPQFCEANGLRYKAMMEIRRLRGQLTTAVNAVCPEAELFVDPKMQPPTESQVTYL RQIVTAGLGDHLARRVQSEEMLEDKWRNAYKTPLLDDPVFIHPSSVLFKELPEFVVYQEIVETTKMYMKGVSSVEVQWIPALLPSYCQFD KPLEEPAPTYCPERGRVLCHRASVFYRVGWPLPAIEVDFPEGIDRYKHFARFLLEGQVFRKLASYRSCLLSSPGTMLKTWARLQPRTESL -------------------------------------------------------------- >57865_57865_2_NCOR2-DHX37_NCOR2_chr12_124950719_ENST00000405201_DHX37_chr12_125450358_ENST00000544745_length(transcript)=3757nt_BP=706nt CATGTCGGGATCCACACAGCCTGTGGCACAGACGTGGAGGGCCACTGAGCCCCGCTACCCGCCCCACAGCCTTTCCTACCCAGTGCAGAT CGCCCGGACGCACACGGACGTCGGGCTCCTGGAGTACCAGCACCACTCCCGCGACTATGCCTCCCACCTGTCGCCCGGCTCCATCATCCA GCCCCAGCGGCGGAGGCCCTCCCTGCTGTCTGAGTTCCAGCCCGGGAATGAACGGTCCCAGGAGCTCCACCTGCGGCCAGAGTCCCACTC ATACCTGCCCGAGCTGGGGAAGTCAGAGATGGAGTTCATTGAAAGCAAGCGCCCTCGGCTAGAGCTGCTGCCTGACCCCCTGCTGCGACC GTCACCCCTGCTGGCCACGGGCCAGCCTGCGGGATCTGAAGACCTCACCAAGGACCGTAGCCTGACGGGCAAGCTGGAACCGGTGTCTCC CCCCAGCCCCCCGCACACTGACCCTGAGCTGGAGCTGGTGCCGCCACGGCTGTCCAAGGAGGAGCTGATCCAGAACATGGACCGCGTGGA CCGAGAGATCACCATGGTAGAGCAGCAGATCTCTAAGCTGAAGAAGAAGCAGCAACAGCTGGAGGAGGAGGCTGCCAAGCCGCCCGAGCC TGAGAAGCCCGTGTCACCGCCGCCCATCGAGTCGAAGCACCGCAGCCTGGTGCAGATCATCTACGACGAGAACCGGGTGCTGCCCCAGAT CAACTTGGATCATTACTCGGTGTTACCGGCAGGCGAAGGCGATGAGGACAGGGAGGCAGAAGTGGATGAGGAAGAGGGGGCCCTGGACTC CGACCTCGATCTGGACCTGGGGGATGGCGGGCAAGATGGAGGTGAGCAGCCGGATGCCTCCCTCCCGCTCCACGTGCTCCCGCTGTACTC TCTGCTGGCCCCAGAGAAGCAAGCACAGGTCTTTAAGCCTCCACCGGAGGGGACTCGGTTGTGTGTTGTGGCCACCAATGTGGCCGAGAC GTCGCTTACCATCCCTGGCATCAAGTACGTGGTGGACTGTGGGAAGGTCAAGAAACGCTACTACGACCGCGTCACTGGCGTATCCTCCTT CCGTGTCACCTGGGTCTCCCAGGCATCAGCTGACCAGCGAGCGGGCAGAGCAGGACGGACGGAGCCCGGCCACTGCTACAGGCTGTATTC ATCTGCGGTTTTTGGTGACTTCGAGCAGTTTCCTCCTCCAGAAATCACCCGGAGGCCTGTTGAAGACTTAATCCTTCAAATGAAGGCGCT CAACGTTGAAAAGGTCATCAACTTCCCCTTCCCGACGCCCCCCTCCGTGGAAGCCCTTCTTGCCGCCGAGGAGCTGTTGATCGCACTGGG TGCCCTGCAACCGCCCCAGAAAGCAGAAAGGGTGAAGCAACTGCAGGAGAACCGGCTGAGCTGCCCCATCACTGCGCTGGGCCGGACAAT GGCCACATTCCCCGTGGCACCCCGCTACGCTAAGATGCTGGCACTGAGCCGACAACACGGCTGCCTGCCCTATGCCATCACCATCGTGGC CAGCATGACGGTGCGGGAGCTGTTTGAGGAGCTGGACAGACCAGCGGCCAGTGACGAGGAGCTCACCAGGCTGAAGAGCAAGCGGGCCCG GGTGGCCCAGATGAAGAGGACCTGGGCAGGGCAGGGGGCTTCTCTGAAGCTCGGCGACCTCATGGTGCTGCTGGGCGCCGTGGGAGCCTG TGAGTATGCCAGCTGCACACCCCAGTTTTGCGAAGCCAACGGGCTGCGGTACAAAGCCATGATGGAGATCCGGCGCCTGCGGGGCCAGCT GACCACCGCAGTCAATGCCGTGTGCCCCGAGGCTGAGCTCTTCGTGGATCCCAAGATGCAGCCGCCCACCGAGAGCCAGGTGACCTACCT GCGACAGATCGTGACGGCAGGCCTGGGGGACCACTTGGCCCGCAGGGTCCAGAGCGAGGAGATGCTGGAGGACAAGTGGAGGAACGCCTA CAAGACCCCTCTCCTCGACGACCCTGTCTTCATCCACCCCAGCTCCGTCCTTTTCAAAGAGCTCCCCGAGTTTGTGGTCTACCAGGAAAT CGTGGAGACCACTAAGATGTACATGAAAGGCGTCTCTAGCGTGGAGGTCCAGTGGATCCCGGCCCTGCTGCCCTCTTACTGCCAGTTTGA CAAGCCCCTGGAGGAACCAGCCCCTACATACTGCCCCGAGCGGGGGCGGGTGCTGTGTCACCGGGCCAGCGTGTTCTATCGCGTGGGCTG GCCGCTCCCCGCCATCGAGGTGGATTTTCCAGAGGGGATTGACCGCTACAAGCACTTTGCCCGGTTCCTGCTGGAAGGGCAGGTCTTCCG CAAGCTGGCCTCATACCGGAGCTGTCTGCTGTCCAGCCCCGGCACCATGCTGAAGACGTGGGCCAGGCTGCAGCCCCGTACGGAGAGCCT TCTGCGAGCCCTGGTTGCAGAGAAGGCTGACTGCCATGAAGCCTTGCTGGCTGCTTGGAAGAAAAACCCCAAATGTGAGTTTGACCAGGG CCAGGGGGTGGGAGTAGACAGGATGGGGTCACTGAGGCAGGGGCTTTGTGCTTTGTGCACGGTCAGCCCTGGCTTGGCAGAAGGATCTGG CCCCACGGCTGCTGGCCAGCTTTTTGCCACCTGAGGAGAGTCCCTGGGCCCAGGGTCACCCTGACACTGTCCTCCTTCCCTCCCCTGCAG ACCTGCTGGCTGAGTACTGTGAGTGGCTTCCACAGGCCATGCACCCCGATATCGAGAAAGCCTGGCCCCCCACCACTGTCCACTGACCAG AAACCTGGCTGCAGGGCCGAGGACTGGTTTGGGGACTGGAGGGCTGGCAGCAGCCTGTCACCGTGCGACCGTGACCACCTGGCATGGGCT TCGTGGCCTGCTCTCAGGAAGTGGGTCAAGCCCTGGGAACCCTCATCCATGAGAGCTCGATCCCGTATGAAGGGTGCTGCCGCCCGTGCC ATCTGGCCCGGGGGTGACTTTTTGAACTGTTTATTATATGGTGGATGATGATTTCATCTCACGTGCTGGACGCTGTTCTGTTCAGTGTGC TCTTTGGACTACATTAGTCCCCTGTGGAGCAGCAGGGCTGGAGATCTCTGCAGTCCCTTCCCCGCCCGCCCTGCCAGAAGGCCGAGGAGG CACGTGGAGGGCCTCCTTCCTGCAATTCTTCCCTCTCCAGAGTCAGGGAGGGCTGCCCAGCCCTGGCCTCACAGCCGTCCCAGATGTTAG GTGAGCCACTGAGCTCTGTGTTGACCTTGAGGGGCCTGGCTGGGGGCCCCCAGGCTCCATGCCTTCTTGGGAGGGTGGCCGCCAACGCCT TTCCTGTGTTATGGCAACAGGGAGTGGGCATCTCATCTGCCTGTGGTCAGCTCTCAGACGGCAGGGAGCGGAGCTGACGTTGGCTGTGCT TGGTCACCGCTGCCATGCCGCAGAGGATGCGCCTAGCTGGGCTGGGGCCACACGACTATTATGTTGGCCTTGAACGGGGACTGCAGAGCC CTCAGTTTGTCTCCCTTGTTCCTCTGTGGCTGAGGTGGGAGGGGGAGGGTGGGGTAGGTCCCCCAGCAAGAAAGAGGGACAGGAGCACCC CAGGCAGGACCAAGGAGTCGGGAGGCCCCTGCCTTCTGTCCTCCATGGTGAGGGCACAGATGTCTCCCCAGAGCCCAGCGCTGGCAGAAT >57865_57865_2_NCOR2-DHX37_NCOR2_chr12_124950719_ENST00000405201_DHX37_chr12_125450358_ENST00000544745_length(amino acids)=880AA_BP=235 MSGSTQPVAQTWRATEPRYPPHSLSYPVQIARTHTDVGLLEYQHHSRDYASHLSPGSIIQPQRRRPSLLSEFQPGNERSQELHLRPESHS YLPELGKSEMEFIESKRPRLELLPDPLLRPSPLLATGQPAGSEDLTKDRSLTGKLEPVSPPSPPHTDPELELVPPRLSKEELIQNMDRVD REITMVEQQISKLKKKQQQLEEEAAKPPEPEKPVSPPPIESKHRSLVQIIYDENRVLPQINLDHYSVLPAGEGDEDREAEVDEEEGALDS DLDLDLGDGGQDGGEQPDASLPLHVLPLYSLLAPEKQAQVFKPPPEGTRLCVVATNVAETSLTIPGIKYVVDCGKVKKRYYDRVTGVSSF RVTWVSQASADQRAGRAGRTEPGHCYRLYSSAVFGDFEQFPPPEITRRPVEDLILQMKALNVEKVINFPFPTPPSVEALLAAEELLIALG ALQPPQKAERVKQLQENRLSCPITALGRTMATFPVAPRYAKMLALSRQHGCLPYAITIVASMTVRELFEELDRPAASDEELTRLKSKRAR VAQMKRTWAGQGASLKLGDLMVLLGAVGACEYASCTPQFCEANGLRYKAMMEIRRLRGQLTTAVNAVCPEAELFVDPKMQPPTESQVTYL RQIVTAGLGDHLARRVQSEEMLEDKWRNAYKTPLLDDPVFIHPSSVLFKELPEFVVYQEIVETTKMYMKGVSSVEVQWIPALLPSYCQFD KPLEEPAPTYCPERGRVLCHRASVFYRVGWPLPAIEVDFPEGIDRYKHFARFLLEGQVFRKLASYRSCLLSSPGTMLKTWARLQPRTESL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NCOR2-DHX37 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | NCOR2 | chr12:124950719 | chr12:125450358 | ENST00000356219 | - | 6 | 48 | 254_312 | 235.0 | 2522.0 | SIN3A/B |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NCOR2-DHX37 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NCOR2-DHX37 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
