
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NF2-EIF3D (FusionGDB2 ID:58722) |
Fusion Gene Summary for NF2-EIF3D |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NF2-EIF3D | Fusion gene ID: 58722 | Hgene | Tgene | Gene symbol | NF2 | EIF3D | Gene ID | 4771 | 8664 |
| Gene name | neurofibromin 2 | eukaryotic translation initiation factor 3 subunit D | |
| Synonyms | ACN|BANF|SCH | EIF3S7|eIF3-p66|eIF3-zeta | |
| Cytomap | 22q12.2 | 22q12.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | merlinmoesin-ezrin-radixin likemoesin-ezrin-radixin-like proteinmoesin-ezrin-radizin-like proteinneurofibromin 2 (bilateral acoustic neuroma)schwannomerlinschwannomin | eukaryotic translation initiation factor 3 subunit DeIF3 p66eukaryotic translation initiation factor 3, subunit 7 zeta, 66/67kDatranslation initiation factor eIF3 p66 subunit | |
| Modification date | 20200322 | 20200313 | |
| UniProtAcc | P35240 | O15371 | |
| Ensembl transtripts involved in fusion gene | ENST00000334961, ENST00000338641, ENST00000347330, ENST00000353887, ENST00000361166, ENST00000361452, ENST00000361676, ENST00000397789, ENST00000403435, ENST00000403999, ENST00000413209, | ENST00000405442, ENST00000478547, ENST00000541106, ENST00000216190, | |
| Fusion gene scores | * DoF score | 15 X 11 X 7=1155 | 14 X 13 X 8=1456 |
| # samples | 18 | 18 | |
| ** MAII score | log2(18/1155*10)=-2.68182403997375 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(18/1456*10)=-3.01594154386902 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NF2 [Title/Abstract] AND EIF3D [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NF2(30000101)-EIF3D(36908649), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | NF2 | GO:0008285 | negative regulation of cell proliferation | 12444102|20178741 |
| Hgene | NF2 | GO:0022408 | negative regulation of cell-cell adhesion | 17210637 |
| Hgene | NF2 | GO:0042532 | negative regulation of tyrosine phosphorylation of STAT protein | 12444102 |
| Hgene | NF2 | GO:0046426 | negative regulation of JAK-STAT cascade | 12444102 |
| Tgene | EIF3D | GO:0002191 | cap-dependent translational initiation | 27462815 |
| Tgene | EIF3D | GO:0006413 | translational initiation | 17581632 |
| Tgene | EIF3D | GO:0075522 | IRES-dependent viral translational initiation | 9573242 |
| Tgene | EIF3D | GO:0075525 | viral translational termination-reinitiation | 21347434 |
 Fusion gene breakpoints across NF2 (5'-gene) Fusion gene breakpoints across NF2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
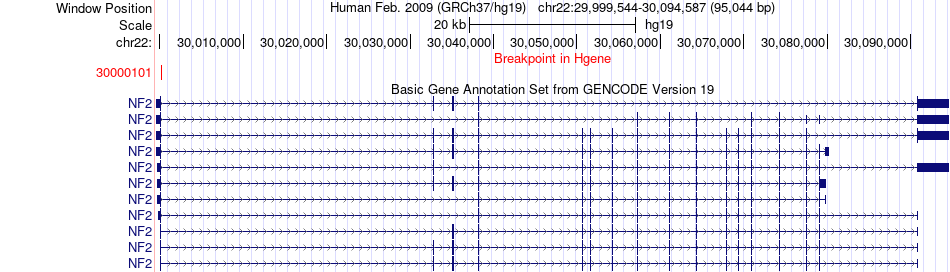 |
 Fusion gene breakpoints across EIF3D (3'-gene) Fusion gene breakpoints across EIF3D (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
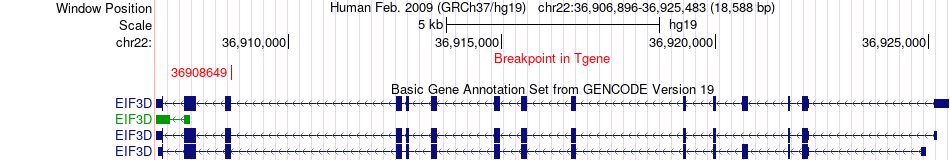 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-V7-A7HQ-01A | NF2 | chr22 | 30000101 | - | EIF3D | chr22 | 36908649 | - |
| ChimerDB4 | BRCA | TCGA-V7-A7HQ-01A | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
Top |
Fusion Gene ORF analysis for NF2-EIF3D |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000334961 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000334961 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000334961 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000338641 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000338641 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000338641 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000347330 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000347330 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000347330 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000353887 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000353887 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000353887 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361166 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361166 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361166 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361452 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361452 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361452 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361676 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361676 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000361676 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000397789 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000397789 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000397789 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000403435 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000403435 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000403435 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000403999 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000403999 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000403999 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000413209 | ENST00000405442 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000413209 | ENST00000478547 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| 5CDS-intron | ENST00000413209 | ENST00000541106 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000334961 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000338641 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000347330 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000353887 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000361166 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000361452 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000361676 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000397789 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000403435 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000403999 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
| In-frame | ENST00000413209 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000347330 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 1135 | 557 | 362 | 997 | 211 |
| ENST00000413209 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 1135 | 557 | 362 | 997 | 211 |
| ENST00000338641 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 1133 | 555 | 360 | 995 | 211 |
| ENST00000403435 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 1103 | 525 | 330 | 965 | 211 |
| ENST00000361452 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 1074 | 496 | 301 | 936 | 211 |
| ENST00000403999 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 1058 | 480 | 285 | 920 | 211 |
| ENST00000334961 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 964 | 386 | 191 | 826 | 211 |
| ENST00000353887 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 944 | 366 | 171 | 806 | 211 |
| ENST00000361166 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 692 | 114 | 0 | 554 | 184 |
| ENST00000397789 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 692 | 114 | 0 | 554 | 184 |
| ENST00000361676 | NF2 | chr22 | 30000101 | + | ENST00000216190 | EIF3D | chr22 | 36908649 | - | 692 | 114 | 0 | 554 | 184 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000347330 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.005323888 | 0.9946761 |
| ENST00000413209 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.005323888 | 0.9946761 |
| ENST00000338641 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.005423156 | 0.9945768 |
| ENST00000403435 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.004869058 | 0.995131 |
| ENST00000361452 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.003848303 | 0.9961516 |
| ENST00000403999 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.004327975 | 0.9956721 |
| ENST00000334961 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.005432279 | 0.99456775 |
| ENST00000353887 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.004933714 | 0.9950663 |
| ENST00000361166 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.003528084 | 0.996472 |
| ENST00000397789 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.003528084 | 0.996472 |
| ENST00000361676 | ENST00000216190 | NF2 | chr22 | 30000101 | + | EIF3D | chr22 | 36908649 | - | 0.003528084 | 0.996472 |
Top |
Fusion Genomic Features for NF2-EIF3D |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for NF2-EIF3D |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr22:30000101/chr22:36908649) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| NF2 | EIF3D |
| FUNCTION: Probable regulator of the Hippo/SWH (Sav/Wts/Hpo) signaling pathway, a signaling pathway that plays a pivotal role in tumor suppression by restricting proliferation and promoting apoptosis. Along with WWC1 can synergistically induce the phosphorylation of LATS1 and LATS2 and can probably function in the regulation of the Hippo/SWH (Sav/Wts/Hpo) signaling pathway. May act as a membrane stabilizing protein. May inhibit PI3 kinase by binding to AGAP2 and impairing its stimulating activity. Suppresses cell proliferation and tumorigenesis by inhibiting the CUL4A-RBX1-DDB1-VprBP/DCAF1 E3 ubiquitin-protein ligase complex. {ECO:0000269|PubMed:20159598, ECO:0000269|PubMed:20178741, ECO:0000269|PubMed:21167305}. | FUNCTION: mRNA cap-binding component of the eukaryotic translation initiation factor 3 (eIF-3) complex, a complex required for several steps in the initiation of protein synthesis of a specialized repertoire of mRNAs (PubMed:27462815). The eIF-3 complex associates with the 40S ribosome and facilitates the recruitment of eIF-1, eIF-1A, eIF-2:GTP:methionyl-tRNAi and eIF-5 to form the 43S pre-initiation complex (43S PIC). The eIF-3 complex stimulates mRNA recruitment to the 43S PIC and scanning of the mRNA for AUG recognition. The eIF-3 complex is also required for disassembly and recycling of post-termination ribosomal complexes and subsequently prevents premature joining of the 40S and 60S ribosomal subunits prior to initiation (PubMed:18599441, PubMed:25849773). The eIF-3 complex specifically targets and initiates translation of a subset of mRNAs involved in cell proliferation, including cell cycling, differentiation and apoptosis, and uses different modes of RNA stem-loop binding to exert either translational activation or repression (PubMed:25849773). In the eIF-3 complex, EIF3D specifically recognizes and binds the 7-methylguanosine cap of a subset of mRNAs (PubMed:27462815). {ECO:0000269|PubMed:18599441, ECO:0000269|PubMed:25849773, ECO:0000269|PubMed:27462815}.; FUNCTION: (Microbial infection) In case of FCV infection, plays a role in the ribosomal termination-reinitiation event leading to the translation of VP2 (PubMed:18056426). {ECO:0000269|PubMed:18056426}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | EIF3D | chr22:30000101 | chr22:36908649 | ENST00000216190 | 11 | 15 | 530_547 | 402 | 549.0 | Compositional bias | Note=Asp/Glu-rich (highly acidic) | |
| Tgene | EIF3D | chr22:30000101 | chr22:36908649 | ENST00000405442 | 11 | 15 | 530_547 | 402 | 549.0 | Compositional bias | Note=Asp/Glu-rich (highly acidic) |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000334961 | + | 1 | 15 | 327_465 | 38 | 451.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000338641 | + | 1 | 16 | 327_465 | 38 | 596.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000347330 | + | 1 | 10 | 327_465 | 38 | 1312.6666666666667 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000353887 | + | 1 | 15 | 327_465 | 38 | 406.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000361166 | + | 1 | 17 | 327_465 | 38 | 571.3333333333334 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000361452 | + | 1 | 16 | 327_465 | 38 | 1665.6666666666667 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000361676 | + | 1 | 16 | 327_465 | 38 | 531.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000397789 | + | 1 | 17 | 327_465 | 38 | 573.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000403435 | + | 1 | 17 | 327_465 | 38 | 541.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000403999 | + | 1 | 16 | 327_465 | 38 | 591.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000413209 | + | 1 | 5 | 327_465 | 38 | 166.0 | Compositional bias | Note=Glu-rich |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000334961 | + | 1 | 15 | 22_311 | 38 | 451.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000338641 | + | 1 | 16 | 22_311 | 38 | 596.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000347330 | + | 1 | 10 | 22_311 | 38 | 1312.6666666666667 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000353887 | + | 1 | 15 | 22_311 | 38 | 406.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000361166 | + | 1 | 17 | 22_311 | 38 | 571.3333333333334 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000361452 | + | 1 | 16 | 22_311 | 38 | 1665.6666666666667 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000361676 | + | 1 | 16 | 22_311 | 38 | 531.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000397789 | + | 1 | 17 | 22_311 | 38 | 573.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000403435 | + | 1 | 17 | 22_311 | 38 | 541.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000403999 | + | 1 | 16 | 22_311 | 38 | 591.0 | Domain | FERM |
| Hgene | NF2 | chr22:30000101 | chr22:36908649 | ENST00000413209 | + | 1 | 5 | 22_311 | 38 | 166.0 | Domain | FERM |
| Tgene | EIF3D | chr22:30000101 | chr22:36908649 | ENST00000216190 | 11 | 15 | 285_299 | 402 | 549.0 | Region | RNA gate | |
| Tgene | EIF3D | chr22:30000101 | chr22:36908649 | ENST00000405442 | 11 | 15 | 285_299 | 402 | 549.0 | Region | RNA gate |
Top |
Fusion Gene Sequence for NF2-EIF3D |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >58722_58722_1_NF2-EIF3D_NF2_chr22_30000101_ENST00000334961_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=964nt_BP=386nt TCCCGGAGTGCCGGGTCGCGCCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCCTTTC CGCTCAGGCAGGGTCCTCGCGGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCATGGT GGCCCTGAGGCCTGTGCAGCAACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCCCGCG CCATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGG ACGCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGG AGCTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTC GGTACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCG TGGAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACA AGCAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAG AAGAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTG >58722_58722_1_NF2-EIF3D_NF2_chr22_30000101_ENST00000334961_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_2_NF2-EIF3D_NF2_chr22_30000101_ENST00000338641_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=1133nt_BP=555nt CGCTTCCCGCGGGCGCGCGGAGTGAGGACGGTGACAGCCACGCGCGCGCGTACGCGCCCGATGCAGCGCGGCCCCGTGACCCTAGTCGGC CGCTGAGAGGCGCGCGGAGTCTGGGCCGCTGCCGTCTAGGGGTCCCGTCCCGAGGCGTCCCCGGCATCTCCGGCCCGAATCCCGGAGTGC CGGGTCGCGCCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCCTTTCCGCTCAGGCAG GGTCCTCGCGGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCATGGTGGCCCTGAGGC CTGTGCAGCAACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCCCGCGCCATGGCCGGG GCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGACGCCGAGATG GAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAGCTGAAGAAC AACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGGTACCACGTG AAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTGGAGAATGCC TGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAGCAGGTCATC CGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAAGAGGAAGAA ACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTCTAACTTGCT >58722_58722_2_NF2-EIF3D_NF2_chr22_30000101_ENST00000338641_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_3_NF2-EIF3D_NF2_chr22_30000101_ENST00000347330_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=1135nt_BP=557nt TGCGCTTCCCGCGGGCGCGCGGAGTGAGGACGGTGACAGCCACGCGCGCGCGTACGCGCCCGATGCAGCGCGGCCCCGTGACCCTAGTCG GCCGCTGAGAGGCGCGCGGAGTCTGGGCCGCTGCCGTCTAGGGGTCCCGTCCCGAGGCGTCCCCGGCATCTCCGGCCCGAATCCCGGAGT GCCGGGTCGCGCCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCCTTTCCGCTCAGGC AGGGTCCTCGCGGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCATGGTGGCCCTGAG GCCTGTGCAGCAACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCCCGCGCCATGGCCG GGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGACGCCGAGA TGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAGCTGAAGA ACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGGTACCACG TGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTGGAGAATG CCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAGCAGGTCA TCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAAGAGGAAG AAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTCTAACTTG >58722_58722_3_NF2-EIF3D_NF2_chr22_30000101_ENST00000347330_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_4_NF2-EIF3D_NF2_chr22_30000101_ENST00000353887_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=944nt_BP=366nt CCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCCTTTCCGCTCAGGCAGGGTCCTCGC GGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCATGGTGGCCCTGAGGCCTGTGCAGC AACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCCCGCGCCATGGCCGGGGCCATCGCT TCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGACGCCGAGATGGAGTTCAAT TGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAGCTGAAGAACAACAGCTAC AAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGGTACCACGTGAAAGACTCC TCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTGGAGAATGCCTGGGGCATT TTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAGCAGGTCATCCGTGTCTAC AGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAAGAGGAAGAAACTTAAACC AGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTCTAACTTGCTCTCTGACAT >58722_58722_4_NF2-EIF3D_NF2_chr22_30000101_ENST00000353887_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_5_NF2-EIF3D_NF2_chr22_30000101_ENST00000361166_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=692nt_BP=114nt ATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGAC GCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAG CTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGG TACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTG GAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAG CAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAA GAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTC >58722_58722_5_NF2-EIF3D_NF2_chr22_30000101_ENST00000361166_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=184AA_BP=38 MAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKNNSYKLARWTCCALLAGSEYLKLGYVSR YHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVIRVYSLPDGTFSSDEDEEEEEEEEEEEE -------------------------------------------------------------- >58722_58722_6_NF2-EIF3D_NF2_chr22_30000101_ENST00000361452_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=1074nt_BP=496nt GATGCAGCGCGGCCCCGTGACCCTAGTCGGCCGCTGAGAGGCGCGCGGAGTCTGGGCCGCTGCCGTCTAGGGGTCCCGTCCCGAGGCGTC CCCGGCATCTCCGGCCCGAATCCCGGAGTGCCGGGTCGCGCCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCC CGTCCCCCAACTCCCCTTTCCGCTCAGGCAGGGTCCTCGCGGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCC GGGCAGCCGGCCACCATGGTGGCCCTGAGGCCTGTGCAGCAACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGG TCCCGGGCCTGAGCCCCGCGCCATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCAC CGTGAGGATCGTCACCATGGACGCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCG AGGGGCTGTCATTGCCACGGAGCTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCT CAAGCTTGGTTATGTGTCTCGGTACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGC CAGCCAGATCAACCTGAGCGTGGAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCT CATCCTCAAGGACCCCAACAAGCAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGA GGAGGAAGAAGAGGAAGAAGAAGAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTT >58722_58722_6_NF2-EIF3D_NF2_chr22_30000101_ENST00000361452_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_7_NF2-EIF3D_NF2_chr22_30000101_ENST00000361676_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=692nt_BP=114nt ATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGAC GCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAG CTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGG TACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTG GAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAG CAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAA GAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTC >58722_58722_7_NF2-EIF3D_NF2_chr22_30000101_ENST00000361676_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=184AA_BP=38 MAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKNNSYKLARWTCCALLAGSEYLKLGYVSR YHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVIRVYSLPDGTFSSDEDEEEEEEEEEEEE -------------------------------------------------------------- >58722_58722_8_NF2-EIF3D_NF2_chr22_30000101_ENST00000397789_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=692nt_BP=114nt ATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGAC GCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAG CTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGG TACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTG GAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAG CAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAA GAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTC >58722_58722_8_NF2-EIF3D_NF2_chr22_30000101_ENST00000397789_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=184AA_BP=38 MAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKNNSYKLARWTCCALLAGSEYLKLGYVSR YHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVIRVYSLPDGTFSSDEDEEEEEEEEEEEE -------------------------------------------------------------- >58722_58722_9_NF2-EIF3D_NF2_chr22_30000101_ENST00000403435_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=1103nt_BP=525nt GTGACAGCCACGCGCGCGCGTACGCGCCCGATGCAGCGCGGCCCCGTGACCCTAGTCGGCCGCTGAGAGGCGCGCGGAGTCTGGGCCGCT GCCGTCTAGGGGTCCCGTCCCGAGGCGTCCCCGGCATCTCCGGCCCGAATCCCGGAGTGCCGGGTCGCGCCTGCACCGAAGGTCCCGGCT CCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCCTTTCCGCTCAGGCAGGGTCCTCGCGGCCCATGCTGGCCGCTGGGG ACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCATGGTGGCCCTGAGGCCTGTGCAGCAACTCCAGGGGGGCTAAAGGG CTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCCCGCGCCATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCT CTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGACGCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTT GACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAGCTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGC TGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGGTACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGC ACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTGGAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATC TGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAGCAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTC AGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAAGAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTT GTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTCTAACTTGCTCTCTGACATTTAGCAGATGAAATAAAATAT >58722_58722_9_NF2-EIF3D_NF2_chr22_30000101_ENST00000403435_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_10_NF2-EIF3D_NF2_chr22_30000101_ENST00000403999_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=1058nt_BP=480nt GTGACCCTAGTCGGCCGCTGAGAGGCGCGCGGAGTCTGGGCCGCTGCCGTCTAGGGGTCCCGTCCCGAGGCGTCCCCGGCATCTCCGGCC CGAATCCCGGAGTGCCGGGTCGCGCCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCC TTTCCGCTCAGGCAGGGTCCTCGCGGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCA TGGTGGCCCTGAGGCCTGTGCAGCAACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCC CGCGCCATGGCCGGGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACC ATGGACGCCGAGATGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCC ACGGAGCTGAAGAACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTG TCTCGGTACCACGTGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTG AGCGTGGAGAATGCCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCC AACAAGCAGGTCATCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAA GAAGAAGAGGAAGAAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATG >58722_58722_10_NF2-EIF3D_NF2_chr22_30000101_ENST00000403999_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- >58722_58722_11_NF2-EIF3D_NF2_chr22_30000101_ENST00000413209_EIF3D_chr22_36908649_ENST00000216190_length(transcript)=1135nt_BP=557nt TGCGCTTCCCGCGGGCGCGCGGAGTGAGGACGGTGACAGCCACGCGCGCGCGTACGCGCCCGATGCAGCGCGGCCCCGTGACCCTAGTCG GCCGCTGAGAGGCGCGCGGAGTCTGGGCCGCTGCCGTCTAGGGGTCCCGTCCCGAGGCGTCCCCGGCATCTCCGGCCCGAATCCCGGAGT GCCGGGTCGCGCCTGCACCGAAGGTCCCGGCTCCTGTGCCCTCCCTGCAGCCGTCAGGGCCCGTCCCCCAACTCCCCTTTCCGCTCAGGC AGGGTCCTCGCGGCCCATGCTGGCCGCTGGGGACCCGCGCAGCCCAGACCGTTCCCGGGCCGGGCAGCCGGCCACCATGGTGGCCCTGAG GCCTGTGCAGCAACTCCAGGGGGGCTAAAGGGCTCAGAGTGCAGGCCGTGGGGCGCGAGGGTCCCGGGCCTGAGCCCCGCGCCATGGCCG GGGCCATCGCTTCCCGCATGAGCTTCAGCTCTCTCAAGAGGAAGCAACCCAAGACGTTCACCGTGAGGATCGTCACCATGGACGCCGAGA TGGAGTTCAATTGCGAGCACTGTAATGGCGTTGACTGGCGTCAGAAGCTGGACTCTCAGCGAGGGGCTGTCATTGCCACGGAGCTGAAGA ACAACAGCTACAAGTTGGCCCGGTGGACCTGCTGTGCTTTGCTGGCTGGATCTGAGTACCTCAAGCTTGGTTATGTGTCTCGGTACCACG TGAAAGACTCCTCACGCCACGTCATCCTAGGCACCCAGCAGTTCAAGCCTAATGAGTTTGCCAGCCAGATCAACCTGAGCGTGGAGAATG CCTGGGGCATTTTACGCTGCGTCATTGACATCTGCATGAAGCTGGAGGAGGGCAAATACCTCATCCTCAAGGACCCCAACAAGCAGGTCA TCCGTGTCTACAGCCTCCCTGATGGCACCTTCAGCTCTGATGAAGATGAGGAGGAAGAGGAGGAGGAAGAAGAGGAAGAAGAAGAGGAAG AAACTTAAACCAGTGATGTGGAGCTGGAGTTTGTCCTTCCACCGAGACTACGAGGGCCTTTGATGCTTAGTGGAATGTGTGTCTAACTTG >58722_58722_11_NF2-EIF3D_NF2_chr22_30000101_ENST00000413209_EIF3D_chr22_36908649_ENST00000216190_length(amino acids)=211AA_BP=65 MCSNSRGAKGLRVQAVGREGPGPEPRAMAGAIASRMSFSSLKRKQPKTFTVRIVTMDAEMEFNCEHCNGVDWRQKLDSQRGAVIATELKN NSYKLARWTCCALLAGSEYLKLGYVSRYHVKDSSRHVILGTQQFKPNEFASQINLSVENAWGILRCVIDICMKLEEGKYLILKDPNKQVI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NF2-EIF3D |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NF2-EIF3D |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NF2-EIF3D |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
