
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NFIX-CALR (FusionGDB2 ID:58938) |
Fusion Gene Summary for NFIX-CALR |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NFIX-CALR | Fusion gene ID: 58938 | Hgene | Tgene | Gene symbol | NFIX | CALR | Gene ID | 4784 | 811 |
| Gene name | nuclear factor I X | calreticulin | |
| Synonyms | CTF|MRSHSS|NF-I/X|NF1-X|NF1A|SOTOS2 | CRT|HEL-S-99n|RO|SSA|cC1qR | |
| Cytomap | 19p13.13 | 19p13.13 | |
| Type of gene | protein-coding | protein-coding | |
| Description | nuclear factor 1 X-typeCCAAT-box-binding transcription factorTGGCA-binding proteinnuclear factor 1/X | calreticulinCRP55ERp60HACBPSicca syndrome antigen A (autoantigen Ro; calreticulin)calregulinendoplasmic reticulum resident protein 60epididymis secretory sperm binding protein Li 99ngrp60 | |
| Modification date | 20200329 | 20200329 | |
| UniProtAcc | . | Q96L12 | |
| Ensembl transtripts involved in fusion gene | ENST00000397661, ENST00000585575, ENST00000587260, ENST00000587760, ENST00000588228, ENST00000592199, ENST00000358552, ENST00000360105, ENST00000588680, | ENST00000316448, | |
| Fusion gene scores | * DoF score | 27 X 14 X 16=6048 | 24 X 30 X 7=5040 |
| # samples | 42 | 38 | |
| ** MAII score | log2(42/6048*10)=-3.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(38/5040*10)=-3.72935241005633 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NFIX [Title/Abstract] AND CALR [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NFIX(13136366)-CALR(13050866), # samples:1 NFIX(13136366)-CALR(13049947), # samples:1 NFIX(13192669)-CALR(13054350), # samples:1 NFIX(13189549)-CALR(13055026), # samples:1 NFIX(13136366)-CALR(13049948), # samples:1 NFIX(13136366)-CALR(13050242), # samples:1 NFIX(13192669)-CALR(13054351), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | NFIX | GO:0000122 | negative regulation of transcription by RNA polymerase II | 19706729 |
| Hgene | NFIX | GO:0045944 | positive regulation of transcription by RNA polymerase II | 19706729 |
| Tgene | CALR | GO:0000122 | negative regulation of transcription by RNA polymerase II | 8107809 |
| Tgene | CALR | GO:0006611 | protein export from nucleus | 11149926 |
| Tgene | CALR | GO:0017148 | negative regulation of translation | 14726956 |
| Tgene | CALR | GO:0033144 | negative regulation of intracellular steroid hormone receptor signaling pathway | 8107809 |
| Tgene | CALR | GO:0034504 | protein localization to nucleus | 15998798 |
| Tgene | CALR | GO:0045665 | negative regulation of neuron differentiation | 8107809 |
| Tgene | CALR | GO:0045892 | negative regulation of transcription, DNA-templated | 8107809 |
| Tgene | CALR | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway | 8107809 |
 Fusion gene breakpoints across NFIX (5'-gene) Fusion gene breakpoints across NFIX (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
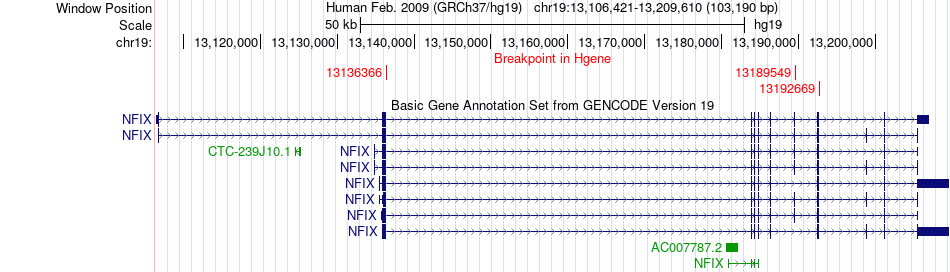 |
 Fusion gene breakpoints across CALR (3'-gene) Fusion gene breakpoints across CALR (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
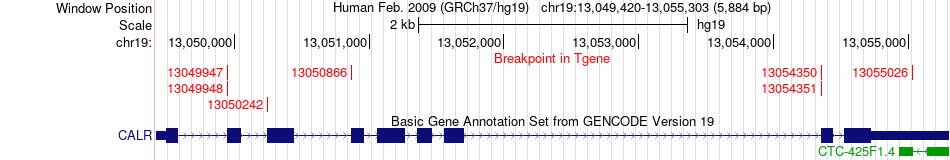 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-AN-A0G0-01A | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| ChimerDB4 | OV | TCGA-13-0887-01A | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| ChimerDB4 | OV | TCGA-13-0887-01A | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| ChimerDB4 | OV | TCGA-13-0887 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| ChimerDB4 | OV | TCGA-25-2396 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| ChimerDB4 | OV | TCGA-36-1577-01A | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| ChimerDB4 | OV | TCGA-36-1577 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
Top |
Fusion Gene ORF analysis for NFIX-CALR |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| 5CDS-3UTR | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| 5CDS-3UTR | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| 5CDS-3UTR | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| 5CDS-3UTR | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| 5CDS-3UTR | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| In-frame | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| In-frame | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| In-frame | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| In-frame | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| In-frame | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| In-frame | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| In-frame | ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| intron-3CDS | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049948 | + |
| intron-3CDS | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050242 | + |
| intron-3CDS | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + |
| intron-3CDS | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + |
| intron-3CDS | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + |
| intron-3CDS | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + |
| intron-3UTR | ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| intron-3UTR | ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
| intron-3UTR | ENST00000588680 | ENST00000316448 | NFIX | chr19 | 13189549 | + | CALR | chr19 | 13055026 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000397661 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2354 | 1484 | 173 | 1777 | 534 |
| ENST00000592199 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2124 | 1254 | 0 | 1547 | 515 |
| ENST00000587760 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2141 | 1271 | 14 | 1564 | 516 |
| ENST00000585575 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2100 | 1230 | 0 | 1523 | 507 |
| ENST00000360105 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2075 | 1205 | 65 | 1498 | 477 |
| ENST00000588228 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2201 | 1331 | 53 | 1624 | 523 |
| ENST00000587260 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 2179 | 1309 | 58 | 1602 | 514 |
| ENST00000358552 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054350 | + | 1998 | 1128 | 0 | 1421 | 473 |
| ENST00000397661 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2354 | 1484 | 173 | 1777 | 534 |
| ENST00000592199 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2124 | 1254 | 0 | 1547 | 515 |
| ENST00000587760 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2141 | 1271 | 14 | 1564 | 516 |
| ENST00000585575 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2100 | 1230 | 0 | 1523 | 507 |
| ENST00000360105 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2075 | 1205 | 65 | 1498 | 477 |
| ENST00000588228 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2201 | 1331 | 53 | 1624 | 523 |
| ENST00000587260 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 2179 | 1309 | 58 | 1602 | 514 |
| ENST00000358552 | NFIX | chr19 | 13192669 | + | ENST00000316448 | CALR | chr19 | 13054351 | + | 1998 | 1128 | 0 | 1421 | 473 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.00724521 | 0.99275476 |
| ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.005153251 | 0.9948467 |
| ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.003311834 | 0.9966882 |
| ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.003115225 | 0.99688476 |
| ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.004845465 | 0.9951545 |
| ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.005771936 | 0.994228 |
| ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.005929697 | 0.99407023 |
| ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 0.003557034 | 0.996443 |
| ENST00000397661 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.00724521 | 0.99275476 |
| ENST00000592199 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.005153251 | 0.9948467 |
| ENST00000587760 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.003311834 | 0.9966882 |
| ENST00000585575 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.003115225 | 0.99688476 |
| ENST00000360105 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.004845465 | 0.9951545 |
| ENST00000588228 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.005771936 | 0.994228 |
| ENST00000587260 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.005929697 | 0.99407023 |
| ENST00000358552 | ENST00000316448 | NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054351 | + | 0.003557034 | 0.996443 |
Top |
Fusion Genomic Features for NFIX-CALR |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050241 | + | 6.62E-05 | 0.99993384 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + | 9.26E-06 | 0.9999907 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + | 9.26E-06 | 0.9999907 |
| NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 3.55E-06 | 0.9999964 |
| NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 3.55E-06 | 0.9999964 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + | 1.65E-06 | 0.99999833 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050241 | + | 6.62E-05 | 0.99993384 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + | 9.26E-06 | 0.9999907 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13049947 | + | 9.26E-06 | 0.9999907 |
| NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 3.55E-06 | 0.9999964 |
| NFIX | chr19 | 13192669 | + | CALR | chr19 | 13054350 | + | 3.55E-06 | 0.9999964 |
| NFIX | chr19 | 13136366 | + | CALR | chr19 | 13050866 | + | 1.65E-06 | 0.99999833 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
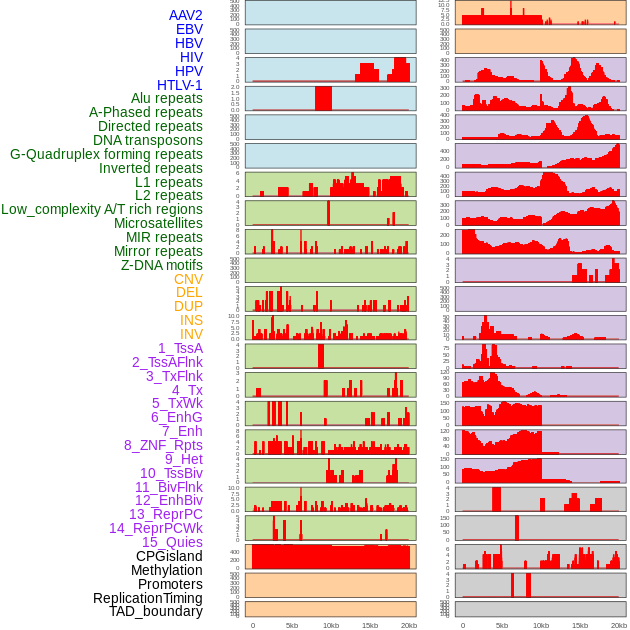 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
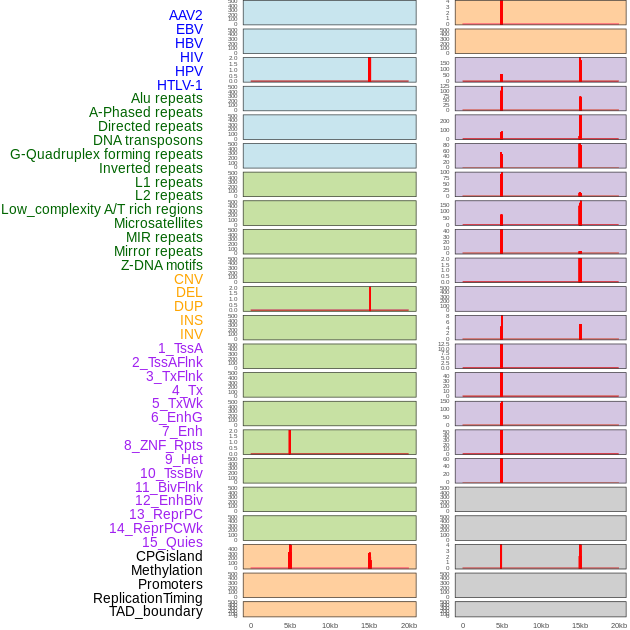 |
Top |
Fusion Protein Features for NFIX-CALR |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:13136366/chr19:13050866) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CALR |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: During spermatogenesis, may act as a lectin-independent chaperone for specific client proteins such as ADAM3. Required for sperm fertility (By similarity). CALR3 capacity for calcium-binding may be absent or much lower than that of CALR. {ECO:0000250, ECO:0000269|PubMed:21590275}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000397661 | + | 8 | 10 | 1_194 | 418 | 878.6666666666666 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000585575 | + | 8 | 11 | 1_194 | 410 | 495.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000587260 | + | 7 | 9 | 1_194 | 417 | 281.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000587760 | + | 8 | 10 | 1_194 | 410 | 460.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000592199 | + | 8 | 11 | 1_194 | 418 | 503.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000397661 | + | 8 | 10 | 1_194 | 418 | 878.6666666666666 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000585575 | + | 8 | 11 | 1_194 | 410 | 495.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000587260 | + | 7 | 9 | 1_194 | 417 | 281.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000587760 | + | 8 | 10 | 1_194 | 410 | 460.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000592199 | + | 8 | 11 | 1_194 | 418 | 503.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000397661 | + | 8 | 10 | 398_406 | 418 | 878.6666666666666 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000585575 | + | 8 | 11 | 398_406 | 410 | 495.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000587260 | + | 7 | 9 | 398_406 | 417 | 281.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000587760 | + | 8 | 10 | 398_406 | 410 | 460.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054350 | ENST00000592199 | + | 8 | 11 | 398_406 | 418 | 503.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000397661 | + | 8 | 10 | 398_406 | 418 | 878.6666666666666 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000585575 | + | 8 | 11 | 398_406 | 410 | 495.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000587260 | + | 7 | 9 | 398_406 | 417 | 281.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000587760 | + | 8 | 10 | 398_406 | 410 | 460.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13192669 | chr19:13054351 | ENST00000592199 | + | 8 | 11 | 398_406 | 418 | 503.0 | Motif | 9aaTAD |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 351_408 | 30 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 351_408 | 30 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 351_408 | 64 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 351_408 | 132 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 351_408 | 320 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 351_408 | 320 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 414_417 | 30 | 418.0 | Motif | Note=Prevents secretion from ER | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 414_417 | 30 | 418.0 | Motif | Note=Prevents secretion from ER | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 414_417 | 64 | 418.0 | Motif | Note=Prevents secretion from ER | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 414_417 | 132 | 418.0 | Motif | Note=Prevents secretion from ER | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 414_417 | 320 | 418.0 | Motif | Note=Prevents secretion from ER | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 414_417 | 320 | 418.0 | Motif | Note=Prevents secretion from ER | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 191_255 | 30 | 418.0 | Region | Note=4 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 198_308 | 30 | 418.0 | Region | Note=P-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 259_297 | 30 | 418.0 | Region | Note=3 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 309_417 | 30 | 418.0 | Region | Note=C-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 191_255 | 30 | 418.0 | Region | Note=4 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 198_308 | 30 | 418.0 | Region | Note=P-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 259_297 | 30 | 418.0 | Region | Note=3 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 309_417 | 30 | 418.0 | Region | Note=C-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 191_255 | 64 | 418.0 | Region | Note=4 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 198_308 | 64 | 418.0 | Region | Note=P-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 259_297 | 64 | 418.0 | Region | Note=3 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 309_417 | 64 | 418.0 | Region | Note=C-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 191_255 | 132 | 418.0 | Region | Note=4 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 198_308 | 132 | 418.0 | Region | Note=P-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 259_297 | 132 | 418.0 | Region | Note=3 X approximate repeats | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 309_417 | 132 | 418.0 | Region | Note=C-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 191_202 | 30 | 418.0 | Repeat | Note=1-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 210_221 | 30 | 418.0 | Repeat | Note=1-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 227_238 | 30 | 418.0 | Repeat | Note=1-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 244_255 | 30 | 418.0 | Repeat | Note=1-4 | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 259_269 | 30 | 418.0 | Repeat | Note=2-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 273_283 | 30 | 418.0 | Repeat | Note=2-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 287_297 | 30 | 418.0 | Repeat | Note=2-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 191_202 | 30 | 418.0 | Repeat | Note=1-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 210_221 | 30 | 418.0 | Repeat | Note=1-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 227_238 | 30 | 418.0 | Repeat | Note=1-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 244_255 | 30 | 418.0 | Repeat | Note=1-4 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 259_269 | 30 | 418.0 | Repeat | Note=2-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 273_283 | 30 | 418.0 | Repeat | Note=2-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 287_297 | 30 | 418.0 | Repeat | Note=2-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 191_202 | 64 | 418.0 | Repeat | Note=1-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 210_221 | 64 | 418.0 | Repeat | Note=1-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 227_238 | 64 | 418.0 | Repeat | Note=1-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 244_255 | 64 | 418.0 | Repeat | Note=1-4 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 259_269 | 64 | 418.0 | Repeat | Note=2-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 273_283 | 64 | 418.0 | Repeat | Note=2-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 287_297 | 64 | 418.0 | Repeat | Note=2-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 191_202 | 132 | 418.0 | Repeat | Note=1-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 210_221 | 132 | 418.0 | Repeat | Note=1-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 227_238 | 132 | 418.0 | Repeat | Note=1-3 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 244_255 | 132 | 418.0 | Repeat | Note=1-4 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 259_269 | 132 | 418.0 | Repeat | Note=2-1 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 273_283 | 132 | 418.0 | Repeat | Note=2-2 | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 287_297 | 132 | 418.0 | Repeat | Note=2-3 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000397661 | + | 2 | 10 | 1_194 | 186 | 878.6666666666666 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000585575 | + | 2 | 11 | 1_194 | 178 | 495.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000587260 | + | 1 | 9 | 1_194 | 185 | 281.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000587760 | + | 2 | 10 | 1_194 | 178 | 460.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000592199 | + | 2 | 11 | 1_194 | 186 | 503.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000397661 | + | 2 | 10 | 1_194 | 186 | 878.6666666666666 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000585575 | + | 2 | 11 | 1_194 | 178 | 495.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000587260 | + | 1 | 9 | 1_194 | 185 | 281.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000587760 | + | 2 | 10 | 1_194 | 178 | 460.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000592199 | + | 2 | 11 | 1_194 | 186 | 503.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000397661 | + | 2 | 10 | 1_194 | 186 | 878.6666666666666 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000585575 | + | 2 | 11 | 1_194 | 178 | 495.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000587260 | + | 1 | 9 | 1_194 | 185 | 281.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000587760 | + | 2 | 10 | 1_194 | 178 | 460.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000592199 | + | 2 | 11 | 1_194 | 186 | 503.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000397661 | + | 2 | 10 | 1_194 | 186 | 878.6666666666666 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000585575 | + | 2 | 11 | 1_194 | 178 | 495.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000587260 | + | 1 | 9 | 1_194 | 185 | 281.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000587760 | + | 2 | 10 | 1_194 | 178 | 460.6666666666667 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000592199 | + | 2 | 11 | 1_194 | 186 | 503.0 | DNA binding | CTF/NF-I |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000397661 | + | 2 | 10 | 398_406 | 186 | 878.6666666666666 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000585575 | + | 2 | 11 | 398_406 | 178 | 495.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000587260 | + | 1 | 9 | 398_406 | 185 | 281.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000587760 | + | 2 | 10 | 398_406 | 178 | 460.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049947 | ENST00000592199 | + | 2 | 11 | 398_406 | 186 | 503.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000397661 | + | 2 | 10 | 398_406 | 186 | 878.6666666666666 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000585575 | + | 2 | 11 | 398_406 | 178 | 495.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000587260 | + | 1 | 9 | 398_406 | 185 | 281.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000587760 | + | 2 | 10 | 398_406 | 178 | 460.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13049948 | ENST00000592199 | + | 2 | 11 | 398_406 | 186 | 503.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000397661 | + | 2 | 10 | 398_406 | 186 | 878.6666666666666 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000585575 | + | 2 | 11 | 398_406 | 178 | 495.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000587260 | + | 1 | 9 | 398_406 | 185 | 281.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000587760 | + | 2 | 10 | 398_406 | 178 | 460.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050242 | ENST00000592199 | + | 2 | 11 | 398_406 | 186 | 503.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000397661 | + | 2 | 10 | 398_406 | 186 | 878.6666666666666 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000585575 | + | 2 | 11 | 398_406 | 178 | 495.0 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000587260 | + | 1 | 9 | 398_406 | 185 | 281.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000587760 | + | 2 | 10 | 398_406 | 178 | 460.6666666666667 | Motif | 9aaTAD |
| Hgene | NFIX | chr19:13136366 | chr19:13050866 | ENST00000592199 | + | 2 | 11 | 398_406 | 186 | 503.0 | Motif | 9aaTAD |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 18_197 | 30 | 418.0 | Region | Note=N-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 18_197 | 30 | 418.0 | Region | Note=N-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 18_197 | 64 | 418.0 | Region | Note=N-domain | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 18_197 | 132 | 418.0 | Region | Note=N-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 18_197 | 320 | 418.0 | Region | Note=N-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 191_255 | 320 | 418.0 | Region | Note=4 X approximate repeats | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 198_308 | 320 | 418.0 | Region | Note=P-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 259_297 | 320 | 418.0 | Region | Note=3 X approximate repeats | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 309_417 | 320 | 418.0 | Region | Note=C-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 18_197 | 320 | 418.0 | Region | Note=N-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 191_255 | 320 | 418.0 | Region | Note=4 X approximate repeats | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 198_308 | 320 | 418.0 | Region | Note=P-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 259_297 | 320 | 418.0 | Region | Note=3 X approximate repeats | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 309_417 | 320 | 418.0 | Region | Note=C-domain | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 191_202 | 320 | 418.0 | Repeat | Note=1-1 | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 210_221 | 320 | 418.0 | Repeat | Note=1-2 | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 227_238 | 320 | 418.0 | Repeat | Note=1-3 | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 244_255 | 320 | 418.0 | Repeat | Note=1-4 | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 259_269 | 320 | 418.0 | Repeat | Note=2-1 | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 273_283 | 320 | 418.0 | Repeat | Note=2-2 | |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 287_297 | 320 | 418.0 | Repeat | Note=2-3 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 191_202 | 320 | 418.0 | Repeat | Note=1-1 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 210_221 | 320 | 418.0 | Repeat | Note=1-2 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 227_238 | 320 | 418.0 | Repeat | Note=1-3 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 244_255 | 320 | 418.0 | Repeat | Note=1-4 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 259_269 | 320 | 418.0 | Repeat | Note=2-1 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 273_283 | 320 | 418.0 | Repeat | Note=2-2 | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 287_297 | 320 | 418.0 | Repeat | Note=2-3 |
Top |
Fusion Gene Sequence for NFIX-CALR |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >58938_58938_1_NFIX-CALR_NFIX_chr19_13192669_ENST00000358552_CALR_chr19_13054350_ENST00000316448_length(transcript)=1998nt_BP=1128nt ATGCTCCCGGCTTGCCGCCTGCAGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTC AACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTG GGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTC GTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGC CAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTAC AAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTAC TTTGTCCACACTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGT TTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCC AACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCC ACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCC CCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCC TCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAG TTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAAC GATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAG CAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGAT GAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGG CTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGC TGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAAC TGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCT CATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGA GGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTG CCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATTA >58938_58938_1_NFIX-CALR_NFIX_chr19_13192669_ENST00000358552_CALR_chr19_13054350_ENST00000316448_length(amino acids)=473AA_BP=375 MLPACRLQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFREDF VLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLAY FVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPPS TSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKE FVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAAEKQMKDKQDEEQRLKEEEEDKKRKEEEEAEDKEDDEDKDED -------------------------------------------------------------- >58938_58938_2_NFIX-CALR_NFIX_chr19_13192669_ENST00000358552_CALR_chr19_13054351_ENST00000316448_length(transcript)=1998nt_BP=1128nt ATGCTCCCGGCTTGCCGCCTGCAGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTC AACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTG GGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTC GTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGC CAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTAC AAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTAC TTTGTCCACACTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGT TTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCC AACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCC ACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCC CCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCC TCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAG TTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAAC GATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAG CAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGAT GAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGG CTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGC TGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAAC TGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCT CATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGA GGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTG CCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATTA >58938_58938_2_NFIX-CALR_NFIX_chr19_13192669_ENST00000358552_CALR_chr19_13054351_ENST00000316448_length(amino acids)=473AA_BP=375 MLPACRLQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFREDF VLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLAY FVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPPS TSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKE FVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAAEKQMKDKQDEEQRLKEEEEDKKRKEEEEAEDKEDDEDKDED -------------------------------------------------------------- >58938_58938_3_NFIX-CALR_NFIX_chr19_13192669_ENST00000360105_CALR_chr19_13054350_ENST00000316448_length(transcript)=2075nt_BP=1205nt CAGACGGACACTGTGCCGGGGCGAGCTGACAGGAGTTCACGGCTGCGATAGAACATGGAGATGTCATGGGCGCGACAGAGCCTGGCGGGG ATACCAGCAGCGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGC GGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCG AGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCA CGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGG TGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGT GCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTC CGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTT TTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGG CGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCA AGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCA GCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCA TCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTG TGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATGAGGCATACG CTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGG AGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGGAGGATGAGG AGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCC TGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTG GGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCT TTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTC CCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGA AGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGG GCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATTAAATTAAATTTTGT >58938_58938_3_NFIX-CALR_NFIX_chr19_13192669_ENST00000360105_CALR_chr19_13054350_ENST00000316448_length(amino acids)=477AA_BP=379 MGATEPGGDTSSDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEF REDFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDL YLAYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSIT SPPSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQD SLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAAEKQMKDKQDEEQRLKEEEEDKKRKEEEEAEDKEDDED -------------------------------------------------------------- >58938_58938_4_NFIX-CALR_NFIX_chr19_13192669_ENST00000360105_CALR_chr19_13054351_ENST00000316448_length(transcript)=2075nt_BP=1205nt CAGACGGACACTGTGCCGGGGCGAGCTGACAGGAGTTCACGGCTGCGATAGAACATGGAGATGTCATGGGCGCGACAGAGCCTGGCGGGG ATACCAGCAGCGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGC GGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCG AGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCA CGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGG TGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGT GCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTC CGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTT TTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGG CGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCA AGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCA GCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCA TCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTG TGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATGAGGCATACG CTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGG AGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGGAGGATGAGG AGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCC TGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTG GGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCT TTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTC CCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGA AGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGG GCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATTAAATTAAATTTTGT >58938_58938_4_NFIX-CALR_NFIX_chr19_13192669_ENST00000360105_CALR_chr19_13054351_ENST00000316448_length(amino acids)=477AA_BP=379 MGATEPGGDTSSDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEF REDFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDL YLAYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSIT SPPSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQD SLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAAEKQMKDKQDEEQRLKEEEEDKKRKEEEEAEDKEDDED -------------------------------------------------------------- >58938_58938_5_NFIX-CALR_NFIX_chr19_13192669_ENST00000397661_CALR_chr19_13054350_ENST00000316448_length(transcript)=2354nt_BP=1484nt GCGAGCGCGGCGGCGGCGGCGGCAGCGGCGGCCCCGGAGCCGGCGGGGCCGAGCTTGCGAGCGGCGAGCGCGGAGCGGCGCCGGGCCGAG CGCGGGGCCGCGGGCCGGGCGGGCGCAGCGCGGCGGAGGCCGGAGGAGCCGAGCCGGAGCCCGAGCCCGAGCGCGGCCGCCGCCTGCCGG GCCTCCCCTCGCCGCGGCCGGCCGCCGCGCTCCCGCCCGGGCGCCCAGCTATGTACTCCCCGTACTGCCTCACCCAGGATGAGTTCCACC CGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGC ATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCC GGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCG TGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGG TGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCC AGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCCGGACAATCAGATAGTT CAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATG TGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACT ACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACA GTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCA ACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCCGCGCA TGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCA CGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTG TGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATG AGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGA GGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGG AGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGG ACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGT TCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGT ATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATC TCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAG GGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCC CAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATTAAATT >58938_58938_5_NFIX-CALR_NFIX_chr19_13192669_ENST00000397661_CALR_chr19_13054350_ENST00000316448_length(amino acids)=534AA_BP=436 MPGLPSPRPAAALPPGRPAMYSPYCLTQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKW ASRLLAKLRKDIRPEFREDFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGL CVQPHHIGVTIKELDLYLAYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESP SYYNINQVTLGRRSITSPPSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSS PRMAFTHHPLPVLAGVRPGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLIT -------------------------------------------------------------- >58938_58938_6_NFIX-CALR_NFIX_chr19_13192669_ENST00000397661_CALR_chr19_13054351_ENST00000316448_length(transcript)=2354nt_BP=1484nt GCGAGCGCGGCGGCGGCGGCGGCAGCGGCGGCCCCGGAGCCGGCGGGGCCGAGCTTGCGAGCGGCGAGCGCGGAGCGGCGCCGGGCCGAG CGCGGGGCCGCGGGCCGGGCGGGCGCAGCGCGGCGGAGGCCGGAGGAGCCGAGCCGGAGCCCGAGCCCGAGCGCGGCCGCCGCCTGCCGG GCCTCCCCTCGCCGCGGCCGGCCGCCGCGCTCCCGCCCGGGCGCCCAGCTATGTACTCCCCGTACTGCCTCACCCAGGATGAGTTCCACC CGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGC ATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCC GGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCG TGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGG TGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCC AGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCCGGACAATCAGATAGTT CAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATG TGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACT ACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACA GTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCA ACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCCGCGCA TGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCA CGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTG TGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATG AGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGA GGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGG AGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGG ACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGT TCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGT ATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATC TCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAG GGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCC CAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATTAAATT >58938_58938_6_NFIX-CALR_NFIX_chr19_13192669_ENST00000397661_CALR_chr19_13054351_ENST00000316448_length(amino acids)=534AA_BP=436 MPGLPSPRPAAALPPGRPAMYSPYCLTQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKW ASRLLAKLRKDIRPEFREDFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGL CVQPHHIGVTIKELDLYLAYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESP SYYNINQVTLGRRSITSPPSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSS PRMAFTHHPLPVLAGVRPGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLIT -------------------------------------------------------------- >58938_58938_7_NFIX-CALR_NFIX_chr19_13192669_ENST00000585575_CALR_chr19_13054350_ENST00000316448_length(transcript)=2100nt_BP=1230nt ATGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGC AAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAG CAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAG AAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGG CTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAAC CCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCC GGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACT TCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTG GAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCC AAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAG TCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAG GGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCC CTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGAC TCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTC CTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAA CAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGAC AAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGC CTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCAT TTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTT TTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCT GTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTC CTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCC CTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACA >58938_58938_7_NFIX-CALR_NFIX_chr19_13192669_ENST00000585575_CALR_chr19_13054350_ENST00000316448_length(amino acids)=507AA_BP=409 MDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFREDFVLTITGK KPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLAYFVHTPES GQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPPSTSTTKRP KSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRPGSPRATASA LHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAAEKQMKDK -------------------------------------------------------------- >58938_58938_8_NFIX-CALR_NFIX_chr19_13192669_ENST00000585575_CALR_chr19_13054351_ENST00000316448_length(transcript)=2100nt_BP=1230nt ATGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGC AAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAG CAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAG AAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGG CTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAAC CCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCC GGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACT TCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTG GAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCC AAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAG TCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAG GGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCC CTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGAC TCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTC CTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAA CAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGAC AAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGC CTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCAT TTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTT TTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCT GTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTC CTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCC CTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACA >58938_58938_8_NFIX-CALR_NFIX_chr19_13192669_ENST00000585575_CALR_chr19_13054351_ENST00000316448_length(amino acids)=507AA_BP=409 MDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFREDFVLTITGK KPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLAYFVHTPES GQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPPSTSTTKRP KSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRPGSPRATASA LHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAAEKQMKDK -------------------------------------------------------------- >58938_58938_9_NFIX-CALR_NFIX_chr19_13192669_ENST00000587260_CALR_chr19_13054350_ENST00000316448_length(transcript)=2179nt_BP=1309nt TCCTTCTCTCTTTCCCCCTCATCCCGTCTTCCCCTCCTCCCGTCCTCCCTCGCCCCGCATGCTCCCGGCTTGCCGCCTGCAGGATGAGTT CCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAA GAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGC ATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTG CTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGT CATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTG CGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCCGGACAATCAGA TAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTG GAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAG CTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCCAAGTCCATCGA TGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTG GCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCC GCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCC CTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGA GTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAA CGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGA GCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGA TGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGG GCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGG CTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAA CTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTC TCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAG AGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCT GCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATT >58938_58938_9_NFIX-CALR_NFIX_chr19_13192669_ENST00000587260_CALR_chr19_13054350_ENST00000316448_length(amino acids)=514AA_BP=416 MLPACRLQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFREDF VLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLAY FVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPPS TSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRPGS PRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAA -------------------------------------------------------------- >58938_58938_10_NFIX-CALR_NFIX_chr19_13192669_ENST00000587260_CALR_chr19_13054351_ENST00000316448_length(transcript)=2179nt_BP=1309nt TCCTTCTCTCTTTCCCCCTCATCCCGTCTTCCCCTCCTCCCGTCCTCCCTCGCCCCGCATGCTCCCGGCTTGCCGCCTGCAGGATGAGTT CCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAA GAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGC ATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTG CTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGT CATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTG CGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCCGGACAATCAGA TAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTG GAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAG CTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCCAAGTCCATCGA TGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTG GCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCC GCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCC CTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGA GTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAA CGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGA GCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGA TGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGG GCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGG CTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAA CTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTC TCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAG AGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCT GCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAGAACTACAAACAAAATTTCTATT >58938_58938_10_NFIX-CALR_NFIX_chr19_13192669_ENST00000587260_CALR_chr19_13054351_ENST00000316448_length(amino acids)=514AA_BP=416 MLPACRLQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFREDF VLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLAY FVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPPS TSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRPGS PRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKAA -------------------------------------------------------------- >58938_58938_11_NFIX-CALR_NFIX_chr19_13192669_ENST00000587760_CALR_chr19_13054350_ENST00000316448_length(transcript)=2141nt_BP=1271nt CTCAGAATCGGGCGTTGGGCTTTGCCGGGTGCTTCAGATCAATGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCG CTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGG CGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGC CCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCC GGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTA CTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAAC TGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCAC TGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTG TTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGT CCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCT ATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGA AGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTG CTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCA CGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGG CCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGT GGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCA AAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGG AAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGG CCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCC TCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCC CTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGT GTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCC AGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGG >58938_58938_11_NFIX-CALR_NFIX_chr19_13192669_ENST00000587760_CALR_chr19_13054350_ENST00000316448_length(amino acids)=516AA_BP=418 MGFAGCFRSMDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFRE DFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYL AYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSP PSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRP GSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTK -------------------------------------------------------------- >58938_58938_12_NFIX-CALR_NFIX_chr19_13192669_ENST00000587760_CALR_chr19_13054351_ENST00000316448_length(transcript)=2141nt_BP=1271nt CTCAGAATCGGGCGTTGGGCTTTGCCGGGTGCTTCAGATCAATGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCG CTTTCTCCTACACCTGGTTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGG CGGTGAAGGACGAGCTGCTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGC CCGAGTTCCGCGAGGACTTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCC GGCGGATTGACTGCCTGCGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTA CTGATGGGGAGCGGCTCTACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAAC TGGATCTTTATCTGGCTTACTTTGTCCACACTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCAC TGCCCAACGGGCACTTAAGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTG TTGCAACAGCATCAGGGCCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGT CCATCACCTCCCCTCCTTCCACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCT ATCCCGGGACAGGCCGTTCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGA AGTCAGGAAAGCTGGACTTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTG CTGGAGTCAGACCAGGGAGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCA CGCACCCGACCATCCGCTACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGG CCACCGGACAGGTCAAGTCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGT GGGGCGTAACAAAGGCAGCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCA AAGAGGAGGAGGAGGCAGAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGG AAGATGTCCCCGGCCAGGCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGG CCGCGCCAAATAATGTCTCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCC TCCTCCACTCTCCCCCACCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCC CTGGTTCTCATCTTTCTTGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGT GTGGAGAAGCCACAGGCCTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCC AGCACTGAGGAAGAACGGGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGG >58938_58938_12_NFIX-CALR_NFIX_chr19_13192669_ENST00000587760_CALR_chr19_13054351_ENST00000316448_length(amino acids)=516AA_BP=418 MGFAGCFRSMDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFRE DFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYL AYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSP PSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRP GSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTK -------------------------------------------------------------- >58938_58938_13_NFIX-CALR_NFIX_chr19_13192669_ENST00000588228_CALR_chr19_13054350_ENST00000316448_length(transcript)=2201nt_BP=1331nt AGACGGACACTGTGCCGGGGCGAGCTGACAGGAGTTCACGGCTGCGATAGAACATGGAGATGTCATGGGCGCGACAGAGCCTGGCGGGGA TACCAGCAGCGTGTGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGG CGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGC CCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCA TCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACA AGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTC AGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACA CTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACT GTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCC TGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCA CCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTG GCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCC TCTCCTCTCAGGGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCA CAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACC ACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCT TTGACAACTTCCTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAA TGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGG ATGATGAGGACAAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGC TGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCG AGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCC CTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTT TTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATC TGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCAT TTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAG >58938_58938_13_NFIX-CALR_NFIX_chr19_13192669_ENST00000588228_CALR_chr19_13054350_ENST00000316448_length(amino acids)=523AA_BP=425 MEMSWARQSLAGIPAACDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKD IRPEFREDFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTI KELDLYLAYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLG RRSITSPPSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLP VLAGVRPGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGN -------------------------------------------------------------- >58938_58938_14_NFIX-CALR_NFIX_chr19_13192669_ENST00000588228_CALR_chr19_13054351_ENST00000316448_length(transcript)=2201nt_BP=1331nt AGACGGACACTGTGCCGGGGCGAGCTGACAGGAGTTCACGGCTGCGATAGAACATGGAGATGTCATGGGCGCGACAGAGCCTGGCGGGGA TACCAGCAGCGTGTGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGGTTCAACCTGCAGG CGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTGCTGGGCGAGAAGC CCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGACTTCGTGCTGACCA TCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTGCGCCAGGCTGACA AGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTCTACAAGTCGCCTC AGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCTTACTTTGTCCACA CTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTAAGTTTCCAGGACT GTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGGCCCAACTTCTCCC TGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCTTCCACCAGCACCA CCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGTTCCCCAGCAGCTG GCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGACTTCTGCAGTGCCC TCTCCTCTCAGGGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGGAGCCCCCGGGCCA CAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGCTACCACCACCACC ACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAGTCTGGCACCATCT TTGACAACTTCCTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCAGCAGAGAAACAAA TGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCAGAGGACAAGGAGG ATGATGAGGACAAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAGGCCAAGGACGAGC TGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTCTCTGTGAGACTCG AGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCACCCCCTCCCCGCC CTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCTTGATCAACATCTT TTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGCCTGAGATTTCATC TGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACGGGGCTCTTCTCAT TTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACACTGAGAATGTAAG >58938_58938_14_NFIX-CALR_NFIX_chr19_13192669_ENST00000588228_CALR_chr19_13054351_ENST00000316448_length(amino acids)=523AA_BP=425 MEMSWARQSLAGIPAACDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKD IRPEFREDFVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTI KELDLYLAYFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLG RRSITSPPSTSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLP VLAGVRPGSPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGN -------------------------------------------------------------- >58938_58938_15_NFIX-CALR_NFIX_chr19_13192669_ENST00000592199_CALR_chr19_13054350_ENST00000316448_length(transcript)=2124nt_BP=1254nt ATGTACTCCCCGTACTGCCTCACCCAGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGG TTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTG CTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGAC TTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTG CGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTC TACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCT TACTTTGTCCACACTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTA AGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGG CCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCT TCCACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGT TCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGAC TTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGG AGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGC TACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAG TCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCA GCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCA GAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAG GCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTC TCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCA CCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCT TGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGC CTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACG GGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACA >58938_58938_15_NFIX-CALR_NFIX_chr19_13192669_ENST00000592199_CALR_chr19_13054350_ENST00000316448_length(amino acids)=515AA_BP=417 MYSPYCLTQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFRED FVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLA YFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPP STSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRPG SPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKA -------------------------------------------------------------- >58938_58938_16_NFIX-CALR_NFIX_chr19_13192669_ENST00000592199_CALR_chr19_13054351_ENST00000316448_length(transcript)=2124nt_BP=1254nt ATGTACTCCCCGTACTGCCTCACCCAGGATGAGTTCCACCCGTTCATCGAGGCACTGCTGCCTCACGTCCGCGCTTTCTCCTACACCTGG TTCAACCTGCAGGCGCGGAAGCGCAAGTACTTCAAGAAGCATGAAAAGCGGATGTCGAAGGACGAGGAGCGGGCGGTGAAGGACGAGCTG CTGGGCGAGAAGCCCGAGATCAAGCAGAAGTGGGCATCCCGGCTGCTGGCCAAGCTGCGCAAGGACATCCGGCCCGAGTTCCGCGAGGAC TTCGTGCTGACCATCACGGGCAAGAAGCCCCCCTGCTGCGTGCTCTCCAACCCCGACCAGAAGGGCAAGATCCGGCGGATTGACTGCCTG CGCCAGGCTGACAAGGTGTGGCGGCTGGACCTGGTCATGGTGATTTTGTTTAAGGGGATCCCCCTGGAAAGTACTGATGGGGAGCGGCTC TACAAGTCGCCTCAGTGCTCGAACCCCGGCCTGTGCGTCCAGCCACATCACATTGGAGTCACAATCAAAGAACTGGATCTTTATCTGGCT TACTTTGTCCACACTCCGGAATCCGGACAATCAGATAGTTCAAACCAGCAAGGAGATGCGGACATCAAACCACTGCCCAACGGGCACTTA AGTTTCCAGGACTGTTTTGTGACTTCCGGGGTCTGGAATGTGACGGAGCTGGTGAGAGTATCACAGACTCCTGTTGCAACAGCATCAGGG CCCAACTTCTCCCTGGCGGACCTGGAGAGTCCCAGCTACTACAACATCAACCAGGTGACCCTGGGGCGGCGGTCCATCACCTCCCCTCCT TCCACCAGCACCACCAAGCGCCCCAAGTCCATCGATGACAGTGAGATGGAGAGCCCTGTTGATGACGTGTTCTATCCCGGGACAGGCCGT TCCCCAGCAGCTGGCAGCAGCCAGTCCAGCGGGTGGCCCAACGATGTGGATGCAGGCCCGGCTTCTCTAAAGAAGTCAGGAAAGCTGGAC TTCTGCAGTGCCCTCTCCTCTCAGGGCAGCTCCCCGCGCATGGCTTTCACCCACCACCCGCTGCCTGTGCTTGCTGGAGTCAGACCAGGG AGCCCCCGGGCCACAGCATCAGCCCTGCACTTCCCCTCCACGTCCATCATCCAGCAGTCGAGCCCGTATTTCACGCACCCGACCATCCGC TACCACCACCACCACGGGCAGGACTCACTGAAGGAGTTTGTGCAGTTTGTGTGCTCGGATGGCTCGGGCCAGGCCACCGGACAGGTCAAG TCTGGCACCATCTTTGACAACTTCCTCATCACCAACGATGAGGCATACGCTGAGGAGTTTGGCAACGAGACGTGGGGCGTAACAAAGGCA GCAGAGAAACAAATGAAGGACAAACAGGACGAGGAGCAGAGGCTTAAGGAGGAGGAAGAAGACAAGAAACGCAAAGAGGAGGAGGAGGCA GAGGACAAGGAGGATGATGAGGACAAAGATGAGGATGAGGAGGATGAGGAGGACAAGGAGGAAGATGAGGAGGAAGATGTCCCCGGCCAG GCCAAGGACGAGCTGTAGAGAGGCCTGCCTCCAGGGCTGGACTGAGGCCTGAGCGCTCCTGCCGCAGAGCTGGCCGCGCCAAATAATGTC TCTGTGAGACTCGAGAACTTTCATTTTTTTCCAGGCTGGTTCGGATTTGGGGTGGATTTTGGTTTTGTTCCCCTCCTCCACTCTCCCCCA CCCCCTCCCCGCCCTTTTTTTTTTTTTTTTTTAAACTGGTATTTTATCTTTGATTCTCCTTCAGCCCTCACCCCTGGTTCTCATCTTTCT TGATCAACATCTTTTCTTGCCTCTGTCCCCTTCTCTCATCTCTTAGCTCCCCTCCAACCTGGGGGGCAGTGGTGTGGAGAAGCCACAGGC CTGAGATTTCATCTGCTCTCCTTCCTGGAGCCCAGAGGAGGGCAGCAGAAGGGGGTGGTGTCTCCAACCCCCCAGCACTGAGGAAGAACG GGGCTCTTCTCATTTCACCCCTCCCTTTCTCCCCTGCCCCCAGGACTGGGCCACTTCTGGGTGGGGCAGTGGGTCCCAGATTGGCTCACA >58938_58938_16_NFIX-CALR_NFIX_chr19_13192669_ENST00000592199_CALR_chr19_13054351_ENST00000316448_length(amino acids)=515AA_BP=417 MYSPYCLTQDEFHPFIEALLPHVRAFSYTWFNLQARKRKYFKKHEKRMSKDEERAVKDELLGEKPEIKQKWASRLLAKLRKDIRPEFRED FVLTITGKKPPCCVLSNPDQKGKIRRIDCLRQADKVWRLDLVMVILFKGIPLESTDGERLYKSPQCSNPGLCVQPHHIGVTIKELDLYLA YFVHTPESGQSDSSNQQGDADIKPLPNGHLSFQDCFVTSGVWNVTELVRVSQTPVATASGPNFSLADLESPSYYNINQVTLGRRSITSPP STSTTKRPKSIDDSEMESPVDDVFYPGTGRSPAAGSSQSSGWPNDVDAGPASLKKSGKLDFCSALSSQGSSPRMAFTHHPLPVLAGVRPG SPRATASALHFPSTSIIQQSSPYFTHPTIRYHHHHGQDSLKEFVQFVCSDGSGQATGQVKSGTIFDNFLITNDEAYAEEFGNETWGVTKA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NFIX-CALR |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | CALR | chr19:13136366 | chr19:13049947 | ENST00000316448 | 0 | 9 | 237_270 | 30.333333333333332 | 418.0 | PPIB | |
| Tgene | CALR | chr19:13136366 | chr19:13049948 | ENST00000316448 | 0 | 9 | 237_270 | 30.333333333333332 | 418.0 | PPIB | |
| Tgene | CALR | chr19:13136366 | chr19:13050242 | ENST00000316448 | 1 | 9 | 237_270 | 64.33333333333333 | 418.0 | PPIB | |
| Tgene | CALR | chr19:13136366 | chr19:13050866 | ENST00000316448 | 2 | 9 | 237_270 | 132.33333333333334 | 418.0 | PPIB |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | CALR | chr19:13192669 | chr19:13054350 | ENST00000316448 | 6 | 9 | 237_270 | 320.0 | 418.0 | PPIB | |
| Tgene | CALR | chr19:13192669 | chr19:13054351 | ENST00000316448 | 6 | 9 | 237_270 | 320.0 | 418.0 | PPIB |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NFIX-CALR |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NFIX-CALR |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
