
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NIN-NEMF (FusionGDB2 ID:59159) |
Fusion Gene Summary for NIN-NEMF |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NIN-NEMF | Fusion gene ID: 59159 | Hgene | Tgene | Gene symbol | NIN | NEMF | Gene ID | 51199 | 9147 |
| Gene name | ninein | nuclear export mediator factor | |
| Synonyms | SCKL7 | NY-CO-1|SDCCAG1 | |
| Cytomap | 14q22.1 | 14q21.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | nineinglycogen synthase kinase 3 beta-interacting proteinhNineinninein (GSK3B interacting protein)ninein centrosomal protein | nuclear export mediator factor NEMFantigen NY-CO-1serologically defined colon cancer antigen 1 | |
| Modification date | 20200328 | 20200313 | |
| UniProtAcc | Q8N4C6 | O60524 | |
| Ensembl transtripts involved in fusion gene | ENST00000245441, ENST00000324330, ENST00000382041, ENST00000382043, ENST00000389868, ENST00000453196, ENST00000530997, ENST00000486200, | ENST00000546046, ENST00000556925, ENST00000382135, ENST00000545773, ENST00000556672, ENST00000298310, | |
| Fusion gene scores | * DoF score | 16 X 18 X 9=2592 | 6 X 8 X 4=192 |
| # samples | 20 | 9 | |
| ** MAII score | log2(20/2592*10)=-3.6959938131099 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/192*10)=-1.09310940439148 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NIN [Title/Abstract] AND NEMF [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NIN(51273455)-NEMF(50292673), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | NIN-NEMF seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. NIN-NEMF seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. NIN-NEMF seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across NIN (5'-gene) Fusion gene breakpoints across NIN (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
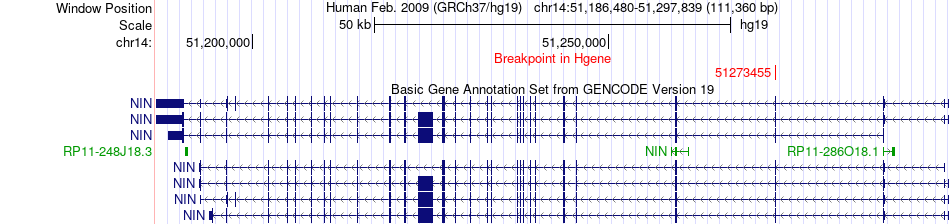 |
 Fusion gene breakpoints across NEMF (3'-gene) Fusion gene breakpoints across NEMF (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
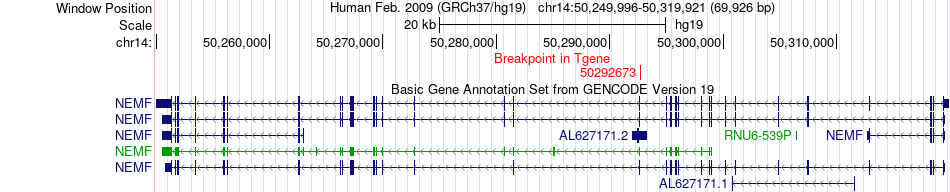 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-BH-A1FE-01A | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
Top |
Fusion Gene ORF analysis for NIN-NEMF |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000245441 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000245441 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000324330 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000324330 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000382041 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000382041 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000382043 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000382043 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000389868 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000389868 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000453196 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000453196 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000530997 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-5UTR | ENST00000530997 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000245441 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000245441 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000245441 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000324330 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000324330 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000324330 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000382041 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000382041 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000382041 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000382043 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000382043 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000382043 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000389868 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000389868 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000389868 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000453196 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000453196 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000453196 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000530997 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000530997 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| 5CDS-intron | ENST00000530997 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| Frame-shift | ENST00000245441 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| Frame-shift | ENST00000382041 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| Frame-shift | ENST00000453196 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| Frame-shift | ENST00000530997 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| In-frame | ENST00000324330 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| In-frame | ENST00000382043 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| In-frame | ENST00000389868 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| intron-3CDS | ENST00000486200 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| intron-5UTR | ENST00000486200 | ENST00000546046 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| intron-5UTR | ENST00000486200 | ENST00000556925 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| intron-intron | ENST00000486200 | ENST00000382135 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| intron-intron | ENST00000486200 | ENST00000545773 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
| intron-intron | ENST00000486200 | ENST00000556672 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000389868 | NIN | chr14 | 51273455 | - | ENST00000298310 | NEMF | chr14 | 50292673 | - | 3556 | 456 | 432 | 2198 | 588 |
| ENST00000382043 | NIN | chr14 | 51273455 | - | ENST00000298310 | NEMF | chr14 | 50292673 | - | 3402 | 302 | 278 | 2044 | 588 |
| ENST00000324330 | NIN | chr14 | 51273455 | - | ENST00000298310 | NEMF | chr14 | 50292673 | - | 3556 | 456 | 432 | 2198 | 588 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000389868 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - | 0.000310878 | 0.9996891 |
| ENST00000382043 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - | 0.000345837 | 0.9996542 |
| ENST00000324330 | ENST00000298310 | NIN | chr14 | 51273455 | - | NEMF | chr14 | 50292673 | - | 0.000310878 | 0.9996891 |
Top |
Fusion Genomic Features for NIN-NEMF |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
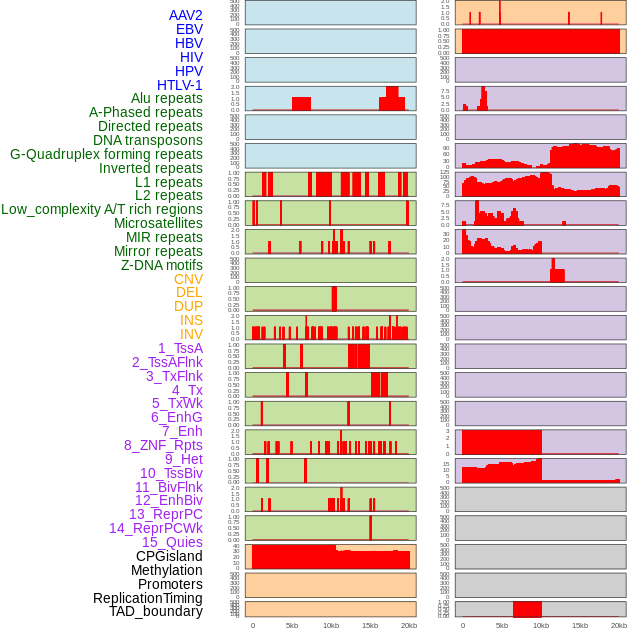 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for NIN-NEMF |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr14:51273455/chr14:50292673) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| NIN | NEMF |
| FUNCTION: Centrosomal protein required in the positioning and anchorage of the microtubule minus-end in epithelial cells (PubMed:15190203, PubMed:23386061). May also act as a centrosome maturation factor (PubMed:11956314). May play a role in microtubule nucleation, by recruiting the gamma-tubulin ring complex to the centrosome (PubMed:15190203). Overexpression does not perturb nucleation or elongation of microtubules but suppresses release of microtubules (PubMed:15190203). Required for centriole organization and microtubule anchoring at the mother centriole (PubMed:23386061). {ECO:0000269|PubMed:11956314, ECO:0000269|PubMed:15190203, ECO:0000269|PubMed:23386061}. | FUNCTION: Component of the ribosome quality control complex (RQC), a ribosome-associated complex that mediates ubiquitination and extraction of incompletely synthesized nascent chains for proteasomal degradation. NEMF is responsible for selective recognition of stalled 60S subunits by recognizing an exposed, nascent chain-conjugated tRNA moiety. NEMF is important for the stable association of LTN1 to the complex (PubMed:25578875). May indirectly play a role in nuclear export (PubMed:16103875). {ECO:0000269|PubMed:16103875, ECO:0000269|PubMed:25578875}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 42_77 | 88 | 2134.0 | Domain | EF-hand 2 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 8_43 | 88 | 2134.0 | Domain | EF-hand 1 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 42_77 | 88 | 2091.0 | Domain | EF-hand 2 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 8_43 | 88 | 2091.0 | Domain | EF-hand 1 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 42_77 | 88 | 1378.0 | Domain | EF-hand 2 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 8_43 | 88 | 1378.0 | Domain | EF-hand 1 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 42_77 | 88 | 2134.0 | Domain | EF-hand 2 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 8_43 | 88 | 2134.0 | Domain | EF-hand 1 |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000298310 | 14 | 33 | 869_894 | 496 | 1077.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000382135 | 0 | 9 | 296_359 | 0 | 277.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000382135 | 0 | 9 | 483_514 | 0 | 277.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000382135 | 0 | 9 | 869_894 | 0 | 277.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000545773 | 13 | 32 | 483_514 | 454 | 1035.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000545773 | 13 | 32 | 869_894 | 454 | 1035.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000546046 | 14 | 32 | 869_894 | 496 | 1056.0 | Coiled coil | Ontology_term=ECO:0000255 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 1068_1099 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 1181_1341 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 1441_1816 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 1854_1885 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 1922_2067 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 357_570 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 625_1027 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 1068_1099 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 1181_1341 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 1441_1816 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 1854_1885 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 1922_2067 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 357_570 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 625_1027 | 88 | 2091.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 1068_1099 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 1181_1341 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 1441_1816 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 1854_1885 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 1922_2067 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 357_570 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 625_1027 | 88 | 1378.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 1068_1099 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 1181_1341 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 1441_1816 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 1854_1885 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 1922_2067 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 357_570 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 625_1027 | 88 | 2134.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 182_217 | 88 | 2134.0 | Domain | EF-hand 3 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 219_252 | 88 | 2134.0 | Domain | EF-hand 4 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 317_352 | 88 | 2134.0 | Domain | EF-hand 5 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 182_217 | 88 | 2091.0 | Domain | EF-hand 3 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 219_252 | 88 | 2091.0 | Domain | EF-hand 4 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 317_352 | 88 | 2091.0 | Domain | EF-hand 5 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 182_217 | 88 | 1378.0 | Domain | EF-hand 3 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 219_252 | 88 | 1378.0 | Domain | EF-hand 4 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 317_352 | 88 | 1378.0 | Domain | EF-hand 5 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 182_217 | 88 | 2134.0 | Domain | EF-hand 3 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 219_252 | 88 | 2134.0 | Domain | EF-hand 4 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 317_352 | 88 | 2134.0 | Domain | EF-hand 5 |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 245_252 | 88 | 2134.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 300_304 | 88 | 2134.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000245441 | - | 4 | 31 | 420_423 | 88 | 2134.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 245_252 | 88 | 2091.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 300_304 | 88 | 2091.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382041 | - | 4 | 30 | 420_423 | 88 | 2091.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 245_252 | 88 | 1378.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 300_304 | 88 | 1378.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000382043 | - | 3 | 28 | 420_423 | 88 | 1378.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 245_252 | 88 | 2134.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 300_304 | 88 | 2134.0 | Nucleotide binding | GTP |
| Hgene | NIN | chr14:51273455 | chr14:50292673 | ENST00000530997 | - | 2 | 29 | 420_423 | 88 | 2134.0 | Nucleotide binding | GTP |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000298310 | 14 | 33 | 296_359 | 496 | 1077.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000298310 | 14 | 33 | 483_514 | 496 | 1077.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000545773 | 13 | 32 | 296_359 | 454 | 1035.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000546046 | 14 | 32 | 296_359 | 496 | 1056.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NEMF | chr14:51273455 | chr14:50292673 | ENST00000546046 | 14 | 32 | 483_514 | 496 | 1056.0 | Coiled coil | Ontology_term=ECO:0000255 |
Top |
Fusion Gene Sequence for NIN-NEMF |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >59159_59159_1_NIN-NEMF_NIN_chr14_51273455_ENST00000324330_NEMF_chr14_50292673_ENST00000298310_length(transcript)=3556nt_BP=456nt CCGAGCCGGGAGCCGAGCGCGCTGGGCGCGGCCGTCCCGCTGCCGCCGCCAAGCCCGGAGTGCGCCAGCGGCCATGGTTCCCGCGGCGGC GCCGGGCGCCTGAGCCCCGGGACGCAAGCGCTGGAGGCGGGCTGCCGGCTGTGCGGGCGCTCCGGAGAGACCGACAGAAGGTGAGCACTG TGGGCTATGGGATGGATGAGGTGGAGCAGGACCAGCATGAGGCCCGACTCAAGGAGCTGTTTGACAGTTTTGACACGACGGGCACAGGGT CCCTGGGGCAGGAGGAACTCACCGACCTTTGCCACATGTTGAGCTTGGAGGAGGTGGCCCCAGTGCTGCAGCAGACATTACTTCAGGACA ACCTCTTGGGCAGGGTACATTTTGACCAATTTAAAGAAGCATTAATACTCATCTTGTCCAGAACTCTGTCAAATGAAGAACACTTTCAAG AACCAGGCATTCAAGTCAGCAGAAAAGAAAACAAAGCAAACATTAAAAGAAGTTCAGACTGTTACCTCTATTCAAAAAGCAAGAAAAGTA TATTGGTTTGAGAAATTTCTGTGGTTCATTAGCTCAGAGAACTATCTAATTATAGGTGGACGAGATCAGCAACAGAATGAAATAATTGTG AAAAGATACTTGACACCAGGAGACATTTATGTACATGCTGATCTTCATGGAGCTACTAGCTGTGTAATTAAGAATCCAACAGGAGAACCC ATCCCCCCACGGACCTTGACTGAAGCTGGCACAATGGCACTTTGCTACAGTGCTGCTTGGGATGCACGAGTTATCACTAGTGCTTGGTGG GTGTACCATCATCAGGTATCTAAAACAGCACCAACTGGAGAATATTTGACAACAGGAAGCTTCATGATAAGAGGAAAAAAGAATTTTCTT CCTCCCTCATATCTAATGATGGGGTTTAGCTTCCTTTTTAAGGTAGATGAGTCTTGTGTTTGGAGACATCAGGGTGAACGAAAAGTCAGA GTACAGGATGAAGACATGGAGACACTGGCAAGTTGTACAAGTGAACTCATATCAGAAGAAATGGAACAATTAGATGGAGGTGACACGAGC AGTGATGAGGATAAAGAAGAACATGAAACTCCTGTGGAAGTAGAACTCATGACTCAGGTTGACCAAGAGGATATCACTCTTCAGAGTGGC AGAGATGAACTAAATGAGGAGCTCATTCAGGAAGAAAGCTCTGAAGACGAAGGAGAATATGAAGAGGTTAGAAAAGATCAGGATTCTGTT GGTGAAATGAAGGATGAAGGGGAAGAGACATTAAATTATCCTGATACTACCATTGACTTGTCTCACCTTCAACCCCAAAGGTCCATCCAG AAATTGGCTTCAAAAGAGGAATCTTCTAATTCTAGTGACAGTAAATCACAGAGCCGGAGACATTTGTCAGCCAAGGAAAGAAGGGAAATG AAAAAGAAAAAACTTCCAAGTGACTCAGGAGATTTAGAAGCGTTAGAGGGAAAGGATAAAGAAAAAGAAAGTACTGTACACATTGAAACT CATCAGAACACAAGCAAAAATGTTGCGGCTGTGCAGCCAATGAAACGAGGACAAAAGAGTAAAATGAAAAAAATGAAAGAAAAATACAAA GACCAGGATGAAGAAGACCGTGAACTTATCATGAAGTTGCTGGGGTCTGCAGGTTCAAACAAAGAAGAAAAAGGGAAGAAGGGGAAGAAA GGAAAAACAAAGGACGAACCTGTGAAGAAACAGCCCCAGAAACCTAGAGGTGGACAGAGGGTCTCTGACAACATTAAGAAAGAAACTCCG TTCCTTGAGGTTATAACTCATGAGTTACAAGACTTTGCTGTAGATGATCCACATGATGACAAGGAAGAGCAAGATCTGGATCAACAGGGA AATGAGGAAAACCTATTTGATTCTTTGACAGGCCAGCCACATCCTGAAGATGTACTACTGTTTGCCATTCCAATATGTGCCCCTTACACC ACCATGACAAACTACAAATATAAAGTGAAACTTACTCCTGGAGTGCAGAAAAAGGGAAAAGCTGCAAAAACAGCCTTGAATAGTTTCATG CATTCCAAAGAAGCAACAGCAAGAGAAAAAGACTTATTCCGCAGCGTAAAGGACACAGATTTATCAAGAAACATTCCTGGCAAAGTGAAA GTGTCTGCACCCAATCTTCTGAATGTAAAAAGGAAATAGCTGAAATGAAATTCTAAAATATTTGAGAAGAGCCAATTTTATAGCCTTTTG GAAGTTCAAAGATGAAAGCACCATGTATCAGGATTTCCGCATTATAAAAATGAACTAAACATTGCCTTGCTATATTCACCAAAAGGACTT ACTTCTTGTTTTTTCCCAGTTTTATATAGAGGAAACACTGTCTATGATAGGATTTCCAAAAGTATTTGTGGACAGTTAAATGCTAATTAA TATACATCTGTAGTTATTCTACATTTTCTTGAAATTTGGGAGGTTAATACCAAGTATTCATTTCATGATGTAAAGAAACTGAACAGTGAA GTGGCTTGATTGCTTAAACTATTGACTTGGTAAGTCTACTGTATATAACATCTAATATATATATTACAGGCCAAATGAACTAAACATTGC CTTGCTATATTCACCAAAAGGACTTAATTCTTGTTTTTTTCCCAGTTTTATATAGAGGAAACACTATGATAGGATTTCCTAAAGTATTTG TGGACAGTTAAATGCTAATTATATACATCTGTAGTTATTCTACATTTTCTTGAAATTTGGGAGGTTAATACCAAGTATTCATTTCATGAT GTAAAGAAACTGAACAGTGAAGTGGCTTGATTGCTTAAACTATTGACTTGGTAAGTCTACTGTATATAACATCTAATATATATATATTAT AGGCCAGCTACAAGGGGTTTAAATATTTAGGATTGTGTCTTGAAAACTAAGTATTGGAGTGGATTTTCTTCTGCTTTCATTGATACTTGT CAGAAAAAAATATTAGACCAAAATGTAAAATATAAGTAATAATTCTCATGACTCAGCTTAATCATTTCTATTCATTTGTAGACCTTCAGG TTTGATCTTAATTTTTTCCTTGTAATCTGATTCAGTTCTGTACCATTTTAATTTCCACCTTTAGTAATTAATTACCAATATAGCACGTTA GAGTATCTTAACCTATAATACCAGAAGCGCATTTTAGGCTTACCACTAAAGACTCTAACCCACTTTTCCTCTGTCTCTTAACAGAATGGA AAATGATGTCATCATTATTTCCATTCCTTCATGAGCTATAAAACCTATTAATTAAAAAAAAACTTTTTAATTGACTGGACAGCACCTGCC TGTATTCAGCATGACCACTTGCTCCAGCCCCTACACTGAAGCATTAAGGTCAAAGTACTGAACAGTTGTTGGTCCCTCCACACTTTGTTT TTCCAGCTTCGTCATAATGTGAATATTAATGTTTAAAAAAAAAAACCTTTATTATATTGTATTTACAACACTAGGTTACATAACTATAGA >59159_59159_1_NIN-NEMF_NIN_chr14_51273455_ENST00000324330_NEMF_chr14_50292673_ENST00000298310_length(amino acids)=588AA_BP=8 MKNTFKNQAFKSAEKKTKQTLKEVQTVTSIQKARKVYWFEKFLWFISSENYLIIGGRDQQQNEIIVKRYLTPGDIYVHADLHGATSCVIK NPTGEPIPPRTLTEAGTMALCYSAAWDARVITSAWWVYHHQVSKTAPTGEYLTTGSFMIRGKKNFLPPSYLMMGFSFLFKVDESCVWRHQ GERKVRVQDEDMETLASCTSELISEEMEQLDGGDTSSDEDKEEHETPVEVELMTQVDQEDITLQSGRDELNEELIQEESSEDEGEYEEVR KDQDSVGEMKDEGEETLNYPDTTIDLSHLQPQRSIQKLASKEESSNSSDSKSQSRRHLSAKERREMKKKKLPSDSGDLEALEGKDKEKES TVHIETHQNTSKNVAAVQPMKRGQKSKMKKMKEKYKDQDEEDRELIMKLLGSAGSNKEEKGKKGKKGKTKDEPVKKQPQKPRGGQRVSDN IKKETPFLEVITHELQDFAVDDPHDDKEEQDLDQQGNEENLFDSLTGQPHPEDVLLFAIPICAPYTTMTNYKYKVKLTPGVQKKGKAAKT -------------------------------------------------------------- >59159_59159_2_NIN-NEMF_NIN_chr14_51273455_ENST00000382043_NEMF_chr14_50292673_ENST00000298310_length(transcript)=3402nt_BP=302nt GAGAGACCGACAGAAGGTGAGCACTGTGGGCTATGGGATGGATGAGGTGGAGCAGGACCAGCATGAGGCCCGACTCAAGGAGCTGTTTGA CAGTTTTGACACGACGGGCACAGGGTCCCTGGGGCAGGAGGAACTCACCGACCTTTGCCACATGTTGAGCTTGGAGGAGGTGGCCCCAGT GCTGCAGCAGACATTACTTCAGGACAACCTCTTGGGCAGGGTACATTTTGACCAATTTAAAGAAGCATTAATACTCATCTTGTCCAGAAC TCTGTCAAATGAAGAACACTTTCAAGAACCAGGCATTCAAGTCAGCAGAAAAGAAAACAAAGCAAACATTAAAAGAAGTTCAGACTGTTA CCTCTATTCAAAAAGCAAGAAAAGTATATTGGTTTGAGAAATTTCTGTGGTTCATTAGCTCAGAGAACTATCTAATTATAGGTGGACGAG ATCAGCAACAGAATGAAATAATTGTGAAAAGATACTTGACACCAGGAGACATTTATGTACATGCTGATCTTCATGGAGCTACTAGCTGTG TAATTAAGAATCCAACAGGAGAACCCATCCCCCCACGGACCTTGACTGAAGCTGGCACAATGGCACTTTGCTACAGTGCTGCTTGGGATG CACGAGTTATCACTAGTGCTTGGTGGGTGTACCATCATCAGGTATCTAAAACAGCACCAACTGGAGAATATTTGACAACAGGAAGCTTCA TGATAAGAGGAAAAAAGAATTTTCTTCCTCCCTCATATCTAATGATGGGGTTTAGCTTCCTTTTTAAGGTAGATGAGTCTTGTGTTTGGA GACATCAGGGTGAACGAAAAGTCAGAGTACAGGATGAAGACATGGAGACACTGGCAAGTTGTACAAGTGAACTCATATCAGAAGAAATGG AACAATTAGATGGAGGTGACACGAGCAGTGATGAGGATAAAGAAGAACATGAAACTCCTGTGGAAGTAGAACTCATGACTCAGGTTGACC AAGAGGATATCACTCTTCAGAGTGGCAGAGATGAACTAAATGAGGAGCTCATTCAGGAAGAAAGCTCTGAAGACGAAGGAGAATATGAAG AGGTTAGAAAAGATCAGGATTCTGTTGGTGAAATGAAGGATGAAGGGGAAGAGACATTAAATTATCCTGATACTACCATTGACTTGTCTC ACCTTCAACCCCAAAGGTCCATCCAGAAATTGGCTTCAAAAGAGGAATCTTCTAATTCTAGTGACAGTAAATCACAGAGCCGGAGACATT TGTCAGCCAAGGAAAGAAGGGAAATGAAAAAGAAAAAACTTCCAAGTGACTCAGGAGATTTAGAAGCGTTAGAGGGAAAGGATAAAGAAA AAGAAAGTACTGTACACATTGAAACTCATCAGAACACAAGCAAAAATGTTGCGGCTGTGCAGCCAATGAAACGAGGACAAAAGAGTAAAA TGAAAAAAATGAAAGAAAAATACAAAGACCAGGATGAAGAAGACCGTGAACTTATCATGAAGTTGCTGGGGTCTGCAGGTTCAAACAAAG AAGAAAAAGGGAAGAAGGGGAAGAAAGGAAAAACAAAGGACGAACCTGTGAAGAAACAGCCCCAGAAACCTAGAGGTGGACAGAGGGTCT CTGACAACATTAAGAAAGAAACTCCGTTCCTTGAGGTTATAACTCATGAGTTACAAGACTTTGCTGTAGATGATCCACATGATGACAAGG AAGAGCAAGATCTGGATCAACAGGGAAATGAGGAAAACCTATTTGATTCTTTGACAGGCCAGCCACATCCTGAAGATGTACTACTGTTTG CCATTCCAATATGTGCCCCTTACACCACCATGACAAACTACAAATATAAAGTGAAACTTACTCCTGGAGTGCAGAAAAAGGGAAAAGCTG CAAAAACAGCCTTGAATAGTTTCATGCATTCCAAAGAAGCAACAGCAAGAGAAAAAGACTTATTCCGCAGCGTAAAGGACACAGATTTAT CAAGAAACATTCCTGGCAAAGTGAAAGTGTCTGCACCCAATCTTCTGAATGTAAAAAGGAAATAGCTGAAATGAAATTCTAAAATATTTG AGAAGAGCCAATTTTATAGCCTTTTGGAAGTTCAAAGATGAAAGCACCATGTATCAGGATTTCCGCATTATAAAAATGAACTAAACATTG CCTTGCTATATTCACCAAAAGGACTTACTTCTTGTTTTTTCCCAGTTTTATATAGAGGAAACACTGTCTATGATAGGATTTCCAAAAGTA TTTGTGGACAGTTAAATGCTAATTAATATACATCTGTAGTTATTCTACATTTTCTTGAAATTTGGGAGGTTAATACCAAGTATTCATTTC ATGATGTAAAGAAACTGAACAGTGAAGTGGCTTGATTGCTTAAACTATTGACTTGGTAAGTCTACTGTATATAACATCTAATATATATAT TACAGGCCAAATGAACTAAACATTGCCTTGCTATATTCACCAAAAGGACTTAATTCTTGTTTTTTTCCCAGTTTTATATAGAGGAAACAC TATGATAGGATTTCCTAAAGTATTTGTGGACAGTTAAATGCTAATTATATACATCTGTAGTTATTCTACATTTTCTTGAAATTTGGGAGG TTAATACCAAGTATTCATTTCATGATGTAAAGAAACTGAACAGTGAAGTGGCTTGATTGCTTAAACTATTGACTTGGTAAGTCTACTGTA TATAACATCTAATATATATATATTATAGGCCAGCTACAAGGGGTTTAAATATTTAGGATTGTGTCTTGAAAACTAAGTATTGGAGTGGAT TTTCTTCTGCTTTCATTGATACTTGTCAGAAAAAAATATTAGACCAAAATGTAAAATATAAGTAATAATTCTCATGACTCAGCTTAATCA TTTCTATTCATTTGTAGACCTTCAGGTTTGATCTTAATTTTTTCCTTGTAATCTGATTCAGTTCTGTACCATTTTAATTTCCACCTTTAG TAATTAATTACCAATATAGCACGTTAGAGTATCTTAACCTATAATACCAGAAGCGCATTTTAGGCTTACCACTAAAGACTCTAACCCACT TTTCCTCTGTCTCTTAACAGAATGGAAAATGATGTCATCATTATTTCCATTCCTTCATGAGCTATAAAACCTATTAATTAAAAAAAAACT TTTTAATTGACTGGACAGCACCTGCCTGTATTCAGCATGACCACTTGCTCCAGCCCCTACACTGAAGCATTAAGGTCAAAGTACTGAACA GTTGTTGGTCCCTCCACACTTTGTTTTTCCAGCTTCGTCATAATGTGAATATTAATGTTTAAAAAAAAAAACCTTTATTATATTGTATTT >59159_59159_2_NIN-NEMF_NIN_chr14_51273455_ENST00000382043_NEMF_chr14_50292673_ENST00000298310_length(amino acids)=588AA_BP=8 MKNTFKNQAFKSAEKKTKQTLKEVQTVTSIQKARKVYWFEKFLWFISSENYLIIGGRDQQQNEIIVKRYLTPGDIYVHADLHGATSCVIK NPTGEPIPPRTLTEAGTMALCYSAAWDARVITSAWWVYHHQVSKTAPTGEYLTTGSFMIRGKKNFLPPSYLMMGFSFLFKVDESCVWRHQ GERKVRVQDEDMETLASCTSELISEEMEQLDGGDTSSDEDKEEHETPVEVELMTQVDQEDITLQSGRDELNEELIQEESSEDEGEYEEVR KDQDSVGEMKDEGEETLNYPDTTIDLSHLQPQRSIQKLASKEESSNSSDSKSQSRRHLSAKERREMKKKKLPSDSGDLEALEGKDKEKES TVHIETHQNTSKNVAAVQPMKRGQKSKMKKMKEKYKDQDEEDRELIMKLLGSAGSNKEEKGKKGKKGKTKDEPVKKQPQKPRGGQRVSDN IKKETPFLEVITHELQDFAVDDPHDDKEEQDLDQQGNEENLFDSLTGQPHPEDVLLFAIPICAPYTTMTNYKYKVKLTPGVQKKGKAAKT -------------------------------------------------------------- >59159_59159_3_NIN-NEMF_NIN_chr14_51273455_ENST00000389868_NEMF_chr14_50292673_ENST00000298310_length(transcript)=3556nt_BP=456nt CCGAGCCGGGAGCCGAGCGCGCTGGGCGCGGCCGTCCCGCTGCCGCCGCCAAGCCCGGAGTGCGCCAGCGGCCATGGTTCCCGCGGCGGC GCCGGGCGCCTGAGCCCCGGGACGCAAGCGCTGGAGGCGGGCTGCCGGCTGTGCGGGCGCTCCGGAGAGACCGACAGAAGGTGAGCACTG TGGGCTATGGGATGGATGAGGTGGAGCAGGACCAGCATGAGGCCCGACTCAAGGAGCTGTTTGACAGTTTTGACACGACGGGCACAGGGT CCCTGGGGCAGGAGGAACTCACCGACCTTTGCCACATGTTGAGCTTGGAGGAGGTGGCCCCAGTGCTGCAGCAGACATTACTTCAGGACA ACCTCTTGGGCAGGGTACATTTTGACCAATTTAAAGAAGCATTAATACTCATCTTGTCCAGAACTCTGTCAAATGAAGAACACTTTCAAG AACCAGGCATTCAAGTCAGCAGAAAAGAAAACAAAGCAAACATTAAAAGAAGTTCAGACTGTTACCTCTATTCAAAAAGCAAGAAAAGTA TATTGGTTTGAGAAATTTCTGTGGTTCATTAGCTCAGAGAACTATCTAATTATAGGTGGACGAGATCAGCAACAGAATGAAATAATTGTG AAAAGATACTTGACACCAGGAGACATTTATGTACATGCTGATCTTCATGGAGCTACTAGCTGTGTAATTAAGAATCCAACAGGAGAACCC ATCCCCCCACGGACCTTGACTGAAGCTGGCACAATGGCACTTTGCTACAGTGCTGCTTGGGATGCACGAGTTATCACTAGTGCTTGGTGG GTGTACCATCATCAGGTATCTAAAACAGCACCAACTGGAGAATATTTGACAACAGGAAGCTTCATGATAAGAGGAAAAAAGAATTTTCTT CCTCCCTCATATCTAATGATGGGGTTTAGCTTCCTTTTTAAGGTAGATGAGTCTTGTGTTTGGAGACATCAGGGTGAACGAAAAGTCAGA GTACAGGATGAAGACATGGAGACACTGGCAAGTTGTACAAGTGAACTCATATCAGAAGAAATGGAACAATTAGATGGAGGTGACACGAGC AGTGATGAGGATAAAGAAGAACATGAAACTCCTGTGGAAGTAGAACTCATGACTCAGGTTGACCAAGAGGATATCACTCTTCAGAGTGGC AGAGATGAACTAAATGAGGAGCTCATTCAGGAAGAAAGCTCTGAAGACGAAGGAGAATATGAAGAGGTTAGAAAAGATCAGGATTCTGTT GGTGAAATGAAGGATGAAGGGGAAGAGACATTAAATTATCCTGATACTACCATTGACTTGTCTCACCTTCAACCCCAAAGGTCCATCCAG AAATTGGCTTCAAAAGAGGAATCTTCTAATTCTAGTGACAGTAAATCACAGAGCCGGAGACATTTGTCAGCCAAGGAAAGAAGGGAAATG AAAAAGAAAAAACTTCCAAGTGACTCAGGAGATTTAGAAGCGTTAGAGGGAAAGGATAAAGAAAAAGAAAGTACTGTACACATTGAAACT CATCAGAACACAAGCAAAAATGTTGCGGCTGTGCAGCCAATGAAACGAGGACAAAAGAGTAAAATGAAAAAAATGAAAGAAAAATACAAA GACCAGGATGAAGAAGACCGTGAACTTATCATGAAGTTGCTGGGGTCTGCAGGTTCAAACAAAGAAGAAAAAGGGAAGAAGGGGAAGAAA GGAAAAACAAAGGACGAACCTGTGAAGAAACAGCCCCAGAAACCTAGAGGTGGACAGAGGGTCTCTGACAACATTAAGAAAGAAACTCCG TTCCTTGAGGTTATAACTCATGAGTTACAAGACTTTGCTGTAGATGATCCACATGATGACAAGGAAGAGCAAGATCTGGATCAACAGGGA AATGAGGAAAACCTATTTGATTCTTTGACAGGCCAGCCACATCCTGAAGATGTACTACTGTTTGCCATTCCAATATGTGCCCCTTACACC ACCATGACAAACTACAAATATAAAGTGAAACTTACTCCTGGAGTGCAGAAAAAGGGAAAAGCTGCAAAAACAGCCTTGAATAGTTTCATG CATTCCAAAGAAGCAACAGCAAGAGAAAAAGACTTATTCCGCAGCGTAAAGGACACAGATTTATCAAGAAACATTCCTGGCAAAGTGAAA GTGTCTGCACCCAATCTTCTGAATGTAAAAAGGAAATAGCTGAAATGAAATTCTAAAATATTTGAGAAGAGCCAATTTTATAGCCTTTTG GAAGTTCAAAGATGAAAGCACCATGTATCAGGATTTCCGCATTATAAAAATGAACTAAACATTGCCTTGCTATATTCACCAAAAGGACTT ACTTCTTGTTTTTTCCCAGTTTTATATAGAGGAAACACTGTCTATGATAGGATTTCCAAAAGTATTTGTGGACAGTTAAATGCTAATTAA TATACATCTGTAGTTATTCTACATTTTCTTGAAATTTGGGAGGTTAATACCAAGTATTCATTTCATGATGTAAAGAAACTGAACAGTGAA GTGGCTTGATTGCTTAAACTATTGACTTGGTAAGTCTACTGTATATAACATCTAATATATATATTACAGGCCAAATGAACTAAACATTGC CTTGCTATATTCACCAAAAGGACTTAATTCTTGTTTTTTTCCCAGTTTTATATAGAGGAAACACTATGATAGGATTTCCTAAAGTATTTG TGGACAGTTAAATGCTAATTATATACATCTGTAGTTATTCTACATTTTCTTGAAATTTGGGAGGTTAATACCAAGTATTCATTTCATGAT GTAAAGAAACTGAACAGTGAAGTGGCTTGATTGCTTAAACTATTGACTTGGTAAGTCTACTGTATATAACATCTAATATATATATATTAT AGGCCAGCTACAAGGGGTTTAAATATTTAGGATTGTGTCTTGAAAACTAAGTATTGGAGTGGATTTTCTTCTGCTTTCATTGATACTTGT CAGAAAAAAATATTAGACCAAAATGTAAAATATAAGTAATAATTCTCATGACTCAGCTTAATCATTTCTATTCATTTGTAGACCTTCAGG TTTGATCTTAATTTTTTCCTTGTAATCTGATTCAGTTCTGTACCATTTTAATTTCCACCTTTAGTAATTAATTACCAATATAGCACGTTA GAGTATCTTAACCTATAATACCAGAAGCGCATTTTAGGCTTACCACTAAAGACTCTAACCCACTTTTCCTCTGTCTCTTAACAGAATGGA AAATGATGTCATCATTATTTCCATTCCTTCATGAGCTATAAAACCTATTAATTAAAAAAAAACTTTTTAATTGACTGGACAGCACCTGCC TGTATTCAGCATGACCACTTGCTCCAGCCCCTACACTGAAGCATTAAGGTCAAAGTACTGAACAGTTGTTGGTCCCTCCACACTTTGTTT TTCCAGCTTCGTCATAATGTGAATATTAATGTTTAAAAAAAAAAACCTTTATTATATTGTATTTACAACACTAGGTTACATAACTATAGA >59159_59159_3_NIN-NEMF_NIN_chr14_51273455_ENST00000389868_NEMF_chr14_50292673_ENST00000298310_length(amino acids)=588AA_BP=8 MKNTFKNQAFKSAEKKTKQTLKEVQTVTSIQKARKVYWFEKFLWFISSENYLIIGGRDQQQNEIIVKRYLTPGDIYVHADLHGATSCVIK NPTGEPIPPRTLTEAGTMALCYSAAWDARVITSAWWVYHHQVSKTAPTGEYLTTGSFMIRGKKNFLPPSYLMMGFSFLFKVDESCVWRHQ GERKVRVQDEDMETLASCTSELISEEMEQLDGGDTSSDEDKEEHETPVEVELMTQVDQEDITLQSGRDELNEELIQEESSEDEGEYEEVR KDQDSVGEMKDEGEETLNYPDTTIDLSHLQPQRSIQKLASKEESSNSSDSKSQSRRHLSAKERREMKKKKLPSDSGDLEALEGKDKEKES TVHIETHQNTSKNVAAVQPMKRGQKSKMKKMKEKYKDQDEEDRELIMKLLGSAGSNKEEKGKKGKKGKTKDEPVKKQPQKPRGGQRVSDN IKKETPFLEVITHELQDFAVDDPHDDKEEQDLDQQGNEENLFDSLTGQPHPEDVLLFAIPICAPYTTMTNYKYKVKLTPGVQKKGKAAKT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NIN-NEMF |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NIN-NEMF |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NIN-NEMF |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
