
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NLK-LGALS9 (FusionGDB2 ID:59350) |
Fusion Gene Summary for NLK-LGALS9 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NLK-LGALS9 | Fusion gene ID: 59350 | Hgene | Tgene | Gene symbol | NLK | LGALS9 | Gene ID | 51701 | 3965 |
| Gene name | nemo like kinase | galectin 9 | |
| Synonyms | - | HUAT|LGALS9A | |
| Cytomap | 17q11.2 | 17q11.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | serine/threonine-protein kinase NLK | galectin-9ecalectingal-9lectin, galactoside-binding, soluble, 9tumor antigen HOM-HD-21urate transporter/channel protein | |
| Modification date | 20200313 | 20200315 | |
| UniProtAcc | Q9UBE8 | Q3B8N2 | |
| Ensembl transtripts involved in fusion gene | ENST00000583517, ENST00000407008, ENST00000582037, | ENST00000302228, ENST00000310394, ENST00000313648, ENST00000395473, ENST00000448970, ENST00000413914, | |
| Fusion gene scores | * DoF score | 18 X 10 X 9=1620 | 8 X 8 X 5=320 |
| # samples | 20 | 9 | |
| ** MAII score | log2(20/1620*10)=-3.01792190799726 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/320*10)=-1.83007499855769 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NLK [Title/Abstract] AND LGALS9 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NLK(26370357)-LGALS9(25967598), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | NLK | GO:0050821 | protein stabilization | 25512613 |
| Tgene | LGALS9 | GO:0006954 | inflammatory response | 23817958 |
| Tgene | LGALS9 | GO:0007565 | female pregnancy | 23242525 |
| Tgene | LGALS9 | GO:0010628 | positive regulation of gene expression | 16116184 |
| Tgene | LGALS9 | GO:0010629 | negative regulation of gene expression | 23408620 |
| Tgene | LGALS9 | GO:0032673 | regulation of interleukin-4 production | 16116184 |
| Tgene | LGALS9 | GO:0032674 | regulation of interleukin-5 production | 16116184 |
| Tgene | LGALS9 | GO:0032689 | negative regulation of interferon-gamma production | 23408620 |
| Tgene | LGALS9 | GO:0032753 | positive regulation of interleukin-4 production | 20209097 |
| Tgene | LGALS9 | GO:0032834 | positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation involved in immune response | 20209097 |
| Tgene | LGALS9 | GO:0034134 | toll-like receptor 2 signaling pathway | 16116184 |
| Tgene | LGALS9 | GO:0034142 | toll-like receptor 4 signaling pathway | 16116184 |
| Tgene | LGALS9 | GO:0038066 | p38MAPK cascade | 16116184 |
| Tgene | LGALS9 | GO:0045953 | negative regulation of natural killer cell mediated cytotoxicity | 23408620 |
| Tgene | LGALS9 | GO:0046598 | positive regulation of viral entry into host cell | 21670307 |
| Tgene | LGALS9 | GO:0050718 | positive regulation of interleukin-1 beta secretion | 20209097 |
| Tgene | LGALS9 | GO:0060135 | maternal process involved in female pregnancy | 25578313 |
| Tgene | LGALS9 | GO:0070241 | positive regulation of activated T cell autonomous cell death | 20209097 |
| Tgene | LGALS9 | GO:0070371 | ERK1 and ERK2 cascade | 16116184 |
| Tgene | LGALS9 | GO:0070555 | response to interleukin-1 | 23817958 |
| Tgene | LGALS9 | GO:0071346 | cellular response to interferon-gamma | 23817958 |
| Tgene | LGALS9 | GO:0071636 | positive regulation of transforming growth factor beta production | 20209097 |
| Tgene | LGALS9 | GO:1902715 | positive regulation of interferon-gamma secretion | 20209097 |
| Tgene | LGALS9 | GO:1904469 | positive regulation of tumor necrosis factor secretion | 20209097 |
| Tgene | LGALS9 | GO:2000562 | negative regulation of CD4-positive, alpha-beta T cell proliferation | 20209097 |
| Tgene | LGALS9 | GO:2000563 | positive regulation of CD4-positive, alpha-beta T cell proliferation | 16116184 |
| Tgene | LGALS9 | GO:2000667 | positive regulation of interleukin-13 secretion | 20209097 |
| Tgene | LGALS9 | GO:2000670 | positive regulation of dendritic cell apoptotic process | 16116184 |
| Tgene | LGALS9 | GO:2001181 | positive regulation of interleukin-10 secretion | 16116184|20209097 |
| Tgene | LGALS9 | GO:2001190 | positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell | 16116184 |
 Fusion gene breakpoints across NLK (5'-gene) Fusion gene breakpoints across NLK (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
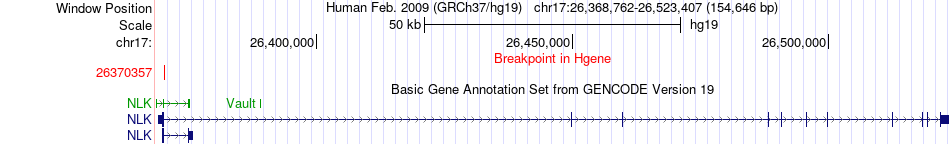 |
 Fusion gene breakpoints across LGALS9 (3'-gene) Fusion gene breakpoints across LGALS9 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
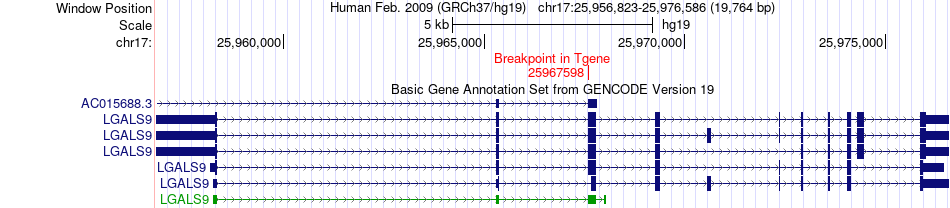 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-AR-A1AJ-01A | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
Top |
Fusion Gene ORF analysis for NLK-LGALS9 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000583517 | ENST00000302228 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 3UTR-3CDS | ENST00000583517 | ENST00000310394 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 3UTR-3CDS | ENST00000583517 | ENST00000313648 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 3UTR-3CDS | ENST00000583517 | ENST00000395473 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 3UTR-3UTR | ENST00000583517 | ENST00000448970 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 3UTR-intron | ENST00000583517 | ENST00000413914 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 5CDS-3UTR | ENST00000407008 | ENST00000448970 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 5CDS-3UTR | ENST00000582037 | ENST00000448970 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 5CDS-intron | ENST00000407008 | ENST00000413914 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| 5CDS-intron | ENST00000582037 | ENST00000413914 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000407008 | ENST00000302228 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000407008 | ENST00000310394 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000407008 | ENST00000313648 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000407008 | ENST00000395473 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000582037 | ENST00000302228 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000582037 | ENST00000310394 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000582037 | ENST00000313648 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
| In-frame | ENST00000582037 | ENST00000395473 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000407008 | NLK | chr17 | 26370357 | + | ENST00000310394 | LGALS9 | chr17 | 25967598 | + | 2552 | 1176 | 718 | 1980 | 420 |
| ENST00000407008 | NLK | chr17 | 26370357 | + | ENST00000395473 | LGALS9 | chr17 | 25967598 | + | 2691 | 1176 | 718 | 2112 | 464 |
| ENST00000407008 | NLK | chr17 | 26370357 | + | ENST00000302228 | LGALS9 | chr17 | 25967598 | + | 2595 | 1176 | 718 | 2016 | 432 |
| ENST00000407008 | NLK | chr17 | 26370357 | + | ENST00000313648 | LGALS9 | chr17 | 25967598 | + | 2305 | 1176 | 718 | 1785 | 355 |
| ENST00000582037 | NLK | chr17 | 26370357 | + | ENST00000310394 | LGALS9 | chr17 | 25967598 | + | 1869 | 493 | 35 | 1297 | 420 |
| ENST00000582037 | NLK | chr17 | 26370357 | + | ENST00000395473 | LGALS9 | chr17 | 25967598 | + | 2008 | 493 | 35 | 1429 | 464 |
| ENST00000582037 | NLK | chr17 | 26370357 | + | ENST00000302228 | LGALS9 | chr17 | 25967598 | + | 1912 | 493 | 35 | 1333 | 432 |
| ENST00000582037 | NLK | chr17 | 26370357 | + | ENST00000313648 | LGALS9 | chr17 | 25967598 | + | 1622 | 493 | 35 | 1102 | 355 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000407008 | ENST00000310394 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.11726748 | 0.8827325 |
| ENST00000407008 | ENST00000395473 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.11993264 | 0.8800674 |
| ENST00000407008 | ENST00000302228 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.112100884 | 0.8878991 |
| ENST00000407008 | ENST00000313648 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.22970979 | 0.7702902 |
| ENST00000582037 | ENST00000310394 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.16752253 | 0.83247745 |
| ENST00000582037 | ENST00000395473 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.17692369 | 0.8230763 |
| ENST00000582037 | ENST00000302228 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.16714405 | 0.8328559 |
| ENST00000582037 | ENST00000313648 | NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967598 | + | 0.1941904 | 0.8058096 |
Top |
Fusion Genomic Features for NLK-LGALS9 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967597 | + | 0.009140295 | 0.9908597 |
| NLK | chr17 | 26370357 | + | LGALS9 | chr17 | 25967597 | + | 0.009140295 | 0.9908597 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
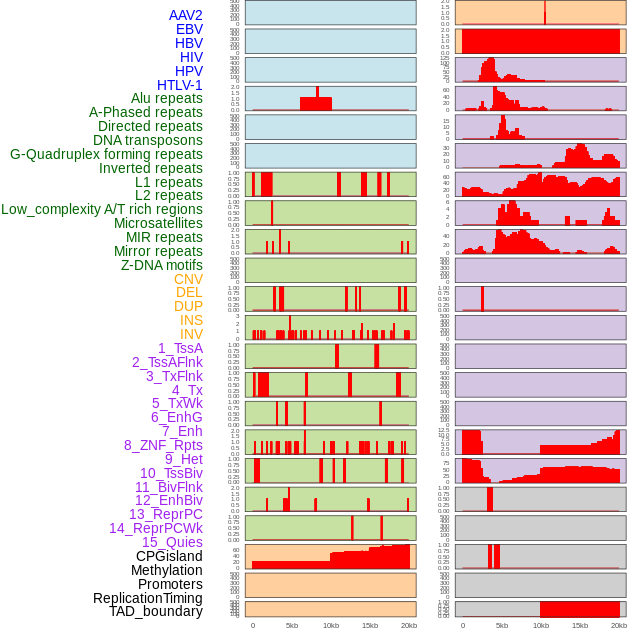 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
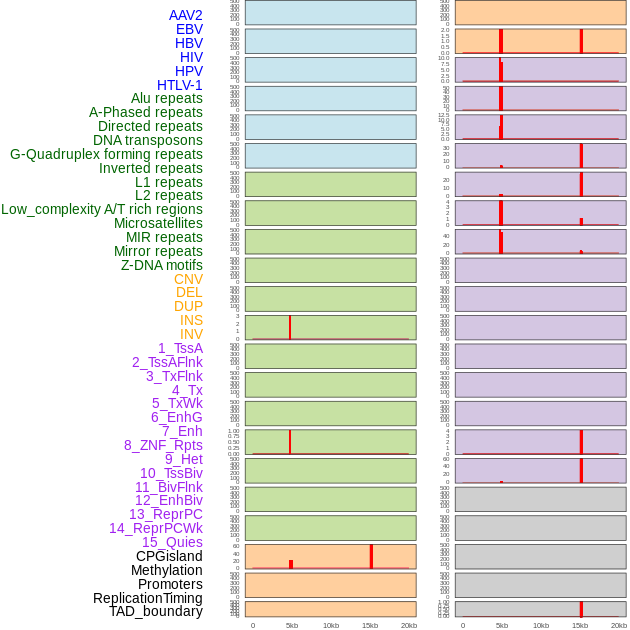 |
Top |
Fusion Protein Features for NLK-LGALS9 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr17:26370357/chr17:25967598) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| NLK | LGALS9 |
| FUNCTION: Serine/threonine-protein kinase that regulates a number of transcription factors with key roles in cell fate determination. Positive effector of the non-canonical Wnt signaling pathway, acting downstream of WNT5A, MAP3K7/TAK1 and HIPK2. Negative regulator of the canonical Wnt/beta-catenin signaling pathway. Binds to and phosphorylates TCF7L2/TCF4 and LEF1, promoting the dissociation of the TCF7L2/LEF1/beta-catenin complex from DNA, as well as the ubiquitination and subsequent proteolysis of LEF1. Together these effects inhibit the transcriptional activation of canonical Wnt/beta-catenin target genes. Negative regulator of the Notch signaling pathway. Binds to and phosphorylates NOTCH1, thereby preventing the formation of a transcriptionally active ternary complex of NOTCH1, RBPJ/RBPSUH and MAML1. Negative regulator of the MYB family of transcription factors. Phosphorylation of MYB leads to its subsequent proteolysis while phosphorylation of MYBL1 and MYBL2 inhibits their interaction with the coactivator CREBBP. Other transcription factors may also be inhibited by direct phosphorylation of CREBBP itself. Acts downstream of IL6 and MAP3K7/TAK1 to phosphorylate STAT3, which is in turn required for activation of NLK by MAP3K7/TAK1. Upon IL1B stimulus, cooperates with ATF5 to activate the transactivation activity of C/EBP subfamily members. Phosphorylates ATF5 but also stabilizes ATF5 protein levels in a kinase-independent manner (PubMed:25512613). {ECO:0000250|UniProtKB:O54949, ECO:0000269|PubMed:12482967, ECO:0000269|PubMed:14960582, ECO:0000269|PubMed:15004007, ECO:0000269|PubMed:15764709, ECO:0000269|PubMed:20061393, ECO:0000269|PubMed:20118921, ECO:0000269|PubMed:20874444, ECO:0000269|PubMed:21454679, ECO:0000269|PubMed:25512613}. | FUNCTION: Binds galactosides. {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 106_111 | 152 | 528.0 | Compositional bias | Note=Poly-Ala |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 115_119 | 152 | 528.0 | Compositional bias | Note=Poly-Ala |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 22_25 | 152 | 528.0 | Compositional bias | Note=Poly-Ala |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 27_34 | 152 | 528.0 | Compositional bias | Note=Poly-His |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 42_48 | 152 | 528.0 | Compositional bias | Note=Poly-His |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 71_83 | 152 | 528.0 | Compositional bias | Note=Poly-Ala |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 144_152 | 152 | 528.0 | Nucleotide binding | ATP |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000302228 | 1 | 10 | 227_355 | 43 | 324.0 | Domain | Galectin 2 | |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000395473 | 1 | 11 | 227_355 | 43 | 356.0 | Domain | Galectin 2 | |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000302228 | 1 | 10 | 287_293 | 43 | 324.0 | Region | Beta-galactoside binding 2 | |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000302228 | 1 | 10 | 82_88 | 43 | 324.0 | Region | Note=Beta-galactoside binding 1 | |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000395473 | 1 | 11 | 287_293 | 43 | 356.0 | Region | Beta-galactoside binding 2 | |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000395473 | 1 | 11 | 82_88 | 43 | 356.0 | Region | Note=Beta-galactoside binding 1 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 138_427 | 152 | 528.0 | Domain | Protein kinase |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 298_300 | 152 | 528.0 | Motif | Note=TQE |
| Hgene | NLK | chr17:26370357 | chr17:25967598 | ENST00000407008 | + | 1 | 11 | 428_527 | 152 | 528.0 | Region | Required for homodimerization and kinase activation and localization to the nucleus |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000302228 | 1 | 10 | 17_148 | 43 | 324.0 | Domain | Galectin 1 | |
| Tgene | LGALS9 | chr17:26370357 | chr17:25967598 | ENST00000395473 | 1 | 11 | 17_148 | 43 | 356.0 | Domain | Galectin 1 |
Top |
Fusion Gene Sequence for NLK-LGALS9 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >59350_59350_1_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000302228_length(transcript)=2595nt_BP=1176nt CCCCGGCTCTCTTTCTCCTCTCTGTCCTGCTGTCGCCTCCTCTCCTCACTCCAGCCTCCTTCCCTGGCTTATTCTGCCCAGGAAAGTTTG GAACCCCCGCCCCCCCTCCCCAATCAGGTCAACCTGACTAGGAGTAGGGGGAGATGTAGGAAGGCCTGAGAGTTCGGGGGGGGGGAAGCT GGAAGCTGGTTTCCCAGAAAATTTCCCCCCATCTTCTCCATTTAGGGTGGTGGGAATTTTTTTTAATCCTGTTTTTGAGGAGGTGGTAGG TGTTTAGAAAAGGATGTGGGAATGGAAGGGGTAGATGAGTCAACCCTCTTTCAAGATTCCAAGGGTGGTGTGTTACCTGATCCTTTTCGT GTGTGGAGTTCAGGGGGTCGGGAGGTTTGGCCTTTATTCCCACATTCAAAGTTGGAACCTCCCCCAAATTAAGAATGGAGTTTTAGTACC CACTTGTGGCAGGAATCCTGGGGGGAAGGGGTCCTTTTTTCCTTTTTCTTTTCTTTTTTCCCTTTTTTTTCCTTTTGGCAACCCACACCC TTCCACACACGCTCACCCCAAAATTAAACACCAAGATCCTCTAACTTGTTTGGATTGACTGATGAAGACATAAAGCTCTATGTTTTTTGA GGTGGAGTGAGTGGTTTTTCTTCATTTTTAAATGGCCAAATGACAGCTTGACCCAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAAT GTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCGGTACATCTGCAGCAGCAGCAGGTCACCACCACCACCA TCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACCCTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGT ACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGGCTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACA GCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAGCCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGT TAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTGAGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGT TGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCTTCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGT GGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACACACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTG CTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCGTGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGA CACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGCCTCCCGGCGTGTGGCCTGCCAACCCGGCTCCCATTACCCA GACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTACTCCCGCCATCCCACCTATGATGTACCCCCACCCCGCCTATCC GATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGTCCATCCTCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGTT CCACATCAACCTGTGCTCTGGGAACCACATCGCCTTCCACCTGAACCCCCGTTTTGATGAGAATGCTGTGGTCCGCAACACCCAGATCGA CAACTCCTGGGGGTCTGAGGAGCGAAGTCTGCCCCGAAAAATGCCCTTCGTCCGTGGCCAGAGCTTCTCAGTGTGGATCTTGTGTGAAGC TCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCTGTTTGAATACTACCATCGCCTGAGGAACCTGCCCACCATCAACAGACTGGAAGT GGGGGGCGACATCCAGCTGACCCATGTGCAGACATAGGCGGCTTCCTGGCCCTGGGGCCGGGGGCTGGGGTGTGGGGCAGTCTGGGTCCT CTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAACCCTGCCTGGGATCTGGGCTTTAATGCAGAGGCCATGTCCTTGTCTGGTCCTGC TTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGCTGACGGGGATTGCCTTCCTCAGCCGCAGCAGCACCTGGGGCTCCAGCTGCTGGA ATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAGAGGGGAGGAGTGGGCAGTGAAGATGAAGCCCCATGCTCAGTCCCCTCCCATCCC CCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTGTCTGCTCCTGGTGGGAGGTGGCCTCCTCAGCCCCTCCTCTCTGACCTTTAACCT CACTCTCACCTTGCACCGTGCACCAACCCTTCACCCCTCCTGGAAAGCAGGCCTGATGGCTTCCCACTGGCCTCCACCACCTGACCAGAG >59350_59350_1_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000302228_length(amino acids)=432AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQPPGVWPANPAPIT QTVIHTVQSAPGQMFSTPAIPPMMYPHPAYPMPFITTILGGLYPSKSILLSGTVLPSAQRFHINLCSGNHIAFHLNPRFDENAVVRNTQI -------------------------------------------------------------- >59350_59350_2_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000310394_length(transcript)=2552nt_BP=1176nt CCCCGGCTCTCTTTCTCCTCTCTGTCCTGCTGTCGCCTCCTCTCCTCACTCCAGCCTCCTTCCCTGGCTTATTCTGCCCAGGAAAGTTTG GAACCCCCGCCCCCCCTCCCCAATCAGGTCAACCTGACTAGGAGTAGGGGGAGATGTAGGAAGGCCTGAGAGTTCGGGGGGGGGGAAGCT GGAAGCTGGTTTCCCAGAAAATTTCCCCCCATCTTCTCCATTTAGGGTGGTGGGAATTTTTTTTAATCCTGTTTTTGAGGAGGTGGTAGG TGTTTAGAAAAGGATGTGGGAATGGAAGGGGTAGATGAGTCAACCCTCTTTCAAGATTCCAAGGGTGGTGTGTTACCTGATCCTTTTCGT GTGTGGAGTTCAGGGGGTCGGGAGGTTTGGCCTTTATTCCCACATTCAAAGTTGGAACCTCCCCCAAATTAAGAATGGAGTTTTAGTACC CACTTGTGGCAGGAATCCTGGGGGGAAGGGGTCCTTTTTTCCTTTTTCTTTTCTTTTTTCCCTTTTTTTTCCTTTTGGCAACCCACACCC TTCCACACACGCTCACCCCAAAATTAAACACCAAGATCCTCTAACTTGTTTGGATTGACTGATGAAGACATAAAGCTCTATGTTTTTTGA GGTGGAGTGAGTGGTTTTTCTTCATTTTTAAATGGCCAAATGACAGCTTGACCCAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAAT GTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCGGTACATCTGCAGCAGCAGCAGGTCACCACCACCACCA TCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACCCTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGT ACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGGCTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACA GCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAGCCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGT TAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTGAGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGT TGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCTTCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGT GGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACACACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTG CTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCGTGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGA CACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGACCCAGACAGTCATCCACACAGTGCAGAGCGCCCCTGGACA GATGTTCTCTACTCCCGCCATCCCACCTATGATGTACCCCCACCCCGCCTATCCGATGCCTTTCATCACCACCATTCTGGGAGGGCTGTA CCCATCCAAGTCCATCCTCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGTTCCACATCAACCTGTGCTCTGGGAACCACATCGCCTT CCACCTGAACCCCCGTTTTGATGAGAATGCTGTGGTCCGCAACACCCAGATCGACAACTCCTGGGGGTCTGAGGAGCGAAGTCTGCCCCG AAAAATGCCCTTCGTCCGTGGCCAGAGCTTCTCAGTGTGGATCTTGTGTGAAGCTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCT GTTTGAATACTACCATCGCCTGAGGAACCTGCCCACCATCAACAGACTGGAAGTGGGGGGCGACATCCAGCTGACCCATGTGCAGACATA GGCGGCTTCCTGGCCCTGGGGCCGGGGGCTGGGGTGTGGGGCAGTCTGGGTCCTCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAA CCCTGCCTGGGATCTGGGCTTTAATGCAGAGGCCATGTCCTTGTCTGGTCCTGCTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGC TGACGGGGATTGCCTTCCTCAGCCGCAGCAGCACCTGGGGCTCCAGCTGCTGGAATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAG AGGGGAGGAGTGGGCAGTGAAGATGAAGCCCCATGCTCAGTCCCCTCCCATCCCCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTG TCTGCTCCTGGTGGGAGGTGGCCTCCTCAGCCCCTCCTCTCTGACCTTTAACCTCACTCTCACCTTGCACCGTGCACCAACCCTTCACCC CTCCTGGAAAGCAGGCCTGATGGCTTCCCACTGGCCTCCACCACCTGACCAGAGTGTTCTCTTCAGAGGACTGGCTCCTTTCCCAGTGTC >59350_59350_2_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000310394_length(amino acids)=420AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQTQTVIHTVQSAPG QMFSTPAIPPMMYPHPAYPMPFITTILGGLYPSKSILLSGTVLPSAQRFHINLCSGNHIAFHLNPRFDENAVVRNTQIDNSWGSEERSLP -------------------------------------------------------------- >59350_59350_3_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000313648_length(transcript)=2305nt_BP=1176nt CCCCGGCTCTCTTTCTCCTCTCTGTCCTGCTGTCGCCTCCTCTCCTCACTCCAGCCTCCTTCCCTGGCTTATTCTGCCCAGGAAAGTTTG GAACCCCCGCCCCCCCTCCCCAATCAGGTCAACCTGACTAGGAGTAGGGGGAGATGTAGGAAGGCCTGAGAGTTCGGGGGGGGGGAAGCT GGAAGCTGGTTTCCCAGAAAATTTCCCCCCATCTTCTCCATTTAGGGTGGTGGGAATTTTTTTTAATCCTGTTTTTGAGGAGGTGGTAGG TGTTTAGAAAAGGATGTGGGAATGGAAGGGGTAGATGAGTCAACCCTCTTTCAAGATTCCAAGGGTGGTGTGTTACCTGATCCTTTTCGT GTGTGGAGTTCAGGGGGTCGGGAGGTTTGGCCTTTATTCCCACATTCAAAGTTGGAACCTCCCCCAAATTAAGAATGGAGTTTTAGTACC CACTTGTGGCAGGAATCCTGGGGGGAAGGGGTCCTTTTTTCCTTTTTCTTTTCTTTTTTCCCTTTTTTTTCCTTTTGGCAACCCACACCC TTCCACACACGCTCACCCCAAAATTAAACACCAAGATCCTCTAACTTGTTTGGATTGACTGATGAAGACATAAAGCTCTATGTTTTTTGA GGTGGAGTGAGTGGTTTTTCTTCATTTTTAAATGGCCAAATGACAGCTTGACCCAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAAT GTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCGGTACATCTGCAGCAGCAGCAGGTCACCACCACCACCA TCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACCCTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGT ACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGGCTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACA GCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAGCCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGT TAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTGAGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGT TGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCTTCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGT GGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACACACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTG CTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCGTGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGA CACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGCCTCCCGGCGTGTGGCCTGCCAACCCGGCTCCCATTACCCA GACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTACTCCCGCCATCCCACCTATGATGTACCCCCACCCCGCCTATCC GATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGTCCATCCTCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGTG TGGATCTTGTGTGAAGCTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCTGTTTGAATACTACCATCGCCTGAGGAACCTGCCCACC ATCAACAGACTGGAAGTGGGGGGCGACATCCAGCTGACCCATGTGCAGACATAGGCGGCTTCCTGGCCCTGGGGCCGGGGGCTGGGGTGT GGGGCAGTCTGGGTCCTCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAACCCTGCCTGGGATCTGGGCTTTAATGCAGAGGCCATG TCCTTGTCTGGTCCTGCTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGCTGACGGGGATTGCCTTCCTCAGCCGCAGCAGCACCTG GGGCTCCAGCTGCTGGAATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAGAGGGGAGGAGTGGGCAGTGAAGATGAAGCCCCATGCT CAGTCCCCTCCCATCCCCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTGTCTGCTCCTGGTGGGAGGTGGCCTCCTCAGCCCCTCC >59350_59350_3_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000313648_length(amino acids)=355AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQPPGVWPANPAPIT -------------------------------------------------------------- >59350_59350_4_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000395473_length(transcript)=2691nt_BP=1176nt CCCCGGCTCTCTTTCTCCTCTCTGTCCTGCTGTCGCCTCCTCTCCTCACTCCAGCCTCCTTCCCTGGCTTATTCTGCCCAGGAAAGTTTG GAACCCCCGCCCCCCCTCCCCAATCAGGTCAACCTGACTAGGAGTAGGGGGAGATGTAGGAAGGCCTGAGAGTTCGGGGGGGGGGAAGCT GGAAGCTGGTTTCCCAGAAAATTTCCCCCCATCTTCTCCATTTAGGGTGGTGGGAATTTTTTTTAATCCTGTTTTTGAGGAGGTGGTAGG TGTTTAGAAAAGGATGTGGGAATGGAAGGGGTAGATGAGTCAACCCTCTTTCAAGATTCCAAGGGTGGTGTGTTACCTGATCCTTTTCGT GTGTGGAGTTCAGGGGGTCGGGAGGTTTGGCCTTTATTCCCACATTCAAAGTTGGAACCTCCCCCAAATTAAGAATGGAGTTTTAGTACC CACTTGTGGCAGGAATCCTGGGGGGAAGGGGTCCTTTTTTCCTTTTTCTTTTCTTTTTTCCCTTTTTTTTCCTTTTGGCAACCCACACCC TTCCACACACGCTCACCCCAAAATTAAACACCAAGATCCTCTAACTTGTTTGGATTGACTGATGAAGACATAAAGCTCTATGTTTTTTGA GGTGGAGTGAGTGGTTTTTCTTCATTTTTAAATGGCCAAATGACAGCTTGACCCAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAAT GTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCGGTACATCTGCAGCAGCAGCAGGTCACCACCACCACCA TCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACCCTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGT ACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGGCTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACA GCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAGCCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGT TAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTGAGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGT TGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCTTCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGT GGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACACACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTG CTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCGTGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGA CACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGAACCCCCGCACAGTCCCTGTTCAGCCTGCCTTCTCCACGGT GCCGTTCTCCCAGCCTGTCTGTTTCCCACCCAGGCCCAGGGGGCGCAGACAAAAACCTCCCGGCGTGTGGCCTGCCAACCCGGCTCCCAT TACCCAGACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTACTCCCGCCATCCCACCTATGATGTACCCCCACCCCGC CTATCCGATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGTCCATCCTCCTGTCAGGCACTGTCCTGCCCAGTGCTCA GAGGTTCCACATCAACCTGTGCTCTGGGAACCACATCGCCTTCCACCTGAACCCCCGTTTTGATGAGAATGCTGTGGTCCGCAACACCCA GATCGACAACTCCTGGGGGTCTGAGGAGCGAAGTCTGCCCCGAAAAATGCCCTTCGTCCGTGGCCAGAGCTTCTCAGTGTGGATCTTGTG TGAAGCTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCTGTTTGAATACTACCATCGCCTGAGGAACCTGCCCACCATCAACAGACT GGAAGTGGGGGGCGACATCCAGCTGACCCATGTGCAGACATAGGCGGCTTCCTGGCCCTGGGGCCGGGGGCTGGGGTGTGGGGCAGTCTG GGTCCTCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAACCCTGCCTGGGATCTGGGCTTTAATGCAGAGGCCATGTCCTTGTCTGG TCCTGCTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGCTGACGGGGATTGCCTTCCTCAGCCGCAGCAGCACCTGGGGCTCCAGCT GCTGGAATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAGAGGGGAGGAGTGGGCAGTGAAGATGAAGCCCCATGCTCAGTCCCCTCC CATCCCCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTGTCTGCTCCTGGTGGGAGGTGGCCTCCTCAGCCCCTCCTCTCTGACCTT TAACCTCACTCTCACCTTGCACCGTGCACCAACCCTTCACCCCTCCTGGAAAGCAGGCCTGATGGCTTCCCACTGGCCTCCACCACCTGA >59350_59350_4_NLK-LGALS9_NLK_chr17_26370357_ENST00000407008_LGALS9_chr17_25967598_ENST00000395473_length(amino acids)=464AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQNPRTVPVQPAFST VPFSQPVCFPPRPRGRRQKPPGVWPANPAPITQTVIHTVQSAPGQMFSTPAIPPMMYPHPAYPMPFITTILGGLYPSKSILLSGTVLPSA QRFHINLCSGNHIAFHLNPRFDENAVVRNTQIDNSWGSEERSLPRKMPFVRGQSFSVWILCEAHCLKVAVDGQHLFEYYHRLRNLPTINR -------------------------------------------------------------- >59350_59350_5_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000302228_length(transcript)=1912nt_BP=493nt CAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAATGTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCG GTACATCTGCAGCAGCAGCAGGTCACCACCACCACCATCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACC CTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGTACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGG CTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACAGCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAG CCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGTTAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTG AGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGTTGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCT TCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGTGGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACAC ACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTGCTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCG TGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGACACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGCCTC CCGGCGTGTGGCCTGCCAACCCGGCTCCCATTACCCAGACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTACTCCCG CCATCCCACCTATGATGTACCCCCACCCCGCCTATCCGATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGTCCATCC TCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGTTCCACATCAACCTGTGCTCTGGGAACCACATCGCCTTCCACCTGAACCCCCGTT TTGATGAGAATGCTGTGGTCCGCAACACCCAGATCGACAACTCCTGGGGGTCTGAGGAGCGAAGTCTGCCCCGAAAAATGCCCTTCGTCC GTGGCCAGAGCTTCTCAGTGTGGATCTTGTGTGAAGCTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCTGTTTGAATACTACCATC GCCTGAGGAACCTGCCCACCATCAACAGACTGGAAGTGGGGGGCGACATCCAGCTGACCCATGTGCAGACATAGGCGGCTTCCTGGCCCT GGGGCCGGGGGCTGGGGTGTGGGGCAGTCTGGGTCCTCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAACCCTGCCTGGGATCTGG GCTTTAATGCAGAGGCCATGTCCTTGTCTGGTCCTGCTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGCTGACGGGGATTGCCTTC CTCAGCCGCAGCAGCACCTGGGGCTCCAGCTGCTGGAATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAGAGGGGAGGAGTGGGCAG TGAAGATGAAGCCCCATGCTCAGTCCCCTCCCATCCCCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTGTCTGCTCCTGGTGGGAG GTGGCCTCCTCAGCCCCTCCTCTCTGACCTTTAACCTCACTCTCACCTTGCACCGTGCACCAACCCTTCACCCCTCCTGGAAAGCAGGCC TGATGGCTTCCCACTGGCCTCCACCACCTGACCAGAGTGTTCTCTTCAGAGGACTGGCTCCTTTCCCAGTGTCCTTAAAATAAAGAAATG >59350_59350_5_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000302228_length(amino acids)=432AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQPPGVWPANPAPIT QTVIHTVQSAPGQMFSTPAIPPMMYPHPAYPMPFITTILGGLYPSKSILLSGTVLPSAQRFHINLCSGNHIAFHLNPRFDENAVVRNTQI -------------------------------------------------------------- >59350_59350_6_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000310394_length(transcript)=1869nt_BP=493nt CAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAATGTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCG GTACATCTGCAGCAGCAGCAGGTCACCACCACCACCATCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACC CTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGTACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGG CTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACAGCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAG CCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGTTAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTG AGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGTTGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCT TCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGTGGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACAC ACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTGCTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCG TGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGACACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGACCC AGACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTACTCCCGCCATCCCACCTATGATGTACCCCCACCCCGCCTATC CGATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGTCCATCCTCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGT TCCACATCAACCTGTGCTCTGGGAACCACATCGCCTTCCACCTGAACCCCCGTTTTGATGAGAATGCTGTGGTCCGCAACACCCAGATCG ACAACTCCTGGGGGTCTGAGGAGCGAAGTCTGCCCCGAAAAATGCCCTTCGTCCGTGGCCAGAGCTTCTCAGTGTGGATCTTGTGTGAAG CTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCTGTTTGAATACTACCATCGCCTGAGGAACCTGCCCACCATCAACAGACTGGAAG TGGGGGGCGACATCCAGCTGACCCATGTGCAGACATAGGCGGCTTCCTGGCCCTGGGGCCGGGGGCTGGGGTGTGGGGCAGTCTGGGTCC TCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAACCCTGCCTGGGATCTGGGCTTTAATGCAGAGGCCATGTCCTTGTCTGGTCCTG CTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGCTGACGGGGATTGCCTTCCTCAGCCGCAGCAGCACCTGGGGCTCCAGCTGCTGG AATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAGAGGGGAGGAGTGGGCAGTGAAGATGAAGCCCCATGCTCAGTCCCCTCCCATCC CCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTGTCTGCTCCTGGTGGGAGGTGGCCTCCTCAGCCCCTCCTCTCTGACCTTTAACC TCACTCTCACCTTGCACCGTGCACCAACCCTTCACCCCTCCTGGAAAGCAGGCCTGATGGCTTCCCACTGGCCTCCACCACCTGACCAGA >59350_59350_6_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000310394_length(amino acids)=420AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQTQTVIHTVQSAPG QMFSTPAIPPMMYPHPAYPMPFITTILGGLYPSKSILLSGTVLPSAQRFHINLCSGNHIAFHLNPRFDENAVVRNTQIDNSWGSEERSLP -------------------------------------------------------------- >59350_59350_7_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000313648_length(transcript)=1622nt_BP=493nt CAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAATGTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCG GTACATCTGCAGCAGCAGCAGGTCACCACCACCACCATCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACC CTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGTACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGG CTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACAGCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAG CCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGTTAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTG AGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGTTGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCT TCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGTGGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACAC ACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTGCTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCG TGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGACACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGCCTC CCGGCGTGTGGCCTGCCAACCCGGCTCCCATTACCCAGACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTACTCCCG CCATCCCACCTATGATGTACCCCCACCCCGCCTATCCGATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGTCCATCC TCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGTGTGGATCTTGTGTGAAGCTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCT GTTTGAATACTACCATCGCCTGAGGAACCTGCCCACCATCAACAGACTGGAAGTGGGGGGCGACATCCAGCTGACCCATGTGCAGACATA GGCGGCTTCCTGGCCCTGGGGCCGGGGGCTGGGGTGTGGGGCAGTCTGGGTCCTCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAA CCCTGCCTGGGATCTGGGCTTTAATGCAGAGGCCATGTCCTTGTCTGGTCCTGCTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGC TGACGGGGATTGCCTTCCTCAGCCGCAGCAGCACCTGGGGCTCCAGCTGCTGGAATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAG AGGGGAGGAGTGGGCAGTGAAGATGAAGCCCCATGCTCAGTCCCCTCCCATCCCCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTG TCTGCTCCTGGTGGGAGGTGGCCTCCTCAGCCCCTCCTCTCTGACCTTTAACCTCACTCTCACCTTGCACCGTGCACCAACCCTTCACCC >59350_59350_7_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000313648_length(amino acids)=355AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQPPGVWPANPAPIT -------------------------------------------------------------- >59350_59350_8_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000395473_length(transcript)=2008nt_BP=493nt CAGTTTGCTTTCCAATCAAAGGGCATTTATTTTGAATGTCTCTTTGTGGCGCAAGAGCCAACGCAAAAATGATGGCGGCTTACAATGGCG GTACATCTGCAGCAGCAGCAGGTCACCACCACCACCATCACCACCACCTTCCACACCTCCCTCCTCCTCACCTGCACCACCACCACCACC CTCAACACCATCTTCATCCGGGGTCGGCTGCCGCTGTACACCCTGTACAGCAGCACACCTCTTCGGCAGCTGCGGCAGCCGCAGCAGCGG CTGCAGCTGCAGCCATGTTAAACCCTGGGCAACAACAGCCATATTTCCCATCACCGGCACCGGGGCAGGCTCCTGGACCAGCTGCAGCAG CCCCAGCTCAGGTACAGGCTGCCGCAGCTGCTACAGTTAAGGCGCACCATCATCAGCACTCGCATCATCCACAGCAGCAGCTGGATATTG AGCCGGATAGACCTATTGGATATGGAGCCTTTGGTGTTGTCTGGTTTGCTGTGAACTTTCAGACTGGCTTCAGTGGAAATGACATTGCCT TCCACTTCAACCCTCGGTTTGAAGATGGAGGGTACGTGGTGTGCAACACGAGGCAGAACGGAAGCTGGGGGCCCGAGGAGAGGAAGACAC ACATGCCTTTCCAGAAGGGGATGCCCTTTGACCTCTGCTTCCTGGTGCAGAGCTCAGATTTCAAGGTGATGGTGAACGGGATCCTCTTCG TGCAGTACTTCCACCGCGTGCCCTTCCACCGTGTGGACACCATCTCCGTCAATGGCTCTGTGCAGCTGTCCTACATCAGCTTCCAGAACC CCCGCACAGTCCCTGTTCAGCCTGCCTTCTCCACGGTGCCGTTCTCCCAGCCTGTCTGTTTCCCACCCAGGCCCAGGGGGCGCAGACAAA AACCTCCCGGCGTGTGGCCTGCCAACCCGGCTCCCATTACCCAGACAGTCATCCACACAGTGCAGAGCGCCCCTGGACAGATGTTCTCTA CTCCCGCCATCCCACCTATGATGTACCCCCACCCCGCCTATCCGATGCCTTTCATCACCACCATTCTGGGAGGGCTGTACCCATCCAAGT CCATCCTCCTGTCAGGCACTGTCCTGCCCAGTGCTCAGAGGTTCCACATCAACCTGTGCTCTGGGAACCACATCGCCTTCCACCTGAACC CCCGTTTTGATGAGAATGCTGTGGTCCGCAACACCCAGATCGACAACTCCTGGGGGTCTGAGGAGCGAAGTCTGCCCCGAAAAATGCCCT TCGTCCGTGGCCAGAGCTTCTCAGTGTGGATCTTGTGTGAAGCTCACTGCCTCAAGGTGGCCGTGGATGGTCAGCACCTGTTTGAATACT ACCATCGCCTGAGGAACCTGCCCACCATCAACAGACTGGAAGTGGGGGGCGACATCCAGCTGACCCATGTGCAGACATAGGCGGCTTCCT GGCCCTGGGGCCGGGGGCTGGGGTGTGGGGCAGTCTGGGTCCTCTCATCATCCCCACTTCCCAGGCCCAGCCTTTCCAACCCTGCCTGGG ATCTGGGCTTTAATGCAGAGGCCATGTCCTTGTCTGGTCCTGCTTCTGGCTACAGCCACCCTGGAACGGAGAAGGCAGCTGACGGGGATT GCCTTCCTCAGCCGCAGCAGCACCTGGGGCTCCAGCTGCTGGAATCCTACCATCCCAGGAGGCAGGCACAGCCAGGGAGAGGGGAGGAGT GGGCAGTGAAGATGAAGCCCCATGCTCAGTCCCCTCCCATCCCCCACGCAGCTCCACCCCAGTCCCAAGCCACCAGCTGTCTGCTCCTGG TGGGAGGTGGCCTCCTCAGCCCCTCCTCTCTGACCTTTAACCTCACTCTCACCTTGCACCGTGCACCAACCCTTCACCCCTCCTGGAAAG CAGGCCTGATGGCTTCCCACTGGCCTCCACCACCTGACCAGAGTGTTCTCTTCAGAGGACTGGCTCCTTTCCCAGTGTCCTTAAAATAAA >59350_59350_8_NLK-LGALS9_NLK_chr17_26370357_ENST00000582037_LGALS9_chr17_25967598_ENST00000395473_length(amino acids)=464AA_BP=153 MSLCGARANAKMMAAYNGGTSAAAAGHHHHHHHHLPHLPPPHLHHHHHPQHHLHPGSAAAVHPVQQHTSSAAAAAAAAAAAAAMLNPGQQ QPYFPSPAPGQAPGPAAAAPAQVQAAAAATVKAHHHQHSHHPQQQLDIEPDRPIGYGAFGVVWFAVNFQTGFSGNDIAFHFNPRFEDGGY VVCNTRQNGSWGPEERKTHMPFQKGMPFDLCFLVQSSDFKVMVNGILFVQYFHRVPFHRVDTISVNGSVQLSYISFQNPRTVPVQPAFST VPFSQPVCFPPRPRGRRQKPPGVWPANPAPITQTVIHTVQSAPGQMFSTPAIPPMMYPHPAYPMPFITTILGGLYPSKSILLSGTVLPSA QRFHINLCSGNHIAFHLNPRFDENAVVRNTQIDNSWGSEERSLPRKMPFVRGQSFSVWILCEAHCLKVAVDGQHLFEYYHRLRNLPTINR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NLK-LGALS9 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NLK-LGALS9 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NLK-LGALS9 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
