
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:NUP188-POLR2A (FusionGDB2 ID:61009) |
Fusion Gene Summary for NUP188-POLR2A |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: NUP188-POLR2A | Fusion gene ID: 61009 | Hgene | Tgene | Gene symbol | NUP188 | POLR2A | Gene ID | 23511 | 5430 |
| Gene name | nucleoporin 188 | RNA polymerase II subunit A | |
| Synonyms | KIAA0169|SANDSTEF|hNup188 | NEDHIB|POLR2|POLRA|RPB1|RPBh1|RPO2|RPOL2|RpIILS|hRPB220|hsRPB1 | |
| Cytomap | 9q34.11 | 17p13.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | nucleoporin NUP188 homolognucleoporin 188kDa | DNA-directed RNA polymerase II subunit RPB1DNA-directed RNA polymerase II largest subunit, RNA polymerase II 220 kd subunitDNA-directed RNA polymerase II subunit ADNA-directed RNA polymerase III largest subunitRNA polymerase II subunit B1RNA-directed | |
| Modification date | 20200314 | 20200327 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000372577, ENST00000550219, | ENST00000572844, ENST00000322644, | |
| Fusion gene scores | * DoF score | 11 X 12 X 8=1056 | 7 X 7 X 2=98 |
| # samples | 14 | 7 | |
| ** MAII score | log2(14/1056*10)=-2.91511110241349 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/98*10)=-0.485426827170242 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: NUP188 [Title/Abstract] AND POLR2A [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | NUP188(131768868)-POLR2A(7407022), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | POLR2A | GO:0006366 | transcription by RNA polymerase II | 9852112 |
| Tgene | POLR2A | GO:0033120 | positive regulation of RNA splicing | 21536736 |
 Fusion gene breakpoints across NUP188 (5'-gene) Fusion gene breakpoints across NUP188 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
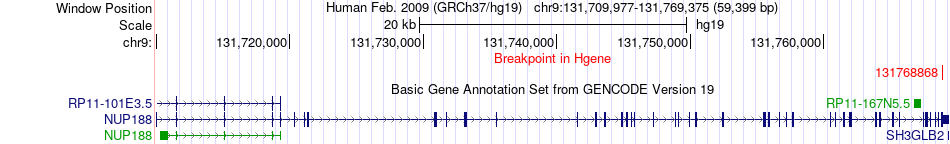 |
 Fusion gene breakpoints across POLR2A (3'-gene) Fusion gene breakpoints across POLR2A (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
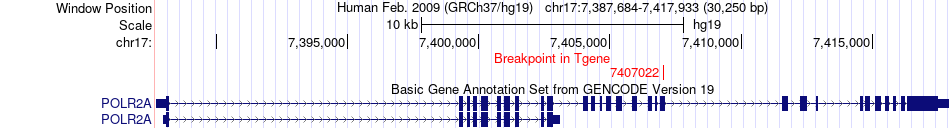 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChiTaRS5.0 | N/A | BE819347 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + |
| ChiTaRS5.0 | N/A | BE819348 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + |
Top |
Fusion Gene ORF analysis for NUP188-POLR2A |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000372577 | ENST00000572844 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + |
| In-frame | ENST00000372577 | ENST00000322644 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + |
| intron-3CDS | ENST00000550219 | ENST00000322644 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + |
| intron-intron | ENST00000550219 | ENST00000572844 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000372577 | NUP188 | chr9 | 131768868 | + | ENST00000322644 | POLR2A | chr17 | 7407022 | + | 8799 | 5601 | 21 | 5141 | 1706 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000372577 | ENST00000322644 | NUP188 | chr9 | 131768868 | + | POLR2A | chr17 | 7407022 | + | 0.000953107 | 0.99904686 |
Top |
Fusion Genomic Features for NUP188-POLR2A |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
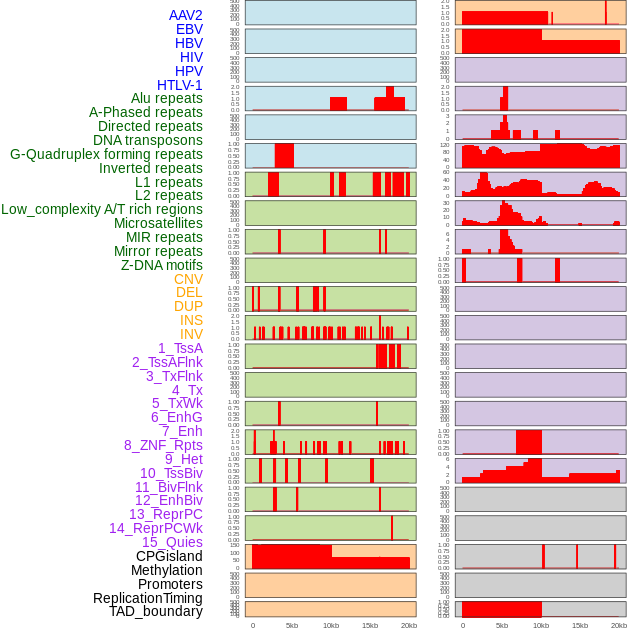 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for NUP188-POLR2A |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr9:131768868/chr17:7407022) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1593_1960 | 0 | 1971.0 | Region | Note=C-terminal domain (CTD)%3B 52 X 7 AA approximate tandem repeats of Y-[ST]-P-[STQ]-[ST]-P-[SRTEVKGN] | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 833_845 | 0 | 1971.0 | Region | Note=Bridging helix | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1593_1599 | 0 | 1971.0 | Repeat | Note=1 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1600_1606 | 0 | 1971.0 | Repeat | Note=2%3B approximate | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1608_1614 | 0 | 1971.0 | Repeat | Note=3 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1615_1621 | 0 | 1971.0 | Repeat | Note=4 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1622_1628 | 0 | 1971.0 | Repeat | Note=5 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1629_1635 | 0 | 1971.0 | Repeat | Note=6 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1636_1642 | 0 | 1971.0 | Repeat | Note=7 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1643_1649 | 0 | 1971.0 | Repeat | Note=8 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1650_1656 | 0 | 1971.0 | Repeat | Note=9 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1657_1663 | 0 | 1971.0 | Repeat | Note=10 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1664_1670 | 0 | 1971.0 | Repeat | Note=11 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1671_1677 | 0 | 1971.0 | Repeat | Note=12 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1678_1684 | 0 | 1971.0 | Repeat | Note=13 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1685_1691 | 0 | 1971.0 | Repeat | Note=14 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1692_1698 | 0 | 1971.0 | Repeat | Note=15 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1699_1705 | 0 | 1971.0 | Repeat | Note=16 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1706_1712 | 0 | 1971.0 | Repeat | Note=17 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1713_1719 | 0 | 1971.0 | Repeat | Note=18 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1720_1726 | 0 | 1971.0 | Repeat | Note=19 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1727_1733 | 0 | 1971.0 | Repeat | Note=20 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1734_1740 | 0 | 1971.0 | Repeat | Note=21 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1741_1747 | 0 | 1971.0 | Repeat | Note=22 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1748_1754 | 0 | 1971.0 | Repeat | Note=23 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1755_1761 | 0 | 1971.0 | Repeat | Note=24 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1762_1768 | 0 | 1971.0 | Repeat | Note=25 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1769_1775 | 0 | 1971.0 | Repeat | Note=26 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1776_1782 | 0 | 1971.0 | Repeat | Note=27 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1783_1789 | 0 | 1971.0 | Repeat | Note=28 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1790_1796 | 0 | 1971.0 | Repeat | Note=29 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1797_1803 | 0 | 1971.0 | Repeat | Note=30 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1804_1810 | 0 | 1971.0 | Repeat | Note=31 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1811_1817 | 0 | 1971.0 | Repeat | Note=32 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1818_1824 | 0 | 1971.0 | Repeat | Note=33 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1825_1831 | 0 | 1971.0 | Repeat | Note=34 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1832_1838 | 0 | 1971.0 | Repeat | Note=35 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1839_1845 | 0 | 1971.0 | Repeat | Note=36 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1846_1852 | 0 | 1971.0 | Repeat | Note=37 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1853_1859 | 0 | 1971.0 | Repeat | Note=38 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1860_1866 | 0 | 1971.0 | Repeat | Note=39 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1867_1873 | 0 | 1971.0 | Repeat | Note=40 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1874_1880 | 0 | 1971.0 | Repeat | Note=41 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1881_1887 | 0 | 1971.0 | Repeat | Note=42 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1888_1894 | 0 | 1971.0 | Repeat | Note=43 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1895_1901 | 0 | 1971.0 | Repeat | Note=44 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1902_1908 | 0 | 1971.0 | Repeat | Note=45 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1909_1915 | 0 | 1971.0 | Repeat | Note=46 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1916_1922 | 0 | 1971.0 | Repeat | Note=47 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1923_1929 | 0 | 1971.0 | Repeat | Note=48 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1930_1936 | 0 | 1971.0 | Repeat | Note=49 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1940_1946 | 0 | 1971.0 | Repeat | Note=50 | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1947_1953 | 0 | 1971.0 | Repeat | Note=51%3B approximate | |
| Tgene | POLR2A | chr9:131768868 | chr17:7407022 | ENST00000322644 | 0 | 29 | 1954_1960 | 0 | 1971.0 | Repeat | Note=52%3B approximate |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for NUP188-POLR2A |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >61009_61009_1_NUP188-POLR2A_NUP188_chr9_131768868_ENST00000372577_POLR2A_chr17_7407022_ENST00000322644_length(transcript)=8799nt_BP=5601nt AGGGCGAGCGGGCGCGCGAAGATGGCGGCGGCCGCCGGCGGGCCGTGTGTGAGGAGCAGTAGAGAACTGTGGACTATTCTGCTTGGAAGG TCAGCTCTGAGAGAGCTGAGTCAGATTGAGGCAGAACTGAATAAACATTGGCGGCGATTGTTAGAGGGGCTTTCTTACTACAAACCTCCC AGTCCAAGTTCAGCTGAAAAAGTGAAAGCTAATAAAGATGTAGCTTCACCATTGAAGGAACTGGGTTTAAGAATCAGCAAGTTTTTGGGT CTTGATGAAGAACAGAGTGTGCAGTTACTCCAGTGTTACCTGCAAGAGGACTACAGGGGTACTCGGGACTCAGTAAAGACAGTACTGCAA GATGAGAGGCAGAGCCAGGCCTTAATCCTGAAGATTGCAGATTATTATTATGAAGAAAGAACCTGTATTCTTCGTTGTGTCTTACACCTT CTCACTTACTTCCAAGATGAAAGACACCCCTATAGGGTTGAATATGCAGACTGTGTTGATAAATTGGAGAAGGAACTAGTTTCAAAATAC AGACAGCAGTTCGAAGAGCTTTATAAAACTGAAGCACCAACTTGGGAGACACATGGAAATCTCATGACAGAGCGCCAAGTGTCTCGCTGG TTTGTTCAGTGCCTTCGGGAACAGTCCATGCTGCTAGAAATTATTTTCCTTTATTATGCATACTTTGAGATGGCACCCAGTGACTTACTT GTATTAACCAAGATGTTTAAAGAGCAAGGATTTGGTAGTAGGCAGACCAATAGGCACCTGGTGGATGAGACTATGGATCCTTTTGTAGAT CGGATTGGCTACTTCAGTGCCCTCATCCTGGTGGAGGGCATGGATATCGAGTCCTTGCATAAGTGTGCTTTGGATGACAGAAGAGAACTG CATCAGTTTGCGCAGGATGGGCTTATTTGTCAGGATATGGACTGTTTAATGTTGACCTTTGGGGACATTCCACATCATGCCCCAGTGCTT TTGGCCTGGGCTCTCCTCCGTCACACTCTGAACCCAGAAGAGACAAGCAGTGTGGTCCGGAAGATAGGTGGCACAGCCATCCAGCTGAAT GTGTTTCAGTACTTGACCCGATTGCTCCAGTCCCTTGCCAGTGGGGGAAATGATTGCACCACCAGCACTGCATGCATGTGTGTCTATGGA CTGCTCTCTTTCGTTCTGACCTCGTTGGAGCTGCACACCCTGGGCAATCAGCAGGATATAATTGATACAGCATGTGAAGTATTGGCCGAC CCTTCTCTTCCGGAACTGTTCTGGGGAACAGAGCCAACTTCTGGCCTTGGGATCATTCTGGACAGTGTGTGTGGAATGTTTCCCCACCTT CTCTCCCCACTCCTGCAACTGCTCCGAGCCCTGGTATCAGGGAAGTCCACAGCCAAAAAGGTGTATAGCTTCTTGGATAAGATGTCTTTC TACAATGAACTTTATAAACACAAGCCTCATGATGTGATCTCCCATGAAGATGGAACTCTTTGGCGGAGACAAACACCCAAACTCCTTTAT CCCCTTGGGGGTCAAACCAACCTTCGCATACCTCAAGGCACTGTGGGCCAAGTAATGTTGGATGATAGGGCATACCTGGTACGCTGGGAA TACTCCTATAGCAGCTGGACCCTCTTTACCTGCGAGATTGAAATGTTGCTTCATGTTGTTTCAACTGCAGATGTGATTCAGCACTGCCAG CGAGTCAAACCCATCATTGATCTCGTCCATAAGGTCATCAGTACAGACCTGTCGATAGCAGACTGTCTCCTGCCCATCACATCTCGCATC TACATGCTGCTGCAGCGGTTAACGACAGTGATCTCCCCACCTGTGGATGTCATTGCTTCTTGTGTCAACTGCTTAACTGTTTTGGCTGCC CGCAATCCAGCAAAGGTCTGGACTGATCTTCGTCACACAGGTTTTTTACCATTTGTGGCCCATCCTGTCTCCAGCCTGAGTCAGATGATT AGTGCGGAAGGGATGAATGCTGGAGGGTACGGAAACCTCTTGATGAACAGTGAACAGCCTCAGGGCGAGTATGGGGTTACTATTGCCTTT CTGCGCTTGATCACCACCCTTGTCAAGGGGCAACTTGGTAGTACCCAGAGCCAAGGACTTGTACCCTGTGTAATGTTTGTGCTGAAGGAG ATGCTTCCCAGCTACCATAAGTGGCGCTACAACTCTCATGGAGTGAGGGAACAGATTGGTTGCCTGATCTTGGAGCTGATTCATGCGATA CTGAACCTGTGCCACGAGACAGACCTGCACAGCAGTCATACTCCCAGCCTGCAGTTTCTCTGCATCTGCAGCCTGGCATACACAGAAGCA GGACAGACAGTTATCAATATCATGGGCATTGGCGTGGACACCATTGACATGGTGATGGCTGCTCAGCCTCGAAGTGATGGGGCAGAGGGC CAGGGGCAGGGCCAGCTGCTGATCAAGACAGTGAAACTGGCATTCTCCGTCACCAACAATGTTATTCGGCTGAAACCTCCTTCTAATGTG GTGTCCCCCCTGGAACAGGCTCTCTCACAACATGGTGCTCATGGAAACAACCTCATTGCTGTTCTAGCCAAATACATCTACCACAAACAT GACCCTGCTTTGCCACGTCTTGCCATTCAGCTGCTGAAACGTCTGGCCACGGTGGCCCCAATGTCAGTGTATGCTTGTCTGGGCAATGAT GCGGCTGCCATTCGTGATGCCTTCCTGACCCGATTGCAGAGCAAAATTGAGGACATGCGCATCAAAGTCATGATTCTAGAGTTCCTCACT GTTGCAGTAGAGACCCAGCCAGGCCTCATCGAACTGTTTCTGAACCTGGAAGTTAAGGATGGCAGTGATGGCTCAAAGGAATTCAGCCTT GGGATGTGGAGCTGTCTCCATGCAGTGCTGGAGCTGATTGATTCCCAACAGCAAGATCGATACTGGTGCCCACCCCTGCTGCATCGTGCC GCCATTGCCTTTTTGCATGCTCTGTGGCAGGATCGGAGGGACAGTGCCATGCTGGTCCTCCGAACCAAACCCAAGTTTTGGGAAAATTTA ACCAGTCCGCTGTTTGGAACCCTTTCTCCTCCCTCTGAAACATCAGAGCCCAGCATCCTGGAAACCTGTGCCCTAATCATGAAGATAATT TGCTTGGAGATATACTATGTAGTAAAGGGTTCATTAGACCAGTCATTAAAGGATACACTGAAGAAATTTTCCATCGAGAAACGCTTTGCC TACTGGTCAGGGTATGTCAAGTCATTGGCAGTTCACGTGGCCGAAACAGAAGGCAGCAGCTGCACCTCCTTGTTAGAGTACCAGATGCTG GTGTCCGCCTGGAGGATGCTTCTCATCATTGCCACCACTCATGCAGATATAATGCACCTGACTGACTCTGTGGTGCGTCGCCAGCTCTTT CTTGACGTGCTTGATGGAACCAAAGCATTACTCCTAGTTCCAGCCTCAGTGAACTGCCTTCGCCTTGGCTCCATGAAGTGCACTCTGCTG CTTATCCTCCTCCGGCAGTGGAAGAGAGAGTTAGGTTCTGTGGATGAAATCCTTGGACCCTTGACGGAGATCCTGGAGGGAGTGCTGCAG GCCGACCAGCAACTCATGGAGAAGACCAAGGCCAAGGTGTTCTCAGCATTCATCACAGTGTTGCAAATGAAGGAGATGAAAGTAAGTGAC ATCCCCCAGTACTCCCAGCTGGTGCTGAATGTCTGTGAGACCCTCCAAGAGGAAGTGATTGCACTCTTCGACCAGACCCGCCACAGTCTG GCATTAGGCAGTGCCACAGAGGACAAGGACAGCATGGAGACTGACGACTGTTCTCGGTCCCGGCACAGGGACCAGCGTGATGGGGTGTGT GTCCTGGGCCTGCACCTGGCCAAGGAGCTGTGTGAGGTAGACGAGGATGGTGACTCCTGGCTGCAGGTAACCCGCAGGCTCCCCATCCTA CCCACCCTCCTCACCACTCTAGAGGTGAGCCTTCGCATGAAGCAGAACCTGCATTTCACTGAGGCCACATTGCATCTGCTCCTCACCCTG GCTCGCACTCAGCAGGGAGCCACAGCAGTGGCTGGAGCTGGCATCACCCAGAGCATTTGTTTGCCCCTTCTGAGTGTGTACCAGCTGAGC ACCAACGGCACAGCACAGACACCTAGTGCCTCTCGGAAGTCCCTGGATGCCCCCTCTTGGCCAGGAGTCTACCGCCTGTCCATGTCCCTG ATGGAGCAGCTGCTCAAAACTCTGCGCTACAACTTCCTGCCTGAGGCCCTGGACTTCGTGGGTGTCCACCAGGAGCGGACCTTACAGTGC CTCAACGCAGTGAGGACAGTGCAGAGTCTGGCCTGCCTGGAGGAGGCGGACCACACCGTGGGTTTTATTCTGCAGCTCTCTAACTTCATG AAGGAGTGGCACTTCCACCTGCCTCAGCTCATGCGTGATATCCAGGTCAACCTGGGTTACTTGTGCCAGGCATGTACCTCTCTCCTGCAC AGTCGAAAGATGCTGCAGCATTACTTACAGAACAAAAATGGGGATGGCCTCCCCTCAGCTGTTGCCCAGCGAGTCCAGAGGCCACCGTCT GCTGCTTCTGCTGCCCCCTCCTCCTCAAAGCAGCCCGCTGCTGACACAGAGGCATCAGAGCAGCAGGCCTTGCACACAGTCCAGTATGGC CTTCTCAAGATCCTCAGCAAGACGCTGGCAGCCCTGCGCCACTTCACCCCAGATGTCTGCCAGATTCTGCTGGATCAGTCCCTGGACCTT GCTGAATACAACTTCCTGTTTGCCCTGAGCTTTACCACTCCCACCTTTGACTCCGAAGTGGCCCCCTCCTTCGGGACCCTTCTGGCCACA GTGAATGTGGCCCTCAACATGCTTGGAGAGCTGGACAAGAAAAAGGAGCCCCTCACCCAGGCAGTGGGGCTCAGCACACAGGCAGAAGGG ACCAGGACGTTAAAGTCCCTCCTGATGTTTACCATGGAAAACTGCTTCTACCTGCTCATCTCTCAGGCGATGCGGTACCTTAGGGACCCG GCTGTGCACCCCCGGGACAAACAGCGGATGAAGCAGGAGCTCAGCTCTGAGTTGAGTCCACCTCTCTCTCCAAAGCCAGCCCTGAGAGTC AGGAGCCTCTGATCCAGTTGGTGCAGGCGTTTGTCCGGCATATGCAAAGATAGGGCAGTGCTGTTCTGCCCACCTACCCCTCTCCACCAG CCTACACTGCACCCTGGCTGGCAGGGGTGCTGCTGGCTGCTAGGGCCTATACAATGGAGGGCACCTCCTGTCACCCCCCTCCCGGAGTAG CCACGACTCCAGCCACCACCCACTGACGTTATTTTTATACTAGATGAAGAGGTCAACAGCAGGCATGGGGAGCCGAGTCTTCTGTGCTCA GGTCCTCACGCTGCAGACGCCCCCTAGAGGAACTTTCCTTCCTTTCCAGCATTCCCCACAGCACTGCCGGCCAGGGGAGAGGCGGCAGCC CAGCAGAGGGCTCTATGCACGGGTTTCAAACCTGTTTTCCACACTCTGTCTTTGCAGTTTTGGTAATTCTGTGGTCTATTTATACAGATA TTAAAATCTTGTTTATAGACACCGCCGCATGGCAGAGGAGTTTCGGCTCAGTGGGGAGGCCTTCGACTGGCTGCTTGGGGAGATTGAGTC CAAGTTCAACCAAGCCATTGCGCATCCCGGGGAAATGGTGGGGGCTCTGGCTGCGCAGTCCCTTGGAGAACCTGCCACCCAGATGACCTT GAATACCTTCCACTATGCTGGTGTGTCTGCCAAGAATGTGACGCTGGGTGTGCCCCGACTTAAGGAGCTCATCAACATTTCCAAGAAGCC AAAGACTCCTTCGCTTACTGTCTTCCTGTTGGGCCAGTCCGCTCGAGATGCTGAGAGAGCCAAGGATATTCTGTGCCGTCTGGAGCATAC AACGTTGAGGAAGGTGACTGCCAACACAGCCATCTACTATGACCCCAACCCCCAGAGCACGGTGGTGGCAGAGGATCAGGAATGGGTGAA TGTCTACTATGAAATGCCTGACTTTGATGTGGCCCGAATCTCCCCCTGGCTGTTGCGGGTGGAGCTGGATCGGAAGCACATGACTGACCG GAAGCTCACCATGGAGCAGATTGCTGAAAAGATCAATGCTGGTTTTGGTGACGACTTGAACTGCATCTTTAATGATGACAATGCAGAGAA GCTGGTGCTCCGTATTCGCATCATGAACAGCGATGAGAACAAGATGCAAGAGGAGGAAGAGGTGGTGGACAAGATGGATGATGATGTCTT CCTGCGCTGCATCGAGTCCAACATGCTGACAGATATGACCCTGCAGGGCATCGAGCAGATCAGCAAGGTGTACATGCACTTGCCACAGAC AGACAACAAGAAGAAGATCATCATCACGGAGGATGGGGAATTCAAGGCCCTGCAGGAGTGGATCCTGGAGACGGACGGCGTGAGCTTGAT GCGGGTGCTGAGTGAGAAGGACGTGGACCCCGTACGCACCACGTCCAATGACATTGTGGAGATCTTCACGGTGCTGGGCATTGAAGCCGT GCGGAAGGCCCTGGAGCGGGAGCTGTACCACGTCATCTCCTTTGATGGCTCCTATGTCAATTACCGACACTTGGCTCTCTTGTGTGATAC CATGACCTGTCGTGGCCACTTGATGGCCATCACCCGACACGGAGTCAACCGCCAGGACACAGGACCACTCATGAAGTGTTCCTTTGAGGA AACGGTGGACGTGCTTATGGAAGCAGCCGCACACGGTGAGAGTGACCCCATGAAGGGGGTCTCTGAGAATATCATGCTGGGCCAGCTGGC TCCGGCCGGCACTGGCTGCTTTGACCTCCTGCTTGATGCAGAGAAGTGCAAGTATGGCATGGAGATCCCCACCAATATCCCCGGCCTGGG GGCTGCTGGACCCACCGGCATGTTCTTTGGTTCAGCACCCAGTCCCATGGGTGGAATCTCTCCTGCCATGACACCTTGGAACCAGGGTGC AACCCCTGCCTATGGCGCCTGGTCCCCCAGTGTTGGGAGTGGAATGACCCCAGGGGCAGCCGGCTTCTCTCCCAGTGCTGCGTCAGATGC CAGCGGCTTCAGCCCAGGTTACTCCCCTGCCTGGTCTCCCACACCGGGCTCCCCGGGGTCCCCAGGTCCCTCAAGCCCCTACATCCCTTC ACCAGGTGGTGCCATGTCTCCCAGCTACTCGCCAACGTCACCTGCCTACGAGCCCCGCTCTCCTGGGGGCTACACACCCCAGAGTCCCTC TTATTCCCCCACTTCACCCTCCTACTCCCCTACCTCTCCATCCTATTCTCCAACCAGTCCCAACTATAGTCCCACATCACCCAGCTATTC GCCAACGTCACCCAGCTACTCACCGACCTCTCCCAGCTACTCACCCACCTCTCCCAGCTACTCGCCCACCTCTCCCAGCTACTCGCCCAC CTCTCCCAGCTACTCACCCACTTCCCCTAGCTACTCGCCCACTTCCCCTAGCTACTCGCCAACGTCTCCCAGCTACTCACCGACATCTCC CAGCTACTCGCCAACTTCACCCAGCTATTCTCCCACTTCTCCCAGCTACTCACCTACCTCTCCAAGCTATTCACCCACCTCCCCCAGCTA CTCACCCACTTCCCCAAGTTACTCACCCACCAGCCCGAACTATTCTCCAACCAGTCCCAATTACACCCCAACATCACCCAGCTACAGCCC GACATCACCCAGCTATTCACCTACTAGTCCCAACTACACACCTACCAGCCCTAACTACAGCCCAACCTCTCCAAGCTACTCTCCAACATC ACCCAGCTATTCCCCGACCTCACCAAGTTACTCCCCTTCCAGCCCACGATACACACCACAGTCTCCAACCTATACCCCAAGCTCACCCAG CTACAGCCCCAGCTCGCCCAGCTACAGCCCAGCCTCACCCAAGTACACCCCAACCAGTCCTTCTTACAGTCCCAGCTCCCCAGAGTATAC CCCAACCTCTCCCAAGTACTCACCTACCAGTCCCAAATATTCACCCACCTCTCCCAAGTACTCGCCTACCAGTCCCACCTATTCACCCAC CACCCCAAAATACTCCCCAACATCTCCTACTTATTCCCCAACCTCTCCAGTCTACACCCCAACCTCTCCCAAGTACTCACCTACTAGCCC CACTTACTCGCCCACTTCCCCCAAGTACTCGCCCACCAGCCCCACCTACTCGCCCACCTCCCCCAAAGGCTCAACCTACTCTCCCACTTC CCCTGGTTACTCGCCCACCAGCCCCACCTACAGTCTCACAAGCCCGGCTATCAGCCCGGATGACAGTGACGAGGAGAACTGAGGGCACGT GGGGTGCGGCAGCGGGCTAGGGCCCAGGGCAGCTTGCCCGTGCTGCTGTGCAGTTCTTGCCTCCCTCACGGGGCGTCACCCCCAGCCCAG CTCCGTTGTACATAAATGCCTTGTGGCAGAGCTCCCGGTGAACTTCTGGATCCCGTTTCTGATGCAGACTCTTGTCTTGTTCTCCACTTG TGCTGTTAGAACTCACTGGCCCAGTGGTGTTCTCACTCCTACCCCACCCACCCCCTGCCTGTCCCCAAATTGAAGATCCTTCCTTGCCTG TGGCTTGATGCGGGGCGGGTAAAGGGTATTTTAACTTAGGGGTAGTTCCTGCTGTGAGTGGTTACAGCTGATCCTCGGGAAGAACAAAGC >61009_61009_1_NUP188-POLR2A_NUP188_chr9_131768868_ENST00000372577_POLR2A_chr17_7407022_ENST00000322644_length(amino acids)=1706AA_BP= MAAAAGGPCVRSSRELWTILLGRSALRELSQIEAELNKHWRRLLEGLSYYKPPSPSSAEKVKANKDVASPLKELGLRISKFLGLDEEQSV QLLQCYLQEDYRGTRDSVKTVLQDERQSQALILKIADYYYEERTCILRCVLHLLTYFQDERHPYRVEYADCVDKLEKELVSKYRQQFEEL YKTEAPTWETHGNLMTERQVSRWFVQCLREQSMLLEIIFLYYAYFEMAPSDLLVLTKMFKEQGFGSRQTNRHLVDETMDPFVDRIGYFSA LILVEGMDIESLHKCALDDRRELHQFAQDGLICQDMDCLMLTFGDIPHHAPVLLAWALLRHTLNPEETSSVVRKIGGTAIQLNVFQYLTR LLQSLASGGNDCTTSTACMCVYGLLSFVLTSLELHTLGNQQDIIDTACEVLADPSLPELFWGTEPTSGLGIILDSVCGMFPHLLSPLLQL LRALVSGKSTAKKVYSFLDKMSFYNELYKHKPHDVISHEDGTLWRRQTPKLLYPLGGQTNLRIPQGTVGQVMLDDRAYLVRWEYSYSSWT LFTCEIEMLLHVVSTADVIQHCQRVKPIIDLVHKVISTDLSIADCLLPITSRIYMLLQRLTTVISPPVDVIASCVNCLTVLAARNPAKVW TDLRHTGFLPFVAHPVSSLSQMISAEGMNAGGYGNLLMNSEQPQGEYGVTIAFLRLITTLVKGQLGSTQSQGLVPCVMFVLKEMLPSYHK WRYNSHGVREQIGCLILELIHAILNLCHETDLHSSHTPSLQFLCICSLAYTEAGQTVINIMGIGVDTIDMVMAAQPRSDGAEGQGQGQLL IKTVKLAFSVTNNVIRLKPPSNVVSPLEQALSQHGAHGNNLIAVLAKYIYHKHDPALPRLAIQLLKRLATVAPMSVYACLGNDAAAIRDA FLTRLQSKIEDMRIKVMILEFLTVAVETQPGLIELFLNLEVKDGSDGSKEFSLGMWSCLHAVLELIDSQQQDRYWCPPLLHRAAIAFLHA LWQDRRDSAMLVLRTKPKFWENLTSPLFGTLSPPSETSEPSILETCALIMKIICLEIYYVVKGSLDQSLKDTLKKFSIEKRFAYWSGYVK SLAVHVAETEGSSCTSLLEYQMLVSAWRMLLIIATTHADIMHLTDSVVRRQLFLDVLDGTKALLLVPASVNCLRLGSMKCTLLLILLRQW KRELGSVDEILGPLTEILEGVLQADQQLMEKTKAKVFSAFITVLQMKEMKVSDIPQYSQLVLNVCETLQEEVIALFDQTRHSLALGSATE DKDSMETDDCSRSRHRDQRDGVCVLGLHLAKELCEVDEDGDSWLQVTRRLPILPTLLTTLEVSLRMKQNLHFTEATLHLLLTLARTQQGA TAVAGAGITQSICLPLLSVYQLSTNGTAQTPSASRKSLDAPSWPGVYRLSMSLMEQLLKTLRYNFLPEALDFVGVHQERTLQCLNAVRTV QSLACLEEADHTVGFILQLSNFMKEWHFHLPQLMRDIQVNLGYLCQACTSLLHSRKMLQHYLQNKNGDGLPSAVAQRVQRPPSAASAAPS SSKQPAADTEASEQQALHTVQYGLLKILSKTLAALRHFTPDVCQILLDQSLDLAEYNFLFALSFTTPTFDSEVAPSFGTLLATVNVALNM -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for NUP188-POLR2A |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for NUP188-POLR2A |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for NUP188-POLR2A |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
