
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PCMTD1-ST18 (FusionGDB2 ID:63379) |
Fusion Gene Summary for PCMTD1-ST18 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PCMTD1-ST18 | Fusion gene ID: 63379 | Hgene | Tgene | Gene symbol | PCMTD1 | ST18 | Gene ID | 115294 | 9705 |
| Gene name | protein-L-isoaspartate (D-aspartate) O-methyltransferase domain containing 1 | ST18 C2H2C-type zinc finger transcription factor | |
| Synonyms | - | NZF-3|NZF3|ZC2H2C3|ZC2HC10|ZNF387 | |
| Cytomap | 8q11.23 | 8q11.23 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein-L-isoaspartate O-methyltransferase domain-containing protein 1 | suppression of tumorigenicity 18 proteinST18, C2H2C-type zinc fingerneural zinc finger transcription factor 3suppression of tumorigenicity 18 (breast carcinoma) (zinc finger protein)suppression of tumorigenicity 18, zinc fingerzinc finger protein 387 | |
| Modification date | 20200313 | 20200320 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000360540, ENST00000522514, ENST00000544451, ENST00000521344, ENST00000519559, | ENST00000276480, ENST00000522102, | |
| Fusion gene scores | * DoF score | 27 X 12 X 15=4860 | 4 X 7 X 5=140 |
| # samples | 40 | 11 | |
| ** MAII score | log2(40/4860*10)=-3.60288440871842 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/140*10)=-0.347923303420307 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PCMTD1 [Title/Abstract] AND ST18 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PCMTD1(52811490)-ST18(53159224), # samples:3 PCMTD1(52811490)-ST18(53079546), # samples:3 PCMTD1(52773405)-ST18(53062537), # samples:3 PCMTD1(52758221)-ST18(53142637), # samples:3 PCMTD1(52811490)-ST18(53085143), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | PCMTD1-ST18 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. PCMTD1-ST18 seems lost the major protein functional domain in Tgene partner, which is a transcription factor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | ST18 | GO:0008285 | negative regulation of cell proliferation | 15489893 |
 Fusion gene breakpoints across PCMTD1 (5'-gene) Fusion gene breakpoints across PCMTD1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
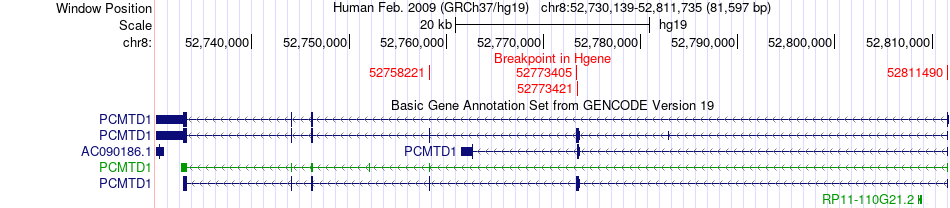 |
 Fusion gene breakpoints across ST18 (3'-gene) Fusion gene breakpoints across ST18 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
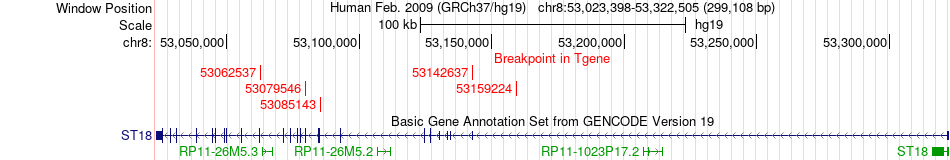 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-A2-A4S3-01A | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| ChimerDB4 | BRCA | TCGA-AO-A128-01A | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| ChimerDB4 | BRCA | TCGA-C8-A12K-01A | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| ChimerDB4 | BRCA | TCGA-C8-A12K-01A | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| ChimerDB4 | PRAD | TCGA-VP-A878-01A | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| ChimerDB4 | PRAD | TCGA-VP-A878-01A | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| ChimerDB4 | PRAD | TCGA-VP-A878 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| ChimerDB4 | PRAD | TCGA-YJ-A8SW-01A | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| ChimerDB4 | PRAD | TCGA-YJ-A8SW | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| ChimerDB4 | PRAD | TCGA-YJ-A8SW | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| ChimerDB4 | UCEC | TCGA-BK-A0CC-01A | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
Top |
Fusion Gene ORF analysis for PCMTD1-ST18 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| 5CDS-5UTR | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| 5CDS-5UTR | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5CDS-intron | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| 5CDS-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| 5CDS-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| 5CDS-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| 5CDS-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| 5CDS-intron | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| 5CDS-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| 5CDS-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| 5CDS-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| 5CDS-intron | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5CDS-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5CDS-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5CDS-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5CDS-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-3CDS | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-3CDS | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-3CDS | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-3CDS | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-3CDS | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-3CDS | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-3CDS | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-3CDS | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-5UTR | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-5UTR | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-5UTR | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-5UTR | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-5UTR | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-intron | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-intron | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-intron | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| 5UTR-intron | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53159224 | - |
| 5UTR-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| 5UTR-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53142637 | - |
| 5UTR-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| Frame-shift | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| Frame-shift | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53079546 | - |
| Frame-shift | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52811490 | - | ST18 | chr8 | 53085143 | - |
| In-frame | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| In-frame | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000519559 | ENST00000276480 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000522514 | ENST00000276480 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| intron-3CDS | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-5UTR | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| intron-5UTR | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| intron-intron | ENST00000360540 | ENST00000522102 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000519559 | ENST00000522102 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| intron-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| intron-intron | ENST00000521344 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| intron-intron | ENST00000522514 | ENST00000522102 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000544451 | ENST00000276480 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
| intron-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - |
| intron-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53142637 | - |
| intron-intron | ENST00000544451 | ENST00000522102 | PCMTD1 | chr8 | 52758221 | - | ST18 | chr8 | 53159224 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000360540 | PCMTD1 | chr8 | 52773405 | - | ENST00000276480 | ST18 | chr8 | 53062537 | - | 4411 | 714 | 747 | 2051 | 434 |
| ENST00000521344 | PCMTD1 | chr8 | 52773421 | - | ENST00000276480 | ST18 | chr8 | 53062537 | - | 4267 | 570 | 267 | 1907 | 546 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000360540 | ENST00000276480 | PCMTD1 | chr8 | 52773405 | - | ST18 | chr8 | 53062537 | - | 0.000287548 | 0.9997124 |
| ENST00000521344 | ENST00000276480 | PCMTD1 | chr8 | 52773421 | - | ST18 | chr8 | 53062537 | - | 0.000492424 | 0.99950755 |
Top |
Fusion Genomic Features for PCMTD1-ST18 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
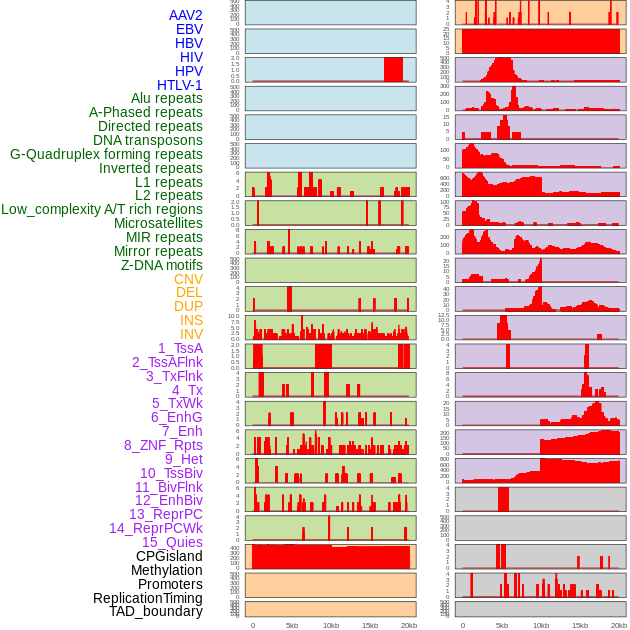 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for PCMTD1-ST18 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr8:52811490/chr8:53159224) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 920_992 | 602 | 1048.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 920_992 | 602 | 1048.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 715_758 | 602 | 1048.0 | Zinc finger | CCHHC-type 3 | |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 759_802 | 602 | 1048.0 | Zinc finger | CCHHC-type 4 | |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 807_850 | 602 | 1048.0 | Zinc finger | CCHHC-type 5 | |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 860_903 | 602 | 1048.0 | Zinc finger | CCHHC-type 6 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 715_758 | 602 | 1048.0 | Zinc finger | CCHHC-type 3 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 759_802 | 602 | 1048.0 | Zinc finger | CCHHC-type 4 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 807_850 | 602 | 1048.0 | Zinc finger | CCHHC-type 5 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 860_903 | 602 | 1048.0 | Zinc finger | CCHHC-type 6 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 359_402 | 602 | 1048.0 | Zinc finger | CCHHC-type 1 | |
| Tgene | ST18 | chr8:52773405 | chr8:53062537 | ENST00000276480 | 14 | 26 | 403_446 | 602 | 1048.0 | Zinc finger | CCHHC-type 2 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 359_402 | 602 | 1048.0 | Zinc finger | CCHHC-type 1 | |
| Tgene | ST18 | chr8:52773421 | chr8:53062537 | ENST00000276480 | 14 | 26 | 403_446 | 602 | 1048.0 | Zinc finger | CCHHC-type 2 |
Top |
Fusion Gene Sequence for PCMTD1-ST18 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >63379_63379_1_PCMTD1-ST18_PCMTD1_chr8_52773405_ENST00000360540_ST18_chr8_53062537_ENST00000276480_length(transcript)=4411nt_BP=714nt GGCGGCTCCGGAGTCCGGGAGGCTCTGGCTGCTCCAGTCGCGGGTCCGGGCCCCGCCTCTCCACTCGGCGGTGGCGTCCGTGTTGTTATT GTGGTGGTAGCGGCGTCTGGCTGCTGCGGACCCCGCCGAGTCCTAGCGCCTGGCCTGCGCGCCGCTGCCCGCGCCACAGCCCCATCACTG AGAAGAATTTTTTCACTGTATAGTTGCCTCTGCACTTAAAGCAGGCTTGGTCCATGAATCCCATGACAAGAGCTTTGAGGACATGAGGAG CATTCTTTGTTCAAACATACTGTGTTTTCCTTTGGATCACTGGATTAATTTATTTTTGGAAATCAAGTGCAATATTGGAAGCCATTTCTA CTAATTTTATTTCACTTTTAAATTTATGATTTGATTTGAATACTATCATGGGAGGAGCTGTGAGTGCTGGGGAAGATAATGATGACTTAA TTGATAATTTAAAAGAAGCTCAGTATATTCGTACTGAAAGAGTGGAGCAAGCCTTCAGAGCGATTGATCGTGGAGATTACTATTTGGAAG GCTACAGAGACAATGCTTACAAAGACTTAGCCTGGAAGCATGGAAACATCCACTTGTCAGCACCTTGCATTTATTCTGAAGTTATGGAAG CATTGAAACTTCAACCAGGATTGTCTTTTCTTAACCTGGGAAGTGGAACCGGATATTTAAGTACAATGGTGGGCTTAATTTTAGGGAGCC GAAATAGAAGTGGATGAAAATGGCACATTGGACTTAAGCATGAAAAAAAATCGAATCCTGGACAAGTCTGCACCCCTAACTTCCTCTAAC ACTTCTATTCCAACTCCTTCCTCTTCCCCATTCAAAACAAGCAGCATTCTGGTCAATGCAGCATTCTATCAGGCTCTTTGTGACCAAGAG GGCTGGGACACTCCTATCAACTATAGCAAAACTCACGGGAAGACAGAGGAGGAGAAAGAGAAAGACCCAGTGAGCTCTCTAGAAAATTTA GAGGAAAAAAAGTTTCCTGGAGAGGCCTCTATACCAAGCCCTAAACCCAAGCTTCATGCAAGAGATCTCAAAAAGGAACTAATCACCTGT CCAACACCAGGATGTGATGGAAGTGGCCACGTGACAGGAAACTATGCATCTCATCGCAGTGTTTCTGGATGTCCTTTAGCAGATAAGACT CTAAAATCCCTCATGGCTGCCAACTCTCAGGAGCTTAAGTGTCCAACCCCAGGCTGCGATGGCTCGGGGCACGTGACTGGAAACTATGCT TCCCACAGAAGCTTGTCCGGATGCCCTCGTGCAAGGAAAGGTGGTGTCAAAATGACCCCTACCAAGGAAGAAAAAGAAGACCCTGAACTG AAATGTCCTGTGATAGGGTGTGATGGCCAAGGTCACATATCAGGTAAATACACATCACACCGCACAGCTTCTGGCTGTCCTCTGGCTGCC AAGAGACAGAAGGAGAATCCTCTCAATGGAGCCTCCCTCTCCTGGAAACTGAACAAACAAGAGCTACCACATTGTCCCTTGCCAGGCTGC AATGGGCTGGGCCATGTAAATAATGTTTTTGTCACCCACCGAAGCTTATCTGGATGTCCTCTCAATGCACAAGTTATCAAAAAGGGCAAG GTTTCTGAAGAACTCATGACCATCAAGCTCAAAGCAACTGGGGGAATAGAGAGTGATGAAGAAATTAGGCATTTGGATGAAGAAATAAAG GAACTGAATGAATCCAACCTTAAAATTGAAGCAGATATGATGAAACTTCAGACCCAGATCACATCTATGGAGAGCAACTTAAAGACGATA GAGGAGGAGAACAAACTCATAGAACAGAACAATGAAAGTCTGCTGAAAGAGCTGGCAGGTCTAAGCCAAGCTCTCATTTCAAGCCTTGCT GACATCCAGCTTCCACAGATGGGACCTATCAGTGAGCAGAATTTTGAAGCATATGTAAATACACTCACAGATATGTACAGCAATCTGGAA CGGGACTATTCCCCGGAATGCAAAGCTCTACTGGAAAGTATCAAACAGGCAGTGAAGGGTATCCATGTGTAGGATCACAGCGCTGCCGGG CAACAGAAGTTACCAACAGCAGTAAACTCCAGATGGATCTGTTAGAGGTTCATGTACTGCTAAGGCGTGGAGGTTGCCGTACTGCATTTA CAATTTGCAACATTGCACTAATTTTATTTTCCCCAGCTGATATAAAAAGGAAAGAAAAACTATGATAGACTTCTTGGATTAAAAGCAATG CAGTCAATTATTAGATCTTATTTATTTTCATATGTTTTTCTTTTATTTCTTCATTGTACTCTTTTTTTTTTTGTAAAGTATATGTAAAAT AAATGTGACATTTTTATAATTTATTTATTACTAATCAAAGAGTTTTTTATCTTTTAACTGCATTTTGAAGTCTGCCGTATTTTTACAAGT GTGTTTATTAATTTATTTTCCAATAGGATTTAAATAGAAATGCTATTCTCAAGTCATCTTTCTTGCTGGGTTTTAATGAGGAAACAGGAA AGGGTGAAGGAAATCCTTGTCTAAGGACTGCACTATAGTTGAGTTTGATTTTTATTGCACACTTCTTCCCCCACCTTTCACTGATTTTTG TATTTATAAATGAATTTGCGGTAAGGTGAGCTGCACGGAAGGAATAAGAAGACAAATGGCGCCCACTAGTGGGGAATCCGCACTCACAAA AGCACAGGATGCTGGAAAACAGCCTGCTCAGAATTTGTTAGCAATAATTAAATATAGCAATCAGCAAAGTATTCGACTTGGCTGGACGGT TTTCGTTAATATGAATTATTTATTTGAAATGTTTTAAAGAAACATAAGCCTTTTTAGTGATGCAGATTTGTCTGTTTGTTTTTCAAGTCA TATCAGATCGTTGGCAACTCGTATCCCAAGATGAAAAATAAGACTTGGTGTGACCAGCCAGGCTTTCCTGCCATATGTTGGTACAATATA CAAGTGACAATATTGGTGTAGATTTGTACTTAGCAAATACAAACACATCCAAATGAAAAATTTTGTAGATACCATATCCCCTGAAATAGC ATTTATCTTACTGGGTTGACTGGAAAGGAATGGAAAATATAGTAACACATGAAAAAATGCTACTCCAATCTGAATGATTACTTCAAACAC TGGCACCTTGGGTCTCACCCACCATAGGAAACAAGACAACATTCAATTTGATAGAAATCTTGCCACAAAACTTCAAATGCTACAAAATAT ACACACACACTCACACACACAGGCATACTCACACACAGACACACACACACACACACACACAGACTCATCCACACTTCAAATTGAGCCCAC AATCTTGAATTTCTGAACGGATCAGAGTTTCATAGTTTCTATAGTAAAGGCAATGTCTATTTCAGGGATTGTAAAGTAGTTAAGCATTGT TTCAAAAGTTTTTTTATATTTATTTTTTTTAAGGAAAAGGTATAGACAACCAGCTAAACTGCCTTTTTGGTGTGCACACACATTTCATGT GCAGACGTGCCTCTGTGTAAATGTACACATGAACTTCATGTGGGCTTAATTTTCTGTGCTATAAACAAAAGTGTTTATTTTTTATTAACC TCATGGATATTTAGATGGAAAGTGATGGCATTCACAGGCTTGATGTATTCCACTGTTATTACTGTTACCTGCACAAATGAAAAACAATAC TCAACAGTAATTCCACTCCCATGAAACTTTGGTCATTGTTATGCATTAAGTGGGGCTTATCTTTGGTTTGGAGTTCATTTGAACTCTTGA ACCTTAGTTTAGTGAAGATGAACTGTCTGTTCTTAGGTAGAAACGGTGTTTATTTAAAAATCAGTTTTAAAAAATGAGCTACCATATGTG CTGTCTATTATAAATGGGACACCAAACAAAATTTTCTATTACAGTTGTGTACTTGCAAACATTTTGCTATACAGTACTTCATAGATGCAT ACAAATGAGCTCACTTATTACAAAGACAAACGTTTAATTTGCTAAATATTTTAACAAGTTTGTTATATATTTTATTTAATTTAAAAGAAA TCTCTTACCAACCTACATATTTATTACTATAATTTGCTATGACTTCAGGTTAATTTATTTGTGTTTGCATAGTTTGAGCAGGATGTTTTG TGAAGTATGTTTGTATTTATTTGCCTACTTTGTACTTGATGTGTTTTGTAATGTGCACTGAATTTGTTTTCTTTTCAACTATGTTAATGA TCAATACTGTAAATTGGGTCTTTTGTAAACAAAAAGGCAATGATGTATGCATTTTTTTTAATTTGAGGTAGTTTGTTTGTATACTGTTTC TCCAAACACTTAATATTTCTTACATCAAAGCAACAAAATTGTGTTCAGTGCTGTACATTTGGTGTATGGTAGGAAATAAAAATTGATAAC >63379_63379_1_PCMTD1-ST18_PCMTD1_chr8_52773405_ENST00000360540_ST18_chr8_53062537_ENST00000276480_length(amino acids)=434AA_BP=110 MDLSMKKNRILDKSAPLTSSNTSIPTPSSSPFKTSSILVNAAFYQALCDQEGWDTPINYSKTHGKTEEEKEKDPVSSLENLEEKKFPGEA SIPSPKPKLHARDLKKELITCPTPGCDGSGHVTGNYASHRSVSGCPLADKTLKSLMAANSQELKCPTPGCDGSGHVTGNYASHRSLSGCP RARKGGVKMTPTKEEKEDPELKCPVIGCDGQGHISGKYTSHRTASGCPLAAKRQKENPLNGASLSWKLNKQELPHCPLPGCNGLGHVNNV FVTHRSLSGCPLNAQVIKKGKVSEELMTIKLKATGGIESDEEIRHLDEEIKELNESNLKIEADMMKLQTQITSMESNLKTIEEENKLIEQ -------------------------------------------------------------- >63379_63379_2_PCMTD1-ST18_PCMTD1_chr8_52773421_ENST00000521344_ST18_chr8_53062537_ENST00000276480_length(transcript)=4267nt_BP=570nt GCATGCCCGGACCCCGGCGGCTCCGGAGTCCGGGAGGCTCTGGCTGCTCCAGTCGCGGGTCCGGGCCCCGCCTCTCCACTCGGCGGTGGC GTCCGTGTTGTTATTGTGGTGGTAGCGGCGTCTGGCTGCTGCGGACCCCGCCGAGTCCTAGCGCCTGGCCTGCGCGCCGCTGCCCGCGCC ACAGGATTAATTTATTTTTGGAAATCAAGTGCAATATTGGAAGCCATTTCTACTAATTTTATTTCACTTTTAAATTTATGATTTGATTTG AATACTATCATGGGAGGAGCTGTGAGTGCTGGGGAAGATAATGATGACTTAATTGATAATTTAAAAGAAGCTCAGTATATTCGTACTGAA AGAGTGGAGCAAGCCTTCAGAGCGATTGATCGTGGAGATTACTATTTGGAAGGCTACAGAGACAATGCTTACAAAGACTTAGCCTGGAAG CATGGAAACATCCACTTGTCAGCACCTTGCATTTATTCTGAAGTTATGGAAGCATTGAAACTTCAACCAGGATTGTCTTTTCTTAACCTG GGAAGTGGAACCGGATATTTAAGTACAATGGGAGCCGAAATAGAAGTGGATGAAAATGGCACATTGGACTTAAGCATGAAAAAAAATCGA ATCCTGGACAAGTCTGCACCCCTAACTTCCTCTAACACTTCTATTCCAACTCCTTCCTCTTCCCCATTCAAAACAAGCAGCATTCTGGTC AATGCAGCATTCTATCAGGCTCTTTGTGACCAAGAGGGCTGGGACACTCCTATCAACTATAGCAAAACTCACGGGAAGACAGAGGAGGAG AAAGAGAAAGACCCAGTGAGCTCTCTAGAAAATTTAGAGGAAAAAAAGTTTCCTGGAGAGGCCTCTATACCAAGCCCTAAACCCAAGCTT CATGCAAGAGATCTCAAAAAGGAACTAATCACCTGTCCAACACCAGGATGTGATGGAAGTGGCCACGTGACAGGAAACTATGCATCTCAT CGCAGTGTTTCTGGATGTCCTTTAGCAGATAAGACTCTAAAATCCCTCATGGCTGCCAACTCTCAGGAGCTTAAGTGTCCAACCCCAGGC TGCGATGGCTCGGGGCACGTGACTGGAAACTATGCTTCCCACAGAAGCTTGTCCGGATGCCCTCGTGCAAGGAAAGGTGGTGTCAAAATG ACCCCTACCAAGGAAGAAAAAGAAGACCCTGAACTGAAATGTCCTGTGATAGGGTGTGATGGCCAAGGTCACATATCAGGTAAATACACA TCACACCGCACAGCTTCTGGCTGTCCTCTGGCTGCCAAGAGACAGAAGGAGAATCCTCTCAATGGAGCCTCCCTCTCCTGGAAACTGAAC AAACAAGAGCTACCACATTGTCCCTTGCCAGGCTGCAATGGGCTGGGCCATGTAAATAATGTTTTTGTCACCCACCGAAGCTTATCTGGA TGTCCTCTCAATGCACAAGTTATCAAAAAGGGCAAGGTTTCTGAAGAACTCATGACCATCAAGCTCAAAGCAACTGGGGGAATAGAGAGT GATGAAGAAATTAGGCATTTGGATGAAGAAATAAAGGAACTGAATGAATCCAACCTTAAAATTGAAGCAGATATGATGAAACTTCAGACC CAGATCACATCTATGGAGAGCAACTTAAAGACGATAGAGGAGGAGAACAAACTCATAGAACAGAACAATGAAAGTCTGCTGAAAGAGCTG GCAGGTCTAAGCCAAGCTCTCATTTCAAGCCTTGCTGACATCCAGCTTCCACAGATGGGACCTATCAGTGAGCAGAATTTTGAAGCATAT GTAAATACACTCACAGATATGTACAGCAATCTGGAACGGGACTATTCCCCGGAATGCAAAGCTCTACTGGAAAGTATCAAACAGGCAGTG AAGGGTATCCATGTGTAGGATCACAGCGCTGCCGGGCAACAGAAGTTACCAACAGCAGTAAACTCCAGATGGATCTGTTAGAGGTTCATG TACTGCTAAGGCGTGGAGGTTGCCGTACTGCATTTACAATTTGCAACATTGCACTAATTTTATTTTCCCCAGCTGATATAAAAAGGAAAG AAAAACTATGATAGACTTCTTGGATTAAAAGCAATGCAGTCAATTATTAGATCTTATTTATTTTCATATGTTTTTCTTTTATTTCTTCAT TGTACTCTTTTTTTTTTTGTAAAGTATATGTAAAATAAATGTGACATTTTTATAATTTATTTATTACTAATCAAAGAGTTTTTTATCTTT TAACTGCATTTTGAAGTCTGCCGTATTTTTACAAGTGTGTTTATTAATTTATTTTCCAATAGGATTTAAATAGAAATGCTATTCTCAAGT CATCTTTCTTGCTGGGTTTTAATGAGGAAACAGGAAAGGGTGAAGGAAATCCTTGTCTAAGGACTGCACTATAGTTGAGTTTGATTTTTA TTGCACACTTCTTCCCCCACCTTTCACTGATTTTTGTATTTATAAATGAATTTGCGGTAAGGTGAGCTGCACGGAAGGAATAAGAAGACA AATGGCGCCCACTAGTGGGGAATCCGCACTCACAAAAGCACAGGATGCTGGAAAACAGCCTGCTCAGAATTTGTTAGCAATAATTAAATA TAGCAATCAGCAAAGTATTCGACTTGGCTGGACGGTTTTCGTTAATATGAATTATTTATTTGAAATGTTTTAAAGAAACATAAGCCTTTT TAGTGATGCAGATTTGTCTGTTTGTTTTTCAAGTCATATCAGATCGTTGGCAACTCGTATCCCAAGATGAAAAATAAGACTTGGTGTGAC CAGCCAGGCTTTCCTGCCATATGTTGGTACAATATACAAGTGACAATATTGGTGTAGATTTGTACTTAGCAAATACAAACACATCCAAAT GAAAAATTTTGTAGATACCATATCCCCTGAAATAGCATTTATCTTACTGGGTTGACTGGAAAGGAATGGAAAATATAGTAACACATGAAA AAATGCTACTCCAATCTGAATGATTACTTCAAACACTGGCACCTTGGGTCTCACCCACCATAGGAAACAAGACAACATTCAATTTGATAG AAATCTTGCCACAAAACTTCAAATGCTACAAAATATACACACACACTCACACACACAGGCATACTCACACACAGACACACACACACACAC ACACACAGACTCATCCACACTTCAAATTGAGCCCACAATCTTGAATTTCTGAACGGATCAGAGTTTCATAGTTTCTATAGTAAAGGCAAT GTCTATTTCAGGGATTGTAAAGTAGTTAAGCATTGTTTCAAAAGTTTTTTTATATTTATTTTTTTTAAGGAAAAGGTATAGACAACCAGC TAAACTGCCTTTTTGGTGTGCACACACATTTCATGTGCAGACGTGCCTCTGTGTAAATGTACACATGAACTTCATGTGGGCTTAATTTTC TGTGCTATAAACAAAAGTGTTTATTTTTTATTAACCTCATGGATATTTAGATGGAAAGTGATGGCATTCACAGGCTTGATGTATTCCACT GTTATTACTGTTACCTGCACAAATGAAAAACAATACTCAACAGTAATTCCACTCCCATGAAACTTTGGTCATTGTTATGCATTAAGTGGG GCTTATCTTTGGTTTGGAGTTCATTTGAACTCTTGAACCTTAGTTTAGTGAAGATGAACTGTCTGTTCTTAGGTAGAAACGGTGTTTATT TAAAAATCAGTTTTAAAAAATGAGCTACCATATGTGCTGTCTATTATAAATGGGACACCAAACAAAATTTTCTATTACAGTTGTGTACTT GCAAACATTTTGCTATACAGTACTTCATAGATGCATACAAATGAGCTCACTTATTACAAAGACAAACGTTTAATTTGCTAAATATTTTAA CAAGTTTGTTATATATTTTATTTAATTTAAAAGAAATCTCTTACCAACCTACATATTTATTACTATAATTTGCTATGACTTCAGGTTAAT TTATTTGTGTTTGCATAGTTTGAGCAGGATGTTTTGTGAAGTATGTTTGTATTTATTTGCCTACTTTGTACTTGATGTGTTTTGTAATGT GCACTGAATTTGTTTTCTTTTCAACTATGTTAATGATCAATACTGTAAATTGGGTCTTTTGTAAACAAAAAGGCAATGATGTATGCATTT TTTTTAATTTGAGGTAGTTTGTTTGTATACTGTTTCTCCAAACACTTAATATTTCTTACATCAAAGCAACAAAATTGTGTTCAGTGCTGT >63379_63379_2_PCMTD1-ST18_PCMTD1_chr8_52773421_ENST00000521344_ST18_chr8_53062537_ENST00000276480_length(amino acids)=546AA_BP=222 MNTIMGGAVSAGEDNDDLIDNLKEAQYIRTERVEQAFRAIDRGDYYLEGYRDNAYKDLAWKHGNIHLSAPCIYSEVMEALKLQPGLSFLN LGSGTGYLSTMGAEIEVDENGTLDLSMKKNRILDKSAPLTSSNTSIPTPSSSPFKTSSILVNAAFYQALCDQEGWDTPINYSKTHGKTEE EKEKDPVSSLENLEEKKFPGEASIPSPKPKLHARDLKKELITCPTPGCDGSGHVTGNYASHRSVSGCPLADKTLKSLMAANSQELKCPTP GCDGSGHVTGNYASHRSLSGCPRARKGGVKMTPTKEEKEDPELKCPVIGCDGQGHISGKYTSHRTASGCPLAAKRQKENPLNGASLSWKL NKQELPHCPLPGCNGLGHVNNVFVTHRSLSGCPLNAQVIKKGKVSEELMTIKLKATGGIESDEEIRHLDEEIKELNESNLKIEADMMKLQ TQITSMESNLKTIEEENKLIEQNNESLLKELAGLSQALISSLADIQLPQMGPISEQNFEAYVNTLTDMYSNLERDYSPECKALLESIKQA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PCMTD1-ST18 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PCMTD1-ST18 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PCMTD1-ST18 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
