
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PDE4D-PLK2 (FusionGDB2 ID:63754) |
Fusion Gene Summary for PDE4D-PLK2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PDE4D-PLK2 | Fusion gene ID: 63754 | Hgene | Tgene | Gene symbol | PDE4D | PLK2 | Gene ID | 5144 | 10769 |
| Gene name | phosphodiesterase 4D | polo like kinase 2 | |
| Synonyms | ACRDYS2|DPDE3|HSPDE4D|PDE43|PDE4DN2|STRK1 | SNK|hPlk2|hSNK | |
| Cytomap | 5q11.2-q12.1 | 5q11.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | cAMP-specific 3',5'-cyclic phosphodiesterase 4DcAMP-specific phosphodiesterase PDE4D6phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila)testicular tissue protein Li 136 | serine/threonine-protein kinase PLK2PLK-2serine/threonine-protein kinase SNKserum-inducible kinase | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000340635, ENST00000358923, ENST00000360047, ENST00000405755, ENST00000502484, ENST00000503258, ENST00000507116, ENST00000546160, ENST00000317118, ENST00000502575, ENST00000503947, | ENST00000502671, ENST00000274289, | |
| Fusion gene scores | * DoF score | 40 X 32 X 12=15360 | 3 X 3 X 2=18 |
| # samples | 56 | 3 | |
| ** MAII score | log2(56/15360*10)=-4.77760757866355 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/18*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: PDE4D [Title/Abstract] AND PLK2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PDE4D(58334686)-PLK2(57754919), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | PDE4D-PLK2 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. PDE4D-PLK2 seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. PDE4D-PLK2 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. PDE4D-PLK2 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. PDE4D-PLK2 seems lost the major protein functional domain in Tgene partner, which is a kinase due to the frame-shifted ORF. PDE4D-PLK2 seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | PLK2 | GO:0006468 | protein phosphorylation | 20352051 |
| Tgene | PLK2 | GO:0007052 | mitotic spindle organization | 19001868 |
| Tgene | PLK2 | GO:0010508 | positive regulation of autophagy | 23983262 |
| Tgene | PLK2 | GO:0018105 | peptidyl-serine phosphorylation | 23983262 |
| Tgene | PLK2 | GO:0045732 | positive regulation of protein catabolic process | 23983262 |
| Tgene | PLK2 | GO:0046599 | regulation of centriole replication | 19001868|20352051|20531387 |
 Fusion gene breakpoints across PDE4D (5'-gene) Fusion gene breakpoints across PDE4D (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
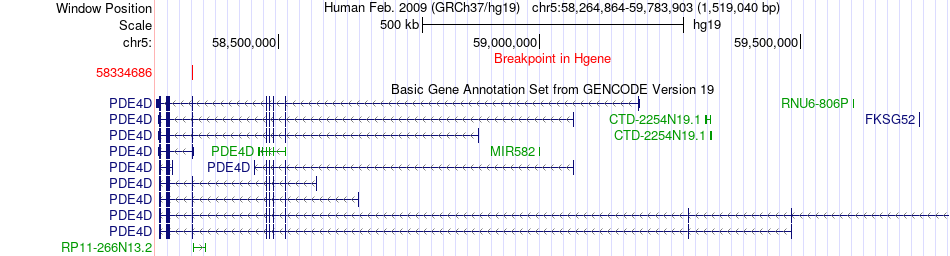 |
 Fusion gene breakpoints across PLK2 (3'-gene) Fusion gene breakpoints across PLK2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
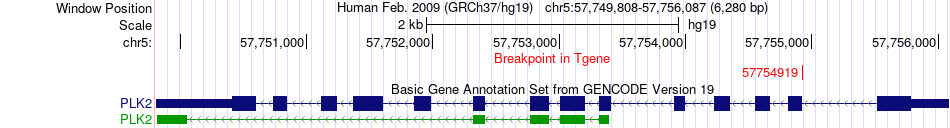 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | MESO | TCGA-TS-A7OY-01A | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
Top |
Fusion Gene ORF analysis for PDE4D-PLK2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000340635 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000358923 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000360047 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000405755 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000502484 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000503258 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000507116 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| 5CDS-intron | ENST00000546160 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| Frame-shift | ENST00000358923 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| Frame-shift | ENST00000502484 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| In-frame | ENST00000340635 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| In-frame | ENST00000360047 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| In-frame | ENST00000405755 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| In-frame | ENST00000503258 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| In-frame | ENST00000507116 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| In-frame | ENST00000546160 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| intron-3CDS | ENST00000317118 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| intron-3CDS | ENST00000502575 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| intron-3CDS | ENST00000503947 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| intron-intron | ENST00000317118 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| intron-intron | ENST00000502575 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
| intron-intron | ENST00000503947 | ENST00000502671 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000340635 | PDE4D | chr5 | 58334686 | - | ENST00000274289 | PLK2 | chr5 | 57754919 | - | 3486 | 1097 | 176 | 2884 | 902 |
| ENST00000360047 | PDE4D | chr5 | 58334686 | - | ENST00000274289 | PLK2 | chr5 | 57754919 | - | 3026 | 637 | 31 | 2424 | 797 |
| ENST00000507116 | PDE4D | chr5 | 58334686 | - | ENST00000274289 | PLK2 | chr5 | 57754919 | - | 3254 | 865 | 7 | 2652 | 881 |
| ENST00000546160 | PDE4D | chr5 | 58334686 | - | ENST00000274289 | PLK2 | chr5 | 57754919 | - | 3127 | 738 | 0 | 2525 | 841 |
| ENST00000503258 | PDE4D | chr5 | 58334686 | - | ENST00000274289 | PLK2 | chr5 | 57754919 | - | 3637 | 1248 | 717 | 3035 | 772 |
| ENST00000405755 | PDE4D | chr5 | 58334686 | - | ENST00000274289 | PLK2 | chr5 | 57754919 | - | 3073 | 684 | 102 | 2471 | 789 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000340635 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - | 0.004528954 | 0.9954711 |
| ENST00000360047 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - | 0.001142968 | 0.998857 |
| ENST00000507116 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - | 0.002743105 | 0.99725693 |
| ENST00000546160 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - | 0.001269901 | 0.9987301 |
| ENST00000503258 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - | 0.001738116 | 0.99826187 |
| ENST00000405755 | ENST00000274289 | PDE4D | chr5 | 58334686 | - | PLK2 | chr5 | 57754919 | - | 0.000960076 | 0.99903995 |
Top |
Fusion Genomic Features for PDE4D-PLK2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
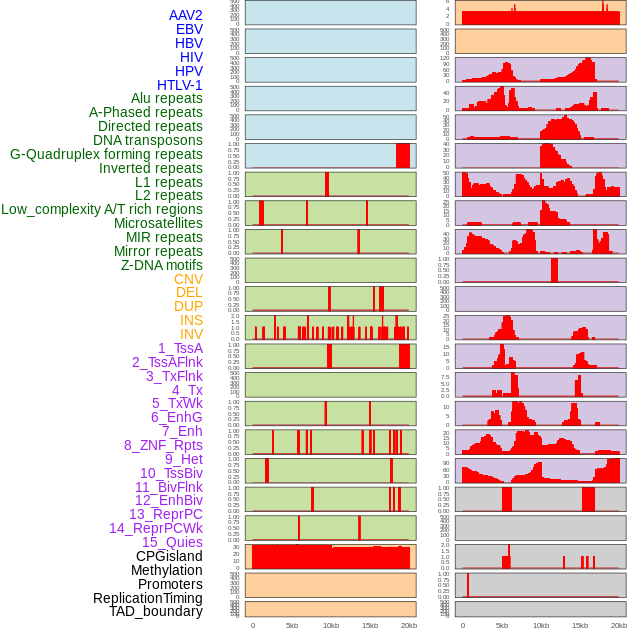 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for PDE4D-PLK2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr5:58334686/chr5:57754919) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000340635 | - | 6 | 15 | 42_88 | 307 | 810.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000360047 | - | 6 | 15 | 42_88 | 171 | 674.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000405755 | - | 6 | 15 | 42_88 | 185 | 688.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000502484 | - | 8 | 17 | 42_88 | 246 | 749.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000503258 | - | 6 | 15 | 42_88 | 177 | 680.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000507116 | - | 6 | 15 | 42_88 | 243 | 746.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000546160 | - | 7 | 16 | 42_88 | 246 | 749.0 | Compositional bias | Note=Pro-rich |
| Tgene | PLK2 | chr5:58334686 | chr5:57754919 | ENST00000274289 | 0 | 14 | 510_573 | 90 | 686.0 | Domain | POLO box 1 | |
| Tgene | PLK2 | chr5:58334686 | chr5:57754919 | ENST00000274289 | 0 | 14 | 606_677 | 90 | 686.0 | Domain | POLO box 2 | |
| Tgene | PLK2 | chr5:58334686 | chr5:57754919 | ENST00000274289 | 0 | 14 | 88_96 | 90 | 686.0 | Nucleotide binding | ATP |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000317118 | - | 1 | 10 | 42_88 | 0 | 519.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000358923 | - | 2 | 11 | 42_88 | 5 | 508.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000502575 | - | 1 | 6 | 42_88 | 0 | 220.0 | Compositional bias | Note=Pro-rich |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000317118 | - | 1 | 10 | 386_715 | 0 | 519.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000340635 | - | 6 | 15 | 386_715 | 307 | 810.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000358923 | - | 2 | 11 | 386_715 | 5 | 508.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000360047 | - | 6 | 15 | 386_715 | 171 | 674.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000405755 | - | 6 | 15 | 386_715 | 185 | 688.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000502484 | - | 8 | 17 | 386_715 | 246 | 749.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000502575 | - | 1 | 6 | 386_715 | 0 | 220.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000503258 | - | 6 | 15 | 386_715 | 177 | 680.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000507116 | - | 6 | 15 | 386_715 | 243 | 746.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000546160 | - | 7 | 16 | 386_715 | 246 | 749.0 | Domain | PDEase |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000317118 | - | 1 | 10 | 462_466 | 0 | 519.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000340635 | - | 6 | 15 | 462_466 | 307 | 810.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000358923 | - | 2 | 11 | 462_466 | 5 | 508.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000360047 | - | 6 | 15 | 462_466 | 171 | 674.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000405755 | - | 6 | 15 | 462_466 | 185 | 688.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000502484 | - | 8 | 17 | 462_466 | 246 | 749.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000502575 | - | 1 | 6 | 462_466 | 0 | 220.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000503258 | - | 6 | 15 | 462_466 | 177 | 680.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000507116 | - | 6 | 15 | 462_466 | 243 | 746.0 | Nucleotide binding | cAMP |
| Hgene | PDE4D | chr5:58334686 | chr5:57754919 | ENST00000546160 | - | 7 | 16 | 462_466 | 246 | 749.0 | Nucleotide binding | cAMP |
| Tgene | PLK2 | chr5:58334686 | chr5:57754919 | ENST00000274289 | 0 | 14 | 57_64 | 90 | 686.0 | Compositional bias | Note=Poly-His | |
| Tgene | PLK2 | chr5:58334686 | chr5:57754919 | ENST00000274289 | 0 | 14 | 82_334 | 90 | 686.0 | Domain | Protein kinase |
Top |
Fusion Gene Sequence for PDE4D-PLK2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >63754_63754_1_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000340635_PLK2_chr5_57754919_ENST00000274289_length(transcript)=3486nt_BP=1097nt GGGCCCCCTCTCGGTAGCCCTGAGGCTCTGGCGCCTTCAAGTGAGAAGCTAAGCACCAGCCTCTGCTGGGCTGCAGAAGCGGCGGCGGCG GCAGCAGCAGCAGCAGCATCAGGAAGGCGCTCGGGCCAGCGCGGTGAACCCGGGCTGGGCAGCAGGTCGCGGAGCCGCGAGCCAGGATGG AGGCAGAGGGCAGCAGCGCGCCGGCCCGGGCGGGCAGCGGAGAGGGCAGCGACAGCGCCGGCGGGGCCACGCTCAAAGCCCCCAAGCATC TCTGGAGGCACGAGCAGCACCACCAGTACCCGCTCCGGCAGCCCCAGTTCCGCCTCCTGCATCCCCATCACCACCTGCCCCCGCCGCCGC CACCCTCGCCCCAGCCCCAGCCCCAGTGTCCGCTACAGCCGCCGCCGCCGCCCCCCCTGCCGCCGCCCCCGCCGCCGCCCGGGGCTGCCC GCGGCCGCTACGCCTCGAGCGGGGCCACCGGCCGCGTCCGGCATCGCGGCTACTCGGACACCGAGCGCTACCTGTACTGTCGCGCCATGG ACCGCACCTCCTACGCGGTGGAGACCGGCCACCGGCCCGGCCTGAAGAAATCCAGGATGTCCTGGCCCTCCTCGTTCCAGGGACTCAGGC GTTTTGATGTGGACAATGGCACATCTGCGGGACGGAGTCCCTTGGATCCCATGACCAGCCCAGGATCCGGGCTAATTCTCCAAGCAAATT TTGTCCACAGTCAACGACGGGAGTCCTTCCTGTATCGATCCGACAGCGATTATGACCTCTCTCCAAAGTCTATGTCCCGGAACTCCTCCA TTGCCAGTGATATACACGGAGATGACTTGATTGTGACTCCATTTGCTCAGGTCTTGGCCAGTCTGCGAACTGTACGAAACAACTTTGCTG CATTAACTAATTTGCAAGATCGAGCACCTAGCAAAAGATCACCCATGTGCAACCAACCATCCATCAACAAAGCCACCATAACAGAGGAGG CCTACCAGAAACTGGCCAGCGAGACCCTGGAGGAGCTGGACTGGTGTCTGGACCAGCTAGAGACCCTACAGACCAGGCACTCCGTCAGTG AGATGGCCTCCAACAAGGGTGGCTTTGCAAAATGTTACGAGATGACAGATTTGACAAATAACAAAGTCTACGCCGCAAAAATTATTCCTC ACAGCAGAGTAGCTAAACCTCATCAAAGGGAAAAGATTGACAAAGAAATAGAGCTTCACAGAATTCTTCATCATAAGCATGTAGTGCAGT TTTACCACTACTTCGAGGACAAAGAAAACATTTACATTCTCTTGGAATACTGCAGTAGAAGGTCAATGGCTCATATTTTGAAAGCAAGAA AGGTGTTGACAGAGCCAGAAGTTCGATACTACCTCAGGCAGATTGTGTCTGGACTGAAATACCTTCATGAACAAGAAATCTTGCACAGAG ATCTCAAACTAGGGAACTTTTTTATTAATGAAGCCATGGAACTAAAAGTTGGGGACTTCGGTCTGGCAGCCAGGCTAGAACCCTTGGAAC ACAGAAGGAGAACGATATGTGGTACCCCAAATTATCTCTCTCCTGAAGTCCTCAACAAACAAGGACATGGCTGTGAATCAGACATTTGGG CCCTGGGCTGTGTAATGTATACAATGTTACTAGGGAGGCCCCCATTTGAAACTACAAATCTCAAAGAAACTTATAGGTGCATAAGGGAAG CAAGGTATACAATGCCGTCCTCATTGCTGGCTCCTGCCAAGCACTTAATTGCTAGTATGTTGTCCAAAAACCCAGAGGATCGTCCCAGTT TGGATGACATCATTCGACATGACTTTTTTTTGCAGGGCTTCACTCCGGACAGACTGTCTTCTAGCTGTTGTCATACAGTTCCAGATTTCC ACTTATCAAGCCCAGCTAAGAATTTCTTTAAGAAAGCAGCTGCTGCTCTTTTTGGTGGCAAAAAAGACAAAGCAAGATATATTGACACAC ATAATAGAGTGTCTAAAGAAGATGAAGACATCTACAAGCTTAGGCATGATTTGAAAAAGACTTCAATAACTCAGCAACCCAGCAAACACA GGACAGATGAGGAGCTCCAGCCACCTACCACCACAGTTGCCAGGTCTGGAACACCCGCAGTAGAAAACAAGCAGCAGATTGGGGATGCTA TTCGGATGATAGTCAGAGGGACTCTTGGCAGCTGTAGCAGCAGCAGTGAATGCCTTGAAGACAGTACCATGGGAAGTGTTGCAGACACAG TGGCAAGGGTTCTTCGGGGATGTCTGGAAAACATGCCGGAAGCTGATTGCATTCCCAAAGAGCAGCTGAGCACATCATTTCAGTGGGTCA CCAAATGGGTTGATTACTCTAACAAATATGGCTTTGGGTACCAGCTCTCAGACCACACCGTCGGTGTCCTTTTCAACAATGGTGCTCACA TGAGCCTCCTTCCAGACAAAAAAACAGTTCACTATTACGCAGAGCTTGGCCAATGCTCAGTTTTCCCAGCAACAGATGCTCCTGAGCAAT TTATTAGTCAAGTGACGGTGCTGAAATACTTTTCTCATTACATGGAGGAGAACCTCATGGATGGTGGAGATCTGCCTAGTGTTACTGATA TTCGAAGACCTCGGCTCTACCTCCTTCAGTGGCTAAAATCTGATAAGGCCCTAATGATGCTCTTTAATGATGGCACCTTTCAGGTGAATT TCTACCATGATCATACAAAAATCATCATCTGTAGCCAAAATGAAGAATACCTTCTCACCTACATCAATGAGGATAGGATATCTACAACTT TCAGGCTGACAACTCTGCTGATGTCTGGCTGTTCATCAGAATTAAAAAATCGAATGGAATATGCCCTGAACATGCTCTTACAAAGATGTA ACTGAAAGACTTTTCGAATGGACCCTATGGGACTCCTCTTTTCCACTGTGAGATCTACAGGGAAGCCAAAAGAATGATCTAGAGTATGTT GAAGAAGATGGACATGTGGTGGTACGAAAACAATTCCCCTGTGGCCTGCTGGACTGGTTGGAACCAGAACAGGCTAAGGCATACAGTTCT TGACTTTGGACAATCCAAGAGTGAACCAGAATGCAGTTTTCCTTGAGATACCTGTTTTAAAAGGTTTTTCAGACAATTTTGCAGAAAGGT GCATTGATTCTTAAATTCTCTCTGTTGAGAGCATTTCAGCCAGAGGACTTTGGAACTGTGAATATACTTCCTGAAGGGGAGGGAGAAGGG AGGAAGCTCCCATGTTGTTTAAAGGCTGTAATTGGAGCAGCTTTTGGCTGCGTAACTGTGAACTATGGCCATATATAATTTTTTTTCATT AATTTTTGAAGATACTTGTGGCTGGAAAAGTGCATTCCTTGTTAATAAACTTTTTATTTATTACAGCCCAAAGAGCAGTATTTATTATCA >63754_63754_1_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000340635_PLK2_chr5_57754919_ENST00000274289_length(amino acids)=902AA_BP=0 MEAEGSSAPARAGSGEGSDSAGGATLKAPKHLWRHEQHHQYPLRQPQFRLLHPHHHLPPPPPPSPQPQPQCPLQPPPPPPLPPPPPPPGA ARGRYASSGATGRVRHRGYSDTERYLYCRAMDRTSYAVETGHRPGLKKSRMSWPSSFQGLRRFDVDNGTSAGRSPLDPMTSPGSGLILQA NFVHSQRRESFLYRSDSDYDLSPKSMSRNSSIASDIHGDDLIVTPFAQVLASLRTVRNNFAALTNLQDRAPSKRSPMCNQPSINKATITE EAYQKLASETLEELDWCLDQLETLQTRHSVSEMASNKGGFAKCYEMTDLTNNKVYAAKIIPHSRVAKPHQREKIDKEIELHRILHHKHVV QFYHYFEDKENIYILLEYCSRRSMAHILKARKVLTEPEVRYYLRQIVSGLKYLHEQEILHRDLKLGNFFINEAMELKVGDFGLAARLEPL EHRRRTICGTPNYLSPEVLNKQGHGCESDIWALGCVMYTMLLGRPPFETTNLKETYRCIREARYTMPSSLLAPAKHLIASMLSKNPEDRP SLDDIIRHDFFLQGFTPDRLSSSCCHTVPDFHLSSPAKNFFKKAAAALFGGKKDKARYIDTHNRVSKEDEDIYKLRHDLKKTSITQQPSK HRTDEELQPPTTTVARSGTPAVENKQQIGDAIRMIVRGTLGSCSSSSECLEDSTMGSVADTVARVLRGCLENMPEADCIPKEQLSTSFQW VTKWVDYSNKYGFGYQLSDHTVGVLFNNGAHMSLLPDKKTVHYYAELGQCSVFPATDAPEQFISQVTVLKYFSHYMEENLMDGGDLPSVT DIRRPRLYLLQWLKSDKALMMLFNDGTFQVNFYHDHTKIIICSQNEEYLLTYINEDRISTTFRLTTLLMSGCSSELKNRMEYALNMLLQR -------------------------------------------------------------- >63754_63754_2_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000360047_PLK2_chr5_57754919_ENST00000274289_length(transcript)=3026nt_BP=637nt CAATACTTGTTGCAATAATTGCCCACGATAGCTGCTCAAACAAGAGAGTTGGAATTCATCTGTAAAAATCACTACATGTAACGTAGGAGA CAAGAAAAATATTAATGACAGAAGATCTGCGAACATGATGCACGTGAATAATTTTCCCTTTAGAAGGCATTCCTGGATATGTTTTGATGT GGACAATGGCACATCTGCGGGACGGAGTCCCTTGGATCCCATGACCAGCCCAGGATCCGGGCTAATTCTCCAAGCAAATTTTGTCCACAG TCAACGACGGGAGTCCTTCCTGTATCGATCCGACAGCGATTATGACCTCTCTCCAAAGTCTATGTCCCGGAACTCCTCCATTGCCAGTGA TATACACGGAGATGACTTGATTGTGACTCCATTTGCTCAGGTCTTGGCCAGTCTGCGAACTGTACGAAACAACTTTGCTGCATTAACTAA TTTGCAAGATCGAGCACCTAGCAAAAGATCACCCATGTGCAACCAACCATCCATCAACAAAGCCACCATAACAGAGGAGGCCTACCAGAA ACTGGCCAGCGAGACCCTGGAGGAGCTGGACTGGTGTCTGGACCAGCTAGAGACCCTACAGACCAGGCACTCCGTCAGTGAGATGGCCTC CAACAAGGGTGGCTTTGCAAAATGTTACGAGATGACAGATTTGACAAATAACAAAGTCTACGCCGCAAAAATTATTCCTCACAGCAGAGT AGCTAAACCTCATCAAAGGGAAAAGATTGACAAAGAAATAGAGCTTCACAGAATTCTTCATCATAAGCATGTAGTGCAGTTTTACCACTA CTTCGAGGACAAAGAAAACATTTACATTCTCTTGGAATACTGCAGTAGAAGGTCAATGGCTCATATTTTGAAAGCAAGAAAGGTGTTGAC AGAGCCAGAAGTTCGATACTACCTCAGGCAGATTGTGTCTGGACTGAAATACCTTCATGAACAAGAAATCTTGCACAGAGATCTCAAACT AGGGAACTTTTTTATTAATGAAGCCATGGAACTAAAAGTTGGGGACTTCGGTCTGGCAGCCAGGCTAGAACCCTTGGAACACAGAAGGAG AACGATATGTGGTACCCCAAATTATCTCTCTCCTGAAGTCCTCAACAAACAAGGACATGGCTGTGAATCAGACATTTGGGCCCTGGGCTG TGTAATGTATACAATGTTACTAGGGAGGCCCCCATTTGAAACTACAAATCTCAAAGAAACTTATAGGTGCATAAGGGAAGCAAGGTATAC AATGCCGTCCTCATTGCTGGCTCCTGCCAAGCACTTAATTGCTAGTATGTTGTCCAAAAACCCAGAGGATCGTCCCAGTTTGGATGACAT CATTCGACATGACTTTTTTTTGCAGGGCTTCACTCCGGACAGACTGTCTTCTAGCTGTTGTCATACAGTTCCAGATTTCCACTTATCAAG CCCAGCTAAGAATTTCTTTAAGAAAGCAGCTGCTGCTCTTTTTGGTGGCAAAAAAGACAAAGCAAGATATATTGACACACATAATAGAGT GTCTAAAGAAGATGAAGACATCTACAAGCTTAGGCATGATTTGAAAAAGACTTCAATAACTCAGCAACCCAGCAAACACAGGACAGATGA GGAGCTCCAGCCACCTACCACCACAGTTGCCAGGTCTGGAACACCCGCAGTAGAAAACAAGCAGCAGATTGGGGATGCTATTCGGATGAT AGTCAGAGGGACTCTTGGCAGCTGTAGCAGCAGCAGTGAATGCCTTGAAGACAGTACCATGGGAAGTGTTGCAGACACAGTGGCAAGGGT TCTTCGGGGATGTCTGGAAAACATGCCGGAAGCTGATTGCATTCCCAAAGAGCAGCTGAGCACATCATTTCAGTGGGTCACCAAATGGGT TGATTACTCTAACAAATATGGCTTTGGGTACCAGCTCTCAGACCACACCGTCGGTGTCCTTTTCAACAATGGTGCTCACATGAGCCTCCT TCCAGACAAAAAAACAGTTCACTATTACGCAGAGCTTGGCCAATGCTCAGTTTTCCCAGCAACAGATGCTCCTGAGCAATTTATTAGTCA AGTGACGGTGCTGAAATACTTTTCTCATTACATGGAGGAGAACCTCATGGATGGTGGAGATCTGCCTAGTGTTACTGATATTCGAAGACC TCGGCTCTACCTCCTTCAGTGGCTAAAATCTGATAAGGCCCTAATGATGCTCTTTAATGATGGCACCTTTCAGGTGAATTTCTACCATGA TCATACAAAAATCATCATCTGTAGCCAAAATGAAGAATACCTTCTCACCTACATCAATGAGGATAGGATATCTACAACTTTCAGGCTGAC AACTCTGCTGATGTCTGGCTGTTCATCAGAATTAAAAAATCGAATGGAATATGCCCTGAACATGCTCTTACAAAGATGTAACTGAAAGAC TTTTCGAATGGACCCTATGGGACTCCTCTTTTCCACTGTGAGATCTACAGGGAAGCCAAAAGAATGATCTAGAGTATGTTGAAGAAGATG GACATGTGGTGGTACGAAAACAATTCCCCTGTGGCCTGCTGGACTGGTTGGAACCAGAACAGGCTAAGGCATACAGTTCTTGACTTTGGA CAATCCAAGAGTGAACCAGAATGCAGTTTTCCTTGAGATACCTGTTTTAAAAGGTTTTTCAGACAATTTTGCAGAAAGGTGCATTGATTC TTAAATTCTCTCTGTTGAGAGCATTTCAGCCAGAGGACTTTGGAACTGTGAATATACTTCCTGAAGGGGAGGGAGAAGGGAGGAAGCTCC CATGTTGTTTAAAGGCTGTAATTGGAGCAGCTTTTGGCTGCGTAACTGTGAACTATGGCCATATATAATTTTTTTTCATTAATTTTTGAA GATACTTGTGGCTGGAAAAGTGCATTCCTTGTTAATAAACTTTTTATTTATTACAGCCCAAAGAGCAGTATTTATTATCAAAATGTCTTT >63754_63754_2_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000360047_PLK2_chr5_57754919_ENST00000274289_length(amino acids)=797AA_BP=1 MLKQESWNSSVKITTCNVGDKKNINDRRSANMMHVNNFPFRRHSWICFDVDNGTSAGRSPLDPMTSPGSGLILQANFVHSQRRESFLYRS DSDYDLSPKSMSRNSSIASDIHGDDLIVTPFAQVLASLRTVRNNFAALTNLQDRAPSKRSPMCNQPSINKATITEEAYQKLASETLEELD WCLDQLETLQTRHSVSEMASNKGGFAKCYEMTDLTNNKVYAAKIIPHSRVAKPHQREKIDKEIELHRILHHKHVVQFYHYFEDKENIYIL LEYCSRRSMAHILKARKVLTEPEVRYYLRQIVSGLKYLHEQEILHRDLKLGNFFINEAMELKVGDFGLAARLEPLEHRRRTICGTPNYLS PEVLNKQGHGCESDIWALGCVMYTMLLGRPPFETTNLKETYRCIREARYTMPSSLLAPAKHLIASMLSKNPEDRPSLDDIIRHDFFLQGF TPDRLSSSCCHTVPDFHLSSPAKNFFKKAAAALFGGKKDKARYIDTHNRVSKEDEDIYKLRHDLKKTSITQQPSKHRTDEELQPPTTTVA RSGTPAVENKQQIGDAIRMIVRGTLGSCSSSSECLEDSTMGSVADTVARVLRGCLENMPEADCIPKEQLSTSFQWVTKWVDYSNKYGFGY QLSDHTVGVLFNNGAHMSLLPDKKTVHYYAELGQCSVFPATDAPEQFISQVTVLKYFSHYMEENLMDGGDLPSVTDIRRPRLYLLQWLKS -------------------------------------------------------------- >63754_63754_3_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000405755_PLK2_chr5_57754919_ENST00000274289_length(transcript)=3073nt_BP=684nt ACTGCACAAAGGAAATGCCCTCACTCAGCTACAAGTGCCTCCTGCTCCGTGCGGGGCCTCAGGGCCCCAGAGCCCCGCACCAGCTCTGAC TTCTCGTGGTTCCTGGGGATCTTGGCATCGTCCTTAAAAATGGCTTTTGTTTGGGATCCTCTGGGAGCCACGGTGCCAGGACCATCTACA AGAGCCAAATCAAGATTGCGTTTCTCAAAGTCCTACAGTTTTGATGTGGACAATGGCACATCTGCGGGACGGAGTCCCTTGGATCCCATG ACCAGCCCAGGATCCGGGCTAATTCTCCAAGCAAATTTTGTCCACAGTCAACGACGGGAGTCCTTCCTGTATCGATCCGACAGCGATTAT GACCTCTCTCCAAAGTCTATGTCCCGGAACTCCTCCATTGCCAGTGATATACACGGAGATGACTTGATTGTGACTCCATTTGCTCAGGTC TTGGCCAGTCTGCGAACTGTACGAAACAACTTTGCTGCATTAACTAATTTGCAAGATCGAGCACCTAGCAAAAGATCACCCATGTGCAAC CAACCATCCATCAACAAAGCCACCATAACAGAGGAGGCCTACCAGAAACTGGCCAGCGAGACCCTGGAGGAGCTGGACTGGTGTCTGGAC CAGCTAGAGACCCTACAGACCAGGCACTCCGTCAGTGAGATGGCCTCCAACAAGGGTGGCTTTGCAAAATGTTACGAGATGACAGATTTG ACAAATAACAAAGTCTACGCCGCAAAAATTATTCCTCACAGCAGAGTAGCTAAACCTCATCAAAGGGAAAAGATTGACAAAGAAATAGAG CTTCACAGAATTCTTCATCATAAGCATGTAGTGCAGTTTTACCACTACTTCGAGGACAAAGAAAACATTTACATTCTCTTGGAATACTGC AGTAGAAGGTCAATGGCTCATATTTTGAAAGCAAGAAAGGTGTTGACAGAGCCAGAAGTTCGATACTACCTCAGGCAGATTGTGTCTGGA CTGAAATACCTTCATGAACAAGAAATCTTGCACAGAGATCTCAAACTAGGGAACTTTTTTATTAATGAAGCCATGGAACTAAAAGTTGGG GACTTCGGTCTGGCAGCCAGGCTAGAACCCTTGGAACACAGAAGGAGAACGATATGTGGTACCCCAAATTATCTCTCTCCTGAAGTCCTC AACAAACAAGGACATGGCTGTGAATCAGACATTTGGGCCCTGGGCTGTGTAATGTATACAATGTTACTAGGGAGGCCCCCATTTGAAACT ACAAATCTCAAAGAAACTTATAGGTGCATAAGGGAAGCAAGGTATACAATGCCGTCCTCATTGCTGGCTCCTGCCAAGCACTTAATTGCT AGTATGTTGTCCAAAAACCCAGAGGATCGTCCCAGTTTGGATGACATCATTCGACATGACTTTTTTTTGCAGGGCTTCACTCCGGACAGA CTGTCTTCTAGCTGTTGTCATACAGTTCCAGATTTCCACTTATCAAGCCCAGCTAAGAATTTCTTTAAGAAAGCAGCTGCTGCTCTTTTT GGTGGCAAAAAAGACAAAGCAAGATATATTGACACACATAATAGAGTGTCTAAAGAAGATGAAGACATCTACAAGCTTAGGCATGATTTG AAAAAGACTTCAATAACTCAGCAACCCAGCAAACACAGGACAGATGAGGAGCTCCAGCCACCTACCACCACAGTTGCCAGGTCTGGAACA CCCGCAGTAGAAAACAAGCAGCAGATTGGGGATGCTATTCGGATGATAGTCAGAGGGACTCTTGGCAGCTGTAGCAGCAGCAGTGAATGC CTTGAAGACAGTACCATGGGAAGTGTTGCAGACACAGTGGCAAGGGTTCTTCGGGGATGTCTGGAAAACATGCCGGAAGCTGATTGCATT CCCAAAGAGCAGCTGAGCACATCATTTCAGTGGGTCACCAAATGGGTTGATTACTCTAACAAATATGGCTTTGGGTACCAGCTCTCAGAC CACACCGTCGGTGTCCTTTTCAACAATGGTGCTCACATGAGCCTCCTTCCAGACAAAAAAACAGTTCACTATTACGCAGAGCTTGGCCAA TGCTCAGTTTTCCCAGCAACAGATGCTCCTGAGCAATTTATTAGTCAAGTGACGGTGCTGAAATACTTTTCTCATTACATGGAGGAGAAC CTCATGGATGGTGGAGATCTGCCTAGTGTTACTGATATTCGAAGACCTCGGCTCTACCTCCTTCAGTGGCTAAAATCTGATAAGGCCCTA ATGATGCTCTTTAATGATGGCACCTTTCAGGTGAATTTCTACCATGATCATACAAAAATCATCATCTGTAGCCAAAATGAAGAATACCTT CTCACCTACATCAATGAGGATAGGATATCTACAACTTTCAGGCTGACAACTCTGCTGATGTCTGGCTGTTCATCAGAATTAAAAAATCGA ATGGAATATGCCCTGAACATGCTCTTACAAAGATGTAACTGAAAGACTTTTCGAATGGACCCTATGGGACTCCTCTTTTCCACTGTGAGA TCTACAGGGAAGCCAAAAGAATGATCTAGAGTATGTTGAAGAAGATGGACATGTGGTGGTACGAAAACAATTCCCCTGTGGCCTGCTGGA CTGGTTGGAACCAGAACAGGCTAAGGCATACAGTTCTTGACTTTGGACAATCCAAGAGTGAACCAGAATGCAGTTTTCCTTGAGATACCT GTTTTAAAAGGTTTTTCAGACAATTTTGCAGAAAGGTGCATTGATTCTTAAATTCTCTCTGTTGAGAGCATTTCAGCCAGAGGACTTTGG AACTGTGAATATACTTCCTGAAGGGGAGGGAGAAGGGAGGAAGCTCCCATGTTGTTTAAAGGCTGTAATTGGAGCAGCTTTTGGCTGCGT AACTGTGAACTATGGCCATATATAATTTTTTTTCATTAATTTTTGAAGATACTTGTGGCTGGAAAAGTGCATTCCTTGTTAATAAACTTT TTATTTATTACAGCCCAAAGAGCAGTATTTATTATCAAAATGTCTTTTTTTTTATGTTGACCATTTTAAACCGTTGGCAATAAAGAGTAT >63754_63754_3_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000405755_PLK2_chr5_57754919_ENST00000274289_length(amino acids)=789AA_BP=1 MGILASSLKMAFVWDPLGATVPGPSTRAKSRLRFSKSYSFDVDNGTSAGRSPLDPMTSPGSGLILQANFVHSQRRESFLYRSDSDYDLSP KSMSRNSSIASDIHGDDLIVTPFAQVLASLRTVRNNFAALTNLQDRAPSKRSPMCNQPSINKATITEEAYQKLASETLEELDWCLDQLET LQTRHSVSEMASNKGGFAKCYEMTDLTNNKVYAAKIIPHSRVAKPHQREKIDKEIELHRILHHKHVVQFYHYFEDKENIYILLEYCSRRS MAHILKARKVLTEPEVRYYLRQIVSGLKYLHEQEILHRDLKLGNFFINEAMELKVGDFGLAARLEPLEHRRRTICGTPNYLSPEVLNKQG HGCESDIWALGCVMYTMLLGRPPFETTNLKETYRCIREARYTMPSSLLAPAKHLIASMLSKNPEDRPSLDDIIRHDFFLQGFTPDRLSSS CCHTVPDFHLSSPAKNFFKKAAAALFGGKKDKARYIDTHNRVSKEDEDIYKLRHDLKKTSITQQPSKHRTDEELQPPTTTVARSGTPAVE NKQQIGDAIRMIVRGTLGSCSSSSECLEDSTMGSVADTVARVLRGCLENMPEADCIPKEQLSTSFQWVTKWVDYSNKYGFGYQLSDHTVG VLFNNGAHMSLLPDKKTVHYYAELGQCSVFPATDAPEQFISQVTVLKYFSHYMEENLMDGGDLPSVTDIRRPRLYLLQWLKSDKALMMLF -------------------------------------------------------------- >63754_63754_4_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000503258_PLK2_chr5_57754919_ENST00000274289_length(transcript)=3637nt_BP=1248nt TTCTCACTGCCCTGCGGTGTTTTGAACTGCCTTCTTACAGACGTCATACAGCCCTTGAGGAATAGTTTCTGCCTGGTGAGATTGAATGAT AGTTCTCATTCACAAAACCCTGGATTCTAAGCAGGGACACACAGAAATTACTTTCGCAGGTAAATCAGCCCACCCAGCCAAAGTGTGGAG AGATTTGTTCCTTGGCTGACTTCTTTGCTCCACGGAGAGGAGTGTTTTCCTGTGCTTGCCCTGAAATGGAACTTCCTTGACAGCTCTCCC GTGTTACAGTACCTCCCGGTCATTTTCTTTTTCTCTCTCTCTACCTGCGCTCTTCGAGTGTCAGAAACCTTTAAAGCTGTTACTATGGAA TTGCAAAAAAGAGATCAAGTGACTCTTTCACTATGCTGGTTTCCCTTGTGACCCAGATGAAGAATCAATTCAGAATTCAGTTCCTCCCTT GGCATTGCAAGACACAGAAGAAACTGTCACTTCCTAACAGCCTAGTACTGGAGTAAATTCAGTATGAAGGAAGAAAGCGCTCCTGCGTGT TAGAACCTTGCCCATGAGCTGGACCGAGGACAGGAGATGGACTCCAGGAAAATTGGATTTCTTCAAGCAGCCTCCCTTGGAAATGGAATA TCTTTAAAATCTTCTTTGCAGAAAGACAGTTAGAATGTATTAATCAGAATAGTTGAAGACTTATTTTCCTTTTTATTTTTTTTCAAAATG AGCATTATTATGAAGCCAAGATCCCGATCTACAAGTTCCCTAAGGACTGCAGAGGCAGTTTGTTTTGATGTGGACAATGGCACATCTGCG GGACGGAGTCCCTTGGATCCCATGACCAGCCCAGGATCCGGGCTAATTCTCCAAGCAAATTTTGTCCACAGTCAACGACGGGAGTCCTTC CTGTATCGATCCGACAGCGATTATGACCTCTCTCCAAAGTCTATGTCCCGGAACTCCTCCATTGCCAGTGATATACACGGAGATGACTTG ATTGTGACTCCATTTGCTCAGGTCTTGGCCAGTCTGCGAACTGTACGAAACAACTTTGCTGCATTAACTAATTTGCAAGATCGAGCACCT AGCAAAAGATCACCCATGTGCAACCAACCATCCATCAACAAAGCCACCATAACAGAGGAGGCCTACCAGAAACTGGCCAGCGAGACCCTG GAGGAGCTGGACTGGTGTCTGGACCAGCTAGAGACCCTACAGACCAGGCACTCCGTCAGTGAGATGGCCTCCAACAAGGGTGGCTTTGCA AAATGTTACGAGATGACAGATTTGACAAATAACAAAGTCTACGCCGCAAAAATTATTCCTCACAGCAGAGTAGCTAAACCTCATCAAAGG GAAAAGATTGACAAAGAAATAGAGCTTCACAGAATTCTTCATCATAAGCATGTAGTGCAGTTTTACCACTACTTCGAGGACAAAGAAAAC ATTTACATTCTCTTGGAATACTGCAGTAGAAGGTCAATGGCTCATATTTTGAAAGCAAGAAAGGTGTTGACAGAGCCAGAAGTTCGATAC TACCTCAGGCAGATTGTGTCTGGACTGAAATACCTTCATGAACAAGAAATCTTGCACAGAGATCTCAAACTAGGGAACTTTTTTATTAAT GAAGCCATGGAACTAAAAGTTGGGGACTTCGGTCTGGCAGCCAGGCTAGAACCCTTGGAACACAGAAGGAGAACGATATGTGGTACCCCA AATTATCTCTCTCCTGAAGTCCTCAACAAACAAGGACATGGCTGTGAATCAGACATTTGGGCCCTGGGCTGTGTAATGTATACAATGTTA CTAGGGAGGCCCCCATTTGAAACTACAAATCTCAAAGAAACTTATAGGTGCATAAGGGAAGCAAGGTATACAATGCCGTCCTCATTGCTG GCTCCTGCCAAGCACTTAATTGCTAGTATGTTGTCCAAAAACCCAGAGGATCGTCCCAGTTTGGATGACATCATTCGACATGACTTTTTT TTGCAGGGCTTCACTCCGGACAGACTGTCTTCTAGCTGTTGTCATACAGTTCCAGATTTCCACTTATCAAGCCCAGCTAAGAATTTCTTT AAGAAAGCAGCTGCTGCTCTTTTTGGTGGCAAAAAAGACAAAGCAAGATATATTGACACACATAATAGAGTGTCTAAAGAAGATGAAGAC ATCTACAAGCTTAGGCATGATTTGAAAAAGACTTCAATAACTCAGCAACCCAGCAAACACAGGACAGATGAGGAGCTCCAGCCACCTACC ACCACAGTTGCCAGGTCTGGAACACCCGCAGTAGAAAACAAGCAGCAGATTGGGGATGCTATTCGGATGATAGTCAGAGGGACTCTTGGC AGCTGTAGCAGCAGCAGTGAATGCCTTGAAGACAGTACCATGGGAAGTGTTGCAGACACAGTGGCAAGGGTTCTTCGGGGATGTCTGGAA AACATGCCGGAAGCTGATTGCATTCCCAAAGAGCAGCTGAGCACATCATTTCAGTGGGTCACCAAATGGGTTGATTACTCTAACAAATAT GGCTTTGGGTACCAGCTCTCAGACCACACCGTCGGTGTCCTTTTCAACAATGGTGCTCACATGAGCCTCCTTCCAGACAAAAAAACAGTT CACTATTACGCAGAGCTTGGCCAATGCTCAGTTTTCCCAGCAACAGATGCTCCTGAGCAATTTATTAGTCAAGTGACGGTGCTGAAATAC TTTTCTCATTACATGGAGGAGAACCTCATGGATGGTGGAGATCTGCCTAGTGTTACTGATATTCGAAGACCTCGGCTCTACCTCCTTCAG TGGCTAAAATCTGATAAGGCCCTAATGATGCTCTTTAATGATGGCACCTTTCAGGTGAATTTCTACCATGATCATACAAAAATCATCATC TGTAGCCAAAATGAAGAATACCTTCTCACCTACATCAATGAGGATAGGATATCTACAACTTTCAGGCTGACAACTCTGCTGATGTCTGGC TGTTCATCAGAATTAAAAAATCGAATGGAATATGCCCTGAACATGCTCTTACAAAGATGTAACTGAAAGACTTTTCGAATGGACCCTATG GGACTCCTCTTTTCCACTGTGAGATCTACAGGGAAGCCAAAAGAATGATCTAGAGTATGTTGAAGAAGATGGACATGTGGTGGTACGAAA ACAATTCCCCTGTGGCCTGCTGGACTGGTTGGAACCAGAACAGGCTAAGGCATACAGTTCTTGACTTTGGACAATCCAAGAGTGAACCAG AATGCAGTTTTCCTTGAGATACCTGTTTTAAAAGGTTTTTCAGACAATTTTGCAGAAAGGTGCATTGATTCTTAAATTCTCTCTGTTGAG AGCATTTCAGCCAGAGGACTTTGGAACTGTGAATATACTTCCTGAAGGGGAGGGAGAAGGGAGGAAGCTCCCATGTTGTTTAAAGGCTGT AATTGGAGCAGCTTTTGGCTGCGTAACTGTGAACTATGGCCATATATAATTTTTTTTCATTAATTTTTGAAGATACTTGTGGCTGGAAAA GTGCATTCCTTGTTAATAAACTTTTTATTTATTACAGCCCAAAGAGCAGTATTTATTATCAAAATGTCTTTTTTTTTATGTTGACCATTT >63754_63754_4_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000503258_PLK2_chr5_57754919_ENST00000274289_length(amino acids)=772AA_BP=0 MSIIMKPRSRSTSSLRTAEAVCFDVDNGTSAGRSPLDPMTSPGSGLILQANFVHSQRRESFLYRSDSDYDLSPKSMSRNSSIASDIHGDD LIVTPFAQVLASLRTVRNNFAALTNLQDRAPSKRSPMCNQPSINKATITEEAYQKLASETLEELDWCLDQLETLQTRHSVSEMASNKGGF AKCYEMTDLTNNKVYAAKIIPHSRVAKPHQREKIDKEIELHRILHHKHVVQFYHYFEDKENIYILLEYCSRRSMAHILKARKVLTEPEVR YYLRQIVSGLKYLHEQEILHRDLKLGNFFINEAMELKVGDFGLAARLEPLEHRRRTICGTPNYLSPEVLNKQGHGCESDIWALGCVMYTM LLGRPPFETTNLKETYRCIREARYTMPSSLLAPAKHLIASMLSKNPEDRPSLDDIIRHDFFLQGFTPDRLSSSCCHTVPDFHLSSPAKNF FKKAAAALFGGKKDKARYIDTHNRVSKEDEDIYKLRHDLKKTSITQQPSKHRTDEELQPPTTTVARSGTPAVENKQQIGDAIRMIVRGTL GSCSSSSECLEDSTMGSVADTVARVLRGCLENMPEADCIPKEQLSTSFQWVTKWVDYSNKYGFGYQLSDHTVGVLFNNGAHMSLLPDKKT VHYYAELGQCSVFPATDAPEQFISQVTVLKYFSHYMEENLMDGGDLPSVTDIRRPRLYLLQWLKSDKALMMLFNDGTFQVNFYHDHTKII -------------------------------------------------------------- >63754_63754_5_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000507116_PLK2_chr5_57754919_ENST00000274289_length(transcript)=3254nt_BP=865nt AGAGAGACTGAGGCAGGCGACTGAATGCACTAACAGCAGCAGGCTCAGACCTGCTTCCCTGGACATTTCCGGGACCGTGAGCGAGGGAAC CACGTTGCCCTGGATTCTTGCCAGCTGTACAAAGTTGACCAGGAAAATGGCTCAGCAGACAAGCCCGGACACTTTAACAGTACCTGAAGT GGATAATCCGCATTGTCCAAACCCGTGGCTGAACGAAGACCTTGTGAAATCCTTGCGAGAAAACCTGTTGCAGCATGAGAAGTCCAAGAC AGCGAGGAAATCGGTTTCTCCCAAGCTCTCTCCAGTGATCTCTCCGAGAAATTCCCCCAGGCTTCTGCGCAGAATGCTTCTCAGCAGCAA CATCCCCAAACAGCGGCGTTTCACGGTGGCACATACATGTTTTGATGTGGACAATGGCACATCTGCGGGACGGAGTCCCTTGGATCCCAT GACCAGCCCAGGATCCGGGCTAATTCTCCAAGCAAATTTTGTCCACAGTCAACGACGGGAGTCCTTCCTGTATCGATCCGACAGCGATTA TGACCTCTCTCCAAAGTCTATGTCCCGGAACTCCTCCATTGCCAGTGATATACACGGAGATGACTTGATTGTGACTCCATTTGCTCAGGT CTTGGCCAGTCTGCGAACTGTACGAAACAACTTTGCTGCATTAACTAATTTGCAAGATCGAGCACCTAGCAAAAGATCACCCATGTGCAA CCAACCATCCATCAACAAAGCCACCATAACAGAGGAGGCCTACCAGAAACTGGCCAGCGAGACCCTGGAGGAGCTGGACTGGTGTCTGGA CCAGCTAGAGACCCTACAGACCAGGCACTCCGTCAGTGAGATGGCCTCCAACAAGGGTGGCTTTGCAAAATGTTACGAGATGACAGATTT GACAAATAACAAAGTCTACGCCGCAAAAATTATTCCTCACAGCAGAGTAGCTAAACCTCATCAAAGGGAAAAGATTGACAAAGAAATAGA GCTTCACAGAATTCTTCATCATAAGCATGTAGTGCAGTTTTACCACTACTTCGAGGACAAAGAAAACATTTACATTCTCTTGGAATACTG CAGTAGAAGGTCAATGGCTCATATTTTGAAAGCAAGAAAGGTGTTGACAGAGCCAGAAGTTCGATACTACCTCAGGCAGATTGTGTCTGG ACTGAAATACCTTCATGAACAAGAAATCTTGCACAGAGATCTCAAACTAGGGAACTTTTTTATTAATGAAGCCATGGAACTAAAAGTTGG GGACTTCGGTCTGGCAGCCAGGCTAGAACCCTTGGAACACAGAAGGAGAACGATATGTGGTACCCCAAATTATCTCTCTCCTGAAGTCCT CAACAAACAAGGACATGGCTGTGAATCAGACATTTGGGCCCTGGGCTGTGTAATGTATACAATGTTACTAGGGAGGCCCCCATTTGAAAC TACAAATCTCAAAGAAACTTATAGGTGCATAAGGGAAGCAAGGTATACAATGCCGTCCTCATTGCTGGCTCCTGCCAAGCACTTAATTGC TAGTATGTTGTCCAAAAACCCAGAGGATCGTCCCAGTTTGGATGACATCATTCGACATGACTTTTTTTTGCAGGGCTTCACTCCGGACAG ACTGTCTTCTAGCTGTTGTCATACAGTTCCAGATTTCCACTTATCAAGCCCAGCTAAGAATTTCTTTAAGAAAGCAGCTGCTGCTCTTTT TGGTGGCAAAAAAGACAAAGCAAGATATATTGACACACATAATAGAGTGTCTAAAGAAGATGAAGACATCTACAAGCTTAGGCATGATTT GAAAAAGACTTCAATAACTCAGCAACCCAGCAAACACAGGACAGATGAGGAGCTCCAGCCACCTACCACCACAGTTGCCAGGTCTGGAAC ACCCGCAGTAGAAAACAAGCAGCAGATTGGGGATGCTATTCGGATGATAGTCAGAGGGACTCTTGGCAGCTGTAGCAGCAGCAGTGAATG CCTTGAAGACAGTACCATGGGAAGTGTTGCAGACACAGTGGCAAGGGTTCTTCGGGGATGTCTGGAAAACATGCCGGAAGCTGATTGCAT TCCCAAAGAGCAGCTGAGCACATCATTTCAGTGGGTCACCAAATGGGTTGATTACTCTAACAAATATGGCTTTGGGTACCAGCTCTCAGA CCACACCGTCGGTGTCCTTTTCAACAATGGTGCTCACATGAGCCTCCTTCCAGACAAAAAAACAGTTCACTATTACGCAGAGCTTGGCCA ATGCTCAGTTTTCCCAGCAACAGATGCTCCTGAGCAATTTATTAGTCAAGTGACGGTGCTGAAATACTTTTCTCATTACATGGAGGAGAA CCTCATGGATGGTGGAGATCTGCCTAGTGTTACTGATATTCGAAGACCTCGGCTCTACCTCCTTCAGTGGCTAAAATCTGATAAGGCCCT AATGATGCTCTTTAATGATGGCACCTTTCAGGTGAATTTCTACCATGATCATACAAAAATCATCATCTGTAGCCAAAATGAAGAATACCT TCTCACCTACATCAATGAGGATAGGATATCTACAACTTTCAGGCTGACAACTCTGCTGATGTCTGGCTGTTCATCAGAATTAAAAAATCG AATGGAATATGCCCTGAACATGCTCTTACAAAGATGTAACTGAAAGACTTTTCGAATGGACCCTATGGGACTCCTCTTTTCCACTGTGAG ATCTACAGGGAAGCCAAAAGAATGATCTAGAGTATGTTGAAGAAGATGGACATGTGGTGGTACGAAAACAATTCCCCTGTGGCCTGCTGG ACTGGTTGGAACCAGAACAGGCTAAGGCATACAGTTCTTGACTTTGGACAATCCAAGAGTGAACCAGAATGCAGTTTTCCTTGAGATACC TGTTTTAAAAGGTTTTTCAGACAATTTTGCAGAAAGGTGCATTGATTCTTAAATTCTCTCTGTTGAGAGCATTTCAGCCAGAGGACTTTG GAACTGTGAATATACTTCCTGAAGGGGAGGGAGAAGGGAGGAAGCTCCCATGTTGTTTAAAGGCTGTAATTGGAGCAGCTTTTGGCTGCG TAACTGTGAACTATGGCCATATATAATTTTTTTTCATTAATTTTTGAAGATACTTGTGGCTGGAAAAGTGCATTCCTTGTTAATAAACTT TTTATTTATTACAGCCCAAAGAGCAGTATTTATTATCAAAATGTCTTTTTTTTTATGTTGACCATTTTAAACCGTTGGCAATAAAGAGTA >63754_63754_5_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000507116_PLK2_chr5_57754919_ENST00000274289_length(amino acids)=881AA_BP=1 MRQATECTNSSRLRPASLDISGTVSEGTTLPWILASCTKLTRKMAQQTSPDTLTVPEVDNPHCPNPWLNEDLVKSLRENLLQHEKSKTAR KSVSPKLSPVISPRNSPRLLRRMLLSSNIPKQRRFTVAHTCFDVDNGTSAGRSPLDPMTSPGSGLILQANFVHSQRRESFLYRSDSDYDL SPKSMSRNSSIASDIHGDDLIVTPFAQVLASLRTVRNNFAALTNLQDRAPSKRSPMCNQPSINKATITEEAYQKLASETLEELDWCLDQL ETLQTRHSVSEMASNKGGFAKCYEMTDLTNNKVYAAKIIPHSRVAKPHQREKIDKEIELHRILHHKHVVQFYHYFEDKENIYILLEYCSR RSMAHILKARKVLTEPEVRYYLRQIVSGLKYLHEQEILHRDLKLGNFFINEAMELKVGDFGLAARLEPLEHRRRTICGTPNYLSPEVLNK QGHGCESDIWALGCVMYTMLLGRPPFETTNLKETYRCIREARYTMPSSLLAPAKHLIASMLSKNPEDRPSLDDIIRHDFFLQGFTPDRLS SSCCHTVPDFHLSSPAKNFFKKAAAALFGGKKDKARYIDTHNRVSKEDEDIYKLRHDLKKTSITQQPSKHRTDEELQPPTTTVARSGTPA VENKQQIGDAIRMIVRGTLGSCSSSSECLEDSTMGSVADTVARVLRGCLENMPEADCIPKEQLSTSFQWVTKWVDYSNKYGFGYQLSDHT VGVLFNNGAHMSLLPDKKTVHYYAELGQCSVFPATDAPEQFISQVTVLKYFSHYMEENLMDGGDLPSVTDIRRPRLYLLQWLKSDKALMM -------------------------------------------------------------- >63754_63754_6_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000546160_PLK2_chr5_57754919_ENST00000274289_length(transcript)=3127nt_BP=738nt ATGAAAAGAAATACCTGTGATTTGCTTTCTCGGAGCAAAAGTGCCTCTGAGGAAACACTACATTCCAGTAATGAAGAGGAAGACCCTTTC CGCGGAATGGAACCCTATCTTGTCCGGAGACTTTCATGTCGCAATATTCAGCTTCCCCCTCTCGCCTTCAGACAGTTGGAACAAGCTGAC TTGAAAAGTGAATCAGAGAACATTCAACGACCAACCAGCCTCCCCCTGAAGATTCTGCCGCTGATTGCTATCACTTCTGCAGAATCCAGT GGTTTTGATGTGGACAATGGCACATCTGCGGGACGGAGTCCCTTGGATCCCATGACCAGCCCAGGATCCGGGCTAATTCTCCAAGCAAAT TTTGTCCACAGTCAACGACGGGAGTCCTTCCTGTATCGATCCGACAGCGATTATGACCTCTCTCCAAAGTCTATGTCCCGGAACTCCTCC ATTGCCAGTGATATACACGGAGATGACTTGATTGTGACTCCATTTGCTCAGGTCTTGGCCAGTCTGCGAACTGTACGAAACAACTTTGCT GCATTAACTAATTTGCAAGATCGAGCACCTAGCAAAAGATCACCCATGTGCAACCAACCATCCATCAACAAAGCCACCATAACAGAGGAG GCCTACCAGAAACTGGCCAGCGAGACCCTGGAGGAGCTGGACTGGTGTCTGGACCAGCTAGAGACCCTACAGACCAGGCACTCCGTCAGT GAGATGGCCTCCAACAAGGGTGGCTTTGCAAAATGTTACGAGATGACAGATTTGACAAATAACAAAGTCTACGCCGCAAAAATTATTCCT CACAGCAGAGTAGCTAAACCTCATCAAAGGGAAAAGATTGACAAAGAAATAGAGCTTCACAGAATTCTTCATCATAAGCATGTAGTGCAG TTTTACCACTACTTCGAGGACAAAGAAAACATTTACATTCTCTTGGAATACTGCAGTAGAAGGTCAATGGCTCATATTTTGAAAGCAAGA AAGGTGTTGACAGAGCCAGAAGTTCGATACTACCTCAGGCAGATTGTGTCTGGACTGAAATACCTTCATGAACAAGAAATCTTGCACAGA GATCTCAAACTAGGGAACTTTTTTATTAATGAAGCCATGGAACTAAAAGTTGGGGACTTCGGTCTGGCAGCCAGGCTAGAACCCTTGGAA CACAGAAGGAGAACGATATGTGGTACCCCAAATTATCTCTCTCCTGAAGTCCTCAACAAACAAGGACATGGCTGTGAATCAGACATTTGG GCCCTGGGCTGTGTAATGTATACAATGTTACTAGGGAGGCCCCCATTTGAAACTACAAATCTCAAAGAAACTTATAGGTGCATAAGGGAA GCAAGGTATACAATGCCGTCCTCATTGCTGGCTCCTGCCAAGCACTTAATTGCTAGTATGTTGTCCAAAAACCCAGAGGATCGTCCCAGT TTGGATGACATCATTCGACATGACTTTTTTTTGCAGGGCTTCACTCCGGACAGACTGTCTTCTAGCTGTTGTCATACAGTTCCAGATTTC CACTTATCAAGCCCAGCTAAGAATTTCTTTAAGAAAGCAGCTGCTGCTCTTTTTGGTGGCAAAAAAGACAAAGCAAGATATATTGACACA CATAATAGAGTGTCTAAAGAAGATGAAGACATCTACAAGCTTAGGCATGATTTGAAAAAGACTTCAATAACTCAGCAACCCAGCAAACAC AGGACAGATGAGGAGCTCCAGCCACCTACCACCACAGTTGCCAGGTCTGGAACACCCGCAGTAGAAAACAAGCAGCAGATTGGGGATGCT ATTCGGATGATAGTCAGAGGGACTCTTGGCAGCTGTAGCAGCAGCAGTGAATGCCTTGAAGACAGTACCATGGGAAGTGTTGCAGACACA GTGGCAAGGGTTCTTCGGGGATGTCTGGAAAACATGCCGGAAGCTGATTGCATTCCCAAAGAGCAGCTGAGCACATCATTTCAGTGGGTC ACCAAATGGGTTGATTACTCTAACAAATATGGCTTTGGGTACCAGCTCTCAGACCACACCGTCGGTGTCCTTTTCAACAATGGTGCTCAC ATGAGCCTCCTTCCAGACAAAAAAACAGTTCACTATTACGCAGAGCTTGGCCAATGCTCAGTTTTCCCAGCAACAGATGCTCCTGAGCAA TTTATTAGTCAAGTGACGGTGCTGAAATACTTTTCTCATTACATGGAGGAGAACCTCATGGATGGTGGAGATCTGCCTAGTGTTACTGAT ATTCGAAGACCTCGGCTCTACCTCCTTCAGTGGCTAAAATCTGATAAGGCCCTAATGATGCTCTTTAATGATGGCACCTTTCAGGTGAAT TTCTACCATGATCATACAAAAATCATCATCTGTAGCCAAAATGAAGAATACCTTCTCACCTACATCAATGAGGATAGGATATCTACAACT TTCAGGCTGACAACTCTGCTGATGTCTGGCTGTTCATCAGAATTAAAAAATCGAATGGAATATGCCCTGAACATGCTCTTACAAAGATGT AACTGAAAGACTTTTCGAATGGACCCTATGGGACTCCTCTTTTCCACTGTGAGATCTACAGGGAAGCCAAAAGAATGATCTAGAGTATGT TGAAGAAGATGGACATGTGGTGGTACGAAAACAATTCCCCTGTGGCCTGCTGGACTGGTTGGAACCAGAACAGGCTAAGGCATACAGTTC TTGACTTTGGACAATCCAAGAGTGAACCAGAATGCAGTTTTCCTTGAGATACCTGTTTTAAAAGGTTTTTCAGACAATTTTGCAGAAAGG TGCATTGATTCTTAAATTCTCTCTGTTGAGAGCATTTCAGCCAGAGGACTTTGGAACTGTGAATATACTTCCTGAAGGGGAGGGAGAAGG GAGGAAGCTCCCATGTTGTTTAAAGGCTGTAATTGGAGCAGCTTTTGGCTGCGTAACTGTGAACTATGGCCATATATAATTTTTTTTCAT TAATTTTTGAAGATACTTGTGGCTGGAAAAGTGCATTCCTTGTTAATAAACTTTTTATTTATTACAGCCCAAAGAGCAGTATTTATTATC >63754_63754_6_PDE4D-PLK2_PDE4D_chr5_58334686_ENST00000546160_PLK2_chr5_57754919_ENST00000274289_length(amino acids)=841AA_BP=0 MKRNTCDLLSRSKSASEETLHSSNEEEDPFRGMEPYLVRRLSCRNIQLPPLAFRQLEQADLKSESENIQRPTSLPLKILPLIAITSAESS GFDVDNGTSAGRSPLDPMTSPGSGLILQANFVHSQRRESFLYRSDSDYDLSPKSMSRNSSIASDIHGDDLIVTPFAQVLASLRTVRNNFA ALTNLQDRAPSKRSPMCNQPSINKATITEEAYQKLASETLEELDWCLDQLETLQTRHSVSEMASNKGGFAKCYEMTDLTNNKVYAAKIIP HSRVAKPHQREKIDKEIELHRILHHKHVVQFYHYFEDKENIYILLEYCSRRSMAHILKARKVLTEPEVRYYLRQIVSGLKYLHEQEILHR DLKLGNFFINEAMELKVGDFGLAARLEPLEHRRRTICGTPNYLSPEVLNKQGHGCESDIWALGCVMYTMLLGRPPFETTNLKETYRCIRE ARYTMPSSLLAPAKHLIASMLSKNPEDRPSLDDIIRHDFFLQGFTPDRLSSSCCHTVPDFHLSSPAKNFFKKAAAALFGGKKDKARYIDT HNRVSKEDEDIYKLRHDLKKTSITQQPSKHRTDEELQPPTTTVARSGTPAVENKQQIGDAIRMIVRGTLGSCSSSSECLEDSTMGSVADT VARVLRGCLENMPEADCIPKEQLSTSFQWVTKWVDYSNKYGFGYQLSDHTVGVLFNNGAHMSLLPDKKTVHYYAELGQCSVFPATDAPEQ FISQVTVLKYFSHYMEENLMDGGDLPSVTDIRRPRLYLLQWLKSDKALMMLFNDGTFQVNFYHDHTKIIICSQNEEYLLTYINEDRISTT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PDE4D-PLK2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PDE4D-PLK2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PDE4D-PLK2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
