
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PDIA3-NAA50 (FusionGDB2 ID:63890) |
Fusion Gene Summary for PDIA3-NAA50 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PDIA3-NAA50 | Fusion gene ID: 63890 | Hgene | Tgene | Gene symbol | PDIA3 | NAA50 | Gene ID | 2923 | 80218 |
| Gene name | protein disulfide isomerase family A member 3 | N-alpha-acetyltransferase 50, NatE catalytic subunit | |
| Synonyms | ER60|ERp57|ERp60|ERp61|GRP57|GRP58|HEL-S-269|HEL-S-93n|HsT17083|P58|PI-PLC | MAK3|NAT13|NAT13P|NAT5|NAT5P|SAN|hNaa50p | |
| Cytomap | 15q15.3 | 3q13.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein disulfide-isomerase A358 kDa glucose-regulated protein58 kDa microsomal proteinER protein 57ER protein 60disulfide isomerase ER-60endoplasmic reticulum P58endoplasmic reticulum resident protein 57endoplasmic reticulum resident protein 60e | N-alpha-acetyltransferase 50N-acetyltransferase 13 (GCN5-related)N-acetyltransferase 5N-acetyltransferase san homologN-epsilon-acetyltransferase 50natE catalytic subunit | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q9GZZ1 | |
| Ensembl transtripts involved in fusion gene | ENST00000469684, ENST00000300289, ENST00000538521, | ENST00000240922, ENST00000477813, ENST00000467022, ENST00000493454, ENST00000493900, ENST00000497255, ENST00000497525, | |
| Fusion gene scores | * DoF score | 13 X 14 X 5=910 | 9 X 5 X 5=225 |
| # samples | 16 | 9 | |
| ** MAII score | log2(16/910*10)=-2.5077946401987 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/225*10)=-1.32192809488736 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PDIA3 [Title/Abstract] AND NAA50 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PDIA3(44038904)-NAA50(113442942), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | NAA50 | GO:0006474 | N-terminal protein amino acid acetylation | 21900231|22311970|27484799 |
| Tgene | NAA50 | GO:0034087 | establishment of mitotic sister chromatid cohesion | 27422821 |
| Tgene | NAA50 | GO:0071962 | mitotic sister chromatid cohesion, centromeric | 17502424 |
 Fusion gene breakpoints across PDIA3 (5'-gene) Fusion gene breakpoints across PDIA3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
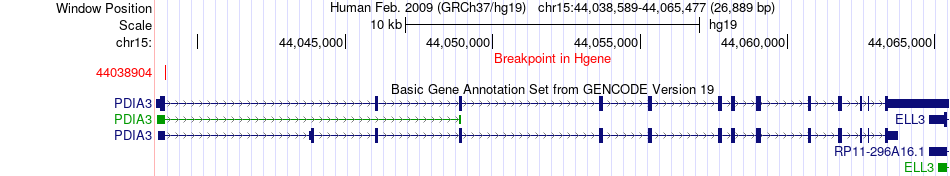 |
 Fusion gene breakpoints across NAA50 (3'-gene) Fusion gene breakpoints across NAA50 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
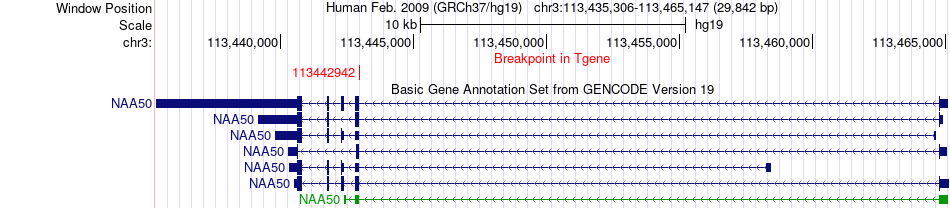 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-BR-A4PE-01A | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
Top |
Fusion Gene ORF analysis for PDIA3-NAA50 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000469684 | ENST00000240922 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 3UTR-3CDS | ENST00000469684 | ENST00000477813 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 3UTR-5UTR | ENST00000469684 | ENST00000467022 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 3UTR-5UTR | ENST00000469684 | ENST00000493454 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 3UTR-5UTR | ENST00000469684 | ENST00000493900 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 3UTR-5UTR | ENST00000469684 | ENST00000497255 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 3UTR-5UTR | ENST00000469684 | ENST00000497525 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5CDS-5UTR | ENST00000300289 | ENST00000467022 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5CDS-5UTR | ENST00000300289 | ENST00000493454 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5CDS-5UTR | ENST00000300289 | ENST00000493900 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5CDS-5UTR | ENST00000300289 | ENST00000497255 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5CDS-5UTR | ENST00000300289 | ENST00000497525 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-3CDS | ENST00000538521 | ENST00000240922 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-3CDS | ENST00000538521 | ENST00000477813 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-5UTR | ENST00000538521 | ENST00000467022 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-5UTR | ENST00000538521 | ENST00000493454 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-5UTR | ENST00000538521 | ENST00000493900 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-5UTR | ENST00000538521 | ENST00000497255 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| 5UTR-5UTR | ENST00000538521 | ENST00000497525 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| In-frame | ENST00000300289 | ENST00000240922 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
| In-frame | ENST00000300289 | ENST00000477813 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000300289 | PDIA3 | chr15 | 44038904 | + | ENST00000240922 | NAA50 | chr3 | 113442942 | - | 6117 | 315 | 7 | 816 | 269 |
| ENST00000300289 | PDIA3 | chr15 | 44038904 | + | ENST00000477813 | NAA50 | chr3 | 113442942 | - | 2142 | 315 | 7 | 696 | 229 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000300289 | ENST00000240922 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - | 0.000244552 | 0.99975544 |
| ENST00000300289 | ENST00000477813 | PDIA3 | chr15 | 44038904 | + | NAA50 | chr3 | 113442942 | - | 0.000356895 | 0.9996431 |
Top |
Fusion Genomic Features for PDIA3-NAA50 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for PDIA3-NAA50 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:44038904/chr3:113442942) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | NAA50 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: N-alpha-acetyltransferase that acetylates the N-terminus of proteins that retain their initiating methionine (PubMed:19744929, PubMed:22311970, PubMed:21900231, PubMed:27484799). Has a broad substrate specificity: able to acetylate the initiator methionine of most peptides, except for those with a proline in second position (PubMed:27484799). Also displays N-epsilon-acetyltransferase activity by mediating acetylation of the side chain of specific lysines on proteins (PubMed:19744929). Autoacetylates in vivo (PubMed:19744929). The relevance of N-epsilon-acetyltransferase activity is however unclear: able to acetylate H4 in vitro, but this result has not been confirmed in vivo (PubMed:19744929). Component of a N-alpha-acetyltransferase complex containing NAA10 and NAA15, but NAA50 does not influence the acetyltransferase activity of NAA10: this multiprotein complex probably constitutes the major contributor for N-terminal acetylation at the ribosome exit tunnel, with NAA10 acetylating all amino termini that are devoid of methionine and NAA50 acetylating other peptides (PubMed:16507339, PubMed:27484799). Required for sister chromatid cohesion during mitosis by promoting binding of CDCA5/sororin to cohesin: may act by counteracting the function of NAA10 (PubMed:17502424, PubMed:27422821). {ECO:0000269|PubMed:16507339, ECO:0000269|PubMed:17502424, ECO:0000269|PubMed:19744929, ECO:0000269|PubMed:21900231, ECO:0000269|PubMed:22311970, ECO:0000269|PubMed:27422821, ECO:0000269|PubMed:27484799}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | NAA50 | chr15:44038904 | chr3:113442942 | ENST00000240922 | 0 | 5 | 6_155 | 2 | 170.0 | Domain | N-acetyltransferase | |
| Tgene | NAA50 | chr15:44038904 | chr3:113442942 | ENST00000240922 | 0 | 5 | 117_126 | 2 | 170.0 | Region | Coenzyme A binding | |
| Tgene | NAA50 | chr15:44038904 | chr3:113442942 | ENST00000240922 | 0 | 5 | 138_141 | 2 | 170.0 | Region | Substrate binding | |
| Tgene | NAA50 | chr15:44038904 | chr3:113442942 | ENST00000240922 | 0 | 5 | 77_90 | 2 | 170.0 | Region | Acetyl-CoA binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PDIA3 | chr15:44038904 | chr3:113442942 | ENST00000300289 | + | 1 | 13 | 25_133 | 55 | 506.0 | Domain | Thioredoxin 1 |
| Hgene | PDIA3 | chr15:44038904 | chr3:113442942 | ENST00000300289 | + | 1 | 13 | 343_485 | 55 | 506.0 | Domain | Thioredoxin 2 |
| Hgene | PDIA3 | chr15:44038904 | chr3:113442942 | ENST00000300289 | + | 1 | 13 | 502_505 | 55 | 506.0 | Motif | Prevents secretion from ER |
Top |
Fusion Gene Sequence for PDIA3-NAA50 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >63890_63890_1_PDIA3-NAA50_PDIA3_chr15_44038904_ENST00000300289_NAA50_chr3_113442942_ENST00000240922_length(transcript)=6117nt_BP=315nt GCCGGGTTTGGGGGTTGGGACCTCCGGCTGCAGGTCCGCCTGGGCCAGACGCGCGAGCGCAAGCAGCGGGTTAGTGGTCGCGCGCCCGAC CTCCGCAGTCCCAGCCGAGCCGCGACCCTTCCGGCCGTCCCCACCCCACCTCGCCGCCATGCGCCTCCGCCGCCTAGCGCTGTTCCCGGG TGTGGCGCTGCTTCTTGCCGCGGCCCGCCTCGCCGCTGCCTCCGACGTGCTAGAACTCACGGACGACAACTTCGAGAGTCGCATCTCCGA CACGGGCTCTGCGGGCCTCATGCTCGTCGAGTTCTTCGCCCCCTGTAGCCGGATCGAGCTGGGAGATGTGACACCACACAATATTAAACA GTTGAAAAGATTGAATCAGGTCATCTTTCCAGTCAGCTACAATGACAAGTTCTACAAGGATGTGCTGGAGGTTGGCGAGCTAGCAAAACT TGCCTATTTCAATGATATTGCTGTAGGTGCAGTATGCTGTAGGGTGGATCATTCACAGAATCAGAAGAGACTTTACATCATGACACTAGG ATGTCTGGCACCTTACCGAAGGCTAGGAATAGGAACTAAAATGTTAAATCATGTCTTAAACATCTGTGAAAAAGATGGTACTTTTGACAA CATTTATCTGCATGTCCAGATCAGCAATGAGTCGGCAATTGACTTCTACAGGAAGTTTGGCTTTGAGATTATTGAGACAAAGAAGAACTA CTATAAGAGGATAGAGCCCGCAGATGCTCATGTGCTGCAGAAAAACCTCAAAGTTCCTTCTGGTCAGAATGCAGATGTGCAAAAGACAGA CAACTGAACAAATTACAAATGAACTTTCTTGCACTTGCTTGTCGCCAAATAAAAGAGAGGCCCATTGATTCCTCCCCCACCCCAACACTT TTCTTTTAAAGCTTTTCTCCCTCCTTGTTCTTGTTTTTCTTTCTTCCTTTCCTTTTCTCTGAGAGTTTTAATACTTTCAAGGACTTTAAA AAAATAATCATGTTTGAATTGTTTTCTCTTATTTTTGTGAGGTGGTTTGAAGGAAGGACAAGGTAGATCTGTTTAGTTTTGCAGTTGAAG TTAGATGGTCCTAAACATTTAATTGTCAAATAATTTCAAATTTAATGTCCTGCTTTCACATTGAAGGGCAGAGCCTACAAAACATTGTAT ATTTCAAAAGACAAAAAGAAGCAGCAGCAGTATCTTGTTCTCTAATTCATAGACAAGTTGAGTGTGTTTGTGGTACTTTGGGTTTTTAAA CACTTTGGGATACTAATCCCTAGACATTGCCTTCACTCCACCTTTAGTCCTTCTGAGCACTCTCTCGGGAGTTGGAACATTGTTATCCTT GTAAGAAATACTAAGCTTATGTTGATTTTTAAGTAATTATATCTTCTCTTCTTGCTGGTGGGTGGGGCAGTTTGGTTTAGTGTTATACTT TGGTCTAAGTATTTGAGTTAAACTGCTTTTTTGCTAATGAGTGGGCTGGTTGTTAGCAGGTTTGTTTTTCCTGCTGTTGATTGTTACTAG TGGCATTAACTTTTAGAATTTGGGCTGGTGAGATTAATTTTTTTTAATATCCCAGCTAGAGATATGGCCTTTAACTGACCTAAAGAGGTG TGTTGTGATTTAATTTTTTCCCGTTCCTTTTTCTTCAGTAAACCCAACAATAGTCTAACCTTAAAAATTGAGTTGATGTCCTTATAGGTC ACTACCCCTAAATAAACCTGAAGCAGGTGTTTTCTCTTGGACATACTAAAAAATACCTAAAAGGAAGCTTAGATGGGCTGTGACACAAAA AATTCAATTACTGTCATCTAATGCCAGCTGTTAAAAGTGTGGCCACTGAGCATTTGATTTTATAGGAAAAAATAGTATTTTTGAGAATAA CATAGCTGTGCTATTGCACATCTGTTGGAGGACATCCCAGATTTGCTTATACTCAGTGCCTGTGATATTGAGTTTAAGGATTTGAGGCAG GGGTAATTATTAAACATATTGCTTCTATTCTTGGAAAAATAGAAGTGTAAAATGTTAATAATACAAATGTCACTGTGACCTCCTCCACTG AGAGGACTGGTTTATGCCAGATCATTTTCCGGCACACACGGAGTGGCTTTGACAGATTGATAACTTTGTAAGATGGGAGACATCTGAAAT ATTCATGTTTTCCTTTTGTAGTCCCATCTCCACTATTTAGAAATGTTCTCAGACTTTAAAATAATGCACAGGGCTTGAGCTTTCTGTCAT TTGACTTTAAAAGGAAGTTTCATTCATATTTATCCTCTTATGTAAAATTGCGGTATAAAGTCTCATTTCCAAATATGTTAAATGACAAAA TTATTTTATAAAATGTTTATGCACACTTTATAACCTTAAGTTTTTATTTGAGAATGTGAAAGTACAAAGTGCAGTAGACTTCAACAATCT TGAGTGCCAAGAATAATACAGAAAAAGAAGACAGTTGATGAATGAGTTTATAGGGTTCTAATCTTAAGATGGTAAAAATGTAGAAAGACC TTGCTGGTTTTTTGGGGGTATTCGTTTCTTAAACAATCCAAATCTAAGCTTAGAAGAAAAGTTTAGCGTTAAGCACCTTTATCTTCATGA ATAAGCTTCAGCTTGCTCTTGGCAAGAGAAGAGTGCTTGAGTTACAGAAGGCATAAGTAGTTTGAAGAATGCAGCAGCCTTTTTGTAAAC TTCCCAGATATCAAAATAGACTTTGATATATAAATGGTTTTCTGAGATGACACTGCCTCTATTTCTATAACCATTTCACCTGGACTATCT AATCAGTCCTATGAATGTATCCCTAAATGTGGTTATTGAAAACCTAATAGCTGCCTCATGACAAGTACATGTTATTTAAGGAGGAAAAAA TATTAAATTTTGAATTGAGTGTGTAGGCTCCCTATCATTATATATAGAGTTTCTTTTTCCACGGTAGTCAGTGACTTAACCTGAATTGTA AATGTTTGTAAAGGGTTAATTGTCCTACATCAAACTTAGTTAAATAATTCCATCCACTTATGGAGGAGGAGGAGAATGTGGAAGAGGTAA AAAGCTGGGCACAAGTTCATATGCCTATGAGTCAGTAAAGACTGAAGTAATGTCCTATGTTGAGCTGGTTATTTTGATATATGATAATAA TTATCTTTGAAGTAGAACAATTCTGTTAACTGGAAAATCACAGGATATATCCATCATATTTTTCAGGACAGATAGTTTTTACTGTGGGGC AAATAGGTTAAAATTACACTATGTTAGTTGCATTTAGGTTTTAAAGCAAAGAATCTGTAGAGAAATCTATGCAATATATAGTTTGTCCAG ATTAGCTTTCATTTGGGGAATGAAGTTCTGAAATATCTAAAGCAGTTTACTCATCAATTGAAAAGTCCTCCAAAAAGAGAACTATTGGGA AACCATGGTGTGGTGGTGGAAAAGAAAAGCTCCCTCAGTTTTTTGGAGGGAATAACTTAAAAAAATACTTAAATGGCTAAGTTTACTTGG TGCAGTTAAGAATTAAACTTGTCAATTTTAACATTGCTGTTACATCTGAAATAAACTTATGTGATGTTCTGGTAGTGATCTGTGATATCT GTAAATGTCAAAACTGTATTGTTGAATTCTGCAGCCAGCAACCGTACATACTCATTGTGTAGTGTTTCCCTTACTGCTTTTATTTACTTT CACATTTAAGATCTAATTTTAAAATCATTAAAATAGGCCAAGTGTAATTAAGGCATCCTAATTCAGATCTTCTGTGACTTGCTAGGTACA GTGCCCAATATCTACAATTCAGTTCCAATGAGAGGGAAAAAGTAAGATCCAAGGAACTGTCTTGCTGCTGTATTTTTAATGTAATTATTA GAAATACTTGACACTTTAGGCTGCAATCTAGTTAGTAATGTTTATTGGCTACAGACAACATATTCAGGTGTTTTGTTTTTCTTCCTTGTA AGGAAGGAGTACGGTAGTTCAATGAGTTGGGAATTGGCTTTTAAGCTTTTTATCAAACATGAATTTAAGATTTTTTCTTTAACAGAATTC AGTATTTAATATTTCTAGTCAAAATTGTTATAATTAGATTTTTGCTTTTTTCATTTAAGAAGCAATGAGTGATCCTTGTCTAGTGTGTCT CAGTTATTACTGTAGTTTAAAACAACAAACAAGTATTACCTTCCACAGTTCTGGATCAGGAATCTAGGAGCTGCTTAACTGGGTGGTTCT GTCTTGAGATCTCAGAAGGTTGCAGTCAAGCTGTTAGCCAGGACTGTACCATCTTGAAGCTTGACAAGGCTAGAGGAGCCATTTCCAAGA TGTCTTGCTTACGTGGCTCTTGGGCTTCTGTTCCTCACTGTGTGGGTATCACACACAGTGCCTGATGCTCACGTGGCTCTTGGGCTTCAG TTCCTTGGGCACCAGTTGCCTGGTGGACAACTGGTATGTGGCTTCCCCTAGAGCAGGTGATATAAGAAAGTGAGCAGAAACCATACTTTT TTATGACGTAGCCTCAGGTGACATTATTTCTGCCATATTCCATGAGTCACACAGACCAACCATTGAATCAGCTTGGTATAGTGTGGGAGG GGACTGCACAAGGATGTGAATACTGGGAGGTGGATATCATTGGGCCCTCTAGGAGACTGGCTATCACTGCTCAGAGAAGATGCTGGGCTT CGTTGACTCCCAAGGTCAGGGTAGTTTTGGTTAGGTTGTGTTTTATTAGTCTCTAAAAGGAGTAGTTTTCTAAATGAGGGTTATAAAGCA CTGCACTTTACAGTTATTCGGGATATAAAAGAAATAGTGGGTCTAAAGGCTGTGCTCTTGTGGTTGGGTTTTCAGTGGGGAGGGGAGACT AGTCTGTCTCAGACTGATGCTGTTCTCACTGATTTGAATATTTCTGGGAAATTGGATACTACAGTCACAAATAGGAACAGTAAGCCTATA GAAGTTTTTCAGGGAGTAAATATTTTCTATGGCAGTGTCCTGAATTGGTTCTCCCTTGCAAGACTGAACTGTAGGAACTTAGTCCTAGTT TATGATACAGCCAAGTAACATAGTCACATGGGAAAAAGAGCTTGAGTCAGATTTCTTAATGTGTGTTGTTAACTTGGTAGAACATTGAGA ATTATTTAAGTCAGAGAACGATCTGTTACTGGGGCAGAAATTCTCAACCTTTTCAGTTCTCCAAAATTTAAGATACTTGATTTCTTAGGT AAAATGTTTTTGTTTTTGTTTTGGAGACAGAGTCTCGCTCTGTCGCCCAGGCTGGAGTGCAGTGGCGCGATCTTGGCTCACTGCAAACTC CGCCTCCCAGATTCAAGCAATTCTGCCTGAGCCTCCCAAGTAGCTGCGACTAGAAAGCGCATGCCACCACGCCTGGCTAATTTTTTGTAT TTTAGTAGAGATGGGGGTTTCACCGTGTTGCCCAGGCTGGTCTCAAACTCCTGAGCTTAGGCAATCCTCCTGGGGCAGCCTCCCAAAGTG CTAGGATTACAGGCGAGCCATGGCGCCTGGCCAGTAAAATGTTTTCTATCTAGAATGAATCAAGGTATTTTCCTTGCTCAGTAGCTTCTA GAATAAGAAAAAAATAGCAGCAAGATCTGATTCAGAAATAGTTGGGAGCAGAAAGTTAATATGAAGGAGTTGCTACTTGTTAACAGCCTA GAGTTGAGATCTAGAAGAATTATTACCTTTTTAAATTTGTGATGAAAGCTTAAATCCAGCATTTGGGAAGTTACTCTATTGGCTGAACTA TTTTGGAGTTTGTAAGCTTTGTATTAGATATTCCTGATTTAACTGAAACTAATTTGCCACATAGCTTTAATTTCATCCCAGTTTTACTTG TTTTACTGTCCTCAAAAACTCAAGACATCTGAACTCAAAGGATCTAAGCAGTATAAATTAAAGCACATGTTGAATCACTGTAGCTTTCGT AGGACATCTGATTATAATGCTTTTCTTTGTTTCTTTGTCCATACTGAACTTGTCTAGTTTATCTTTGAGAAACATTTGCTAGTATCAAAA >63890_63890_1_PDIA3-NAA50_PDIA3_chr15_44038904_ENST00000300289_NAA50_chr3_113442942_ENST00000240922_length(amino acids)=269AA_BP=103 MGVGTSGCRSAWARRASASSGLVVARPTSAVPAEPRPFRPSPPHLAAMRLRRLALFPGVALLLAAARLAAASDVLELTDDNFESRISDTG SAGLMLVEFFAPCSRIELGDVTPHNIKQLKRLNQVIFPVSYNDKFYKDVLEVGELAKLAYFNDIAVGAVCCRVDHSQNQKRLYIMTLGCL -------------------------------------------------------------- >63890_63890_2_PDIA3-NAA50_PDIA3_chr15_44038904_ENST00000300289_NAA50_chr3_113442942_ENST00000477813_length(transcript)=2142nt_BP=315nt GCCGGGTTTGGGGGTTGGGACCTCCGGCTGCAGGTCCGCCTGGGCCAGACGCGCGAGCGCAAGCAGCGGGTTAGTGGTCGCGCGCCCGAC CTCCGCAGTCCCAGCCGAGCCGCGACCCTTCCGGCCGTCCCCACCCCACCTCGCCGCCATGCGCCTCCGCCGCCTAGCGCTGTTCCCGGG TGTGGCGCTGCTTCTTGCCGCGGCCCGCCTCGCCGCTGCCTCCGACGTGCTAGAACTCACGGACGACAACTTCGAGAGTCGCATCTCCGA CACGGGCTCTGCGGGCCTCATGCTCGTCGAGTTCTTCGCCCCCTGTAGCCGGATCGAGCTGGGAGATGTGACACCACACAATATTAAACA GTTGAAAAGATTGAATCAGGTCATCTTTCCAGTCAGCTACAATGACAAGTTCTACAAGGATGTGCTGGAGGTTGGCGAGCTAGCAAAACT TGGAACTAAAATGTTAAATCATGTCTTAAACATCTGTGAAAAAGATGGTACTTTTGACAACATTTATCTGCATGTCCAGATCAGCAATGA GTCGGCAATTGACTTCTACAGGAAGTTTGGCTTTGAGATTATTGAGACAAAGAAGAACTACTATAAGAGGATAGAGCCCGCAGATGCTCA TGTGCTGCAGAAAAACCTCAAAGTTCCTTCTGGTCAGAATGCAGATGTGCAAAAGACAGACAACTGAACAAATTACAAATGAACTTTCTT GCACTTGCTTGTCGCCAAATAAAAGAGAGGCCCATTGATTCCTCCCCCACCCCAACACTTTTCTTTTAAAGCTTTTCTCCCTCCTTGTTC TTGTTTTTCTTTCTTCCTTTCCTTTTCTCTGAGAGTTTTAATACTTTCAAGGACTTTAAAAAAATAATCATGTTTGAATTGTTTTCTCTT ATTTTTGTGAGGTGGTTTGAAGGAAGGACAAGGTAGATCTGTTTAGTTTTGCAGTTGAAGTTAGATGGTCCTAAACATTTAATTGTCAAA TAATTTCAAATTTAATGTCCTGCTTTCACATTGAAGGGCAGAGCCTACAAAACATTGTATATTTCAAAAGACAAAAAGAAGCAGCAGCAG TATCTTGTTCTCTAATTCATAGACAAGTTGAGTGTGTTTGTGGTACTTTGGGTTTTTAAACACTTTGGGATACTAATCCCTAGACATTGC CTTCACTCCACCTTTAGTCCTTCTGAGCACTCTCTCGGGAGTTGGAACATTGTTATCCTTGTAAGAAATACTAAGCTTATGTTGATTTTT AAGTAATTATATCTTCTCTTCTTGCTGGTGGGTGGGGCAGTTTGGTTTAGTGTTATACTTTGGTCTAAGTATTTGAGTTAAACTGCTTTT TTGCTAATGAGTGGGCTGGTTGTTAGCAGGTTTGTTTTTCCTGCTGTTGATTGTTACTAGTGGCATTAACTTTTAGAATTTGGGCTGGTG AGATTAATTTTTTTTAATATCCCAGCTAGAGATATGGCCTTTAACTGACCTAAAGAGGTGTGTTGTGATTTAATTTTTTCCCGTTCCTTT TTCTTCAGTAAACCCAACAATAGTCTAACCTTAAAAATTGAGTTGATGTCCTTATAGGTCACTACCCCTAAATAAACCTGAAGCAGGTGT TTTCTCTTGGACATACTAAAAAATACCTAAAAGGAAGCTTAGATGGGCTGTGACACAAAAAATTCAATTACTGTCATCTAATGCCAGCTG TTAAAAGTGTGGCCACTGAGCATTTGATTTTATAGGAAAAAATAGTATTTTTGAGAATAACATAGCTGTGCTATTGCACATCTGTTGGAG GACATCCCAGATTTGCTTATACTCAGTGCCTGTGATATTGAGTTTAAGGATTTGAGGCAGGGGTAATTATTAAACATATTGCTTCTATTC TTGGAAAAATAGAAGTGTAAAATGTTAATAATACAAATGTCACTGTGACCTCCTCCACTGAGAGGACTGGTTTATGCCAGATCATTTTCC GGCACACACGGAGTGGCTTTGACAGATTGATAACTTTGTAAGATGGGAGACATCTGAAATATTCATGTTTTCCTTTTGTAGTCCCATCTC >63890_63890_2_PDIA3-NAA50_PDIA3_chr15_44038904_ENST00000300289_NAA50_chr3_113442942_ENST00000477813_length(amino acids)=229AA_BP=103 MGVGTSGCRSAWARRASASSGLVVARPTSAVPAEPRPFRPSPPHLAAMRLRRLALFPGVALLLAAARLAAASDVLELTDDNFESRISDTG SAGLMLVEFFAPCSRIELGDVTPHNIKQLKRLNQVIFPVSYNDKFYKDVLEVGELAKLGTKMLNHVLNICEKDGTFDNIYLHVQISNESA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PDIA3-NAA50 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PDIA3-NAA50 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PDIA3-NAA50 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
