
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PGAP1-ZDBF2 (FusionGDB2 ID:64542) |
Fusion Gene Summary for PGAP1-ZDBF2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PGAP1-ZDBF2 | Fusion gene ID: 64542 | Hgene | Tgene | Gene symbol | PGAP1 | ZDBF2 | Gene ID | 80055 | 57683 |
| Gene name | post-GPI attachment to proteins inositol deacylase 1 | zinc finger DBF-type containing 2 | |
| Synonyms | Bst1|ISPD3024|MRT42|SPG67 | Slx9 | |
| Cytomap | 2q33.1 | 2q33.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | GPI inositol-deacylasepost-GPI attachment to proteins 1post-GPI attachment to proteins factor 1 | DBF4-type zinc finger-containing protein 2 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000354764, ENST00000409475, ENST00000409188, ENST00000485830, | ENST00000374423, | |
| Fusion gene scores | * DoF score | 9 X 12 X 7=756 | 2 X 3 X 2=12 |
| # samples | 10 | 3 | |
| ** MAII score | log2(10/756*10)=-2.91838623444635 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/12*10)=1.32192809488736 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: PGAP1 [Title/Abstract] AND ZDBF2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PGAP1(197735665)-ZDBF2(207169441), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | PGAP1-ZDBF2 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. PGAP1-ZDBF2 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across PGAP1 (5'-gene) Fusion gene breakpoints across PGAP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
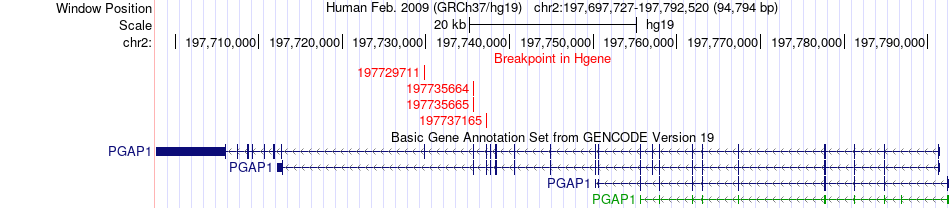 |
 Fusion gene breakpoints across ZDBF2 (3'-gene) Fusion gene breakpoints across ZDBF2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
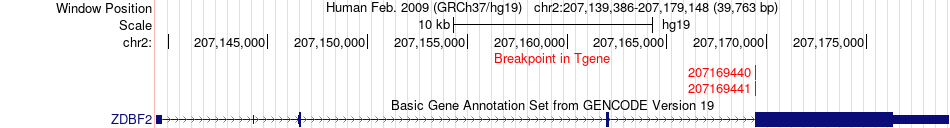 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | HNSC | TCGA-CV-7568-01A | PGAP1 | chr2 | 197735665 | - | ZDBF2 | chr2 | 207169441 | + |
| ChimerDB4 | HNSC | TCGA-CV-7568 | PGAP1 | chr2 | 197729711 | - | ZDBF2 | chr2 | 207169441 | + |
| ChimerDB4 | HNSC | TCGA-CV-7568 | PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + |
| ChimerDB4 | HNSC | TCGA-CV-7568 | PGAP1 | chr2 | 197735665 | - | ZDBF2 | chr2 | 207169441 | + |
| ChimerDB4 | HNSC | TCGA-CV-7568 | PGAP1 | chr2 | 197737165 | - | ZDBF2 | chr2 | 207169441 | + |
Top |
Fusion Gene ORF analysis for PGAP1-ZDBF2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000354764 | ENST00000374423 | PGAP1 | chr2 | 197735665 | - | ZDBF2 | chr2 | 207169441 | + |
| Frame-shift | ENST00000354764 | ENST00000374423 | PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + |
| Frame-shift | ENST00000354764 | ENST00000374423 | PGAP1 | chr2 | 197737165 | - | ZDBF2 | chr2 | 207169441 | + |
| Frame-shift | ENST00000409475 | ENST00000374423 | PGAP1 | chr2 | 197735665 | - | ZDBF2 | chr2 | 207169441 | + |
| Frame-shift | ENST00000409475 | ENST00000374423 | PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + |
| Frame-shift | ENST00000409475 | ENST00000374423 | PGAP1 | chr2 | 197737165 | - | ZDBF2 | chr2 | 207169441 | + |
| In-frame | ENST00000354764 | ENST00000374423 | PGAP1 | chr2 | 197729711 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000409188 | ENST00000374423 | PGAP1 | chr2 | 197735665 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000409188 | ENST00000374423 | PGAP1 | chr2 | 197729711 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000409188 | ENST00000374423 | PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + |
| intron-3CDS | ENST00000409188 | ENST00000374423 | PGAP1 | chr2 | 197737165 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000409475 | ENST00000374423 | PGAP1 | chr2 | 197729711 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000485830 | ENST00000374423 | PGAP1 | chr2 | 197735665 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000485830 | ENST00000374423 | PGAP1 | chr2 | 197729711 | - | ZDBF2 | chr2 | 207169441 | + |
| intron-3CDS | ENST00000485830 | ENST00000374423 | PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + |
| intron-3CDS | ENST00000485830 | ENST00000374423 | PGAP1 | chr2 | 197737165 | - | ZDBF2 | chr2 | 207169441 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000354764 | PGAP1 | chr2 | 197729711 | - | ENST00000374423 | ZDBF2 | chr2 | 207169441 | + | 11684 | 1976 | 2031 | 8852 | 2273 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000354764 | ENST00000374423 | PGAP1 | chr2 | 197729711 | - | ZDBF2 | chr2 | 207169441 | + | 0.001968285 | 0.9980317 |
Top |
Fusion Genomic Features for PGAP1-ZDBF2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + | 2.65E-07 | 0.99999976 |
| PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + | 2.65E-07 | 0.99999976 |
| PGAP1 | chr2 | 197729710 | - | ZDBF2 | chr2 | 207169440 | + | 1.53E-07 | 0.9999999 |
| PGAP1 | chr2 | 197737164 | - | ZDBF2 | chr2 | 207169440 | + | 4.79E-09 | 1 |
| PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + | 2.65E-07 | 0.99999976 |
| PGAP1 | chr2 | 197735664 | - | ZDBF2 | chr2 | 207169440 | + | 2.65E-07 | 0.99999976 |
| PGAP1 | chr2 | 197729710 | - | ZDBF2 | chr2 | 207169440 | + | 1.53E-07 | 0.9999999 |
| PGAP1 | chr2 | 197737164 | - | ZDBF2 | chr2 | 207169440 | + | 4.79E-09 | 1 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
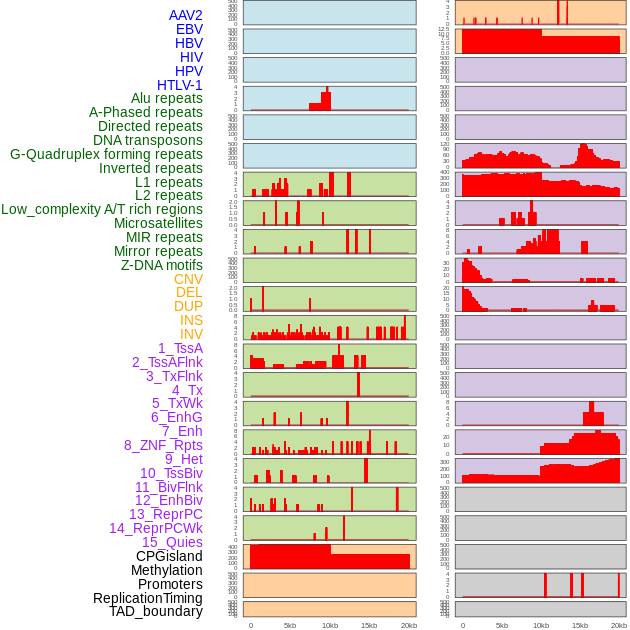 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
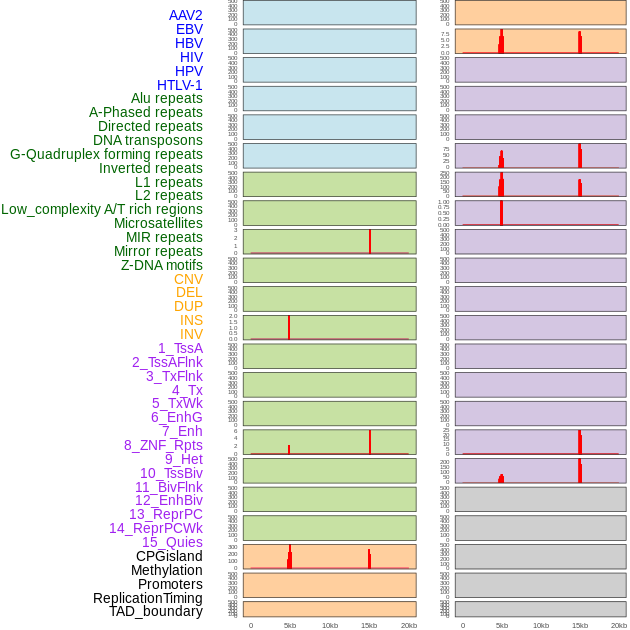 |
Top |
Fusion Protein Features for PGAP1-ZDBF2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:197735665/chr2:207169441) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 1_11 | 620 | 923.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 12_32 | 620 | 923.0 | Transmembrane | Helical |
| Tgene | ZDBF2 | chr2:197729711 | chr2:207169441 | ENST00000374423 | 3 | 5 | 1080_1110 | 62 | 2355.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | ZDBF2 | chr2:197729711 | chr2:207169441 | ENST00000374423 | 3 | 5 | 636_660 | 62 | 2355.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | ZDBF2 | chr2:197729711 | chr2:207169441 | ENST00000374423 | 3 | 5 | 2250_2253 | 62 | 2355.0 | Compositional bias | Note=Poly-Lys | |
| Tgene | ZDBF2 | chr2:197729711 | chr2:207169441 | ENST00000374423 | 3 | 5 | 88_94 | 62 | 2355.0 | Compositional bias | Note=Poly-Glu |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 33_641 | 620 | 923.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 663_668 | 620 | 923.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 690_733 | 620 | 923.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 755_817 | 620 | 923.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 839_853 | 620 | 923.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 875_893 | 620 | 923.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 915_922 | 620 | 923.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 1_11 | 0 | 593.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 33_641 | 0 | 593.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 663_668 | 0 | 593.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 690_733 | 0 | 593.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 755_817 | 0 | 593.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 839_853 | 0 | 593.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 875_893 | 0 | 593.0 | Topological domain | Cytoplasmic |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 915_922 | 0 | 593.0 | Topological domain | Lumenal |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 642_662 | 620 | 923.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 669_689 | 620 | 923.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 734_754 | 620 | 923.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 818_838 | 620 | 923.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 854_874 | 620 | 923.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000354764 | - | 20 | 27 | 894_914 | 620 | 923.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 12_32 | 0 | 593.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 642_662 | 0 | 593.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 669_689 | 0 | 593.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 734_754 | 0 | 593.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 818_838 | 0 | 593.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 854_874 | 0 | 593.0 | Transmembrane | Helical |
| Hgene | PGAP1 | chr2:197729711 | chr2:207169441 | ENST00000409475 | - | 1 | 20 | 894_914 | 0 | 593.0 | Transmembrane | Helical |
| Tgene | ZDBF2 | chr2:197729711 | chr2:207169441 | ENST00000374423 | 3 | 5 | 1_50 | 62 | 2355.0 | Zinc finger | DBF4-type |
Top |
Fusion Gene Sequence for PGAP1-ZDBF2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >64542_64542_1_PGAP1-ZDBF2_PGAP1_chr2_197729711_ENST00000354764_ZDBF2_chr2_207169441_ENST00000374423_length(transcript)=11684nt_BP=1976nt CGGCTCCGGGGGTGGCGGTCGGACAGTGTCTAGCACGCTCAGTCCGGGCTTGGGGCCCCGCGGCGGAGAAGGAGGTAGAGGGGGCGGCGG CGGCGGCGGCGGTGGTGGCGGCACCATGTTTCTTCACTCAGTTAATCTCTGGAACCTGGCGTTTTATGTCTTCATGGTCTTTCTGGCAAC CCTGGGGCTGTGGGATGTCTTCTTCGGCTTCGAGGAGAATAAGTGCAGTATGAGCTACATGTTTGAGTACCCGGAGTATCAGAAAATAGA ACTTCCAAAGAAACTGGCAAAACGCTATCCCGCATATGAGTTGTATCTTTATGGAGAGGGATCCTATGCTGAAGAACACAAAATTCTCCC TTTGACGGGTATTCCAGTTCTCTTTCTTCCTGGTAATGCTGGAAGTTATAAGCAAGTTCGTTCTATTGGCTCCATTGCACTTAGAAAAGC AGAGGACATTGACTTCAAGTACCACTTTGACTTCTTTAGTGTGAACTTCAATGAAGAACTGGTGGCTTTGTATGGTGGAAGTCTTCAGAA GCAGACCAAGTTTGTACATGAATGTATTAAAACAATTCTCAAACTCTATAAGGGTCAAGAATTTGCTCCAAAAAGTGTGGCAATAATTGG TCATTCTATGGGTGGCCTTGTTGCAAGAGCATTGCTTACACTGAAAAATTTTAAGCATGATCTGATAAATCTTCTTATTACACAAGCCAC ACCTCATGTTGCTCCTGTGATGCCATTAGATCGTTTCATTACAGATTTTTATACGACTGTAAACAACTATTGGATTCTAAATGCTCGACA CATAAATTTAACCACACTTTCTGTAGCTGGAGGATTCCGGGATTACCAAGTTCGTTCAGGATTGACTTTTCTACCAAAATTAAGCCATCA TACCAGTGCCTTATCTGTTGTGAGTTCAGCAGTGCCTAAGACCTGGGTCTCAACAGACCACCTCTCCATTGTGTGGTGTAAACAATTGCA GTTGACTACAGTTCGAGCATTCTTTGATCTTATTGATGCTGATACTAAACAAATAACTCAAAATTCCAAGAAGAAACTGTCAGTTTTGTA TCACCACTTTATAAGACACCCATCAAAACATTTTGAGGAAAATCCAGCTATAATTTCTGACTTAACAGGGACATCTATGTGGGTTCTAGT AAAAGTGTCCAAATGGACCTATGTAGCTTACAACGAATCTGAGAAGATATATTTTACATTTCCTCTTGAAAATCATAGAAAAATCTACAC TCATGTCTATTGTCAGAGCACTATGCTGGATACAAATAGTTGGATTTTTGCTTGCATAAACAGCACTTCTATGTGCCTGCAAGGGGTTGA TTTATCATGGAAAGCTGAACTGCTGCCAACAATTAAGTATCTGACATTAAGACTTCAAGACTATCCATCTTTGTCTCATCTTGTTGTTTA TGTACCATCTGTTCGTGGAAGTAAGTTTGTTGTAGATTGTGAATTCTTTAAAAAAGAGAAAAGATACATACAGCTTCCTGTAACTCATCT TTTTTCCTTTGGATTGTCTTCAAGGAAAGTGGTGTTAAATACAAATGGCCTATACTACAATCTAGAGCTTCTGAACTTTGGACAGATATA CCAAGCTTTTAAAATCAACGTGGTAAGCAAGTGCTCAGCAGTCAAAGAAGAAATAACCAGTATCTATAGACTTCATATTCCTTGGTCTTA TGAAGATTCACTAACCATTGCACAGGCTCCATCTTCCACAGAAATTTCTCTGAAACTCCATATTGCTCAACCAGAAAACAATACCCATGT GGCATTATTTAAAATGTACACGTCATCAGACTGTCGGTACGAGGTGACAGTTAAGACTTCCTTTTCACAAATACTTGGACAGGTAGTTAG ATTTCATGGTGGAGCTCTTCCTGCTTATGTCGTATCTAATATCCTTCTTGCTTATAGAGGACAGTTATATTCTCTTTTCTCAACAGTTCA ACACAAGATGAGACACATGTGAATACTGGGTCATCGTCTGAAGTGGTGCATTTGGATGATGCTTTTTCTGAAGAAGAGGAAGAGGATGAG GATAAGGTTGAGGATGAGGATGCTACCGAAGAGAGACCATCCGAGGTTTCAGAACCTATTGAAGAGTTACATTCCAGACCTCATAAATCT CAGGAAGGCACGCAGGAGGTTTCAGTTCGACCATCAGTTATTCAAAAACTGGAGAAGGGACAGCAGCAGCCCTTGGAGTTTGTTCATAAA ATTGGGGCCAGTGTGAGAAAATGTAACCTAGTAGATATTGGTCAGGCTACAAATAATAGAAGCAACTTGGTACGCCCCCCAGTGATTTGT AATGCTCCTGCTAGTTGTTTACCTGAAAGCTCTAACGATAGACCAGTTACAGCTAATACAACTAGTTTACCACCAGCAGCTCATTTGGAT TCAGTTAGCAAATGTGACCCAAACAAAGTTGAGAAATATCTTGAACAGCCAGATGGGGCCTCTAGAAATCCTGTGCCATCATCCCATGTA GAAACTACTTCATTTTCGTATCAGAAACATAAAGAATCAAATAGGAAATCTTTACGCATGAATTCAGATAAGTTGGTTTTGTGGAAAGAT GTAAAATCTCAGGGTAAAACTTTATCAGCTGGCTTGAAATTCCATGAACGCATGGGTACTAAGGGCTCCTTAAGAGTTAAATCTCCTTCC AAATTAGCAGTAAACCCGAATAAAACTGACATGCCTTCTAATAAAGGAATCTTTGAAGATACTATTGCAAAGAACCATGAGGAATTCTTT TCTAACATGGATTGTACCCAAGAAGAAAAGCATTTGGTTTTTAACAAGACAGCCTTTTGGGAACAGAAGTGCTCAGTGAGTTCTGAAATG AAGTTTGATTGTATCTCTCTTCAGTCAGCATCTGATCAGCCCCAAGAGACTGCACAAGACTTAAGTCTTTGGAAGGAGGAGCAAATTGAC CAAGAAGATAACTATGAGTCTAGAGGTTCAGAAATGAGTTTTGATTGCAGTTCCTCTTTTCATTCACTGACTGACCAATCTAAAGTGAGT GCCAAAGAAGTAAACCTTTCCAAGGAAGTACGTACTGATGTACAGTATAAGAATAATAAATCTTATGTTTCTAAAATAAGTTCTGATTGT GATGACATTCTTCACTTGGTTACCAACCAATCCCAAATGATTGTTAAAGAAATAAGTCTTCAGAATGCAAGGCATATTAGCCTGGTTGAC CAAAGCTATGAATCTAGTAGTTCTGAAACGAATTTTGATTGTGATGCTTCACCTCAGTCCACTAGTGACTACCCCCAACAATCTGTAACA GAAGTAAACCTTCCTAAGGAAGTGCACATTGGTTTGGTTGATAAGAACTATGGTTCCAGTAGCTCTGAAGTAAGTGCTGATTCTGTTTTC CCACTGCAGTCAGTGGTTGACAGACCCCCAGTGGCTGTCACAGAAACAAAACTTCGGAAGAAGGCTCATACCAGCTTGGTTGATAACTAT GGATCGAGTTGTTCTGAAACAAGTTTTGATTGTGATGTTTCTCTTGAGTCAGTAGTTGATCATCCCCAACTGACTGTCAAAGGAAGAAAC CTGAAAGGTAGACAAGTCCACCTAAAACATAAGAAGCGTAAACCCAGTAGTGCTAAAGCACATCTTGATTGTGATGTCTCACTTGGGACA GTTGCAGATGAATCCCAGAGGGCTGTTGAAAAGATAAATCTTCTGAAGGAGAAGAATGCTGACCTTATGGATATGAACTGTGAATCCCAT GGTCCTGAAATGGGTTTTCAGGCTGATGCTCAATTAGCTGACCAGTCTCAAGTAGCCGAAATAGAGCGTCAGAAAGTGGATGTTGACCTT GAGAATAAGAGTGTTCAGTCTAGCCGTTCTTCTCTGAGTTCTGATTCTCCGGCTTCTCTTTATCATTCAGCTCATGATGAGCCTCAAGAA GCTTTGGATGAAGTAAATCTTAAAGAGTTAAATATTGACATGGAAGTTAGGAGCTATGATTGCTCCAGCTCTGAGTTGACTTTTGATTCT GACCCGCCTCTTCTGTCAGTTACTGAGCAGTCTCATCTGGATGCTGAAGGAAAAGAACGGCACATTGACCTGGAAGATGAGAGCTGTGAG TCAGATAGTTCTGAAATAACTTTTGATTCTGATATTCCTCTTTATTCAGTAATTGACCAACCTGAAGTAGCTGTTTATGAGGAAGAAACT GTTGATCTGGAAAGTAAAAGTAATGAATCTTGTGTCTCTGAAATAACTTTTGATTCTGATATTCCTCTTCATTCAGGAAATGATCACCCT GAAGTAGCTGTTAAAGAAGTAATTCAGAAAGAAGAGTACATTCACTTAGAAAGGAAGAATGATGAACCCAGTGGTTCTGAAATAAGTTCG GATTCCCATGCCCCTCTTCATTCAGTGACTAATTCTCCCGAAGTAGCTGTTAAAAAGCTAAATCCTCAAAAAGAAGAGCAGGTACACTTA GAAAATAAGGAAAATGAACCTATTGATTCTGAAGTAAGTTTGGATTATAATATCATTTTTCATTCAGTGACTGGACGTTCTGAAGATCCC ATTAAAGAAATAAGCCTTCACACAAAAGAGCACATGTACTTAGAAAATAAGAGTGTTTTTGAAACAAGTTTGGATTCTGATGTCCCTCTT CAGGCAGCGACTCACAAACCTGAAGTAATTGTCAAAGAAACATGGCTTCAAAGAGAAAAGCACGCTGAATTCCAAGGTAGAAGTACTGAA TTCAGTGGTTCAAAAACAAGTTTAGATTCTGGTGTCCCTCATTATTCAGTAACTGAACCTCAAGTAGCTGTTAACAAAATAAACAGAAAG AAGCAATATGTTCTAGAAAACAAGAATGATAAATGTAGTGGTTCTGAAATAATTTTGGATTCTAATGTTCCACCTCAGTCAATGACTGAC CAACCTCAACTAGCTTTTTTGAAGGAAAAACATGTTAATCTGAAGGACAAAAACAGTAAATCAGGTGATTCTAAAATAACTTTTGATTCT GAACAACTTCAGGAAGCGGTTAAAAAAATAGACCAATGGAAGGAAGAGGTTATTGGCCTGAAAAATAAGATTAATGAACCTAGTACTTAT AAATTAATACATCATCCTGATGTTTCTGTCCAATCTGTGGCTGATCAACCCAAAGTAGCTATTAAACATGTGAACCTTGGGAATGAAAAC CATATGTACTTGGAAGTTAAGAACAGCCAATATAGTTGTTCAGAAATGAATTTGGATTCTGGTTTCTTGGGTCAGTCAATAGTCAATCGA CCTCAAATAACTATTTTGGAGCAGGAGCACATTGAACTAGAAGGTAAGCACAATCAATGTTGTGGTTCTGAAGTAAGTTTTGATTCTGAT GACCCTCTTCAGTCAGTGGCTGACCGGCTGAGAGAAACCGTTAAAGAAATAAGCCTTTGGAAGGATGAAGAAGTTGACACGGAAGATAGG AGAAATGAAGCTAAGGGTTTTGAAATTATGTATGATTCTGATGTTCTTCAGCCAGTGGCTGGCCAACCTGAAGAAGTAGTTAAGGAGGTC AGTCTTTGGAAAGAGCATGTTGACTTGGAAAATAAGATTGTCAAACCTACAGATTCCAGAATAAATTTTGATTCTCATGAACCCCTTCAG TCCGTAACTAATAAAATTCCAGGGGCGAATAAAGAAATAAATCTTTTGAGGGAGGAACATGTTTGTCTGGATGATAAGGGCTATGTGCCC AGTGATTCTGAAATAATTTATGTTTCAAATATCCCTCTTCAGTCAGTGATAAAACAACCACACATTTTGGAAGAGGAGCATGCCAGTCTG GAAGATAAGAGCAGTAATTCTTATAGTCCTGAAGAAAGTTCTGATTCCAATGACTCTTTTCAGGCAGCAGCAGATGAGCTTCAAAAACCT GTCAAAGAAATAAATCTTTGGAAGGAAGACCATATTTACCTGGAAGATAAGAGCTATAAATTAGGTGATTTTGATGTAAGTTATGCTTCT CATATTCCTGTTCAGTTTGTGACTGATCAATCTTCTGTACCTGTCAAAGAAATAAACTTGCAAAAGAAGGATCATAATGATCTAGAAAAT AAGAACTGTGAAGTCTGTGGTTCTGAAATAAAATGTCATTCTTGTGTTCATCTTCAGTCAGAAGTTGACCAACCTCAAGTGTCTTACAAA GAGGCAGACCTTCAGAAGGAAGAGCATGTTGTCATGGAAGAAAAGACCGATCAACCTAGTGATTCAGAAATGATGTATGATTCTGATGTT CCTTTTCAAATAGTAGTTAACCAATTTCCAGGATCAGTCAAAGAAACCCACCTTCCAAAGGTGGTACTTGTGGATCTGGTGCCCGGTGAT AGTGATTATGAAGTAATTTCAGATGATATTCCCCTTCAGTTAGTGACTGACCCACCTCAGTTGACTGTCAAAGATATCAGCTGTATAAAT ACAGAATGTATTGATATAGAAGATAAGAGCTGTGACTTTTTTGGTTCTGAAGTCAGATGTAATTGTAAAGCCTCTACTCCCTCAATGACA AACCAATGCAAAGAGACTTTCAAAATAATAAACCGGAAGAAGGACTATATTATTCTGGGAGAGCCAAGTTGTCAATCTTGTGGTTCTGAA ATGAATTTTAATGTTGATGCCTCTGATCAGTCCATGACTTACGAGTCACAAGGACCTGATGAGAAAATGGTGAAATATATTGATTCAGAA GATAAGAGCTGTGGATATAATGGTTCTAAAGGAAAATTTAATTTGGAAGACACTTCTCATCGAACGACTCACCGACTGCAGAAAGCTCAC AAAGAAGCCAGTCTTCGGAAGGATCCAAGAAATGCTGGCCTAAAAGGTAAGAGCTGTCAGTCTAGTGCTTCTGCAGTGGATTTTGGTGCC TCTTCCAAGTCAGCGCTCCATCGAAGGGCTGATAAAAAAAAACGTTCGAAGCTAAAACATAGAGATCTAGAAGTGAGCTGTGAACCGGAT GGTTTTGAGATGAATTTTCAGTGTGCTCCCCCTCTTCCATCTGATACTGATCAGCCTCAAGAAACTGTTAAGAAAAGACACCCTTGTAAG AAGGTATCTTCTGACTTGAAAGAAAAGAACCATGATTCCCAGTCAAGCTCTGTTCTCAAGGTTGATTCTGTAAGGAACCTGAAAAAAGCA AAGGATGTCATAGAGGATAATCCTGATGAACCAGTTCTTGAAGCCTTGCCTCATGTACCTCCTTCATTTGTGGGGAAAACATGGTCTCAG ATAATGAGAGAAGATGACATAAAAATTAATGCTCTGGTGAAGGAGTTTAGGGAAGGTCGTTTCCACTGTTACTTTGATGATGACTGTGAG ACCAAAAAAGTTTCTTCGAAGGGGAAAAAAAAGGTTACCTGGGCTGACTTGCAAGGTAAGGAGGACACTGCACCAACTCAAGCTGTGTCA GAGAGTGATGATATTGTCTGTGGTATTTCAGATATTGATGACTTGTCAGTGGCCTTAGATAAACCATGCCATCGTCATCCTCCAGCAGAG AGGCCTCCTAAGCAAAAGGGGCGTGTGGCTTCTCAATGCCAGACAGCGAAAATCAGCCATAGTACTCAGACCAGTTGTAAGAATTACCCA GTGATGAAAAGAAAAATAATTAGACAAGAGGAAGACCCACCAAAAAGTAAGTGTTCACGTTTACAGGATGACAGAAAAACCAAAAAGAAA GTCAAAATTGGGACAGTTGAATTTCCTGCATCATGTACTAAAGTTTTGAAGCCTATGCAACCCAAAGCCTTAGTCTGTGTTCTTTCTTCT TTAAATATTAAACTGAAAGAGGGTGAAGGCCTTCCTTTCCCTAAAATGAGGCACCATAGTTGGGATAATGATATTCGGTTTATATGCAAA TATAAACGGAATATCTTTGATTATTATGAGCCCTTGATTAAGCAAATTGTAATTAGTCCTCCCCTGAGTGTAATAGTACCAGAGTTTGAG AGGCGTAACTGGGTTAAAATTCATTTTAATAGGAGCAACCAAAACTCCAGTGCAGGAGATAATGATGCTGATGGACAAGGCTCTGCTTCA GCGCCTTTAATGGCAGTGCCGGCAAGATATGGATTTAATTCACATCAGGGAACCAGTGACTCTTCTCTGTTTCTGGAAGAATCAAAGGTT CTGCATGCTCGTGAGCTTCCAAAGAAAAGAAATTTCCAGCTAACATTTTTAAATCATGATGTTGTCAAAATCTCTCCAAAATCAGTTAGA AATAAGCTTTTGGAAAGTCAAAGTAAAAAGAAAATTCATGGAAAGAGGGTGACAACTAGTAGTAATAAGCTAGGTTTTCCCAAAAAGGTT TATAAACCAATTATTCTCCAGCAAAAACCCAGAAAAGCTTCAGAGAAACAGTCAATTTGGATTCGGACCAAACCAAGTGATATCATTAGA AAGTATATTTCGAAATACTCTGTCTTTTTACGTCATAGATATCAGTCCAGGAGCGCTTTTCTTGGAAGGTATCTGAAGAAGAAAAAATCT GTTGTCAGTAGGCTAAAGAAGGCGAAGAGAACAGCTAAAGTGCTTTTGAACTCTTCAGTTCCACCAGCTGGTGCCGAAGAGCTGTCAAGC GCTATGGCAAATCCTCCTCCAAAGCGACCTGTGCGGGCTTCTTGCCGCGTTGCAAGAAGGAGGAAGAAGACTGATGAAAGCTACCATGGC CGACAGAAAGGTCCTTCTACACCTGTGAGAGCATATGATCTGAGAAGCTCATCTTGTTTACAACAACGTGAGAGAATGATGACTCGGCTA GCAAACAAACTGAGAGGTAATGAGGTAAAATAGAAGTTGGTTTTGTGTTCAGGCTAGTTGAGGATTCAGTACTTCAGTTGAAATATATTT GGTACTTTGTGCACACAGCTTTCCCAGCTTTGGTGAGAAAACTAATCTTGAACTATTTTGCTATAAATATTATTTTTCAGAATTTTTATA TTCATTTAAAATTAGTGTTTAGAGTCCTTTCATTTTAAGATTCAGAAACAGCCTTTGTCCAACATTTTCTGTAGAAGAACTTCATCTTAA TTTATGTCACAAAATAATTTAGCACCGTATATAGAATTTTGTTCCTCAAATCATATCCCTCTTATTTTTTATGATTATCTCTCTCATGCC AGCAAATTTAACATTGTGAGAATGAATATAGCAAACTTATTAAAAAGAAGGAATTATAATGAATTATGTCAACTTTTGAATGGGGTTGAC TTAAATGAATACAAACAAAATATAAGTTTTCTTTCTTTTTAAAGCATTCCAGAAAATGAGATTCCATAATCTTTAATACAGTTTTTAAAA ACTGGTGATTTTTTTTATGCATAGCACAAACTCCTCCTGCTTCATTTAAATTGGATCTGCAAGTCTCTAACTTTTATCATTTGGACACAT GCACACATACAGTCTCTAGGTTTTTTGGGTTAGCCTTATTAGATCTTAAAGGGGAAAATAATTTTGCCCATGTGTTTAGGCATTCCCCCC CTTTTTTTTAACCACCTTTACTCTCTTCTACATGAAGAGTTGGTGGCTTAATATGAAAATCCAGGTTATTTTGTGGCCTTATCCCCTAAA AATATTTTAGGCATTACCATATCAGTTTTTATTTTTATGTTTATTTATTATTATTATTTTTTGAGACAGAGTCTCACTGTCTTGCCCAGG CTGGAGTGCAGTGGCCTGAACTTAGCTCACCGCAACCTCCACCTCCCGGGTTCAAGCAATTCTCCTGCCTAAGCCTCTCGAGTAGCTGGG ACTACAGGCGTGCGCCACCATGCCCGGCTAATTTTTGTATTTTTAGTAGAGTTGGGGGTTTCACTATGTTGGCCAGGCTGGTCTCAAACT CCTGACCTTGTGGTCCACCCGCCTCAGCCTCCCAAAGTGCTGGGATTACAGTCATGAGCCACCATGCCCAGCCTACCATATCAGTTTTTA ATAACTTGAACTGAAATTATTGGCTTGGTAATATTTGGGCAGGCCATTAATTCTCATTAGAAAATTACTTAATATTTCAAAAAATTGAAT AAAACTGGTAATCTTTGTGGAGGCCCATTTTGCAAGTGGAAACCTGTATTTATACATTAGTTTGTCTTTAACTATTTGTTAACAGATTTT AACAGTTCAAATGAAAGATGTTAACGAGATGCAGTTTGAGTCTGGGATACTCAGAATAGATCCAGAACCTTTATTCAGGGACACTGGTTG AGTTGGTACAGATGCATTCAGTTGTCATTCATTCGTGTTAAGCCAAGATCCTGGATCTTCTATCATTAATGTTAAGAGGAAACATGAATT ACAACTATATTGTATGACTGCCATATTTTATTGAAACTAAGGTCAATGTTGAACCATATCTTGATTTCAGAATTTAAAATGTGTTACCTT AGAAATAATGAAATATAGTGTAACAGTTGGTTTCTAAATGTGGTTTTATTTCAGGGGCCATTATTTAACCTGTGTGTATGCTATGATGGA GTTCTTTTGTGATTTCTTTACTTCTAAATTTAAATAGTAATTATGTTTAAAAACAAATGTTTTTGTCTTTAGTTGCAAAGCAGCATATAT CTCATTTCCTTCAACCTTGCCTCCTCATACACAAATATATTTTCATTTCTTAAAAGATCACTGGGCCAGTAGTTCAAAATAATCCTCCAA TTTTAATTTAGGAACAAAGCTTTGTCCTACTGACTAGGAAGTAAACAAACAGCTGATATATAAGATATTTGTGGGTATTGGTGTCTTTTT TCCCTATGGAATAGACACATAGTTCCCTTAAATATTATTAATAGACATTCTACAGTCTTTCAGTGTAAGTTTAAAAGCAAGAGATCAAAT GAAAATGTGTACTATGTACCTATGATGTGTCATATCGTGATTACATCATGCAGAGAGAACAGTTCCTGTAATTCAAGTAATTAAAATTTC AAGTTGTAAAAATGTTACCATTTTCACATTGGAAGCCCTTCTAAGTACCATTTTCATTGGGGGCAGGGAGAAGGTTTATATTTAAATTTT ATATTACCAGACATAAAAGTACAGTGCAGTACTGATGATTTTGTTTAAAGTTTTTGAAATGTTTTGTTTACTAAGTTCTTGTAATCATTA TTTTAGACAAGCTTGAAAACTTGAAATTGTTGAAACTTATAATTTAAATTAAATTGGAAATTACTTTCAGAATTTGGTAGTATACAAGAT GTGCCCCTTTGTCTAAATAAATGTTGGCTTAATTTGCAGAATAGATAAAACAATTGATGTTTTTGTGTGTTCAAGAATGCATTAAATTCT TTTAATAGTAAAAAGTCAAATTACGACCATAATGAGTATTTTAATCTTGTTCTTGTTCAGTATTTTGCGTAGATTTTTATTGTAATTGTC CTGAGTTCTACAGTATGTGAACAATATCGTGTGAAGTGTGTTTTTGCATTTGTGCATTTAAATTATTTATGTAACTAGGGCTGTGAGTAA CACTTTTTTTTTTTTTTTTTGGTTAAAGAAAAGCTATATTTAAATATTCAAATAATTATATATTTCCATGTAAAACTAATGTGAAGACTA >64542_64542_1_PGAP1-ZDBF2_PGAP1_chr2_197729711_ENST00000354764_ZDBF2_chr2_207169441_ENST00000374423_length(amino acids)=2273AA_BP= MDDAFSEEEEEDEDKVEDEDATEERPSEVSEPIEELHSRPHKSQEGTQEVSVRPSVIQKLEKGQQQPLEFVHKIGASVRKCNLVDIGQAT NNRSNLVRPPVICNAPASCLPESSNDRPVTANTTSLPPAAHLDSVSKCDPNKVEKYLEQPDGASRNPVPSSHVETTSFSYQKHKESNRKS LRMNSDKLVLWKDVKSQGKTLSAGLKFHERMGTKGSLRVKSPSKLAVNPNKTDMPSNKGIFEDTIAKNHEEFFSNMDCTQEEKHLVFNKT AFWEQKCSVSSEMKFDCISLQSASDQPQETAQDLSLWKEEQIDQEDNYESRGSEMSFDCSSSFHSLTDQSKVSAKEVNLSKEVRTDVQYK NNKSYVSKISSDCDDILHLVTNQSQMIVKEISLQNARHISLVDQSYESSSSETNFDCDASPQSTSDYPQQSVTEVNLPKEVHIGLVDKNY GSSSSEVSADSVFPLQSVVDRPPVAVTETKLRKKAHTSLVDNYGSSCSETSFDCDVSLESVVDHPQLTVKGRNLKGRQVHLKHKKRKPSS AKAHLDCDVSLGTVADESQRAVEKINLLKEKNADLMDMNCESHGPEMGFQADAQLADQSQVAEIERQKVDVDLENKSVQSSRSSLSSDSP ASLYHSAHDEPQEALDEVNLKELNIDMEVRSYDCSSSELTFDSDPPLLSVTEQSHLDAEGKERHIDLEDESCESDSSEITFDSDIPLYSV IDQPEVAVYEEETVDLESKSNESCVSEITFDSDIPLHSGNDHPEVAVKEVIQKEEYIHLERKNDEPSGSEISSDSHAPLHSVTNSPEVAV KKLNPQKEEQVHLENKENEPIDSEVSLDYNIIFHSVTGRSEDPIKEISLHTKEHMYLENKSVFETSLDSDVPLQAATHKPEVIVKETWLQ REKHAEFQGRSTEFSGSKTSLDSGVPHYSVTEPQVAVNKINRKKQYVLENKNDKCSGSEIILDSNVPPQSMTDQPQLAFLKEKHVNLKDK NSKSGDSKITFDSEQLQEAVKKIDQWKEEVIGLKNKINEPSTYKLIHHPDVSVQSVADQPKVAIKHVNLGNENHMYLEVKNSQYSCSEMN LDSGFLGQSIVNRPQITILEQEHIELEGKHNQCCGSEVSFDSDDPLQSVADRLRETVKEISLWKDEEVDTEDRRNEAKGFEIMYDSDVLQ PVAGQPEEVVKEVSLWKEHVDLENKIVKPTDSRINFDSHEPLQSVTNKIPGANKEINLLREEHVCLDDKGYVPSDSEIIYVSNIPLQSVI KQPHILEEEHASLEDKSSNSYSPEESSDSNDSFQAAADELQKPVKEINLWKEDHIYLEDKSYKLGDFDVSYASHIPVQFVTDQSSVPVKE INLQKKDHNDLENKNCEVCGSEIKCHSCVHLQSEVDQPQVSYKEADLQKEEHVVMEEKTDQPSDSEMMYDSDVPFQIVVNQFPGSVKETH LPKVVLVDLVPGDSDYEVISDDIPLQLVTDPPQLTVKDISCINTECIDIEDKSCDFFGSEVRCNCKASTPSMTNQCKETFKIINRKKDYI ILGEPSCQSCGSEMNFNVDASDQSMTYESQGPDEKMVKYIDSEDKSCGYNGSKGKFNLEDTSHRTTHRLQKAHKEASLRKDPRNAGLKGK SCQSSASAVDFGASSKSALHRRADKKKRSKLKHRDLEVSCEPDGFEMNFQCAPPLPSDTDQPQETVKKRHPCKKVSSDLKEKNHDSQSSS VLKVDSVRNLKKAKDVIEDNPDEPVLEALPHVPPSFVGKTWSQIMREDDIKINALVKEFREGRFHCYFDDDCETKKVSSKGKKKVTWADL QGKEDTAPTQAVSESDDIVCGISDIDDLSVALDKPCHRHPPAERPPKQKGRVASQCQTAKISHSTQTSCKNYPVMKRKIIRQEEDPPKSK CSRLQDDRKTKKKVKIGTVEFPASCTKVLKPMQPKALVCVLSSLNIKLKEGEGLPFPKMRHHSWDNDIRFICKYKRNIFDYYEPLIKQIV ISPPLSVIVPEFERRNWVKIHFNRSNQNSSAGDNDADGQGSASAPLMAVPARYGFNSHQGTSDSSLFLEESKVLHARELPKKRNFQLTFL NHDVVKISPKSVRNKLLESQSKKKIHGKRVTTSSNKLGFPKKVYKPIILQQKPRKASEKQSIWIRTKPSDIIRKYISKYSVFLRHRYQSR SAFLGRYLKKKKSVVSRLKKAKRTAKVLLNSSVPPAGAEELSSAMANPPPKRPVRASCRVARRRKKTDESYHGRQKGPSTPVRAYDLRSS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PGAP1-ZDBF2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PGAP1-ZDBF2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PGAP1-ZDBF2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
