
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PNKP-FZR1 (FusionGDB2 ID:66727) |
Fusion Gene Summary for PNKP-FZR1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PNKP-FZR1 | Fusion gene ID: 66727 | Hgene | Tgene | Gene symbol | PNKP | FZR1 | Gene ID | 11284 | 51343 |
| Gene name | polynucleotide kinase 3'-phosphatase | fizzy and cell division cycle 20 related 1 | |
| Synonyms | AOA4|EIEE10|MCSZ|PNK | CDC20C|CDH1|FZR|FZR2|HCDH|HCDH1 | |
| Cytomap | 19q13.33 | 19p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | bifunctional polynucleotide phosphatase/kinaseDNA 5'-kinase/3'-phosphataseHomo sapiens polynucleotide kinase 3'-phosphatase (PNKP) | fizzy-related protein homologCDC20 homolog 1CDC20-like 1bCDC20-like protein 1cdh1/Hct1 homologfizzy/cell division cycle 20 related 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q9UM11 | |
| Ensembl transtripts involved in fusion gene | ENST00000322344, ENST00000600573, ENST00000600910, ENST00000596014, ENST00000595792, | ENST00000313639, ENST00000395095, ENST00000441788, | |
| Fusion gene scores | * DoF score | 6 X 3 X 4=72 | 6 X 5 X 4=120 |
| # samples | 6 | 6 | |
| ** MAII score | log2(6/72*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/120*10)=-1 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PNKP [Title/Abstract] AND FZR1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PNKP(50369656)-FZR1(3525866), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | PNKP-FZR1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. PNKP-FZR1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. PNKP-FZR1 seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | PNKP | GO:0006979 | response to oxidative stress | 10446192 |
| Hgene | PNKP | GO:0016311 | dephosphorylation | 10446193 |
| Hgene | PNKP | GO:0042769 | DNA damage response, detection of DNA damage | 10446192 |
| Hgene | PNKP | GO:0046939 | nucleotide phosphorylation | 10446193 |
| Tgene | FZR1 | GO:0031145 | anaphase-promoting complex-dependent catabolic process | 18662541|21596315 |
| Tgene | FZR1 | GO:0072425 | signal transduction involved in G2 DNA damage checkpoint | 18662541 |
| Tgene | FZR1 | GO:1904668 | positive regulation of ubiquitin protein ligase activity | 11459826 |
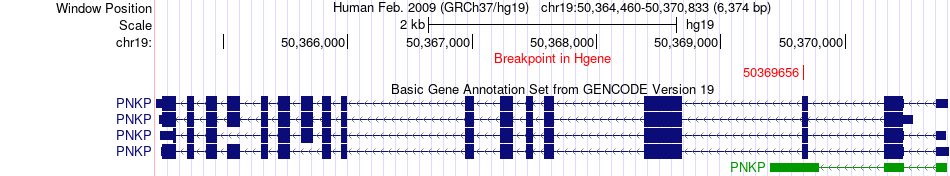
 Fusion gene breakpoints across PNKP (5'-gene) Fusion gene breakpoints across PNKP (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
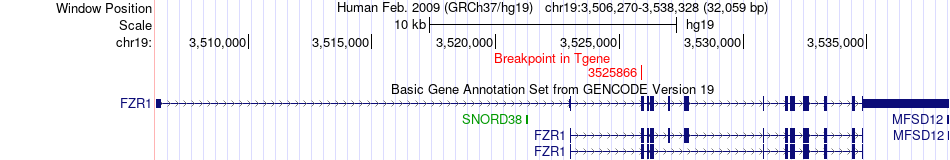
 Fusion gene breakpoints across FZR1 (3'-gene) Fusion gene breakpoints across FZR1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-BR-A4PD-01A | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
Top |
Fusion Gene ORF analysis for PNKP-FZR1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000322344 | ENST00000313639 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000322344 | ENST00000395095 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000322344 | ENST00000441788 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000600573 | ENST00000313639 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000600573 | ENST00000395095 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000600573 | ENST00000441788 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000600910 | ENST00000313639 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000600910 | ENST00000395095 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| Frame-shift | ENST00000600910 | ENST00000441788 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| In-frame | ENST00000596014 | ENST00000313639 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| In-frame | ENST00000596014 | ENST00000395095 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| In-frame | ENST00000596014 | ENST00000441788 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| intron-3CDS | ENST00000595792 | ENST00000313639 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| intron-3CDS | ENST00000595792 | ENST00000395095 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
| intron-3CDS | ENST00000595792 | ENST00000441788 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000596014 | PNKP | chr19 | 50369656 | - | ENST00000441788 | FZR1 | chr19 | 3525866 | + | 5187 | 280 | 58 | 1692 | 544 |
| ENST00000596014 | PNKP | chr19 | 50369656 | - | ENST00000313639 | FZR1 | chr19 | 3525866 | + | 1426 | 280 | 58 | 1425 | 456 |
| ENST00000596014 | PNKP | chr19 | 50369656 | - | ENST00000395095 | FZR1 | chr19 | 3525866 | + | 1702 | 280 | 58 | 1701 | 548 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000596014 | ENST00000441788 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + | 0.03441717 | 0.96558285 |
| ENST00000596014 | ENST00000313639 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + | 0.12959912 | 0.87040085 |
| ENST00000596014 | ENST00000395095 | PNKP | chr19 | 50369656 | - | FZR1 | chr19 | 3525866 | + | 0.06925461 | 0.93074536 |
Top |
Fusion Genomic Features for PNKP-FZR1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| PNKP | chr19 | 50369655 | - | FZR1 | chr19 | 3525865 | + | 0.001547228 | 0.9984528 |
| PNKP | chr19 | 50369655 | - | FZR1 | chr19 | 3525865 | + | 0.001547228 | 0.9984528 |
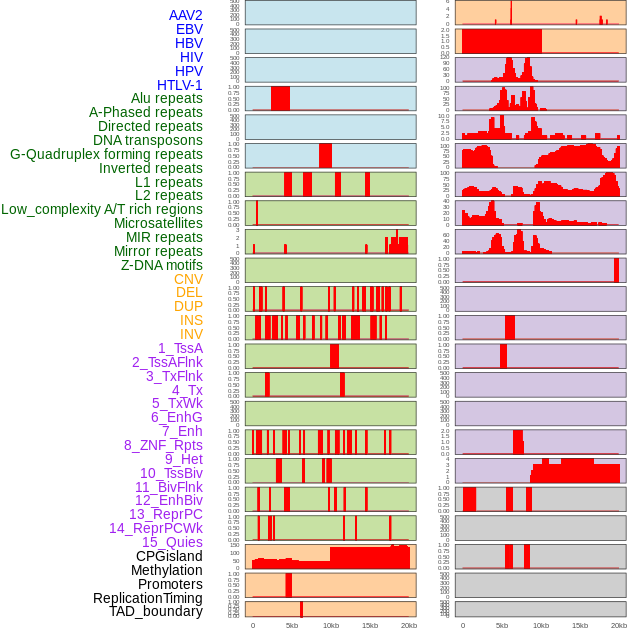
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for PNKP-FZR1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:50369656/chr19:3525866) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | FZR1 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Substrate-specific adapter for the anaphase promoting complex/cyclosome (APC/C) E3 ubiquitin-protein ligase complex. Associates with the APC/C in late mitosis, in replacement of CDC20, and activates the APC/C during anaphase and telophase. The APC/C remains active in degrading substrates to ensure that positive regulators of the cell cycle do not accumulate prematurely. At the G1/S transition FZR1 is phosphorylated, leading to its dissociation from the APC/C. Following DNA damage, it is required for the G2 DNA damage checkpoint: its dephosphorylation and reassociation with the APC/C leads to the ubiquitination of PLK1, preventing entry into mitosis. Acts as an adapter for APC/C to target the DNA-end resection factor RBBP8/CtIP for ubiquitination and subsequent proteasomal degradation. Through the regulation of RBBP8/CtIP protein turnover, may play a role in DNA damage response, favoring DNA double-strand repair through error-prone non-homologous end joining (NHEJ) over error-free, RBBP8-mediated homologous recombination (HR) (PubMed:25349192). {ECO:0000269|PubMed:18662541, ECO:0000269|PubMed:21596315, ECO:0000269|PubMed:25349192, ECO:0000269|PubMed:9734353}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 47_52 | 23 | 405.0 | Region | Involved in APC/FZR1 E3 ubiquitin-protein ligase complex activity | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 47_52 | 23 | 497.0 | Region | Involved in APC/FZR1 E3 ubiquitin-protein ligase complex activity | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 47_52 | 23 | 494.0 | Region | Involved in APC/FZR1 E3 ubiquitin-protein ligase complex activity | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 182_222 | 23 | 405.0 | Repeat | Note=WD 1 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 227_266 | 23 | 405.0 | Repeat | Note=WD 2 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 269_306 | 23 | 405.0 | Repeat | Note=WD 3 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 311_350 | 23 | 405.0 | Repeat | Note=WD 4 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 353_395 | 23 | 405.0 | Repeat | Note=WD 5 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 397_438 | 23 | 405.0 | Repeat | Note=WD 6 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000313639 | 0 | 11 | 441_480 | 23 | 405.0 | Repeat | Note=WD 7 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 182_222 | 23 | 497.0 | Repeat | Note=WD 1 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 227_266 | 23 | 497.0 | Repeat | Note=WD 2 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 269_306 | 23 | 497.0 | Repeat | Note=WD 3 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 311_350 | 23 | 497.0 | Repeat | Note=WD 4 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 353_395 | 23 | 497.0 | Repeat | Note=WD 5 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 397_438 | 23 | 497.0 | Repeat | Note=WD 6 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000395095 | 0 | 13 | 441_480 | 23 | 497.0 | Repeat | Note=WD 7 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 182_222 | 23 | 494.0 | Repeat | Note=WD 1 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 227_266 | 23 | 494.0 | Repeat | Note=WD 2 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 269_306 | 23 | 494.0 | Repeat | Note=WD 3 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 311_350 | 23 | 494.0 | Repeat | Note=WD 4 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 353_395 | 23 | 494.0 | Repeat | Note=WD 5 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 397_438 | 23 | 494.0 | Repeat | Note=WD 6 | |
| Tgene | FZR1 | chr19:50369656 | chr19:3525866 | ENST00000441788 | 1 | 14 | 441_480 | 23 | 494.0 | Repeat | Note=WD 7 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000322344 | - | 3 | 17 | 6_110 | 66 | 522.0 | Domain | Note=FHA |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000596014 | - | 2 | 16 | 6_110 | 66 | 522.0 | Domain | Note=FHA |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000322344 | - | 3 | 17 | 372_379 | 66 | 522.0 | Nucleotide binding | ATP |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000596014 | - | 2 | 16 | 372_379 | 66 | 522.0 | Nucleotide binding | ATP |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000322344 | - | 3 | 17 | 146_337 | 66 | 522.0 | Region | Phosphatase |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000322344 | - | 3 | 17 | 341_516 | 66 | 522.0 | Region | Kinase |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000596014 | - | 2 | 16 | 146_337 | 66 | 522.0 | Region | Phosphatase |
| Hgene | PNKP | chr19:50369656 | chr19:3525866 | ENST00000596014 | - | 2 | 16 | 341_516 | 66 | 522.0 | Region | Kinase |
Top |
Fusion Gene Sequence for PNKP-FZR1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >66727_66727_1_PNKP-FZR1_PNKP_chr19_50369656_ENST00000596014_FZR1_chr19_3525866_ENST00000313639_length(transcript)=1426nt_BP=280nt GGAGGAGGCGGGACTTCCTGCAGCAAATTGGGGCTGTGCGCCGCTCAAGCCCGTTTACCTGCTCCCCAGGCCGGCACCCAGGATGGGCGA GGTGGAGGCCCCGGGCCGCTTGTGGCTCGAGAGCCCCCCTGGGGGAGCGCCCCCCATCTTCCTGCCCTCGGACGGGCAAGCCCTGGTCCT GGGCAGGGGACCCCTGACCCAGGTTACGGACCGGAAGTGCTCCAGAACTCAAGTGGAGCTGGTCGCAGATCCTGAGACCCGGACAGTGGC AGTGAAACAGGTCACAGAGATGCGGCGGACCCTGACGCCTGCCAGCTCCCCAGTGTCCTCGCCCAGCAAGCACGGAGACCGCTTCATCCC CTCCAGAGCCGGAGCCAACTGGAGCGTGAACTTCCACAGGATTAACGAGAATGAGAAGTCTCCCAGTCAGAACCGGAAAGCCAAGGACGC CACCTCAGACAACGGCAAAGACGGCCTGGCCTACTCTGCCCTGCTCAAGAATGAGCTGCTGGGTGCCGGCATCGAGAAGGTGCAGGACCC GCAGACTGAGGACCGCAGGCTGCAGCCCTCCACGCCTGAGAAGAAGGGTCTGTTCACGGTGACGCGGCTCTGTGACCTCTCAGTGGAAGG GGACTCAGTGACCTCCGTGGGCTGGTCTGAGCGGGGGAACCTGGTGGCGGTGGGCACACACAAGGGCTTCGTGCAGATCTGGGACGCAGC CGCAGGGAAGAAGCTGTCCATGTTGGAGGGCCACACGGCACGCGTCGGGGCGCTGGCCTGGAATGCTGAGCAGCTGTCGTCCGGGAGCCG CGACCGCATGATCCTGCAGAGGGACATCCGCACCCCGCCACTGCAGTCGGAGCGGCGGCTGCAGGGCCACCGGCAGGAGGTGTGCGGGCT CAAGTGGTCCACAGACCACCAGCTCCTCGCCTCGGGGGGCAACGACAACAAGCTGCTGGTCTGGAATCACTCGAGCCTGAGCCCCGTGCA GCAGTACACGGAGCACCTGGCGGCCGTGAAGGCCATCGCCTGGTCCCCACATCAGCACGGGCTGCTGGCCTCGGGGGGCGGCACAGCTGA CCGCTGTATCCGCTTCTGGAACACGCTGACAGGACAACCACTGCAGTGTATCGACACGGGCTCCCAAGTGTGCAATCTGGCCTGGTCCAA GCACGCCAACGAGCTGGTGAGCACGCACGGCTACTCACAGAACCAGATCCTTGTCTGGAAGTACCCCTCCCTGACCCAGGTGGCCAAGCT GACCGGGCACTCCTACCGCGTGCTGTACCTGGCAATGTCCCCTGATGGGGAGGCCATCGTCACTGGTGCTGGAGACGAGACCCTGAGGTT >66727_66727_1_PNKP-FZR1_PNKP_chr19_50369656_ENST00000596014_FZR1_chr19_3525866_ENST00000313639_length(amino acids)=456AA_BP=73 MLPRPAPRMGEVEAPGRLWLESPPGGAPPIFLPSDGQALVLGRGPLTQVTDRKCSRTQVELVADPETRTVAVKQVTEMRRTLTPASSPVS SPSKHGDRFIPSRAGANWSVNFHRINENEKSPSQNRKAKDATSDNGKDGLAYSALLKNELLGAGIEKVQDPQTEDRRLQPSTPEKKGLFT VTRLCDLSVEGDSVTSVGWSERGNLVAVGTHKGFVQIWDAAAGKKLSMLEGHTARVGALAWNAEQLSSGSRDRMILQRDIRTPPLQSERR LQGHRQEVCGLKWSTDHQLLASGGNDNKLLVWNHSSLSPVQQYTEHLAAVKAIAWSPHQHGLLASGGGTADRCIRFWNTLTGQPLQCIDT GSQVCNLAWSKHANELVSTHGYSQNQILVWKYPSLTQVAKLTGHSYRVLYLAMSPDGEAIVTGAGDETLRFWNVFSKTRSTKESVSVLNL -------------------------------------------------------------- >66727_66727_2_PNKP-FZR1_PNKP_chr19_50369656_ENST00000596014_FZR1_chr19_3525866_ENST00000395095_length(transcript)=1702nt_BP=280nt GGAGGAGGCGGGACTTCCTGCAGCAAATTGGGGCTGTGCGCCGCTCAAGCCCGTTTACCTGCTCCCCAGGCCGGCACCCAGGATGGGCGA GGTGGAGGCCCCGGGCCGCTTGTGGCTCGAGAGCCCCCCTGGGGGAGCGCCCCCCATCTTCCTGCCCTCGGACGGGCAAGCCCTGGTCCT GGGCAGGGGACCCCTGACCCAGGTTACGGACCGGAAGTGCTCCAGAACTCAAGTGGAGCTGGTCGCAGATCCTGAGACCCGGACAGTGGC AGTGAAACAGGTCACAGAGATGCGGCGGACCCTGACGCCTGCCAGCTCCCCAGTGTCCTCGCCCAGCAAGCACGGAGACCGCTTCATCCC CTCCAGAGCCGGAGCCAACTGGAGCGTGAACTTCCACAGGATTAACGAGAATGAGAAGTCTCCCAGTCAGAACCGGAAAGCCAAGGACGC CACCTCAGACAACGGCAAAGACGGCCTGGCCTACTCTGCCCTGCTCAAGAATGAGCTGCTGGGTGCCGGCATCGAGAAGGTGCAGGACCC GCAGACTGAGGACCGCAGGCTGCAGCCCTCCACGCCTGAGAAGAAGGGTCTGTTCACGTATTCCCTTAGCACCAAGCGCTCCAGCCCCGA TGACGGCAACGATGTGTCTCCCTACTCCCTGTCTCCCGTCAGCAACAAGAGCCAGAAGCTGCTCCGGTCCCCCCGGAAACCCACCCGCAA GATCTCCAAGATCCCCTTCAAGGTGCTGGACGCGCCCGAGCTGCAGGACGACTTCTACCTCAATCTGGTGGACTGGTCGTCCCTCAATGT GCTCAGCGTGGGGCTAGGCACCTGCGTGTACCTGTGGAGTGCCTGTACCAGCCAGGTGACGCGGCTCTGTGACCTCTCAGTGGAAGGGGA CTCAGTGACCTCCGTGGGCTGGTCTGAGCGGGGGAACCTGGTGGCGGTGGGCACACACAAGGGCTTCGTGCAGATCTGGGACGCAGCCGC AGGGAAGAAGCTGTCCATGTTGGAGGGCCACACGGCACGCGTCGGGGCGCTGGCCTGGAATGCTGAGCAGCTGTCGTCCGGGAGCCGCGA CCGCATGATCCTGCAGAGGGACATCCGCACCCCGCCACTGCAGTCGGAGCGGCGGCTGCAGGGCCACCGGCAGGAGGTGTGCGGGCTCAA GTGGTCCACAGACCACCAGCTCCTCGCCTCGGGGGGCAACGACAACAAGCTGCTGGTCTGGAATCACTCGAGCCTGAGCCCCGTGCAGCA GTACACGGAGCACCTGGCGGCCGTGAAGGCCATCGCCTGGTCCCCACATCAGCACGGGCTGCTGGCCTCGGGGGGCGGCACAGCTGACCG CTGTATCCGCTTCTGGAACACGCTGACAGGACAACCACTGCAGTGTATCGACACGGGCTCCCAAGTGTGCAATCTGGCCTGGTCCAAGCA CGCCAACGAGCTGGTGAGCACGCACGGCTACTCACAGAACCAGATCCTTGTCTGGAAGTACCCCTCCCTGACCCAGGTGGCCAAGCTGAC CGGGCACTCCTACCGCGTGCTGTACCTGGCAATGTCCCCTGATGGGGAGGCCATCGTCACTGGTGCTGGAGACGAGACCCTGAGGTTCTG >66727_66727_2_PNKP-FZR1_PNKP_chr19_50369656_ENST00000596014_FZR1_chr19_3525866_ENST00000395095_length(amino acids)=548AA_BP=73 MLPRPAPRMGEVEAPGRLWLESPPGGAPPIFLPSDGQALVLGRGPLTQVTDRKCSRTQVELVADPETRTVAVKQVTEMRRTLTPASSPVS SPSKHGDRFIPSRAGANWSVNFHRINENEKSPSQNRKAKDATSDNGKDGLAYSALLKNELLGAGIEKVQDPQTEDRRLQPSTPEKKGLFT YSLSTKRSSPDDGNDVSPYSLSPVSNKSQKLLRSPRKPTRKISKIPFKVLDAPELQDDFYLNLVDWSSLNVLSVGLGTCVYLWSACTSQV TRLCDLSVEGDSVTSVGWSERGNLVAVGTHKGFVQIWDAAAGKKLSMLEGHTARVGALAWNAEQLSSGSRDRMILQRDIRTPPLQSERRL QGHRQEVCGLKWSTDHQLLASGGNDNKLLVWNHSSLSPVQQYTEHLAAVKAIAWSPHQHGLLASGGGTADRCIRFWNTLTGQPLQCIDTG SQVCNLAWSKHANELVSTHGYSQNQILVWKYPSLTQVAKLTGHSYRVLYLAMSPDGEAIVTGAGDETLRFWNVFSKTRSTKVKWESVSVL -------------------------------------------------------------- >66727_66727_3_PNKP-FZR1_PNKP_chr19_50369656_ENST00000596014_FZR1_chr19_3525866_ENST00000441788_length(transcript)=5187nt_BP=280nt GGAGGAGGCGGGACTTCCTGCAGCAAATTGGGGCTGTGCGCCGCTCAAGCCCGTTTACCTGCTCCCCAGGCCGGCACCCAGGATGGGCGA GGTGGAGGCCCCGGGCCGCTTGTGGCTCGAGAGCCCCCCTGGGGGAGCGCCCCCCATCTTCCTGCCCTCGGACGGGCAAGCCCTGGTCCT GGGCAGGGGACCCCTGACCCAGGTTACGGACCGGAAGTGCTCCAGAACTCAAGTGGAGCTGGTCGCAGATCCTGAGACCCGGACAGTGGC AGTGAAACAGGTCACAGAGATGCGGCGGACCCTGACGCCTGCCAGCTCCCCAGTGTCCTCGCCCAGCAAGCACGGAGACCGCTTCATCCC CTCCAGAGCCGGAGCCAACTGGAGCGTGAACTTCCACAGGATTAACGAGAATGAGAAGTCTCCCAGTCAGAACCGGAAAGCCAAGGACGC CACCTCAGACAACGGCAAAGACGGCCTGGCCTACTCTGCCCTGCTCAAGAATGAGCTGCTGGGTGCCGGCATCGAGAAGGTGCAGGACCC GCAGACTGAGGACCGCAGGCTGCAGCCCTCCACGCCTGAGAAGAAGGGTCTGTTCACGTATTCCCTTAGCACCAAGCGCTCCAGCCCCGA TGACGGCAACGATGTGTCTCCCTACTCCCTGTCTCCCGTCAGCAACAAGAGCCAGAAGCTGCTCCGGTCCCCCCGGAAACCCACCCGCAA GATCTCCAAGATCCCCTTCAAGGTGCTGGACGCGCCCGAGCTGCAGGACGACTTCTACCTCAATCTGGTGGACTGGTCGTCCCTCAATGT GCTCAGCGTGGGGCTAGGCACCTGCGTGTACCTGTGGAGTGCCTGTACCAGCCAGGTGACGCGGCTCTGTGACCTCTCAGTGGAAGGGGA CTCAGTGACCTCCGTGGGCTGGTCTGAGCGGGGGAACCTGGTGGCGGTGGGCACACACAAGGGCTTCGTGCAGATCTGGGACGCAGCCGC AGGGAAGAAGCTGTCCATGTTGGAGGGCCACACGGCACGCGTCGGGGCGCTGGCCTGGAATGCTGAGCAGCTGTCGTCCGGGAGCCGCGA CCGCATGATCCTGCAGAGGGACATCCGCACCCCGCCACTGCAGTCGGAGCGGCGGCTGCAGGGCCACCGGCAGGAGGTGTGCGGGCTCAA GTGGTCCACAGACCACCAGCTCCTCGCCTCGGGGGGCAACGACAACAAGCTGCTGGTCTGGAATCACTCGAGCCTGAGCCCCGTGCAGCA GTACACGGAGCACCTGGCGGCCGTGAAGGCCATCGCCTGGTCCCCACATCAGCACGGGCTGCTGGCCTCGGGGGGCGGCACAGCTGACCG CTGTATCCGCTTCTGGAACACGCTGACAGGACAACCACTGCAGTGTATCGACACGGGCTCCCAAGTGTGCAATCTGGCCTGGTCCAAGCA CGCCAACGAGCTGGTGAGCACGCACGGCTACTCACAGAACCAGATCCTTGTCTGGAAGTACCCCTCCCTGACCCAGGTGGCCAAGCTGAC CGGGCACTCCTACCGCGTGCTGTACCTGGCAATGTCCCCTGATGGGGAGGCCATCGTCACTGGTGCTGGAGACGAGACCCTGAGGTTCTG GAACGTCTTTAGCAAAACCCGTTCGACAAAGGAGTCTGTGTCTGTGCTCAACCTCTTCACCAGGATCCGGTAAACCTGCCGGGCAGGACC GTGCCACACCAGCTGTCCAGAGTCGGAGGACCCCAGCTCCTCAGCTTGCATGGACTCTGCCTTCCCAGCGCTTGTCCCCCGAGGAAGGCG GCTGGGCGGGCGGGGAGCTGGGCCTGGAGGATCCTGGAGTCTCATTAAATGCCTGATTGTGAACCATGTCCACCAGTATCTGGGGTGGGC ACGTGGTCGGGGACCCTCAGCAGCAGGGGCTCTGTCTCCCTTCCCAAAGGGCGAGAACCACATTGGACGGTCCCGGCTCAGACCGTCTGT ACTCAGAGCGACGGATGCCCCCTGGGACCCTCACTGCCTCCGTCTGTTCATCACCTGCCCACCGGAGCCGCATGCTCTTCCTGGAACTGC CCACGTCTGCACAGAACAGACCACCAGACGCCAGGGCTGATTGGTGGGGGCCTGAGACCCCGGTTGCCCATTCATGGCTGCACCCCACCA TGTCAAACCCAAGACCAGCCCCAAGGCCAGACCAAGGCATGTAGGCCTGGGCAGGTGGCTCGGGGCCACTGGCGGAGCCAGCCTGTGGAT CCAAGAGACAGTCCCCACCTGGGCTTCACGGCATCCTTGCAGCCACCTCTGCTGTCACTGCTCGAAGCAGCAGTCTCTCTGGAAGCATCT GTGTCATGGCCATCGCCCGGCGGTCAGTGGGCTTCAGATGGGCCTGTGCATCCTGGCCAAGCGTCACCCTCACACTGGAGGAGGATGTCT GCTCTGGACTTATCACCCCAGGAGAACTGAACCCGGACCTGCTCACTGCCCTGGCTGGAGAGGAGCACAACAGATGCCACGTCTTCGTGC ATTCGCCAACACGTGCCCTCACAGGGCCAGCGTCCTCCTTCCCTGCGCAAGACTTGCGTCCCCCATGCCTGCTGGGTGGCTGGGTCCTGT GGAGGCCAGCAGCGGTGTGGCCCCCGCCCCCAGGCTGCCTGTGTCTTCACCTGTCCTGTCCACCAGCGCCAACAGCCGTGGGGAAGCCAA GGAGACCCAAGGGGTCCAGGAGGTGGGCGCCCTCCATCCTTCGAGAAGCTTCCCAGGCTCCTCTGCTTCTCTGTCTCATGCTCCCAGGCT GCACAGCAGGCAGGGAGGGAGGCAAGGCAGGGGAGTGGGGCCTGAGCTGAGCACTGCCCCCTCACCCCCCCACCACCCCTTCCCATTTCA TCGGTGGGGACGTGGAGAGGGTGGGGCGGGCTGGGGTTGGAGGGTCCCACCCACCACCCTGCTGTGCTTGGGAACCCCCACTCCCCACTC CCCACATCCCAACATCCTGGTGTCTGTCCCCAGTGGGGTTGGCGTGCATGTGTACATATGTATTTGTGACTTTTCTTTGGATTTGTTTTG TGTTTTTGTTGACTAGTCCTGGAAATGTTTGAGGCTAGACGGGGAGGGGCCAGGACCCACCCACTGCTCCTGGGGGATGAGGTCCTGGTT TTAAAGCCCCGTCATTTCAAGCGGGTCGATCTTCCACATTCACTGGAGAGACTCTCCCCACCTCTGTCTGGGTGGGGCGCGGACCCCTCA CTGTGCGCCTGTGCAGGGGGTGCTGGTGCACGTGGCAGTGTGGATTTCCAGTGGTCACGGTCTTACTGTTTCAAGGTTTTTAAATAAGAA AACCAACCCTGCCTTCGCCCATGCCCGCCCCTGCCCGCAGTTGCCAAAGAGCCGCCTTGTCGCTGTGGGCGTCAGGGCTTGGCTGGCTCA GTGCACAACCCACAGTGGCCTTCAGAGGCTCCTCCTGGGACTGGGAACCGCCGCAGGGCCAGGCGGACGGCGTGAGGTTTGTGTTGGGGC TGGTTCTGCCCATGCTAGGGGGTGGGGGAGCTCCCAGGACAGACCAGCCTTGTTTCTCATGTAATGCAGTGACGCTGTCATTAAACACGT GGATTCATGTGTGGCCGGGACTGGCTGGCTCTAGGTCCCCGGCTCGGGTGGGGTCACACGGTCCTGCCCTAGAGTCCCCATCTGGCCCTG GAGCTGCAGAAGCAGCTTCTGAGGGGCTTCCCAGGCCTGCATTTCACAGATGGGGAGCTCAGCCCTCGAAGGCCGCAGAGACGCCTCCCA GGCCCGTCTGCCAGGGCGCCGGCCACAATCCTGCAGGGCCAAGGACTGGACTCCAGGCAAGTCCCTGCGCTCCAGCTGGACGGCCCTGTT CCAGGGAGGAGGTGCTCGGTTGACACCATCAGGGAGGGAGGGTGGGCACTGCTGGGCTGAGTTCACCCCCAGGGCTGGCCAGATGGGGCC AGGAGGGACAGAGCAAGGGGGGTGAAGGCCGTGGTGGGAGGGTCCCATGATGATGGGCCAGGGCTCGTGTAGAAATGGGGGAATTGGTTC CCCATGGCCCAGGACAGCTGAGAGGAGGTGGAGGGGCCCCAGGGGAGTGTACGTCAGGCTTTGCGGGGCACGGGGGCCACTCAGCAGCGC TGGGGCAGGTGCCTCTGCTGTCAGCTCCACCCGACAGGCAGACGAAGGCCAGTGGGGCCATCGCTTCCTGGGGCGACCCTGGCAGTGGTT GGGAGACGCCCAGATGGAGGGGGAGGCTGACCAAGGGCCCCGCAGGGCGGGCTGCAACTTTTCTGTTGATCCTGGAATGTAGCTGGTGCA GTGAGAGGGAAAGAGAATTGAAAAACTCAGGCTGCCATAGGTTCTGCGATGAGAGGTGCAGGAGGCAGGAGCCTGGCCCAGGGGGTGCTG GTGCCTCCCCGGGGTCTGGGCGGAGAGAACAGGAGGAATGGCTGGGAAGTGGCTGAGGGAGCCAGGAGGCCGGGGGGCCGGGGGCTGCAG GGGAGGCTGTGGGGGTCCTGGCAGCCAGGAGGCCCCAGGTGGTTTTGAGGCTCGCTCTTGCGCGGTGCCTGAGAAGAGGGTGAAGGAGCT GGGGCAGGCCCCATCCTGGGCATTGGAGATGATGAAACCGAGCAGACCTGGCCCATGTGGAGCTGGCATGGGGGACACAGCCCAGAGACA GAGAAGCTTATGAGGAAGTGAGGAGGTGGCGTCACAAGGGTGGGGAGGGGGCCTTGGGGAAGGGCGGCCTTGGATCAGAGGCTCACCACA AGCCTGGCATTTCAGCCAGGGCTGGAGAAGGCAGGGACGCCTGGGTGAGAGGCAAAGGGCACAGCCATGCAAAGGCCCTGGGGCAGGACG GCACCTGGTATGCGGGAGGAACAGAGTGAGGAGAGGAGGGCAGGGCGTGCAGGGCCTTGTGGGCCTCAGGGAGGACTTGGGCACCTACGC CGAGGGAGTGGAGCTCCTGGGTGCGTGTCCAGATGGGAAAGGCAGGGTCGTATCTGTGGGGACCTGACAAGGGCAGGGGAAGCGGAGACC AGGGTGCAGGCTCCGCCCCCACCCAAGGCCGGGCCCAGCCAGAGGAGGGGCAGGGCAGGGCAGGAGGTTTCTGGATGTTTGTTGGGTTTG >66727_66727_3_PNKP-FZR1_PNKP_chr19_50369656_ENST00000596014_FZR1_chr19_3525866_ENST00000441788_length(amino acids)=544AA_BP=73 MLPRPAPRMGEVEAPGRLWLESPPGGAPPIFLPSDGQALVLGRGPLTQVTDRKCSRTQVELVADPETRTVAVKQVTEMRRTLTPASSPVS SPSKHGDRFIPSRAGANWSVNFHRINENEKSPSQNRKAKDATSDNGKDGLAYSALLKNELLGAGIEKVQDPQTEDRRLQPSTPEKKGLFT YSLSTKRSSPDDGNDVSPYSLSPVSNKSQKLLRSPRKPTRKISKIPFKVLDAPELQDDFYLNLVDWSSLNVLSVGLGTCVYLWSACTSQV TRLCDLSVEGDSVTSVGWSERGNLVAVGTHKGFVQIWDAAAGKKLSMLEGHTARVGALAWNAEQLSSGSRDRMILQRDIRTPPLQSERRL QGHRQEVCGLKWSTDHQLLASGGNDNKLLVWNHSSLSPVQQYTEHLAAVKAIAWSPHQHGLLASGGGTADRCIRFWNTLTGQPLQCIDTG SQVCNLAWSKHANELVSTHGYSQNQILVWKYPSLTQVAKLTGHSYRVLYLAMSPDGEAIVTGAGDETLRFWNVFSKTRSTKESVSVLNLF -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PNKP-FZR1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PNKP-FZR1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PNKP-FZR1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
