
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PNPLA6-EXOSC10 (FusionGDB2 ID:66774) |
Fusion Gene Summary for PNPLA6-EXOSC10 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PNPLA6-EXOSC10 | Fusion gene ID: 66774 | Hgene | Tgene | Gene symbol | PNPLA6 | EXOSC10 | Gene ID | 10908 | 5394 |
| Gene name | patatin like phospholipase domain containing 6 | exosome component 10 | |
| Synonyms | BNHS|LNMS|NTE|NTEMND|OMCS|SPG39|iPLA2delta|sws | PM-Scl|PM/Scl-100|PMSCL|PMSCL2|RRP6|Rrp6p|p2|p3|p4 | |
| Cytomap | 19p13.2 | 1p36.22 | |
| Type of gene | protein-coding | protein-coding | |
| Description | neuropathy target esterasepatatin-like phospholipase domain-containing protein 6 | exosome component 10P100 polymyositis-scleroderma overlap syndrome-associated autoantigenautoantigen PM-SCLpolymyositis/scleroderma autoantigen 100 kDapolymyositis/scleroderma autoantigen 2 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q01780 | |
| Ensembl transtripts involved in fusion gene | ENST00000221249, ENST00000414982, ENST00000450331, ENST00000545201, ENST00000600737, ENST00000594864, | ENST00000544779, ENST00000485606, ENST00000304457, ENST00000376936, | |
| Fusion gene scores | * DoF score | 5 X 5 X 4=100 | 10 X 11 X 7=770 |
| # samples | 5 | 16 | |
| ** MAII score | log2(5/100*10)=-1 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(16/770*10)=-2.2667865406949 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PNPLA6 [Title/Abstract] AND EXOSC10 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PNPLA6(7606953)-EXOSC10(11137005), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across PNPLA6 (5'-gene) Fusion gene breakpoints across PNPLA6 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
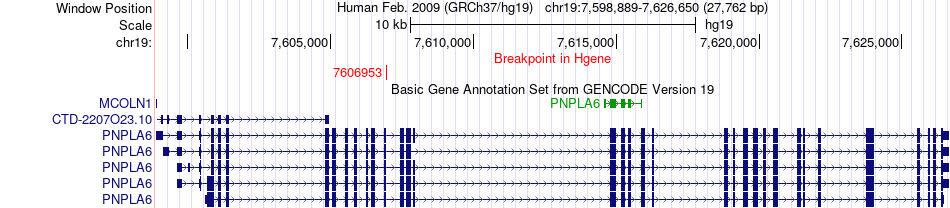 |
 Fusion gene breakpoints across EXOSC10 (3'-gene) Fusion gene breakpoints across EXOSC10 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
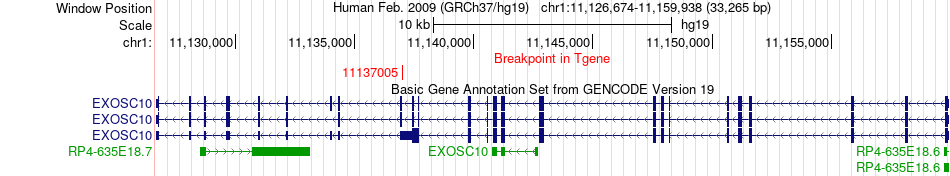 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | GBM | TCGA-16-0846-01A | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
Top |
Fusion Gene ORF analysis for PNPLA6-EXOSC10 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000221249 | ENST00000544779 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-3UTR | ENST00000414982 | ENST00000544779 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-3UTR | ENST00000450331 | ENST00000544779 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-3UTR | ENST00000545201 | ENST00000544779 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-3UTR | ENST00000600737 | ENST00000544779 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-intron | ENST00000221249 | ENST00000485606 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-intron | ENST00000414982 | ENST00000485606 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-intron | ENST00000450331 | ENST00000485606 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-intron | ENST00000545201 | ENST00000485606 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| 5CDS-intron | ENST00000600737 | ENST00000485606 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000221249 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000221249 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000414982 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000414982 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000450331 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000450331 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000545201 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000545201 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000600737 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| In-frame | ENST00000600737 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| intron-3CDS | ENST00000594864 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| intron-3CDS | ENST00000594864 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| intron-3UTR | ENST00000594864 | ENST00000544779 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
| intron-intron | ENST00000594864 | ENST00000485606 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000221249 | PNPLA6 | chr19 | 7606953 | + | ENST00000304457 | EXOSC10 | chr1 | 11137005 | - | 2370 | 1566 | 431 | 2269 | 612 |
| ENST00000221249 | PNPLA6 | chr19 | 7606953 | + | ENST00000376936 | EXOSC10 | chr1 | 11137005 | - | 2445 | 1566 | 431 | 2344 | 637 |
| ENST00000545201 | PNPLA6 | chr19 | 7606953 | + | ENST00000304457 | EXOSC10 | chr1 | 11137005 | - | 2339 | 1535 | 400 | 2238 | 612 |
| ENST00000545201 | PNPLA6 | chr19 | 7606953 | + | ENST00000376936 | EXOSC10 | chr1 | 11137005 | - | 2414 | 1535 | 400 | 2313 | 637 |
| ENST00000414982 | PNPLA6 | chr19 | 7606953 | + | ENST00000304457 | EXOSC10 | chr1 | 11137005 | - | 2278 | 1474 | 195 | 2177 | 660 |
| ENST00000414982 | PNPLA6 | chr19 | 7606953 | + | ENST00000376936 | EXOSC10 | chr1 | 11137005 | - | 2353 | 1474 | 195 | 2252 | 685 |
| ENST00000450331 | PNPLA6 | chr19 | 7606953 | + | ENST00000304457 | EXOSC10 | chr1 | 11137005 | - | 2210 | 1406 | 199 | 2109 | 636 |
| ENST00000450331 | PNPLA6 | chr19 | 7606953 | + | ENST00000376936 | EXOSC10 | chr1 | 11137005 | - | 2285 | 1406 | 199 | 2184 | 661 |
| ENST00000600737 | PNPLA6 | chr19 | 7606953 | + | ENST00000304457 | EXOSC10 | chr1 | 11137005 | - | 2139 | 1335 | 14 | 2038 | 674 |
| ENST00000600737 | PNPLA6 | chr19 | 7606953 | + | ENST00000376936 | EXOSC10 | chr1 | 11137005 | - | 2214 | 1335 | 14 | 2113 | 699 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000221249 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.011733515 | 0.9882665 |
| ENST00000221249 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.014080802 | 0.98591924 |
| ENST00000545201 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.009505034 | 0.99049497 |
| ENST00000545201 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.011731857 | 0.9882682 |
| ENST00000414982 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.013980352 | 0.98601973 |
| ENST00000414982 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.013351858 | 0.98664814 |
| ENST00000450331 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.012209203 | 0.9877908 |
| ENST00000450331 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.014904432 | 0.98509556 |
| ENST00000600737 | ENST00000304457 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.009334105 | 0.99066585 |
| ENST00000600737 | ENST00000376936 | PNPLA6 | chr19 | 7606953 | + | EXOSC10 | chr1 | 11137005 | - | 0.009230932 | 0.9907691 |
Top |
Fusion Genomic Features for PNPLA6-EXOSC10 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
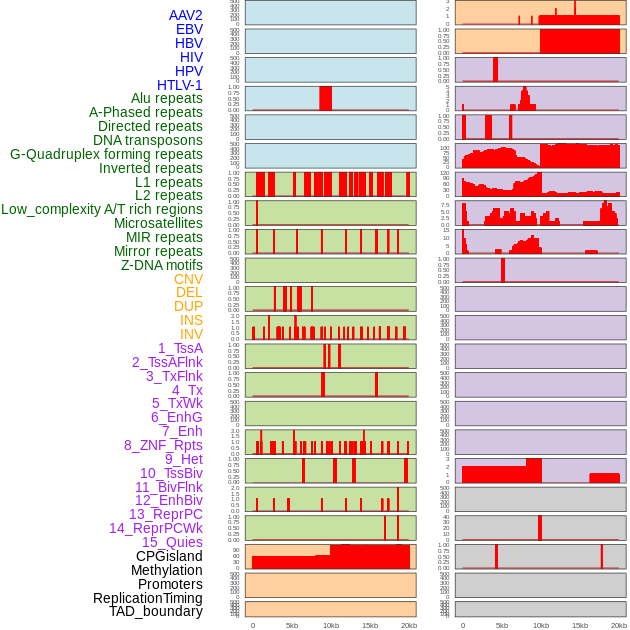 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for PNPLA6-EXOSC10 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:7606953/chr1:11137005) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | EXOSC10 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Putative catalytic component of the RNA exosome complex which has 3'->5' exoribonuclease activity and participates in a multitude of cellular RNA processing and degradation events. In the nucleus, the RNA exosome complex is involved in proper maturation of stable RNA species such as rRNA, snRNA and snoRNA, in the elimination of RNA processing by-products and non-coding 'pervasive' transcripts, such as antisense RNA species and promoter-upstream transcripts (PROMPTs), and of mRNAs with processing defects, thereby limiting or excluding their export to the cytoplasm. The RNA exosome may be involved in Ig class switch recombination (CSR) and/or Ig variable region somatic hypermutation (SHM) by targeting AICDA deamination activity to transcribed dsDNA substrates. In the cytoplasm, the RNA exosome complex is involved in general mRNA turnover and specifically degrades inherently unstable mRNAs containing AU-rich elements (AREs) within their 3' untranslated regions, and in RNA surveillance pathways, preventing translation of aberrant mRNAs. It seems to be involved in degradation of histone mRNA. EXOSC10 has 3'-5' exonuclease activity (By similarity). EXOSC10 is required for nucleolar localization of C1D and probably mediates the association of MTREX, C1D and MPP6 wth the RNA exosome involved in the maturation of 5.8S rRNA. {ECO:0000250, ECO:0000269|PubMed:14527413, ECO:0000269|PubMed:16455498, ECO:0000269|PubMed:17412707, ECO:0000269|PubMed:17545563, ECO:0000269|PubMed:18172165, ECO:0000269|PubMed:19056938, ECO:0000269|PubMed:20368444, ECO:0000269|PubMed:20699273}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 195_322 | 378 | 1328.0 | Nucleotide binding | Note=cNMP 1 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 195_322 | 426 | 1376.0 | Nucleotide binding | Note=cNMP 1 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 195_322 | 378 | 1328.0 | Nucleotide binding | Note=cNMP 1 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 195_322 | 378 | 1301.0 | Nucleotide binding | Note=cNMP 1 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 195_322 | 417 | 1366.0 | Nucleotide binding | Note=cNMP 1 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 1_59 | 378 | 1328.0 | Topological domain | Lumenal |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 1_59 | 426 | 1376.0 | Topological domain | Lumenal |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 1_59 | 378 | 1328.0 | Topological domain | Lumenal |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 1_59 | 378 | 1301.0 | Topological domain | Lumenal |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 1_59 | 417 | 1366.0 | Topological domain | Lumenal |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 60_80 | 378 | 1328.0 | Transmembrane | Helical |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 60_80 | 426 | 1376.0 | Transmembrane | Helical |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 60_80 | 378 | 1328.0 | Transmembrane | Helical |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 60_80 | 378 | 1301.0 | Transmembrane | Helical |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 60_80 | 417 | 1366.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 981_1147 | 378 | 1328.0 | Domain | PNPLA |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 981_1147 | 426 | 1376.0 | Domain | PNPLA |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 981_1147 | 378 | 1328.0 | Domain | PNPLA |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 981_1147 | 378 | 1301.0 | Domain | PNPLA |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 981_1147 | 417 | 1366.0 | Domain | PNPLA |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 1012_1016 | 378 | 1328.0 | Motif | GXSXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 1134_1136 | 378 | 1328.0 | Motif | DGA/G |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 985_990 | 378 | 1328.0 | Motif | GXGXXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 1012_1016 | 426 | 1376.0 | Motif | GXSXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 1134_1136 | 426 | 1376.0 | Motif | DGA/G |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 985_990 | 426 | 1376.0 | Motif | GXGXXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 1012_1016 | 378 | 1328.0 | Motif | GXSXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 1134_1136 | 378 | 1328.0 | Motif | DGA/G |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 985_990 | 378 | 1328.0 | Motif | GXGXXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 1012_1016 | 378 | 1301.0 | Motif | GXSXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 1134_1136 | 378 | 1301.0 | Motif | DGA/G |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 985_990 | 378 | 1301.0 | Motif | GXGXXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 1012_1016 | 417 | 1366.0 | Motif | GXSXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 1134_1136 | 417 | 1366.0 | Motif | DGA/G |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 985_990 | 417 | 1366.0 | Motif | GXGXXG |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 511_633 | 378 | 1328.0 | Nucleotide binding | Note=cNMP 2 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 629_749 | 378 | 1328.0 | Nucleotide binding | Note=cNMP 3 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 511_633 | 426 | 1376.0 | Nucleotide binding | Note=cNMP 2 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 629_749 | 426 | 1376.0 | Nucleotide binding | Note=cNMP 3 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 511_633 | 378 | 1328.0 | Nucleotide binding | Note=cNMP 2 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 629_749 | 378 | 1328.0 | Nucleotide binding | Note=cNMP 3 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 511_633 | 378 | 1301.0 | Nucleotide binding | Note=cNMP 2 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 629_749 | 378 | 1301.0 | Nucleotide binding | Note=cNMP 3 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 511_633 | 417 | 1366.0 | Nucleotide binding | Note=cNMP 2 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 629_749 | 417 | 1366.0 | Nucleotide binding | Note=cNMP 3 |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000221249 | + | 13 | 35 | 81_1375 | 378 | 1328.0 | Topological domain | Cytoplasmic |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000414982 | + | 12 | 34 | 81_1375 | 426 | 1376.0 | Topological domain | Cytoplasmic |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000450331 | + | 13 | 35 | 81_1375 | 378 | 1328.0 | Topological domain | Cytoplasmic |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000545201 | + | 13 | 34 | 81_1375 | 378 | 1301.0 | Topological domain | Cytoplasmic |
| Hgene | PNPLA6 | chr19:7606953 | chr1:11137005 | ENST00000600737 | + | 10 | 32 | 81_1375 | 417 | 1366.0 | Topological domain | Cytoplasmic |
| Tgene | EXOSC10 | chr19:7606953 | chr1:11137005 | ENST00000304457 | 15 | 24 | 503_583 | 626 | 861.0 | Domain | HRDC | |
| Tgene | EXOSC10 | chr19:7606953 | chr1:11137005 | ENST00000376936 | 15 | 25 | 503_583 | 626 | 886.0 | Domain | HRDC |
Top |
Fusion Gene Sequence for PNPLA6-EXOSC10 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >66774_66774_1_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000221249_EXOSC10_chr1_11137005_ENST00000304457_length(transcript)=2370nt_BP=1566nt TTTTTAGCGGGAGGAGCAGTCCTTGCGTTAAGCGGTGTGAGGCCCTTTAAGGCGCGGCCACACTCAGCATGGCGGCCTCAGTCGGCCTTC CAAGCATGGCGCGGGGAGGAGGGGTGGGAGGGTCGGGAGGGACTGCGGTGCGACTAGGAGTGAATAATTTAAAGGGGCCGTGCCTGCGGA GCCGGGCGGAACGCTAGCGGTGTTGGCGCGGAGTGGACCCCGGCTGCGGCCCCTGGCAATGGCGCCACCATCGGTCCCGGAGTCCCAGTG ATGCTCTGTGCCATAGAGCCCCCATAACTTCACTACTACGTGATAGTAAATCCCCGGCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCA CCCCAAGCATCCCAGGACTCTTCTGAAACGACTCCGGGCTACCAGATCGGCCGTCCAGCTGGAATCAACCGATGGAGGCTCCGCTGCAAA CTGGAATGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAG TGCCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGG TGTCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTG CCAAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGG GCGACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCC TGGAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGG TGCAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACA GCCTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGC GCCTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGC GAGTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCC CCGGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGC GGCCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGC TGAGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATA ACTTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGA CAGTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCATTTAGGATGATCAGCAACCGCTGGAAGCTGGCCCAGGTAC AAGTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTG CAGCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCA CAGAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACT ACAGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCA AGAAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGT ACAACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTGGTGCTGCTTTTGTACATACATATT >66774_66774_1_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000221249_EXOSC10_chr1_11137005_ENST00000304457_length(amino acids)=612AA_BP=378 MEAPLQTGMVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKK LKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQP DASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQI IMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLE TPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMISNR WKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEK -------------------------------------------------------------- >66774_66774_2_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000221249_EXOSC10_chr1_11137005_ENST00000376936_length(transcript)=2445nt_BP=1566nt TTTTTAGCGGGAGGAGCAGTCCTTGCGTTAAGCGGTGTGAGGCCCTTTAAGGCGCGGCCACACTCAGCATGGCGGCCTCAGTCGGCCTTC CAAGCATGGCGCGGGGAGGAGGGGTGGGAGGGTCGGGAGGGACTGCGGTGCGACTAGGAGTGAATAATTTAAAGGGGCCGTGCCTGCGGA GCCGGGCGGAACGCTAGCGGTGTTGGCGCGGAGTGGACCCCGGCTGCGGCCCCTGGCAATGGCGCCACCATCGGTCCCGGAGTCCCAGTG ATGCTCTGTGCCATAGAGCCCCCATAACTTCACTACTACGTGATAGTAAATCCCCGGCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCA CCCCAAGCATCCCAGGACTCTTCTGAAACGACTCCGGGCTACCAGATCGGCCGTCCAGCTGGAATCAACCGATGGAGGCTCCGCTGCAAA CTGGAATGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAG TGCCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGG TGTCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTG CCAAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGG GCGACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCC TGGAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGG TGCAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACA GCCTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGC GCCTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGC GAGTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCC CCGGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGC GGCCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGC TGAGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATA ACTTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGA CAGTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCATTTAGGATGTTTCTGCCCTCACTGGGACACCGTGCTCCCG TCTCTCAGGCAGCGAAGTTCGATCCATCAACCAAAATCTATGAAATCAGCAACCGCTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACT CTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCA TCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCACAGAACAGAAACAAG AGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACTACAGCCAGTCAGACT TCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCAAGAAATGCATTGCAG CCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGTACAACTGGCCACAGA GATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTGGTGCTGCTTTTGTACATACATATTTTTAAACCATTAAAA >66774_66774_2_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000221249_EXOSC10_chr1_11137005_ENST00000376936_length(amino acids)=637AA_BP=378 MEAPLQTGMVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKK LKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQP DASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQI IMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLE TPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMFLPS LGHRAPVSQAAKFDPSTKIYEISNRWKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPR TTEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAGNSKSKVSSQFDPNKQTPSGKKCIAAKKIKQSVGNKSMSFPTGKSDRGF -------------------------------------------------------------- >66774_66774_3_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000414982_EXOSC10_chr1_11137005_ENST00000304457_length(transcript)=2278nt_BP=1474nt CAATGGCGCCACCATCGGTCCCGGAGTCCCAGTGATGCTCTGTGCCATAGAGCCCCCATAACTTCACTACTACGTGATAGTAAATCCCCG GCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCACCCCAAGCATCCCAGGACTCTTCTGAAACGACTCCGGGCTACCAGATCGGCCGTCC AGCTGGAATCAACCGATGGAGGCTCCGCTGCAAACTGGAATGATGGGGACATCGAGTCACGGGCTGGCTACGAACTCCTCGGGGGCGAAG GTGGCGGAGAGGGATGGGTTCCAGGACGTCCTGGCGCCCGGGGAAGGCTCGGCGGGACGGATTTGCGGTGCGCAGCCAGTGCCGTTCGTC CCTCAGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAGTG CCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGGTG TCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTGCC AAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGGGC GACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCCTG GAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGGTG CAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACAGC CTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGCGC CTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGCGA GTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCCCC GGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGCGG CCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGCTG AGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATAAC TTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGACA GTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCATTTAGGATGATCAGCAACCGCTGGAAGCTGGCCCAGGTACAA GTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTGCA GCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCACA GAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACTAC AGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCAAG AAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGTAC AACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTGGTGCTGCTTTTGTACATACATATTTT >66774_66774_3_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000414982_EXOSC10_chr1_11137005_ENST00000304457_length(amino acids)=660AA_BP=426 MEAPLQTGMMGTSSHGLATNSSGAKVAERDGFQDVLAPGEGSAGRICGAQPVPFVPQVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPA PDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKKLKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANS HLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQPDASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSIL DVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQIIMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTR TSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLETPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTT CLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMISNRWKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAI SVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAGNSKSKVSSQFDPNKQTPSGKKCIAA -------------------------------------------------------------- >66774_66774_4_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000414982_EXOSC10_chr1_11137005_ENST00000376936_length(transcript)=2353nt_BP=1474nt CAATGGCGCCACCATCGGTCCCGGAGTCCCAGTGATGCTCTGTGCCATAGAGCCCCCATAACTTCACTACTACGTGATAGTAAATCCCCG GCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCACCCCAAGCATCCCAGGACTCTTCTGAAACGACTCCGGGCTACCAGATCGGCCGTCC AGCTGGAATCAACCGATGGAGGCTCCGCTGCAAACTGGAATGATGGGGACATCGAGTCACGGGCTGGCTACGAACTCCTCGGGGGCGAAG GTGGCGGAGAGGGATGGGTTCCAGGACGTCCTGGCGCCCGGGGAAGGCTCGGCGGGACGGATTTGCGGTGCGCAGCCAGTGCCGTTCGTC CCTCAGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAGTG CCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGGTG TCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTGCC AAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGGGC GACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCCTG GAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGGTG CAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACAGC CTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGCGC CTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGCGA GTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCCCC GGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGCGG CCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGCTG AGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATAAC TTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGACA GTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCATTTAGGATGTTTCTGCCCTCACTGGGACACCGTGCTCCCGTC TCTCAGGCAGCGAAGTTCGATCCATCAACCAAAATCTATGAAATCAGCAACCGCTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACTCT AAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCATC TCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCACAGAACAGAAACAAGAG AAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACTACAGCCAGTCAGACTTC AAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCAAGAAATGCATTGCAGCC AAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGTACAACTGGCCACAGAGA TAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTGGTGCTGCTTTTGTACATACATATTTTTAAACCATTAAAATT >66774_66774_4_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000414982_EXOSC10_chr1_11137005_ENST00000376936_length(amino acids)=685AA_BP=426 MEAPLQTGMMGTSSHGLATNSSGAKVAERDGFQDVLAPGEGSAGRICGAQPVPFVPQVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPA PDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKKLKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANS HLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQPDASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSIL DVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQIIMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTR TSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLETPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTT CLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMFLPSLGHRAPVSQAAKFDPSTKIYEISNRWKLAQVQVQKDSKEAVK KKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAG -------------------------------------------------------------- >66774_66774_5_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000450331_EXOSC10_chr1_11137005_ENST00000304457_length(transcript)=2210nt_BP=1406nt CAATGGCGCCACCATCGGTCCCGGAGTCCCAGTGATGCTCTGTGCCATAGAGCCCCCATAACTTCACTACTACGTGATAGTAAATCCCCG GCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCACCCCAAGCATCCCAGGACTCTTCTGAAACAGGGCAGTGACCCGGGAGGAAGTCACC AGGTGCAGAGGACCGTGCCCTGTGGGGACCTGCCTGCAGGCTGCTGACAGACTCCGGGCTACCAGATCGGCCGTCCAGCTGGAATCAACC GATGGAGGCTCCGCTGCAAACTGGAATGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCT GGTGGTGCGGAGGCTGCGAGTGCCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGG CCGGAAGATTATGCGGAAGGTGTCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAA ACTGAAGATGCTCAACATTGCCAAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCT AGAAGCTGACCTGACCGAGGGCGACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCA CTTCGAGAAGCCACTCTTCCTGGAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCC AGATGCCAGCATCTACGTGGTGCAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGT TCCTGGGGACAGCGTCAACAGCCTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGC CCGGGACTCCACGGTGCTGCGCCTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGAT CATCATGGTGCGGCTGCAGCGAGTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCC CCTGCGTCTGTTCCCCAGCCCCGGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGA GCCCAGGGAGACCCCAGGGCGGCCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGA AACCCCCTCGGCCCCTCTGCTGAGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTT CCCTGATGAAAAAGAAGATAACTTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGA CAGTAAAAAGGGTCCATTGACAGTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCATTTAGGATGATCAGCAACCG CTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGC AAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGC AACAAGCGACCCAAGGACCACAGAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAA AGAGTTTACGCCTTACGACTACAGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAA TAAACAGACCCCGTCTGGCAAGAAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAA GTCAGACAGAGGCTTCAGGTACAACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTGGTGC >66774_66774_5_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000450331_EXOSC10_chr1_11137005_ENST00000304457_length(amino acids)=636AA_BP=402 MWGPACRLLTDSGLPDRPSSWNQPMEAPLQTGMVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRK VSQSTSSLVDTSVSATSRPRMRKKLKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLF LELCRHMVFQRLGQGDYVFRPGQPDASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVL RLPVEAFSAVFTKYPESLVRVVQIIMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPG RPPDPTGAPLPGPTGDPVKPTSLETPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPL TVAQKKAQNIMESFENPFRMISNRWKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPRT TEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAGNSKSKVSSQFDPNKQTPSGKKCIAAKKIKQSVGNKSMSFPTGKSDRGFR -------------------------------------------------------------- >66774_66774_6_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000450331_EXOSC10_chr1_11137005_ENST00000376936_length(transcript)=2285nt_BP=1406nt CAATGGCGCCACCATCGGTCCCGGAGTCCCAGTGATGCTCTGTGCCATAGAGCCCCCATAACTTCACTACTACGTGATAGTAAATCCCCG GCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCACCCCAAGCATCCCAGGACTCTTCTGAAACAGGGCAGTGACCCGGGAGGAAGTCACC AGGTGCAGAGGACCGTGCCCTGTGGGGACCTGCCTGCAGGCTGCTGACAGACTCCGGGCTACCAGATCGGCCGTCCAGCTGGAATCAACC GATGGAGGCTCCGCTGCAAACTGGAATGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCT GGTGGTGCGGAGGCTGCGAGTGCCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGG CCGGAAGATTATGCGGAAGGTGTCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAA ACTGAAGATGCTCAACATTGCCAAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCT AGAAGCTGACCTGACCGAGGGCGACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCA CTTCGAGAAGCCACTCTTCCTGGAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCC AGATGCCAGCATCTACGTGGTGCAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGT TCCTGGGGACAGCGTCAACAGCCTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGC CCGGGACTCCACGGTGCTGCGCCTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGAT CATCATGGTGCGGCTGCAGCGAGTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCC CCTGCGTCTGTTCCCCAGCCCCGGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGA GCCCAGGGAGACCCCAGGGCGGCCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGA AACCCCCTCGGCCCCTCTGCTGAGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTT CCCTGATGAAAAAGAAGATAACTTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGA CAGTAAAAAGGGTCCATTGACAGTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCATTTAGGATGTTTCTGCCCTC ACTGGGACACCGTGCTCCCGTCTCTCAGGCAGCGAAGTTCGATCCATCAACCAAAATCTATGAAATCAGCAACCGCTGGAAGCTGGCCCA GGTACAAGTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAA AGCTGCAGCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAG GACCACAGAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTA CGACTACAGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTC TGGCAAGAAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTT CAGGTACAACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTGGTGCTGCTTTTGTACATAC >66774_66774_6_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000450331_EXOSC10_chr1_11137005_ENST00000376936_length(amino acids)=661AA_BP=402 MWGPACRLLTDSGLPDRPSSWNQPMEAPLQTGMVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRK VSQSTSSLVDTSVSATSRPRMRKKLKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLF LELCRHMVFQRLGQGDYVFRPGQPDASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVL RLPVEAFSAVFTKYPESLVRVVQIIMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPG RPPDPTGAPLPGPTGDPVKPTSLETPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPL TVAQKKAQNIMESFENPFRMFLPSLGHRAPVSQAAKFDPSTKIYEISNRWKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQA ISVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAGNSKSKVSSQFDPNKQTPSGKKCIA -------------------------------------------------------------- >66774_66774_7_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000545201_EXOSC10_chr1_11137005_ENST00000304457_length(transcript)=2339nt_BP=1535nt GAGTCTGGGTTTCCGTTAGCCTCGCAGGGGTGTCCCTTCGAGGGTCGTTAGCGAGCCTCCGCTTTCCCACGATCTGTCCTCCGATTCCTG TTAACTCTAGACTTTCTGATGTTCCCATACCCCCCACGTCTCGGCAGGTGTTTCCACACCGGTAGCCAGCTGTGCCCTGAGGTGGAAGAG GACCGGCCACCCAGGAATTTTCCAACAATGGCGCCACCATCGGTCCCGGAGTCCCAGTGATGCTCTGTGCCATAGAGCCCCCATAACTTC ACTACTACGTGATAGTAAATCCCCGGCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCACCCCAAGCATCCCAGGACTCTTCTGAAACGA CTCCGGGCTACCAGATCGGCCGTCCAGCTGGAATCAACCGATGGAGGCTCCGCTGCAAACTGGAATGGTGCTTGGCGTGATGATCGGGGC CGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAGTGCCAAAAACCCCAGCCCCGGATGGCCCCCG GTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGGTGTCACAATCCACCTCCTCCCTCGTGGATAC CTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTGCCAAGAAGATCCTGCGCATCCAGAAAGAGAC GCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGGGCGACCTGGCTAACTCCCATCTGCCCTCTGA AGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCCTGGAGCTCTGCCGCCACATGGTCTTCCAGCG GCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGGTGCAGGACGGGCTGCTGGAGCTCTGTCTGCC AGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACAGCCTTCTCAGCATCCTGGATGTCATCACCGG TCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGCGCCTGCCGGTGGAAGCATTCTCCGCGGTCTT CACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGCGAGTCACCTTCCTGGCACTGCACAACTACCT GGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCCCCGGCCTCCCAACTCGCACCAGCCCTGTGCG GGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGCGGCCACCCGATCCCACCGGGGCCCCGCTGCC TGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGCTGAGCCGCTGCGTCTCCATGCCAGGGGACAT CTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATAACTTGCTGGGTACCACATGCCTGATTGCCAC AGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGACAGTTGCACAGAAAAAAGCCCAGAACATCAT GGAGTCCTTTGAAAATCCATTTAGGATGATCAGCAACCGCTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACTCTAAAGAAGCTGTCAA GAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCATCTCCGTCCGACAGCA GGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCACAGAACAGAAACAAGAGAAGAAACGACTCAA AATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACTACAGCCAGTCAGACTTCAAGGCTTTTGCTGG AAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCAAGAAATGCATTGCAGCCAAAAAAATTAAACA GTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGTACAACTGGCCACAGAGATAGTCCTGGAAGAC >66774_66774_7_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000545201_EXOSC10_chr1_11137005_ENST00000304457_length(amino acids)=612AA_BP=378 MEAPLQTGMVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKK LKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQP DASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQI IMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLE TPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMISNR WKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEK -------------------------------------------------------------- >66774_66774_8_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000545201_EXOSC10_chr1_11137005_ENST00000376936_length(transcript)=2414nt_BP=1535nt GAGTCTGGGTTTCCGTTAGCCTCGCAGGGGTGTCCCTTCGAGGGTCGTTAGCGAGCCTCCGCTTTCCCACGATCTGTCCTCCGATTCCTG TTAACTCTAGACTTTCTGATGTTCCCATACCCCCCACGTCTCGGCAGGTGTTTCCACACCGGTAGCCAGCTGTGCCCTGAGGTGGAAGAG GACCGGCCACCCAGGAATTTTCCAACAATGGCGCCACCATCGGTCCCGGAGTCCCAGTGATGCTCTGTGCCATAGAGCCCCCATAACTTC ACTACTACGTGATAGTAAATCCCCGGCAAAAACCAGCAGCGCCTTGCAAGCCCACGCCACCCCAAGCATCCCAGGACTCTTCTGAAACGA CTCCGGGCTACCAGATCGGCCGTCCAGCTGGAATCAACCGATGGAGGCTCCGCTGCAAACTGGAATGGTGCTTGGCGTGATGATCGGGGC CGGAGTGGCGGTGGTGGTCACGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAGTGCCAAAAACCCCAGCCCCGGATGGCCCCCG GTATCGGTTCCGGAAGAGGGACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGGTGTCACAATCCACCTCCTCCCTCGTGGATAC CTCTGTCTCCGCCACCTCCCGGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTGCCAAGAAGATCCTGCGCATCCAGAAAGAGAC GCCCACGCTGCAGCGGAAGGAGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGGGCGACCTGGCTAACTCCCATCTGCCCTCTGA AGTGCTTTATATGCTCAAGAACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCCTGGAGCTCTGCCGCCACATGGTCTTCCAGCG GCTGGGCCAGGGTGACTACGTCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGGTGCAGGACGGGCTGCTGGAGCTCTGTCTGCC AGGGCCTGACGGGAAGGAGTGTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACAGCCTTCTCAGCATCCTGGATGTCATCACCGG TCACCAGCATCCCCAGCGGACCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGCGCCTGCCGGTGGAAGCATTCTCCGCGGTCTT CACCAAGTACCCGGAGAGCTTGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGCGAGTCACCTTCCTGGCACTGCACAACTACCT GGGTCTGACCAATGAGCTCTTCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCCCCGGCCTCCCAACTCGCACCAGCCCTGTGCG GGGCTCCAAGAGAATGGTCAGCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGCGGCCACCCGATCCCACCGGGGCCCCGCTGCC TGGACCTACAGGGGACCCTGTGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGCTGAGCCGCTGCGTCTCCATGCCAGGGGACAT CTCAGGATCTGTGCCAGTTCAGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATAACTTGCTGGGTACCACATGCCTGATTGCCAC AGCTGTCATCACGTTATTTAATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGACAGTTGCACAGAAAAAAGCCCAGAACATCAT GGAGTCCTTTGAAAATCCATTTAGGATGTTTCTGCCCTCACTGGGACACCGTGCTCCCGTCTCTCAGGCAGCGAAGTTCGATCCATCAAC CAAAATCTATGAAATCAGCAACCGCTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGA GCAAACAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAA TGCTGCAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCACAGAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCC AAAGGACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACTACAGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAA AGTTTCTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCAAGAAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAA AAGCATGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGTACAACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGG >66774_66774_8_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000545201_EXOSC10_chr1_11137005_ENST00000376936_length(amino acids)=637AA_BP=378 MEAPLQTGMVLGVMIGAGVAVVVTAVLILLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKK LKMLNIAKKILRIQKETPTLQRKEPPPAVLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQP DASIYVVQDGLLELCLPGPDGKECVVKEVVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQI IMVRLQRVTFLALHNYLGLTNELFSHEIQPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLE TPSAPLLSRCVSMPGDISGSVPVQKQASLFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMFLPS LGHRAPVSQAAKFDPSTKIYEISNRWKLAQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPR TTEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAGNSKSKVSSQFDPNKQTPSGKKCIAAKKIKQSVGNKSMSFPTGKSDRGF -------------------------------------------------------------- >66774_66774_9_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000600737_EXOSC10_chr1_11137005_ENST00000304457_length(transcript)=2139nt_BP=1335nt TTTCCCGGCATGCACTGCGGGCCGCCGGGCCTCAGGGAAGAGTCGCGCCCCCGGGGAGGGAGCAGCACTGGCCCATTCTGCAGATGGGGA CATCGAGTCACGGGCTGGCTACGAACTCCTCGGGGGCGAAGGTGGCGGAGAGGGATGGGTTCCAGGACGTCCTGGCGCCCGGGGAAGGCT CGGCGGGACGGATTTGCGGTGCGCAGCCAGTGCCGTTCGTCCCTCAGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCA CGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAGTGCCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGG ACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGGTGTCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCC GGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTGCCAAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGG AGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGGGCGACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGA ACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCCTGGAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACG TCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGGTGCAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGT GTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACAGCCTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGA CCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGCGCCTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCT TGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGCGAGTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCT TCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCCCCGGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCA GCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGCGGCCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTG TGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGCTGAGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTC AGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATAACTTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTA ATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGACAGTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCAT TTAGGATGATCAGCAACCGCTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAA CAGCTGCCCGGGAACAGGCAAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTG CAAAGAAGAGAGAGCGAGCAACAAGCGACCCAAGGACCACAGAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGG ACCCAGAGCCACCAGAAAAAGAGTTTACGCCTTACGACTACAGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTT CTTCTCAGTTTGATCCAAATAAACAGACCCCGTCTGGCAAGAAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCA TGTCCTTTCCAACTGGAAAGTCAGACAGAGGCTTCAGGTACAACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGG >66774_66774_9_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000600737_EXOSC10_chr1_11137005_ENST00000304457_length(amino acids)=674AA_BP=440 MRAAGPQGRVAPPGREQHWPILQMGTSSHGLATNSSGAKVAERDGFQDVLAPGEGSAGRICGAQPVPFVPQVLGVMIGAGVAVVVTAVLI LLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKKLKMLNIAKKILRIQKETPTLQRKEPPPA VLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQPDASIYVVQDGLLELCLPGPDGKECVVKE VVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQIIMVRLQRVTFLALHNYLGLTNELFSHEI QPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLETPSAPLLSRCVSMPGDISGSVPVQKQAS LFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMISNRWKLAQVQVQKDSKEAVKKKAAEQTAARE QAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEKEFTPYDYSQSDFKAFAGNSKSKVSSQFD -------------------------------------------------------------- >66774_66774_10_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000600737_EXOSC10_chr1_11137005_ENST00000376936_length(transcript)=2214nt_BP=1335nt TTTCCCGGCATGCACTGCGGGCCGCCGGGCCTCAGGGAAGAGTCGCGCCCCCGGGGAGGGAGCAGCACTGGCCCATTCTGCAGATGGGGA CATCGAGTCACGGGCTGGCTACGAACTCCTCGGGGGCGAAGGTGGCGGAGAGGGATGGGTTCCAGGACGTCCTGGCGCCCGGGGAAGGCT CGGCGGGACGGATTTGCGGTGCGCAGCCAGTGCCGTTCGTCCCTCAGGTGCTTGGCGTGATGATCGGGGCCGGAGTGGCGGTGGTGGTCA CGGCCGTGCTCATCCTCCTGGTGGTGCGGAGGCTGCGAGTGCCAAAAACCCCAGCCCCGGATGGCCCCCGGTATCGGTTCCGGAAGAGGG ACAAAGTGCTCTTCTATGGCCGGAAGATTATGCGGAAGGTGTCACAATCCACCTCCTCCCTCGTGGATACCTCTGTCTCCGCCACCTCCC GGCCACGCATGAGGAAGAAACTGAAGATGCTCAACATTGCCAAGAAGATCCTGCGCATCCAGAAAGAGACGCCCACGCTGCAGCGGAAGG AGCCCCCGCCCGCAGTGCTAGAAGCTGACCTGACCGAGGGCGACCTGGCTAACTCCCATCTGCCCTCTGAAGTGCTTTATATGCTCAAGA ACGTCCGGGTGCTGGGCCACTTCGAGAAGCCACTCTTCCTGGAGCTCTGCCGCCACATGGTCTTCCAGCGGCTGGGCCAGGGTGACTACG TCTTCCGGCCGGGCCAGCCAGATGCCAGCATCTACGTGGTGCAGGACGGGCTGCTGGAGCTCTGTCTGCCAGGGCCTGACGGGAAGGAGT GTGTGGTGAAGGAAGTGGTTCCTGGGGACAGCGTCAACAGCCTTCTCAGCATCCTGGATGTCATCACCGGTCACCAGCATCCCCAGCGGA CCGTGTCTGCCCGGGCGGCCCGGGACTCCACGGTGCTGCGCCTGCCGGTGGAAGCATTCTCCGCGGTCTTCACCAAGTACCCGGAGAGCT TGGTGCGGGTCGTGCAGATCATCATGGTGCGGCTGCAGCGAGTCACCTTCCTGGCACTGCACAACTACCTGGGTCTGACCAATGAGCTCT TCAGCCACGAGATCCAGCCCCTGCGTCTGTTCCCCAGCCCCGGCCTCCCAACTCGCACCAGCCCTGTGCGGGGCTCCAAGAGAATGGTCA GCACCTCAGCTACAGACGAGCCCAGGGAGACCCCAGGGCGGCCACCCGATCCCACCGGGGCCCCGCTGCCTGGACCTACAGGGGACCCTG TGAAGCCCACATCCCTGGAAACCCCCTCGGCCCCTCTGCTGAGCCGCTGCGTCTCCATGCCAGGGGACATCTCAGGATCTGTGCCAGTTC AGAAGCAGGCGAGCCTCTTCCCTGATGAAAAAGAAGATAACTTGCTGGGTACCACATGCCTGATTGCCACAGCTGTCATCACGTTATTTA ATGAACCTAGTGCTGAAGACAGTAAAAAGGGTCCATTGACAGTTGCACAGAAAAAAGCCCAGAACATCATGGAGTCCTTTGAAAATCCAT TTAGGATGTTTCTGCCCTCACTGGGACACCGTGCTCCCGTCTCTCAGGCAGCGAAGTTCGATCCATCAACCAAAATCTATGAAATCAGCA ACCGCTGGAAGCTGGCCCAGGTACAAGTACAAAAAGACTCTAAAGAAGCTGTCAAGAAGAAGGCAGCTGAGCAAACAGCTGCCCGGGAAC AGGCAAAGGAGGCGTGCAAAGCTGCAGCAGAACAGGCCATCTCCGTCCGACAGCAGGTCGTGCTAGAAAATGCTGCAAAGAAGAGAGAGC GAGCAACAAGCGACCCAAGGACCACAGAACAGAAACAAGAGAAGAAACGACTCAAAATTTCCAAGAAGCCAAAGGACCCAGAGCCACCAG AAAAAGAGTTTACGCCTTACGACTACAGCCAGTCAGACTTCAAGGCTTTTGCTGGAAACAGCAAATCCAAAGTTTCTTCTCAGTTTGATC CAAATAAACAGACCCCGTCTGGCAAGAAATGCATTGCAGCCAAAAAAATTAAACAGTCGGTGGGAAACAAAAGCATGTCCTTTCCAACTG GAAAGTCAGACAGAGGCTTCAGGTACAACTGGCCACAGAGATAGTCCTGGAAGACACGTGGCGCCTGTGGACCGGAAGCACCAAATGCTG >66774_66774_10_PNPLA6-EXOSC10_PNPLA6_chr19_7606953_ENST00000600737_EXOSC10_chr1_11137005_ENST00000376936_length(amino acids)=699AA_BP=440 MRAAGPQGRVAPPGREQHWPILQMGTSSHGLATNSSGAKVAERDGFQDVLAPGEGSAGRICGAQPVPFVPQVLGVMIGAGVAVVVTAVLI LLVVRRLRVPKTPAPDGPRYRFRKRDKVLFYGRKIMRKVSQSTSSLVDTSVSATSRPRMRKKLKMLNIAKKILRIQKETPTLQRKEPPPA VLEADLTEGDLANSHLPSEVLYMLKNVRVLGHFEKPLFLELCRHMVFQRLGQGDYVFRPGQPDASIYVVQDGLLELCLPGPDGKECVVKE VVPGDSVNSLLSILDVITGHQHPQRTVSARAARDSTVLRLPVEAFSAVFTKYPESLVRVVQIIMVRLQRVTFLALHNYLGLTNELFSHEI QPLRLFPSPGLPTRTSPVRGSKRMVSTSATDEPRETPGRPPDPTGAPLPGPTGDPVKPTSLETPSAPLLSRCVSMPGDISGSVPVQKQAS LFPDEKEDNLLGTTCLIATAVITLFNEPSAEDSKKGPLTVAQKKAQNIMESFENPFRMFLPSLGHRAPVSQAAKFDPSTKIYEISNRWKL AQVQVQKDSKEAVKKKAAEQTAAREQAKEACKAAAEQAISVRQQVVLENAAKKRERATSDPRTTEQKQEKKRLKISKKPKDPEPPEKEFT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PNPLA6-EXOSC10 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PNPLA6-EXOSC10 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PNPLA6-EXOSC10 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
