
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PPME1-ELP3 (FusionGDB2 ID:67684) |
Fusion Gene Summary for PPME1-ELP3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PPME1-ELP3 | Fusion gene ID: 67684 | Hgene | Tgene | Gene symbol | PPME1 | ELP3 | Gene ID | 51400 | 55140 |
| Gene name | protein phosphatase methylesterase 1 | elongator acetyltransferase complex subunit 3 | |
| Synonyms | ABDH19|PME-1 | KAT9 | |
| Cytomap | 11q13.4 | 8p21.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein phosphatase methylesterase 1testicular secretory protein Li 39 | elongator complex protein 3elongation protein 3 homologprotein lysine acetyltransferase ELP3tRNA uridine(34) acetyltransferase | |
| Modification date | 20200313 | 20200322 | |
| UniProtAcc | . | Q9H9T3 | |
| Ensembl transtripts involved in fusion gene | ENST00000542710, ENST00000328257, ENST00000398427, ENST00000543525, | ENST00000256398, ENST00000521015, ENST00000542181, ENST00000380353, ENST00000523760, ENST00000524103, ENST00000537665, | |
| Fusion gene scores | * DoF score | 8 X 3 X 5=120 | 5 X 5 X 4=100 |
| # samples | 8 | 5 | |
| ** MAII score | log2(8/120*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(5/100*10)=-1 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PPME1 [Title/Abstract] AND ELP3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PPME1(73882567)-ELP3(27987019), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | PPME1 | GO:0006482 | protein demethylation | 10318862 |
| Tgene | ELP3 | GO:0006357 | regulation of transcription by RNA polymerase II | 11818576 |
 Fusion gene breakpoints across PPME1 (5'-gene) Fusion gene breakpoints across PPME1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
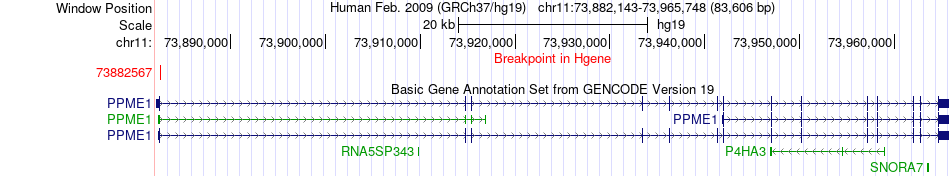 |
 Fusion gene breakpoints across ELP3 (3'-gene) Fusion gene breakpoints across ELP3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
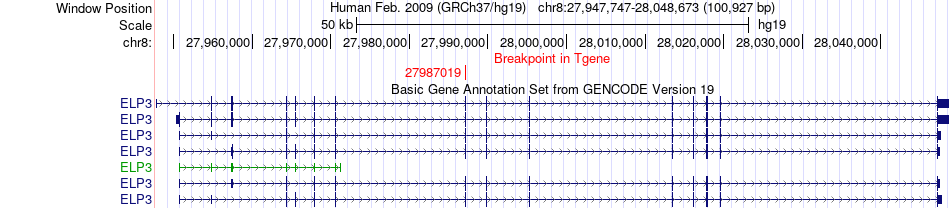 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-CD-8531-01A | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
Top |
Fusion Gene ORF analysis for PPME1-ELP3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000542710 | ENST00000256398 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 3UTR-3CDS | ENST00000542710 | ENST00000521015 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 3UTR-3CDS | ENST00000542710 | ENST00000542181 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 3UTR-intron | ENST00000542710 | ENST00000380353 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 3UTR-intron | ENST00000542710 | ENST00000523760 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 3UTR-intron | ENST00000542710 | ENST00000524103 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 3UTR-intron | ENST00000542710 | ENST00000537665 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000328257 | ENST00000380353 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000328257 | ENST00000523760 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000328257 | ENST00000524103 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000328257 | ENST00000537665 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000398427 | ENST00000380353 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000398427 | ENST00000523760 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000398427 | ENST00000524103 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| 5CDS-intron | ENST00000398427 | ENST00000537665 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| In-frame | ENST00000328257 | ENST00000256398 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| In-frame | ENST00000328257 | ENST00000521015 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| In-frame | ENST00000328257 | ENST00000542181 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| In-frame | ENST00000398427 | ENST00000256398 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| In-frame | ENST00000398427 | ENST00000521015 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| In-frame | ENST00000398427 | ENST00000542181 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-3CDS | ENST00000543525 | ENST00000256398 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-3CDS | ENST00000543525 | ENST00000521015 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-3CDS | ENST00000543525 | ENST00000542181 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-intron | ENST00000543525 | ENST00000380353 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-intron | ENST00000543525 | ENST00000523760 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-intron | ENST00000543525 | ENST00000524103 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
| intron-intron | ENST00000543525 | ENST00000537665 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000328257 | PPME1 | chr11 | 73882567 | + | ENST00000521015 | ELP3 | chr8 | 27987019 | + | 2882 | 424 | 323 | 1450 | 375 |
| ENST00000328257 | PPME1 | chr11 | 73882567 | + | ENST00000256398 | ELP3 | chr8 | 27987019 | + | 2882 | 424 | 323 | 1450 | 375 |
| ENST00000328257 | PPME1 | chr11 | 73882567 | + | ENST00000542181 | ELP3 | chr8 | 27987019 | + | 1820 | 424 | 323 | 1450 | 375 |
| ENST00000398427 | PPME1 | chr11 | 73882567 | + | ENST00000521015 | ELP3 | chr8 | 27987019 | + | 2659 | 201 | 100 | 1227 | 375 |
| ENST00000398427 | PPME1 | chr11 | 73882567 | + | ENST00000256398 | ELP3 | chr8 | 27987019 | + | 2659 | 201 | 100 | 1227 | 375 |
| ENST00000398427 | PPME1 | chr11 | 73882567 | + | ENST00000542181 | ELP3 | chr8 | 27987019 | + | 1597 | 201 | 100 | 1227 | 375 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000328257 | ENST00000521015 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + | 0.001278604 | 0.9987214 |
| ENST00000328257 | ENST00000256398 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + | 0.001278604 | 0.9987214 |
| ENST00000328257 | ENST00000542181 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + | 0.002656855 | 0.9973431 |
| ENST00000398427 | ENST00000521015 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + | 0.001463176 | 0.9985368 |
| ENST00000398427 | ENST00000256398 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + | 0.001463176 | 0.9985368 |
| ENST00000398427 | ENST00000542181 | PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987019 | + | 0.00354945 | 0.99645054 |
Top |
Fusion Genomic Features for PPME1-ELP3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987018 | + | 0.000124341 | 0.99987566 |
| PPME1 | chr11 | 73882567 | + | ELP3 | chr8 | 27987018 | + | 0.000124341 | 0.99987566 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
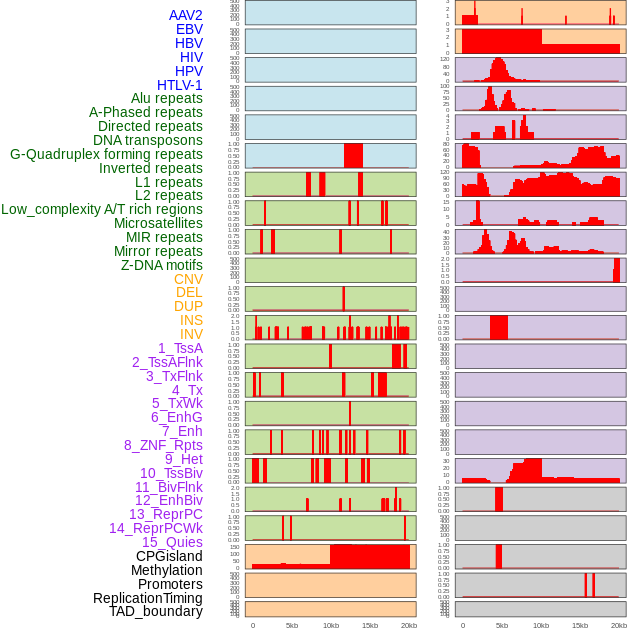 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
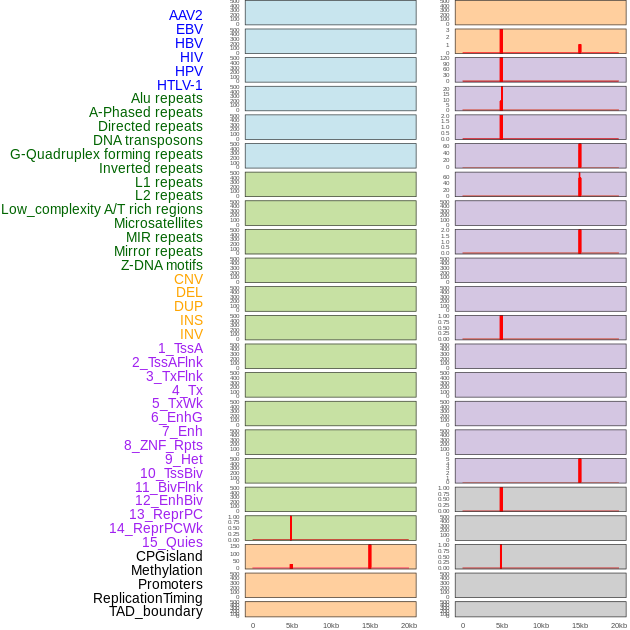 |
Top |
Fusion Protein Features for PPME1-ELP3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:73882567/chr8:27987019) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | ELP3 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Catalytic tRNA acetyltransferase subunit of the RNA polymerase II elongator complex, which is a component of the RNA polymerase II (Pol II) holoenzyme and is involved in transcriptional elongation (PubMed:11714725, PubMed:11818576, PubMed:15902492, PubMed:16713582). The elongator complex is required for multiple tRNA modifications, including mcm5U (5-methoxycarbonylmethyl uridine), mcm5s2U (5-methoxycarbonylmethyl-2-thiouridine), and ncm5U (5-carbamoylmethyl uridine) (PubMed:29415125). In the elongator complex, acts as a tRNA uridine(34) acetyltransferase by mediating formation of carboxymethyluridine in the wobble base at position 34 in tRNAs (By similarity). May also act as a protein lysine acetyltransferase by mediating acetylation of target proteins; such activity is however unclear in vivo and recent evidences suggest that ELP3 primarily acts as a tRNA acetyltransferase (PubMed:29415125). Involved in neurogenesis: regulates the migration and branching of projection neurons in the developing cerebral cortex, through a process depending on alpha-tubulin acetylation (PubMed:19185337). Required for acetylation of GJA1 in the developing cerebral cortex (By similarity). {ECO:0000250|UniProtKB:D5VRB9, ECO:0000250|UniProtKB:Q9CZX0, ECO:0000269|PubMed:11714725, ECO:0000269|PubMed:11818576, ECO:0000269|PubMed:15902492, ECO:0000269|PubMed:16713582, ECO:0000269|PubMed:19185337, ECO:0000269|PubMed:29415125}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000256398 | 6 | 15 | 396_547 | 205 | 548.0 | Domain | N-acetyltransferase | |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000521015 | 6 | 15 | 396_547 | 191 | 534.0 | Domain | N-acetyltransferase | |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000256398 | 6 | 15 | 474_477 | 205 | 548.0 | Region | Acetyl-CoA binding | |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000256398 | 6 | 15 | 497_499 | 205 | 548.0 | Region | Acetyl-CoA binding | |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000521015 | 6 | 15 | 474_477 | 191 | 534.0 | Region | Acetyl-CoA binding | |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000521015 | 6 | 15 | 497_499 | 191 | 534.0 | Region | Acetyl-CoA binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PPME1 | chr11:73882567 | chr8:27987019 | ENST00000328257 | + | 1 | 14 | 256_263 | 33 | 387.0 | Compositional bias | Note=Poly-Glu |
| Hgene | PPME1 | chr11:73882567 | chr8:27987019 | ENST00000543525 | + | 1 | 8 | 256_263 | 0 | 200.0 | Compositional bias | Note=Poly-Glu |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000256398 | 6 | 15 | 82_372 | 205 | 548.0 | Domain | Radical SAM core | |
| Tgene | ELP3 | chr11:73882567 | chr8:27987019 | ENST00000521015 | 6 | 15 | 82_372 | 191 | 534.0 | Domain | Radical SAM core |
Top |
Fusion Gene Sequence for PPME1-ELP3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >67684_67684_1_PPME1-ELP3_PPME1_chr11_73882567_ENST00000328257_ELP3_chr8_27987019_ENST00000256398_length(transcript)=2882nt_BP=424nt ATTGCGCAGGGTGCATTGCCCTGAAGAGCGCTCCTCCTTCCGGTGAAAGTCGTCGGGTTCCCAGCCGGGAGTCTGCGGAGTTCTCATACT CTGTTACCCAGCTTCCTTTGTATGAGTGCTGTTGCGCACTTCCGGTGGAAGCGCTGAAGCGAGTGGGTGGAGTTTACGGAGCCGGTGGGC GGTAGGCGGTGCTACGGGTAGCTGGGTGCTGTCCAAAGGCGACAGGGCGTCGTTAGGGGAGCGAGTCGTGACCGGTTGGGCCACACTCAA CGTGGGACGAAGCTTCGCCTACTGTTTGACTACGTGCGTGCAGCCTCCCCTCGATGTCGGCCCTCGAAAAGAGCATGCACCTCGGCCGCC TTCCCTCTCGCCCACCTCTACCCGGCAGCGGGGGCAGTCAGAGCGGAGCCAAGATGCGAATGGGGTATTCTGAGAGAAGCCTCACAAAGT GTATTGGAATTACTATTGAAACCAGACCAGATTACTGCATGAAGCGACATTTAAGTGACATGTTGACCTATGGCTGCACAAGGCTGGAGA TTGGGGTGCAGAGTGTTTATGAAGATGTGGCTAGAGACACCAACAGGGGCCACACTGTGAAGGCAGTGTGTGAGTCATTTCACCTGGCCA AAGATTCCGGTTTTAAAGTGGTGGCCCATATGATGCCTGACCTGCCAAACGTGGGACTAGAAAGAGACATTGAACAGTTCACAGAGTTTT TTGAGAACCCTGCTTTTCGTCCCGATGGGCTGAAACTCTATCCTACCCTGGTGATTCGTGGGACCGGGCTTTATGAGCTTTGGAAATCAG GAAGATATAAGAGTTACTCTCCTAGTGACCTGGTTGAATTGGTGGCTCGGATCCTAGCCCTCGTGCCTCCATGGACTCGAGTGTACCGAG TACAGAGGGATATTCCAATGCCTTTAGTTAGCTCAGGAGTAGAGCATGGTAACCTGAGAGAGCTGGCACTTGCAAGAATGAAAGACCTCG GAATACAGTGTCGAGATGTGAGAACCAGAGAAGTTGGAATCCAAGAAATTCATCACAAAGTACGGCCATACCAGGTTGAATTGGTAAGGA GAGATTATGTTGCAAATGGTGGCTGGGAAACATTCTTGTCATACGAAGACCCAGATCAAGACATTTTGATTGGCCTCCTACGATTACGCA AGTGTTCAGAAGAAACTTTCCGTTTCGAATTGGGTGGAGGTGTCTCCATAGTACGAGAGCTGCATGTGTATGGGAGTGTGGTCCCTGTGA GCAGCCGGGATCCTACTAAATTTCAGCATCAGGGATTTGGCATGCTGCTGATGGAGGAAGCAGAAAGAATAGCTAGAGAAGAACATGGGT CTGGGAAAATCGCTGTGATATCAGGGGTCGGCACCAGGAATTATTATAGAAAGATCGGCTACAGATTACAAGGCCCGTACATGGTGAAGA TGCTGAAATAATGGCCACACCAGTCCACTCTTCTGCAGTATCCTCCCTGGCAGAACACGGAGAATCAGGATTTCTTAAATACTCAACAGA GAGGCTGAGCAGAGCAAATGGGGGGCTTCACCCTCATCCCGCAGCTGCAGAGACTGGAAACTGCCTTCAAGGCCACGGCTGGTCATCTGC TGACCACACCCCAGATCCGCCCTCTCCTGCGTGCACCCCAAAAAATCACTTGCGTTTTTGAGGCTTAAATCATCTATCCAGTTTCTACAT TTTGCATGAGGCCTGCAGGTGGCCTATTTTGACTCAGACGGTGAAAAAAGCAAATTAACTCATTTGGACACCATAACTCATGCAATAAAA CTGATTGTCATTCGAGGAGCAAACTTAAGAGTAGTTTATTTATATACCCTGGGGACAGAAAGTCAGGTTGAAACAGGAAAACCACCAGAC TCTAATCTCAGCCCTTTAACGACATACGCATTGGAGCGCAAGTTAGGAAAATGAGCTTTTGTTTTCATGGAAATCATTCTGATTACAGTG CTGATGTTTAGAAATAAATAGCAGTGTGACTGGGAAAGAGGAATTGCAGTTGTGGGGGTGTGAGCCTGGCAGCAGCCAGCCAGCAGCCTC TCCCAGGCGGGAGTCTACCATCCGAGACGGCGATGACAAAGAGCTTCATTCCACATTCTTTGTTATCTCTACTTCCCACCCTCTTGGCAA CTACAGAGCAGTGTGGGCAGCCCCAAGTGTGGTCCCCAGAGAGCGTTTGGCTTTCCTGTCTGTCTATCCTGAGCGGGTGGAGTCTCAGGT TGTGTGCCCCTAAATCAAGATTTGCTTCCACAGAAGCCATTACTTGCAATTTTTTTTTTTTTTCTGAGAAAGTCTCGCTGTGTCACCCAG GCTGGAGTGCAGTGGCGCAATCTCACTGCATCCTCCGCCTCCCGGGTTCAAGCGATTCTCCCGCCTCAGCCTCCTGAGTAGCTGGGATTA CAGGCACCCGCCGCTGCTAATTTTTGTATTTTTAGTAGAGATGGGGGTTTCACCATATTGGTCAGGCTGGTCTCGAACTCCTGACCTCAG GTGATCAACCCACCTTGGCCTCCCTAAATGCCGGGATTACAGGCATGAGCCACCGCTCCCAGCCTTTGATTTTTTAAGGTGGATTTTGGT TGTTATAAATGGAGAAAGGTAAGAGTTCAAGTTCAACCCGTGTGTGAAAGCAAAACAATGGAAAACAGGATTGGCTTCTTCAAAGGCTCC TCTTGTAGAACTGCCTCTTTGAAATTTCGAGGTAATCTACTTTGGAGACTCTGCCTGGAGAGGGTCAGTTCCTAAGTTAAAAGCATCGCT TAACCTTGGCTCCTGTGGCATTTTACAAAGGTTTAAAGGAATTGATTCCTCTGAAAGGGCCTGAAAATAAAAAGTCTTTAACATACAAAG >67684_67684_1_PPME1-ELP3_PPME1_chr11_73882567_ENST00000328257_ELP3_chr8_27987019_ENST00000256398_length(amino acids)=375AA_BP=34 MSALEKSMHLGRLPSRPPLPGSGGSQSGAKMRMGYSERSLTKCIGITIETRPDYCMKRHLSDMLTYGCTRLEIGVQSVYEDVARDTNRGH TVKAVCESFHLAKDSGFKVVAHMMPDLPNVGLERDIEQFTEFFENPAFRPDGLKLYPTLVIRGTGLYELWKSGRYKSYSPSDLVELVARI LALVPPWTRVYRVQRDIPMPLVSSGVEHGNLRELALARMKDLGIQCRDVRTREVGIQEIHHKVRPYQVELVRRDYVANGGWETFLSYEDP DQDILIGLLRLRKCSEETFRFELGGGVSIVRELHVYGSVVPVSSRDPTKFQHQGFGMLLMEEAERIAREEHGSGKIAVISGVGTRNYYRK -------------------------------------------------------------- >67684_67684_2_PPME1-ELP3_PPME1_chr11_73882567_ENST00000328257_ELP3_chr8_27987019_ENST00000521015_length(transcript)=2882nt_BP=424nt ATTGCGCAGGGTGCATTGCCCTGAAGAGCGCTCCTCCTTCCGGTGAAAGTCGTCGGGTTCCCAGCCGGGAGTCTGCGGAGTTCTCATACT CTGTTACCCAGCTTCCTTTGTATGAGTGCTGTTGCGCACTTCCGGTGGAAGCGCTGAAGCGAGTGGGTGGAGTTTACGGAGCCGGTGGGC GGTAGGCGGTGCTACGGGTAGCTGGGTGCTGTCCAAAGGCGACAGGGCGTCGTTAGGGGAGCGAGTCGTGACCGGTTGGGCCACACTCAA CGTGGGACGAAGCTTCGCCTACTGTTTGACTACGTGCGTGCAGCCTCCCCTCGATGTCGGCCCTCGAAAAGAGCATGCACCTCGGCCGCC TTCCCTCTCGCCCACCTCTACCCGGCAGCGGGGGCAGTCAGAGCGGAGCCAAGATGCGAATGGGGTATTCTGAGAGAAGCCTCACAAAGT GTATTGGAATTACTATTGAAACCAGACCAGATTACTGCATGAAGCGACATTTAAGTGACATGTTGACCTATGGCTGCACAAGGCTGGAGA TTGGGGTGCAGAGTGTTTATGAAGATGTGGCTAGAGACACCAACAGGGGCCACACTGTGAAGGCAGTGTGTGAGTCATTTCACCTGGCCA AAGATTCCGGTTTTAAAGTGGTGGCCCATATGATGCCTGACCTGCCAAACGTGGGACTAGAAAGAGACATTGAACAGTTCACAGAGTTTT TTGAGAACCCTGCTTTTCGTCCCGATGGGCTGAAACTCTATCCTACCCTGGTGATTCGTGGGACCGGGCTTTATGAGCTTTGGAAATCAG GAAGATATAAGAGTTACTCTCCTAGTGACCTGGTTGAATTGGTGGCTCGGATCCTAGCCCTCGTGCCTCCATGGACTCGAGTGTACCGAG TACAGAGGGATATTCCAATGCCTTTAGTTAGCTCAGGAGTAGAGCATGGTAACCTGAGAGAGCTGGCACTTGCAAGAATGAAAGACCTCG GAATACAGTGTCGAGATGTGAGAACCAGAGAAGTTGGAATCCAAGAAATTCATCACAAAGTACGGCCATACCAGGTTGAATTGGTAAGGA GAGATTATGTTGCAAATGGTGGCTGGGAAACATTCTTGTCATACGAAGACCCAGATCAAGACATTTTGATTGGCCTCCTACGATTACGCA AGTGTTCAGAAGAAACTTTCCGTTTCGAATTGGGTGGAGGTGTCTCCATAGTACGAGAGCTGCATGTGTATGGGAGTGTGGTCCCTGTGA GCAGCCGGGATCCTACTAAATTTCAGCATCAGGGATTTGGCATGCTGCTGATGGAGGAAGCAGAAAGAATAGCTAGAGAAGAACATGGGT CTGGGAAAATCGCTGTGATATCAGGGGTCGGCACCAGGAATTATTATAGAAAGATCGGCTACAGATTACAAGGCCCGTACATGGTGAAGA TGCTGAAATAATGGCCACACCAGTCCACTCTTCTGCAGTATCCTCCCTGGCAGAACACGGAGAATCAGGATTTCTTAAATACTCAACAGA GAGGCTGAGCAGAGCAAATGGGGGGCTTCACCCTCATCCCGCAGCTGCAGAGACTGGAAACTGCCTTCAAGGCCACGGCTGGTCATCTGC TGACCACACCCCAGATCCGCCCTCTCCTGCGTGCACCCCAAAAAATCACTTGCGTTTTTGAGGCTTAAATCATCTATCCAGTTTCTACAT TTTGCATGAGGCCTGCAGGTGGCCTATTTTGACTCAGACGGTGAAAAAAGCAAATTAACTCATTTGGACACCATAACTCATGCAATAAAA CTGATTGTCATTCGAGGAGCAAACTTAAGAGTAGTTTATTTATATACCCTGGGGACAGAAAGTCAGGTTGAAACAGGAAAACCACCAGAC TCTAATCTCAGCCCTTTAACGACATACGCATTGGAGCGCAAGTTAGGAAAATGAGCTTTTGTTTTCATGGAAATCATTCTGATTACAGTG CTGATGTTTAGAAATAAATAGCAGTGTGACTGGGAAAGAGGAATTGCAGTTGTGGGGGTGTGAGCCTGGCAGCAGCCAGCCAGCAGCCTC TCCCAGGCGGGAGTCTACCATCCGAGACGGCGATGACAAAGAGCTTCATTCCACATTCTTTGTTATCTCTACTTCCCACCCTCTTGGCAA CTACAGAGCAGTGTGGGCAGCCCCAAGTGTGGTCCCCAGAGAGCGTTTGGCTTTCCTGTCTGTCTATCCTGAGCGGGTGGAGTCTCAGGT TGTGTGCCCCTAAATCAAGATTTGCTTCCACAGAAGCCATTACTTGCAATTTTTTTTTTTTTTCTGAGAAAGTCTCGCTGTGTCACCCAG GCTGGAGTGCAGTGGCGCAATCTCACTGCATCCTCCGCCTCCCGGGTTCAAGCGATTCTCCCGCCTCAGCCTCCTGAGTAGCTGGGATTA CAGGCACCCGCCGCTGCTAATTTTTGTATTTTTAGTAGAGATGGGGGTTTCACCATATTGGTCAGGCTGGTCTCGAACTCCTGACCTCAG GTGATCAACCCACCTTGGCCTCCCTAAATGCCGGGATTACAGGCATGAGCCACCGCTCCCAGCCTTTGATTTTTTAAGGTGGATTTTGGT TGTTATAAATGGAGAAAGGTAAGAGTTCAAGTTCAACCCGTGTGTGAAAGCAAAACAATGGAAAACAGGATTGGCTTCTTCAAAGGCTCC TCTTGTAGAACTGCCTCTTTGAAATTTCGAGGTAATCTACTTTGGAGACTCTGCCTGGAGAGGGTCAGTTCCTAAGTTAAAAGCATCGCT TAACCTTGGCTCCTGTGGCATTTTACAAAGGTTTAAAGGAATTGATTCCTCTGAAAGGGCCTGAAAATAAAAAGTCTTTAACATACAAAG >67684_67684_2_PPME1-ELP3_PPME1_chr11_73882567_ENST00000328257_ELP3_chr8_27987019_ENST00000521015_length(amino acids)=375AA_BP=34 MSALEKSMHLGRLPSRPPLPGSGGSQSGAKMRMGYSERSLTKCIGITIETRPDYCMKRHLSDMLTYGCTRLEIGVQSVYEDVARDTNRGH TVKAVCESFHLAKDSGFKVVAHMMPDLPNVGLERDIEQFTEFFENPAFRPDGLKLYPTLVIRGTGLYELWKSGRYKSYSPSDLVELVARI LALVPPWTRVYRVQRDIPMPLVSSGVEHGNLRELALARMKDLGIQCRDVRTREVGIQEIHHKVRPYQVELVRRDYVANGGWETFLSYEDP DQDILIGLLRLRKCSEETFRFELGGGVSIVRELHVYGSVVPVSSRDPTKFQHQGFGMLLMEEAERIAREEHGSGKIAVISGVGTRNYYRK -------------------------------------------------------------- >67684_67684_3_PPME1-ELP3_PPME1_chr11_73882567_ENST00000328257_ELP3_chr8_27987019_ENST00000542181_length(transcript)=1820nt_BP=424nt ATTGCGCAGGGTGCATTGCCCTGAAGAGCGCTCCTCCTTCCGGTGAAAGTCGTCGGGTTCCCAGCCGGGAGTCTGCGGAGTTCTCATACT CTGTTACCCAGCTTCCTTTGTATGAGTGCTGTTGCGCACTTCCGGTGGAAGCGCTGAAGCGAGTGGGTGGAGTTTACGGAGCCGGTGGGC GGTAGGCGGTGCTACGGGTAGCTGGGTGCTGTCCAAAGGCGACAGGGCGTCGTTAGGGGAGCGAGTCGTGACCGGTTGGGCCACACTCAA CGTGGGACGAAGCTTCGCCTACTGTTTGACTACGTGCGTGCAGCCTCCCCTCGATGTCGGCCCTCGAAAAGAGCATGCACCTCGGCCGCC TTCCCTCTCGCCCACCTCTACCCGGCAGCGGGGGCAGTCAGAGCGGAGCCAAGATGCGAATGGGGTATTCTGAGAGAAGCCTCACAAAGT GTATTGGAATTACTATTGAAACCAGACCAGATTACTGCATGAAGCGACATTTAAGTGACATGTTGACCTATGGCTGCACAAGGCTGGAGA TTGGGGTGCAGAGTGTTTATGAAGATGTGGCTAGAGACACCAACAGGGGCCACACTGTGAAGGCAGTGTGTGAGTCATTTCACCTGGCCA AAGATTCCGGTTTTAAAGTGGTGGCCCATATGATGCCTGACCTGCCAAACGTGGGACTAGAAAGAGACATTGAACAGTTCACAGAGTTTT TTGAGAACCCTGCTTTTCGTCCCGATGGGCTGAAACTCTATCCTACCCTGGTGATTCGTGGGACCGGGCTTTATGAGCTTTGGAAATCAG GAAGATATAAGAGTTACTCTCCTAGTGACCTGGTTGAATTGGTGGCTCGGATCCTAGCCCTCGTGCCTCCATGGACTCGAGTGTACCGAG TACAGAGGGATATTCCAATGCCTTTAGTTAGCTCAGGAGTAGAGCATGGTAACCTGAGAGAGCTGGCACTTGCAAGAATGAAAGACCTCG GAATACAGTGTCGAGATGTGAGAACCAGAGAAGTTGGAATCCAAGAAATTCATCACAAAGTACGGCCATACCAGGTTGAATTGGTAAGGA GAGATTATGTTGCAAATGGTGGCTGGGAAACATTCTTGTCATACGAAGACCCAGATCAAGACATTTTGATTGGCCTCCTACGATTACGCA AGTGTTCAGAAGAAACTTTCCGTTTCGAATTGGGTGGAGGTGTCTCCATAGTACGAGAGCTGCATGTGTATGGGAGTGTGGTCCCTGTGA GCAGCCGGGATCCTACTAAATTTCAGCATCAGGGATTTGGCATGCTGCTGATGGAGGAAGCAGAAAGAATAGCTAGAGAAGAACATGGGT CTGGGAAAATCGCTGTGATATCAGGGGTCGGCACCAGGAATTATTATAGAAAGATCGGCTACAGATTACAAGGCCCGTACATGGTGAAGA TGCTGAAATAATGGCCACACCAGTCCACTCTTCTGCAGTATCCTCCCTGGCAGAACACGGAGAATCAGGATTTCTTAAATACTCAACAGA GAGGCTGAGCAGAGCAAATGGGGGGCTTCACCCTCATCCCGCAGCTGCAGAGACTGGAAACTGCCTTCAAGGCCACGGCTGGTCATCTGC TGACCACACCCCAGATCCGCCCTCTCCTGCGTGCACCCCAAAAAATCACTTGCGTTTTTGAGGCTTAAATCATCTATCCAGTTTCTACAT TTTGCATGAGGCCTGCAGGTGGCCTATTTTGACTCAGACGGTGAAAAAAGCAAATTAACTCATTTGGACACCATAACTCATGCAATAAAA >67684_67684_3_PPME1-ELP3_PPME1_chr11_73882567_ENST00000328257_ELP3_chr8_27987019_ENST00000542181_length(amino acids)=375AA_BP=34 MSALEKSMHLGRLPSRPPLPGSGGSQSGAKMRMGYSERSLTKCIGITIETRPDYCMKRHLSDMLTYGCTRLEIGVQSVYEDVARDTNRGH TVKAVCESFHLAKDSGFKVVAHMMPDLPNVGLERDIEQFTEFFENPAFRPDGLKLYPTLVIRGTGLYELWKSGRYKSYSPSDLVELVARI LALVPPWTRVYRVQRDIPMPLVSSGVEHGNLRELALARMKDLGIQCRDVRTREVGIQEIHHKVRPYQVELVRRDYVANGGWETFLSYEDP DQDILIGLLRLRKCSEETFRFELGGGVSIVRELHVYGSVVPVSSRDPTKFQHQGFGMLLMEEAERIAREEHGSGKIAVISGVGTRNYYRK -------------------------------------------------------------- >67684_67684_4_PPME1-ELP3_PPME1_chr11_73882567_ENST00000398427_ELP3_chr8_27987019_ENST00000256398_length(transcript)=2659nt_BP=201nt AGGGCGTCGTTAGGGGAGCGAGTCGTGACCGGTTGGGCCACACTCAACGTGGGACGAAGCTTCGCCTACTGTTTGACTACGTGCGTGCAG CCTCCCCTCGATGTCGGCCCTCGAAAAGAGCATGCACCTCGGCCGCCTTCCCTCTCGCCCACCTCTACCCGGCAGCGGGGGCAGTCAGAG CGGAGCCAAGATGCGAATGGGGTATTCTGAGAGAAGCCTCACAAAGTGTATTGGAATTACTATTGAAACCAGACCAGATTACTGCATGAA GCGACATTTAAGTGACATGTTGACCTATGGCTGCACAAGGCTGGAGATTGGGGTGCAGAGTGTTTATGAAGATGTGGCTAGAGACACCAA CAGGGGCCACACTGTGAAGGCAGTGTGTGAGTCATTTCACCTGGCCAAAGATTCCGGTTTTAAAGTGGTGGCCCATATGATGCCTGACCT GCCAAACGTGGGACTAGAAAGAGACATTGAACAGTTCACAGAGTTTTTTGAGAACCCTGCTTTTCGTCCCGATGGGCTGAAACTCTATCC TACCCTGGTGATTCGTGGGACCGGGCTTTATGAGCTTTGGAAATCAGGAAGATATAAGAGTTACTCTCCTAGTGACCTGGTTGAATTGGT GGCTCGGATCCTAGCCCTCGTGCCTCCATGGACTCGAGTGTACCGAGTACAGAGGGATATTCCAATGCCTTTAGTTAGCTCAGGAGTAGA GCATGGTAACCTGAGAGAGCTGGCACTTGCAAGAATGAAAGACCTCGGAATACAGTGTCGAGATGTGAGAACCAGAGAAGTTGGAATCCA AGAAATTCATCACAAAGTACGGCCATACCAGGTTGAATTGGTAAGGAGAGATTATGTTGCAAATGGTGGCTGGGAAACATTCTTGTCATA CGAAGACCCAGATCAAGACATTTTGATTGGCCTCCTACGATTACGCAAGTGTTCAGAAGAAACTTTCCGTTTCGAATTGGGTGGAGGTGT CTCCATAGTACGAGAGCTGCATGTGTATGGGAGTGTGGTCCCTGTGAGCAGCCGGGATCCTACTAAATTTCAGCATCAGGGATTTGGCAT GCTGCTGATGGAGGAAGCAGAAAGAATAGCTAGAGAAGAACATGGGTCTGGGAAAATCGCTGTGATATCAGGGGTCGGCACCAGGAATTA TTATAGAAAGATCGGCTACAGATTACAAGGCCCGTACATGGTGAAGATGCTGAAATAATGGCCACACCAGTCCACTCTTCTGCAGTATCC TCCCTGGCAGAACACGGAGAATCAGGATTTCTTAAATACTCAACAGAGAGGCTGAGCAGAGCAAATGGGGGGCTTCACCCTCATCCCGCA GCTGCAGAGACTGGAAACTGCCTTCAAGGCCACGGCTGGTCATCTGCTGACCACACCCCAGATCCGCCCTCTCCTGCGTGCACCCCAAAA AATCACTTGCGTTTTTGAGGCTTAAATCATCTATCCAGTTTCTACATTTTGCATGAGGCCTGCAGGTGGCCTATTTTGACTCAGACGGTG AAAAAAGCAAATTAACTCATTTGGACACCATAACTCATGCAATAAAACTGATTGTCATTCGAGGAGCAAACTTAAGAGTAGTTTATTTAT ATACCCTGGGGACAGAAAGTCAGGTTGAAACAGGAAAACCACCAGACTCTAATCTCAGCCCTTTAACGACATACGCATTGGAGCGCAAGT TAGGAAAATGAGCTTTTGTTTTCATGGAAATCATTCTGATTACAGTGCTGATGTTTAGAAATAAATAGCAGTGTGACTGGGAAAGAGGAA TTGCAGTTGTGGGGGTGTGAGCCTGGCAGCAGCCAGCCAGCAGCCTCTCCCAGGCGGGAGTCTACCATCCGAGACGGCGATGACAAAGAG CTTCATTCCACATTCTTTGTTATCTCTACTTCCCACCCTCTTGGCAACTACAGAGCAGTGTGGGCAGCCCCAAGTGTGGTCCCCAGAGAG CGTTTGGCTTTCCTGTCTGTCTATCCTGAGCGGGTGGAGTCTCAGGTTGTGTGCCCCTAAATCAAGATTTGCTTCCACAGAAGCCATTAC TTGCAATTTTTTTTTTTTTTCTGAGAAAGTCTCGCTGTGTCACCCAGGCTGGAGTGCAGTGGCGCAATCTCACTGCATCCTCCGCCTCCC GGGTTCAAGCGATTCTCCCGCCTCAGCCTCCTGAGTAGCTGGGATTACAGGCACCCGCCGCTGCTAATTTTTGTATTTTTAGTAGAGATG GGGGTTTCACCATATTGGTCAGGCTGGTCTCGAACTCCTGACCTCAGGTGATCAACCCACCTTGGCCTCCCTAAATGCCGGGATTACAGG CATGAGCCACCGCTCCCAGCCTTTGATTTTTTAAGGTGGATTTTGGTTGTTATAAATGGAGAAAGGTAAGAGTTCAAGTTCAACCCGTGT GTGAAAGCAAAACAATGGAAAACAGGATTGGCTTCTTCAAAGGCTCCTCTTGTAGAACTGCCTCTTTGAAATTTCGAGGTAATCTACTTT GGAGACTCTGCCTGGAGAGGGTCAGTTCCTAAGTTAAAAGCATCGCTTAACCTTGGCTCCTGTGGCATTTTACAAAGGTTTAAAGGAATT >67684_67684_4_PPME1-ELP3_PPME1_chr11_73882567_ENST00000398427_ELP3_chr8_27987019_ENST00000256398_length(amino acids)=375AA_BP=34 MSALEKSMHLGRLPSRPPLPGSGGSQSGAKMRMGYSERSLTKCIGITIETRPDYCMKRHLSDMLTYGCTRLEIGVQSVYEDVARDTNRGH TVKAVCESFHLAKDSGFKVVAHMMPDLPNVGLERDIEQFTEFFENPAFRPDGLKLYPTLVIRGTGLYELWKSGRYKSYSPSDLVELVARI LALVPPWTRVYRVQRDIPMPLVSSGVEHGNLRELALARMKDLGIQCRDVRTREVGIQEIHHKVRPYQVELVRRDYVANGGWETFLSYEDP DQDILIGLLRLRKCSEETFRFELGGGVSIVRELHVYGSVVPVSSRDPTKFQHQGFGMLLMEEAERIAREEHGSGKIAVISGVGTRNYYRK -------------------------------------------------------------- >67684_67684_5_PPME1-ELP3_PPME1_chr11_73882567_ENST00000398427_ELP3_chr8_27987019_ENST00000521015_length(transcript)=2659nt_BP=201nt AGGGCGTCGTTAGGGGAGCGAGTCGTGACCGGTTGGGCCACACTCAACGTGGGACGAAGCTTCGCCTACTGTTTGACTACGTGCGTGCAG CCTCCCCTCGATGTCGGCCCTCGAAAAGAGCATGCACCTCGGCCGCCTTCCCTCTCGCCCACCTCTACCCGGCAGCGGGGGCAGTCAGAG CGGAGCCAAGATGCGAATGGGGTATTCTGAGAGAAGCCTCACAAAGTGTATTGGAATTACTATTGAAACCAGACCAGATTACTGCATGAA GCGACATTTAAGTGACATGTTGACCTATGGCTGCACAAGGCTGGAGATTGGGGTGCAGAGTGTTTATGAAGATGTGGCTAGAGACACCAA CAGGGGCCACACTGTGAAGGCAGTGTGTGAGTCATTTCACCTGGCCAAAGATTCCGGTTTTAAAGTGGTGGCCCATATGATGCCTGACCT GCCAAACGTGGGACTAGAAAGAGACATTGAACAGTTCACAGAGTTTTTTGAGAACCCTGCTTTTCGTCCCGATGGGCTGAAACTCTATCC TACCCTGGTGATTCGTGGGACCGGGCTTTATGAGCTTTGGAAATCAGGAAGATATAAGAGTTACTCTCCTAGTGACCTGGTTGAATTGGT GGCTCGGATCCTAGCCCTCGTGCCTCCATGGACTCGAGTGTACCGAGTACAGAGGGATATTCCAATGCCTTTAGTTAGCTCAGGAGTAGA GCATGGTAACCTGAGAGAGCTGGCACTTGCAAGAATGAAAGACCTCGGAATACAGTGTCGAGATGTGAGAACCAGAGAAGTTGGAATCCA AGAAATTCATCACAAAGTACGGCCATACCAGGTTGAATTGGTAAGGAGAGATTATGTTGCAAATGGTGGCTGGGAAACATTCTTGTCATA CGAAGACCCAGATCAAGACATTTTGATTGGCCTCCTACGATTACGCAAGTGTTCAGAAGAAACTTTCCGTTTCGAATTGGGTGGAGGTGT CTCCATAGTACGAGAGCTGCATGTGTATGGGAGTGTGGTCCCTGTGAGCAGCCGGGATCCTACTAAATTTCAGCATCAGGGATTTGGCAT GCTGCTGATGGAGGAAGCAGAAAGAATAGCTAGAGAAGAACATGGGTCTGGGAAAATCGCTGTGATATCAGGGGTCGGCACCAGGAATTA TTATAGAAAGATCGGCTACAGATTACAAGGCCCGTACATGGTGAAGATGCTGAAATAATGGCCACACCAGTCCACTCTTCTGCAGTATCC TCCCTGGCAGAACACGGAGAATCAGGATTTCTTAAATACTCAACAGAGAGGCTGAGCAGAGCAAATGGGGGGCTTCACCCTCATCCCGCA GCTGCAGAGACTGGAAACTGCCTTCAAGGCCACGGCTGGTCATCTGCTGACCACACCCCAGATCCGCCCTCTCCTGCGTGCACCCCAAAA AATCACTTGCGTTTTTGAGGCTTAAATCATCTATCCAGTTTCTACATTTTGCATGAGGCCTGCAGGTGGCCTATTTTGACTCAGACGGTG AAAAAAGCAAATTAACTCATTTGGACACCATAACTCATGCAATAAAACTGATTGTCATTCGAGGAGCAAACTTAAGAGTAGTTTATTTAT ATACCCTGGGGACAGAAAGTCAGGTTGAAACAGGAAAACCACCAGACTCTAATCTCAGCCCTTTAACGACATACGCATTGGAGCGCAAGT TAGGAAAATGAGCTTTTGTTTTCATGGAAATCATTCTGATTACAGTGCTGATGTTTAGAAATAAATAGCAGTGTGACTGGGAAAGAGGAA TTGCAGTTGTGGGGGTGTGAGCCTGGCAGCAGCCAGCCAGCAGCCTCTCCCAGGCGGGAGTCTACCATCCGAGACGGCGATGACAAAGAG CTTCATTCCACATTCTTTGTTATCTCTACTTCCCACCCTCTTGGCAACTACAGAGCAGTGTGGGCAGCCCCAAGTGTGGTCCCCAGAGAG CGTTTGGCTTTCCTGTCTGTCTATCCTGAGCGGGTGGAGTCTCAGGTTGTGTGCCCCTAAATCAAGATTTGCTTCCACAGAAGCCATTAC TTGCAATTTTTTTTTTTTTTCTGAGAAAGTCTCGCTGTGTCACCCAGGCTGGAGTGCAGTGGCGCAATCTCACTGCATCCTCCGCCTCCC GGGTTCAAGCGATTCTCCCGCCTCAGCCTCCTGAGTAGCTGGGATTACAGGCACCCGCCGCTGCTAATTTTTGTATTTTTAGTAGAGATG GGGGTTTCACCATATTGGTCAGGCTGGTCTCGAACTCCTGACCTCAGGTGATCAACCCACCTTGGCCTCCCTAAATGCCGGGATTACAGG CATGAGCCACCGCTCCCAGCCTTTGATTTTTTAAGGTGGATTTTGGTTGTTATAAATGGAGAAAGGTAAGAGTTCAAGTTCAACCCGTGT GTGAAAGCAAAACAATGGAAAACAGGATTGGCTTCTTCAAAGGCTCCTCTTGTAGAACTGCCTCTTTGAAATTTCGAGGTAATCTACTTT GGAGACTCTGCCTGGAGAGGGTCAGTTCCTAAGTTAAAAGCATCGCTTAACCTTGGCTCCTGTGGCATTTTACAAAGGTTTAAAGGAATT >67684_67684_5_PPME1-ELP3_PPME1_chr11_73882567_ENST00000398427_ELP3_chr8_27987019_ENST00000521015_length(amino acids)=375AA_BP=34 MSALEKSMHLGRLPSRPPLPGSGGSQSGAKMRMGYSERSLTKCIGITIETRPDYCMKRHLSDMLTYGCTRLEIGVQSVYEDVARDTNRGH TVKAVCESFHLAKDSGFKVVAHMMPDLPNVGLERDIEQFTEFFENPAFRPDGLKLYPTLVIRGTGLYELWKSGRYKSYSPSDLVELVARI LALVPPWTRVYRVQRDIPMPLVSSGVEHGNLRELALARMKDLGIQCRDVRTREVGIQEIHHKVRPYQVELVRRDYVANGGWETFLSYEDP DQDILIGLLRLRKCSEETFRFELGGGVSIVRELHVYGSVVPVSSRDPTKFQHQGFGMLLMEEAERIAREEHGSGKIAVISGVGTRNYYRK -------------------------------------------------------------- >67684_67684_6_PPME1-ELP3_PPME1_chr11_73882567_ENST00000398427_ELP3_chr8_27987019_ENST00000542181_length(transcript)=1597nt_BP=201nt AGGGCGTCGTTAGGGGAGCGAGTCGTGACCGGTTGGGCCACACTCAACGTGGGACGAAGCTTCGCCTACTGTTTGACTACGTGCGTGCAG CCTCCCCTCGATGTCGGCCCTCGAAAAGAGCATGCACCTCGGCCGCCTTCCCTCTCGCCCACCTCTACCCGGCAGCGGGGGCAGTCAGAG CGGAGCCAAGATGCGAATGGGGTATTCTGAGAGAAGCCTCACAAAGTGTATTGGAATTACTATTGAAACCAGACCAGATTACTGCATGAA GCGACATTTAAGTGACATGTTGACCTATGGCTGCACAAGGCTGGAGATTGGGGTGCAGAGTGTTTATGAAGATGTGGCTAGAGACACCAA CAGGGGCCACACTGTGAAGGCAGTGTGTGAGTCATTTCACCTGGCCAAAGATTCCGGTTTTAAAGTGGTGGCCCATATGATGCCTGACCT GCCAAACGTGGGACTAGAAAGAGACATTGAACAGTTCACAGAGTTTTTTGAGAACCCTGCTTTTCGTCCCGATGGGCTGAAACTCTATCC TACCCTGGTGATTCGTGGGACCGGGCTTTATGAGCTTTGGAAATCAGGAAGATATAAGAGTTACTCTCCTAGTGACCTGGTTGAATTGGT GGCTCGGATCCTAGCCCTCGTGCCTCCATGGACTCGAGTGTACCGAGTACAGAGGGATATTCCAATGCCTTTAGTTAGCTCAGGAGTAGA GCATGGTAACCTGAGAGAGCTGGCACTTGCAAGAATGAAAGACCTCGGAATACAGTGTCGAGATGTGAGAACCAGAGAAGTTGGAATCCA AGAAATTCATCACAAAGTACGGCCATACCAGGTTGAATTGGTAAGGAGAGATTATGTTGCAAATGGTGGCTGGGAAACATTCTTGTCATA CGAAGACCCAGATCAAGACATTTTGATTGGCCTCCTACGATTACGCAAGTGTTCAGAAGAAACTTTCCGTTTCGAATTGGGTGGAGGTGT CTCCATAGTACGAGAGCTGCATGTGTATGGGAGTGTGGTCCCTGTGAGCAGCCGGGATCCTACTAAATTTCAGCATCAGGGATTTGGCAT GCTGCTGATGGAGGAAGCAGAAAGAATAGCTAGAGAAGAACATGGGTCTGGGAAAATCGCTGTGATATCAGGGGTCGGCACCAGGAATTA TTATAGAAAGATCGGCTACAGATTACAAGGCCCGTACATGGTGAAGATGCTGAAATAATGGCCACACCAGTCCACTCTTCTGCAGTATCC TCCCTGGCAGAACACGGAGAATCAGGATTTCTTAAATACTCAACAGAGAGGCTGAGCAGAGCAAATGGGGGGCTTCACCCTCATCCCGCA GCTGCAGAGACTGGAAACTGCCTTCAAGGCCACGGCTGGTCATCTGCTGACCACACCCCAGATCCGCCCTCTCCTGCGTGCACCCCAAAA AATCACTTGCGTTTTTGAGGCTTAAATCATCTATCCAGTTTCTACATTTTGCATGAGGCCTGCAGGTGGCCTATTTTGACTCAGACGGTG >67684_67684_6_PPME1-ELP3_PPME1_chr11_73882567_ENST00000398427_ELP3_chr8_27987019_ENST00000542181_length(amino acids)=375AA_BP=34 MSALEKSMHLGRLPSRPPLPGSGGSQSGAKMRMGYSERSLTKCIGITIETRPDYCMKRHLSDMLTYGCTRLEIGVQSVYEDVARDTNRGH TVKAVCESFHLAKDSGFKVVAHMMPDLPNVGLERDIEQFTEFFENPAFRPDGLKLYPTLVIRGTGLYELWKSGRYKSYSPSDLVELVARI LALVPPWTRVYRVQRDIPMPLVSSGVEHGNLRELALARMKDLGIQCRDVRTREVGIQEIHHKVRPYQVELVRRDYVANGGWETFLSYEDP DQDILIGLLRLRKCSEETFRFELGGGVSIVRELHVYGSVVPVSSRDPTKFQHQGFGMLLMEEAERIAREEHGSGKIAVISGVGTRNYYRK -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PPME1-ELP3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PPME1-ELP3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PPME1-ELP3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
