
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:PPP3CA-SLC39A8 (FusionGDB2 ID:68132) |
Fusion Gene Summary for PPP3CA-SLC39A8 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: PPP3CA-SLC39A8 | Fusion gene ID: 68132 | Hgene | Tgene | Gene symbol | PPP3CA | SLC39A8 | Gene ID | 5530 | 64116 |
| Gene name | protein phosphatase 3 catalytic subunit alpha | solute carrier family 39 member 8 | |
| Synonyms | ACCIID|CALN|CALNA|CALNA1|CCN1|CNA1|IECEE|IECEE1|PPP2B | BIGM103|CDG2N|LZT-Hs6|PP3105|ZIP8 | |
| Cytomap | 4q24 | 4q24 | |
| Type of gene | protein-coding | protein-coding | |
| Description | serine/threonine-protein phosphatase 2B catalytic subunit alpha isoformCAM-PRP catalytic subunitcalcineurin A alphacalmodulin-dependent calcineurin A subunit alpha isoformprotein phosphatase 2B, catalytic subunit, alpha isoformprotein phosphatase 3 ( | zinc transporter ZIP8BCG induced integral membrane protein BIGM103BCG-induced integral membrane protein in monocyte clone 103 proteinLIV-1 subfamily of ZIP zinc transporter 6ZIP-8Zrt- and Irt-like protein 8solute carrier family 39 (metal ion transpo | |
| Modification date | 20200315 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000323055, ENST00000394853, ENST00000394854, ENST00000507176, ENST00000523694, ENST00000510292, ENST00000512215, | ENST00000510255, ENST00000356736, ENST00000424970, ENST00000394833, | |
| Fusion gene scores | * DoF score | 25 X 13 X 8=2600 | 6 X 6 X 3=108 |
| # samples | 23 | 6 | |
| ** MAII score | log2(23/2600*10)=-3.49880585697144 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/108*10)=-0.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: PPP3CA [Title/Abstract] AND SLC39A8 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | PPP3CA(102014932)-SLC39A8(103184350), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | PPP3CA-SLC39A8 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | PPP3CA | GO:0006470 | protein dephosphorylation | 18815128|19154138 |
| Hgene | PPP3CA | GO:0033173 | calcineurin-NFAT signaling cascade | 19154138|22688515 |
| Hgene | PPP3CA | GO:0035690 | cellular response to drug | 11005320 |
| Hgene | PPP3CA | GO:0045944 | positive regulation of transcription by RNA polymerase II | 22688515 |
| Hgene | PPP3CA | GO:0051592 | response to calcium ion | 18815128 |
| Tgene | SLC39A8 | GO:0006882 | cellular zinc ion homeostasis | 19401385 |
| Tgene | SLC39A8 | GO:0071577 | zinc ion transmembrane transport | 19401385 |
| Tgene | SLC39A8 | GO:0097080 | plasma membrane selenite transport | 27166256 |
| Tgene | SLC39A8 | GO:0098711 | iron ion import across plasma membrane | 22898811 |
 Fusion gene breakpoints across PPP3CA (5'-gene) Fusion gene breakpoints across PPP3CA (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
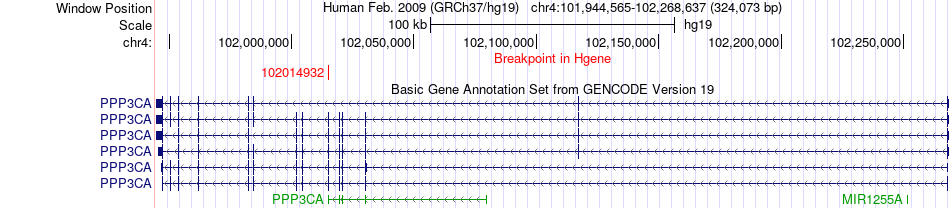 |
 Fusion gene breakpoints across SLC39A8 (3'-gene) Fusion gene breakpoints across SLC39A8 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
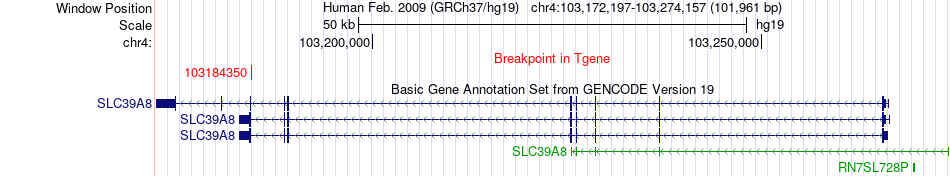 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | TCGA-56-8201-11A | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
Top |
Fusion Gene ORF analysis for PPP3CA-SLC39A8 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000323055 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| 5CDS-intron | ENST00000394853 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| 5CDS-intron | ENST00000394854 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| 5CDS-intron | ENST00000507176 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| 5CDS-intron | ENST00000523694 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000323055 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000323055 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000394853 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000394853 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000394854 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000394854 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000507176 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000507176 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000523694 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| Frame-shift | ENST00000523694 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| In-frame | ENST00000323055 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| In-frame | ENST00000394853 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| In-frame | ENST00000394854 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| In-frame | ENST00000507176 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| In-frame | ENST00000523694 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-3CDS | ENST00000510292 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-3CDS | ENST00000510292 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-3CDS | ENST00000510292 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-3CDS | ENST00000512215 | ENST00000356736 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-3CDS | ENST00000512215 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-3CDS | ENST00000512215 | ENST00000424970 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-intron | ENST00000510292 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
| intron-intron | ENST00000512215 | ENST00000510255 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000394854 | PPP3CA | chr4 | 102014932 | - | ENST00000394833 | SLC39A8 | chr4 | 103184350 | - | 2994 | 1466 | 603 | 1478 | 291 |
| ENST00000323055 | PPP3CA | chr4 | 102014932 | - | ENST00000394833 | SLC39A8 | chr4 | 103184350 | - | 2985 | 1457 | 594 | 1469 | 291 |
| ENST00000394853 | PPP3CA | chr4 | 102014932 | - | ENST00000394833 | SLC39A8 | chr4 | 103184350 | - | 2841 | 1313 | 450 | 1325 | 291 |
| ENST00000507176 | PPP3CA | chr4 | 102014932 | - | ENST00000394833 | SLC39A8 | chr4 | 103184350 | - | 2155 | 627 | 40 | 639 | 199 |
| ENST00000523694 | PPP3CA | chr4 | 102014932 | - | ENST00000394833 | SLC39A8 | chr4 | 103184350 | - | 2109 | 581 | 0 | 593 | 197 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000394854 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - | 0.000693279 | 0.9993067 |
| ENST00000323055 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - | 0.000684066 | 0.999316 |
| ENST00000394853 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - | 0.000730521 | 0.9992694 |
| ENST00000507176 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - | 0.001548872 | 0.9984511 |
| ENST00000523694 | ENST00000394833 | PPP3CA | chr4 | 102014932 | - | SLC39A8 | chr4 | 103184350 | - | 0.000929728 | 0.99907035 |
Top |
Fusion Genomic Features for PPP3CA-SLC39A8 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
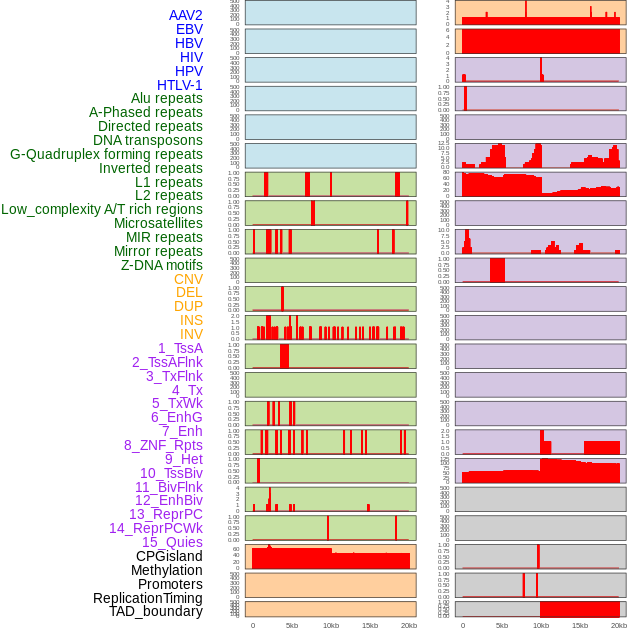 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for PPP3CA-SLC39A8 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr4:102014932/chr4:103184350) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 410_429 | 411 | 461.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 451_460 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 410_429 | 411 | 461.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 451_460 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 410_429 | 411 | 445.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 451_460 | 411 | 445.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 430_450 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 430_450 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 430_450 | 411 | 445.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 307_311 | 260 | 470.0 | Motif | SAPNY motif |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 307_311 | 260 | 512.0 | Motif | SAPNY motif |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 307_311 | 260 | 522.0 | Motif | SAPNY motif |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 307_311 | 0 | 290.0 | Motif | SAPNY motif |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 341_369 | 260 | 470.0 | Region | Calcineurin B binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 392_406 | 260 | 470.0 | Region | Calmodulin-binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 407_414 | 260 | 470.0 | Region | Autoinhibitory segment |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 465_487 | 260 | 470.0 | Region | Autoinhibitory domain |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 56_340 | 260 | 470.0 | Region | Catalytic |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 341_369 | 260 | 512.0 | Region | Calcineurin B binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 392_406 | 260 | 512.0 | Region | Calmodulin-binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 407_414 | 260 | 512.0 | Region | Autoinhibitory segment |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 465_487 | 260 | 512.0 | Region | Autoinhibitory domain |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 56_340 | 260 | 512.0 | Region | Catalytic |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 341_369 | 260 | 522.0 | Region | Calcineurin B binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 392_406 | 260 | 522.0 | Region | Calmodulin-binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 407_414 | 260 | 522.0 | Region | Autoinhibitory segment |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 465_487 | 260 | 522.0 | Region | Autoinhibitory domain |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 56_340 | 260 | 522.0 | Region | Catalytic |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 341_369 | 0 | 290.0 | Region | Calcineurin B binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 392_406 | 0 | 290.0 | Region | Calmodulin-binding |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 407_414 | 0 | 290.0 | Region | Autoinhibitory segment |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 465_487 | 0 | 290.0 | Region | Autoinhibitory domain |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 56_340 | 0 | 290.0 | Region | Catalytic |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 343_348 | 411 | 461.0 | Motif | XEXPHE-motif | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 343_348 | 411 | 461.0 | Motif | XEXPHE-motif | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 343_348 | 411 | 445.0 | Motif | XEXPHE-motif | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 154_160 | 411 | 461.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 182_191 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 213_365 | 411 | 461.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 23_132 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 387_388 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 154_160 | 411 | 461.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 182_191 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 213_365 | 411 | 461.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 23_132 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 387_388 | 411 | 461.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 154_160 | 411 | 445.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 182_191 | 411 | 445.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 213_365 | 411 | 445.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 23_132 | 411 | 445.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 387_388 | 411 | 445.0 | Topological domain | Extracellular | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 133_153 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 161_181 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 192_212 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 366_386 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000356736 | 7 | 9 | 389_409 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 133_153 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 161_181 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 192_212 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 366_386 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000394833 | 6 | 8 | 389_409 | 411 | 461.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 133_153 | 411 | 445.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 161_181 | 411 | 445.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 192_212 | 411 | 445.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 366_386 | 411 | 445.0 | Transmembrane | Helical | |
| Tgene | SLC39A8 | chr4:102014932 | chr4:103184350 | ENST00000424970 | 7 | 11 | 389_409 | 411 | 445.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for PPP3CA-SLC39A8 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >68132_68132_1_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000323055_SLC39A8_chr4_103184350_ENST00000394833_length(transcript)=2985nt_BP=1457nt AGGAGCGGCGCGGGGAGGCAGAGGCGAGCGCGGGCGATCGCGGCGGCGGGACGTCTCCGGCGGCCGCGGCACCAGCGGCGGCGCTCTGTG TGGAGAAGCAGGGGCAGCGGCAGCAGCAACAGCGGCAGCGGCAGCAGCAGCGGCAGCAGCGGCGGCCGCCTCTCCGCCGCCGGGGACGCC AAGGTGCGGCTGGTGGCACTGACATCGGCGGCCCTGCCTCTCCTCCCTCGCCCCCGGCCTTCCCCATGTGATCGGTCTTAATCCTGGCCG TGTGTGCGCGCGTGGGGCTCCATTCCGCGGTGCTGGGTCTCCGTGCCGGGGTGGGTGCTCGTGTGTGCGCTTCTCCTCCCCATCCCCCTT CCCCCAAGAATAAAAGAAGAACCGGGAGGCGTGCTCAGAAAATAAATAAATAACCACCACACACGCGCAGCCCGGAGCGAGTCGGCGGGG CTGGCGGCAGCGGCGGAGGAGGAGTGAAGGCGGCGGCGGCGGAGGAGGGACGCGCGGAAAAGGCAGCAACTTTAAAGCCAGCTCAGAGCC TAGACCTCCAGCCGAGCGGTTTGCAGCGCGGCGGCGGCGGCGGCGGCGGCGGCGTTGAGTGTCTGGCCCGCCGGTCCGGTCGGGGTGTGC AGTCGGACGGACGAGCAGCGCGTCGCTGTCCTCCGGCAGCTGGAGATGTCCGAGCCCAAGGCAATTGATCCCAAGTTGTCGACGACCGAC AGGGTGGTGAAAGCTGTTCCATTTCCTCCAAGTCACCGGCTTACAGCAAAAGAAGTGTTTGATAATGATGGAAAACCTCGTGTGGATATC TTAAAGGCGCATCTTATGAAGGAGGGAAGGCTGGAAGAGAGTGTTGCATTGAGAATAATAACAGAGGGTGCATCAATTCTTCGACAGGAA AAAAATTTGCTGGATATTGATGCGCCAGTCACTGTTTGTGGGGACATTCATGGACAATTCTTTGATTTGATGAAGCTCTTTGAAGTCGGG GGATCTCCTGCCAACACTCGCTACCTCTTCTTAGGGGACTATGTTGACAGAGGGTACTTCAGTATTGAATGTGTGCTGTATTTGTGGGCC TTGAAAATTCTCTACCCCAAAACACTGTTTTTACTTCGTGGAAATCATGAATGTAGACATCTAACAGAGTATTTCACATTTAAACAAGAA TGTAAAATAAAGTATTCAGAACGCGTATATGATGCCTGTATGGATGCCTTTGACTGCCTTCCCCTGGCTGCCCTGATGAACCAACAGTTC CTGTGTGTGCATGGTGGTTTGTCTCCAGAGATTAACACTTTAGATGATATCAGAAAATTAGACCGATTCAAAGAACCACCTGCATATGGA CCTATGTGTGATATCCTGTGGTCAGACCCCCTGGAAGATTTTGGAAATGAGAAGACTCAGGAACATTTCACTCACAACACAGTCAGGGGG TGTTCATACTTCTACAGTTTCCAGAGATGAATGATATGCTGAGAGAAAAGGTAACTGGAAGAAAAACCGATTTCACCTTCTTCATGATTC AGAATGCTGGAATGTTAACTGGATTCACAGCCATTCTACTCATTACCTTGTATGCAGGAGAAATCGAATTGGAGTAATAGAAAATGGAAG ATGGTGTTGTTAATAAAGGCATTTAATAGATAAAAACATCTCCAAAAAGGATTTTGAAGCTGATCCTATTTAGTTAAAAAGATAATTTTG CTTTCAACTGTAGGTCCAGAAAACTAATTATTGGCATCAGTCTGTGAAATAGTCCATTATTTGTTGTTAAAAATGCTTCAAAAGGTTTTC AGTGTCAGTCTGAGATGCCTGGTATATAGGAGCCTTTGGGAAATACCTATTTTTCAGTATTCCATGCATATTAGATATCACCATGAAGCA AGAGACATGCATTCTATAATCATGTAGACACTCAGACTCAGGGGAAAATACAAGTTATATCCTGAAAGCCTTTAAAACTCTATGGTAGGA TCAAAGATTCAAATGGTTTCAGAGAGGTTTTATTTCAATTAATTTGTTCTAGTGCTTTCAAGAGCAAGTACATCAAAATGTAGAAGGTAA AATGTATGCAACACTAATATAAATTATTCCAAGTCTTTAAGGAGCCAAAGAAAAAAAAGATTTCTCACAGCTTTTTGTTCTGTTTTGTAT TTCAATTAGGAACTTGCAGTATTATTTTGAAAACCATTCTAAAATAATAGGAGTTAGGAAATAAATAAAGTTTTGCTAGCCCTGCTAAGT TCAGGCTTAGAGGCTTATCGCTAAGTATAAACTTCACCAGATTCCACGAAAAGCTGGATAGCTTTTTTTCTGACTTATGTTGTGGTTGCA CCCCTCACAAATGGCAGAACAGTATGTAAAGCTGGTAACACCTCGGTTTCAGTGCACCATGTGTTTGCTTTGTGAAGGTGAAGAATATGT TGGTTTAGAGAAAGAAATTGGATGTAATTTTATGCAATTTACTTTTAAAGACAAACATAACTATTTAGCAGAGAATATTTTAATAAATGC AAAACAACAGCTGGACTGCTGTACATCAAGGACAGATTAACTGGAAAACATATGTTCCTTATGTGTGATCGAGAGCCATTCAGAAAAGAC TTCCTTTGTGTTCAGCCTATACTTTTCCATATGGTATACCTTGAAAAAAATTAGCACACCATGGTTATTTTTCTACCTTTTATAAAAGAC AGAGCCTGTTTACTCATTTAGAAGATAGAGAAAATTGGTCTAAAATTGAACATCCTAGATTCACACTCCCAAGTCACTTAAGGTGATTTG ATGGTGAGGAAAATGATTGACAAAGCCCAACAATGATCTCAGGAATTACATTTTCCAACAGACCAAAAAATGTTTTCATGTAGCAGCAAT GCAGATTTGGTGAATATTTAATATATATTTTAGTATGTATTTCACTTTATGACTGACAATTAAAAAATATTGTTTGGCCAAATAGTAAAC >68132_68132_1_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000323055_SLC39A8_chr4_103184350_ENST00000394833_length(amino acids)=291AA_BP= MSVWPAGPVGVCSRTDEQRVAVLRQLEMSEPKAIDPKLSTTDRVVKAVPFPPSHRLTAKEVFDNDGKPRVDILKAHLMKEGRLEESVALR IITEGASILRQEKNLLDIDAPVTVCGDIHGQFFDLMKLFEVGGSPANTRYLFLGDYVDRGYFSIECVLYLWALKILYPKTLFLLRGNHEC RHLTEYFTFKQECKIKYSERVYDACMDAFDCLPLAALMNQQFLCVHGGLSPEINTLDDIRKLDRFKEPPAYGPMCDILWSDPLEDFGNEK -------------------------------------------------------------- >68132_68132_2_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000394853_SLC39A8_chr4_103184350_ENST00000394833_length(transcript)=2841nt_BP=1313nt AGCAGCGGCGGCCGCCTCTCCGCCGCCGGGGACGCCAAGGTGCGGCTGGTGGCACTGACATCGGCGGCCCTGCCTCTCCTCCCTCGCCCC CGGCCTTCCCCATGTGATCGGTCTTAATCCTGGCCGTGTGTGCGCGCGTGGGGCTCCATTCCGCGGTGCTGGGTCTCCGTGCCGGGGTGG GTGCTCGTGTGTGCGCTTCTCCTCCCCATCCCCCTTCCCCCAAGAATAAAAGAAGAACCGGGAGGCGTGCTCAGAAAATAAATAAATAAC CACCACACACGCGCAGCCCGGAGCGAGTCGGCGGGGCTGGCGGCAGCGGCGGAGGAGGAGTGAAGGCGGCGGCGGCGGAGGAGGGACGCG CGGAAAAGGCAGCAACTTTAAAGCCAGCTCAGAGCCTAGACCTCCAGCCGAGCGGTTTGCAGCGCGGCGGCGGCGGCGGCGGCGGCGGCG TTGAGTGTCTGGCCCGCCGGTCCGGTCGGGGTGTGCAGTCGGACGGACGAGCAGCGCGTCGCTGTCCTCCGGCAGCTGGAGATGTCCGAG CCCAAGGCAATTGATCCCAAGTTGTCGACGACCGACAGGGTGGTGAAAGCTGTTCCATTTCCTCCAAGTCACCGGCTTACAGCAAAAGAA GTGTTTGATAATGATGGAAAACCTCGTGTGGATATCTTAAAGGCGCATCTTATGAAGGAGGGAAGGCTGGAAGAGAGTGTTGCATTGAGA ATAATAACAGAGGGTGCATCAATTCTTCGACAGGAAAAAAATTTGCTGGATATTGATGCGCCAGTCACTGTTTGTGGGGACATTCATGGA CAATTCTTTGATTTGATGAAGCTCTTTGAAGTCGGGGGATCTCCTGCCAACACTCGCTACCTCTTCTTAGGGGACTATGTTGACAGAGGG TACTTCAGTATTGAATGTGTGCTGTATTTGTGGGCCTTGAAAATTCTCTACCCCAAAACACTGTTTTTACTTCGTGGAAATCATGAATGT AGACATCTAACAGAGTATTTCACATTTAAACAAGAATGTAAAATAAAGTATTCAGAACGCGTATATGATGCCTGTATGGATGCCTTTGAC TGCCTTCCCCTGGCTGCCCTGATGAACCAACAGTTCCTGTGTGTGCATGGTGGTTTGTCTCCAGAGATTAACACTTTAGATGATATCAGA AAATTAGACCGATTCAAAGAACCACCTGCATATGGACCTATGTGTGATATCCTGTGGTCAGACCCCCTGGAAGATTTTGGAAATGAGAAG ACTCAGGAACATTTCACTCACAACACAGTCAGGGGGTGTTCATACTTCTACAGTTTCCAGAGATGAATGATATGCTGAGAGAAAAGGTAA CTGGAAGAAAAACCGATTTCACCTTCTTCATGATTCAGAATGCTGGAATGTTAACTGGATTCACAGCCATTCTACTCATTACCTTGTATG CAGGAGAAATCGAATTGGAGTAATAGAAAATGGAAGATGGTGTTGTTAATAAAGGCATTTAATAGATAAAAACATCTCCAAAAAGGATTT TGAAGCTGATCCTATTTAGTTAAAAAGATAATTTTGCTTTCAACTGTAGGTCCAGAAAACTAATTATTGGCATCAGTCTGTGAAATAGTC CATTATTTGTTGTTAAAAATGCTTCAAAAGGTTTTCAGTGTCAGTCTGAGATGCCTGGTATATAGGAGCCTTTGGGAAATACCTATTTTT CAGTATTCCATGCATATTAGATATCACCATGAAGCAAGAGACATGCATTCTATAATCATGTAGACACTCAGACTCAGGGGAAAATACAAG TTATATCCTGAAAGCCTTTAAAACTCTATGGTAGGATCAAAGATTCAAATGGTTTCAGAGAGGTTTTATTTCAATTAATTTGTTCTAGTG CTTTCAAGAGCAAGTACATCAAAATGTAGAAGGTAAAATGTATGCAACACTAATATAAATTATTCCAAGTCTTTAAGGAGCCAAAGAAAA AAAAGATTTCTCACAGCTTTTTGTTCTGTTTTGTATTTCAATTAGGAACTTGCAGTATTATTTTGAAAACCATTCTAAAATAATAGGAGT TAGGAAATAAATAAAGTTTTGCTAGCCCTGCTAAGTTCAGGCTTAGAGGCTTATCGCTAAGTATAAACTTCACCAGATTCCACGAAAAGC TGGATAGCTTTTTTTCTGACTTATGTTGTGGTTGCACCCCTCACAAATGGCAGAACAGTATGTAAAGCTGGTAACACCTCGGTTTCAGTG CACCATGTGTTTGCTTTGTGAAGGTGAAGAATATGTTGGTTTAGAGAAAGAAATTGGATGTAATTTTATGCAATTTACTTTTAAAGACAA ACATAACTATTTAGCAGAGAATATTTTAATAAATGCAAAACAACAGCTGGACTGCTGTACATCAAGGACAGATTAACTGGAAAACATATG TTCCTTATGTGTGATCGAGAGCCATTCAGAAAAGACTTCCTTTGTGTTCAGCCTATACTTTTCCATATGGTATACCTTGAAAAAAATTAG CACACCATGGTTATTTTTCTACCTTTTATAAAAGACAGAGCCTGTTTACTCATTTAGAAGATAGAGAAAATTGGTCTAAAATTGAACATC CTAGATTCACACTCCCAAGTCACTTAAGGTGATTTGATGGTGAGGAAAATGATTGACAAAGCCCAACAATGATCTCAGGAATTACATTTT CCAACAGACCAAAAAATGTTTTCATGTAGCAGCAATGCAGATTTGGTGAATATTTAATATATATTTTAGTATGTATTTCACTTTATGACT >68132_68132_2_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000394853_SLC39A8_chr4_103184350_ENST00000394833_length(amino acids)=291AA_BP= MSVWPAGPVGVCSRTDEQRVAVLRQLEMSEPKAIDPKLSTTDRVVKAVPFPPSHRLTAKEVFDNDGKPRVDILKAHLMKEGRLEESVALR IITEGASILRQEKNLLDIDAPVTVCGDIHGQFFDLMKLFEVGGSPANTRYLFLGDYVDRGYFSIECVLYLWALKILYPKTLFLLRGNHEC RHLTEYFTFKQECKIKYSERVYDACMDAFDCLPLAALMNQQFLCVHGGLSPEINTLDDIRKLDRFKEPPAYGPMCDILWSDPLEDFGNEK -------------------------------------------------------------- >68132_68132_3_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000394854_SLC39A8_chr4_103184350_ENST00000394833_length(transcript)=2994nt_BP=1466nt GGTGAGTGGAGGAGCGGCGCGGGGAGGCAGAGGCGAGCGCGGGCGATCGCGGCGGCGGGACGTCTCCGGCGGCCGCGGCACCAGCGGCGG CGCTCTGTGTGGAGAAGCAGGGGCAGCGGCAGCAGCAACAGCGGCAGCGGCAGCAGCAGCGGCAGCAGCGGCGGCCGCCTCTCCGCCGCC GGGGACGCCAAGGTGCGGCTGGTGGCACTGACATCGGCGGCCCTGCCTCTCCTCCCTCGCCCCCGGCCTTCCCCATGTGATCGGTCTTAA TCCTGGCCGTGTGTGCGCGCGTGGGGCTCCATTCCGCGGTGCTGGGTCTCCGTGCCGGGGTGGGTGCTCGTGTGTGCGCTTCTCCTCCCC ATCCCCCTTCCCCCAAGAATAAAAGAAGAACCGGGAGGCGTGCTCAGAAAATAAATAAATAACCACCACACACGCGCAGCCCGGAGCGAG TCGGCGGGGCTGGCGGCAGCGGCGGAGGAGGAGTGAAGGCGGCGGCGGCGGAGGAGGGACGCGCGGAAAAGGCAGCAACTTTAAAGCCAG CTCAGAGCCTAGACCTCCAGCCGAGCGGTTTGCAGCGCGGCGGCGGCGGCGGCGGCGGCGGCGTTGAGTGTCTGGCCCGCCGGTCCGGTC GGGGTGTGCAGTCGGACGGACGAGCAGCGCGTCGCTGTCCTCCGGCAGCTGGAGATGTCCGAGCCCAAGGCAATTGATCCCAAGTTGTCG ACGACCGACAGGGTGGTGAAAGCTGTTCCATTTCCTCCAAGTCACCGGCTTACAGCAAAAGAAGTGTTTGATAATGATGGAAAACCTCGT GTGGATATCTTAAAGGCGCATCTTATGAAGGAGGGAAGGCTGGAAGAGAGTGTTGCATTGAGAATAATAACAGAGGGTGCATCAATTCTT CGACAGGAAAAAAATTTGCTGGATATTGATGCGCCAGTCACTGTTTGTGGGGACATTCATGGACAATTCTTTGATTTGATGAAGCTCTTT GAAGTCGGGGGATCTCCTGCCAACACTCGCTACCTCTTCTTAGGGGACTATGTTGACAGAGGGTACTTCAGTATTGAATGTGTGCTGTAT TTGTGGGCCTTGAAAATTCTCTACCCCAAAACACTGTTTTTACTTCGTGGAAATCATGAATGTAGACATCTAACAGAGTATTTCACATTT AAACAAGAATGTAAAATAAAGTATTCAGAACGCGTATATGATGCCTGTATGGATGCCTTTGACTGCCTTCCCCTGGCTGCCCTGATGAAC CAACAGTTCCTGTGTGTGCATGGTGGTTTGTCTCCAGAGATTAACACTTTAGATGATATCAGAAAATTAGACCGATTCAAAGAACCACCT GCATATGGACCTATGTGTGATATCCTGTGGTCAGACCCCCTGGAAGATTTTGGAAATGAGAAGACTCAGGAACATTTCACTCACAACACA GTCAGGGGGTGTTCATACTTCTACAGTTTCCAGAGATGAATGATATGCTGAGAGAAAAGGTAACTGGAAGAAAAACCGATTTCACCTTCT TCATGATTCAGAATGCTGGAATGTTAACTGGATTCACAGCCATTCTACTCATTACCTTGTATGCAGGAGAAATCGAATTGGAGTAATAGA AAATGGAAGATGGTGTTGTTAATAAAGGCATTTAATAGATAAAAACATCTCCAAAAAGGATTTTGAAGCTGATCCTATTTAGTTAAAAAG ATAATTTTGCTTTCAACTGTAGGTCCAGAAAACTAATTATTGGCATCAGTCTGTGAAATAGTCCATTATTTGTTGTTAAAAATGCTTCAA AAGGTTTTCAGTGTCAGTCTGAGATGCCTGGTATATAGGAGCCTTTGGGAAATACCTATTTTTCAGTATTCCATGCATATTAGATATCAC CATGAAGCAAGAGACATGCATTCTATAATCATGTAGACACTCAGACTCAGGGGAAAATACAAGTTATATCCTGAAAGCCTTTAAAACTCT ATGGTAGGATCAAAGATTCAAATGGTTTCAGAGAGGTTTTATTTCAATTAATTTGTTCTAGTGCTTTCAAGAGCAAGTACATCAAAATGT AGAAGGTAAAATGTATGCAACACTAATATAAATTATTCCAAGTCTTTAAGGAGCCAAAGAAAAAAAAGATTTCTCACAGCTTTTTGTTCT GTTTTGTATTTCAATTAGGAACTTGCAGTATTATTTTGAAAACCATTCTAAAATAATAGGAGTTAGGAAATAAATAAAGTTTTGCTAGCC CTGCTAAGTTCAGGCTTAGAGGCTTATCGCTAAGTATAAACTTCACCAGATTCCACGAAAAGCTGGATAGCTTTTTTTCTGACTTATGTT GTGGTTGCACCCCTCACAAATGGCAGAACAGTATGTAAAGCTGGTAACACCTCGGTTTCAGTGCACCATGTGTTTGCTTTGTGAAGGTGA AGAATATGTTGGTTTAGAGAAAGAAATTGGATGTAATTTTATGCAATTTACTTTTAAAGACAAACATAACTATTTAGCAGAGAATATTTT AATAAATGCAAAACAACAGCTGGACTGCTGTACATCAAGGACAGATTAACTGGAAAACATATGTTCCTTATGTGTGATCGAGAGCCATTC AGAAAAGACTTCCTTTGTGTTCAGCCTATACTTTTCCATATGGTATACCTTGAAAAAAATTAGCACACCATGGTTATTTTTCTACCTTTT ATAAAAGACAGAGCCTGTTTACTCATTTAGAAGATAGAGAAAATTGGTCTAAAATTGAACATCCTAGATTCACACTCCCAAGTCACTTAA GGTGATTTGATGGTGAGGAAAATGATTGACAAAGCCCAACAATGATCTCAGGAATTACATTTTCCAACAGACCAAAAAATGTTTTCATGT AGCAGCAATGCAGATTTGGTGAATATTTAATATATATTTTAGTATGTATTTCACTTTATGACTGACAATTAAAAAATATTGTTTGGCCAA >68132_68132_3_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000394854_SLC39A8_chr4_103184350_ENST00000394833_length(amino acids)=291AA_BP= MSVWPAGPVGVCSRTDEQRVAVLRQLEMSEPKAIDPKLSTTDRVVKAVPFPPSHRLTAKEVFDNDGKPRVDILKAHLMKEGRLEESVALR IITEGASILRQEKNLLDIDAPVTVCGDIHGQFFDLMKLFEVGGSPANTRYLFLGDYVDRGYFSIECVLYLWALKILYPKTLFLLRGNHEC RHLTEYFTFKQECKIKYSERVYDACMDAFDCLPLAALMNQQFLCVHGGLSPEINTLDDIRKLDRFKEPPAYGPMCDILWSDPLEDFGNEK -------------------------------------------------------------- >68132_68132_4_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000507176_SLC39A8_chr4_103184350_ENST00000394833_length(transcript)=2155nt_BP=627nt CAGTCGGACGGACGAGCAGCGCGTCGCTGTCCTCCGGCAGCTGGAGATGTCCGAGCCCAAGGCAATTGATCCCAAGTTGTCGACGACCGA CAGGGTGGTGAAAGTTTGTGGGGACATTCATGGACAATTCTTTGATTTGATGAAGCTCTTTGAAGTCGGGGGATCTCCTGCCAACACTCG CTACCTCTTCTTAGGGGACTATGTTGACAGAGGGTACTTCAGTATTGAATGTGTGCTGTATTTGTGGGCCTTGAAAATTCTCTACCCCAA AACACTGTTTTTACTTCGTGGAAATCATGAATGTAGACATCTAACAGAGTATTTCACATTTAAACAAGAATGTAAAATAAAGTATTCAGA ACGCGTATATGATGCCTGTATGGATGCCTTTGACTGCCTTCCCCTGGCTGCCCTGATGAACCAACAGTTCCTGTGTGTGCATGGTGGTTT GTCTCCAGAGATTAACACTTTAGATGATATCAGAAAATTAGACCGATTCAAAGAACCACCTGCATATGGACCTATGTGTGATATCCTGTG GTCAGACCCCCTGGAAGATTTTGGAAATGAGAAGACTCAGGAACATTTCACTCACAACACAGTCAGGGGGTGTTCATACTTCTACAGTTT CCAGAGATGAATGATATGCTGAGAGAAAAGGTAACTGGAAGAAAAACCGATTTCACCTTCTTCATGATTCAGAATGCTGGAATGTTAACT GGATTCACAGCCATTCTACTCATTACCTTGTATGCAGGAGAAATCGAATTGGAGTAATAGAAAATGGAAGATGGTGTTGTTAATAAAGGC ATTTAATAGATAAAAACATCTCCAAAAAGGATTTTGAAGCTGATCCTATTTAGTTAAAAAGATAATTTTGCTTTCAACTGTAGGTCCAGA AAACTAATTATTGGCATCAGTCTGTGAAATAGTCCATTATTTGTTGTTAAAAATGCTTCAAAAGGTTTTCAGTGTCAGTCTGAGATGCCT GGTATATAGGAGCCTTTGGGAAATACCTATTTTTCAGTATTCCATGCATATTAGATATCACCATGAAGCAAGAGACATGCATTCTATAAT CATGTAGACACTCAGACTCAGGGGAAAATACAAGTTATATCCTGAAAGCCTTTAAAACTCTATGGTAGGATCAAAGATTCAAATGGTTTC AGAGAGGTTTTATTTCAATTAATTTGTTCTAGTGCTTTCAAGAGCAAGTACATCAAAATGTAGAAGGTAAAATGTATGCAACACTAATAT AAATTATTCCAAGTCTTTAAGGAGCCAAAGAAAAAAAAGATTTCTCACAGCTTTTTGTTCTGTTTTGTATTTCAATTAGGAACTTGCAGT ATTATTTTGAAAACCATTCTAAAATAATAGGAGTTAGGAAATAAATAAAGTTTTGCTAGCCCTGCTAAGTTCAGGCTTAGAGGCTTATCG CTAAGTATAAACTTCACCAGATTCCACGAAAAGCTGGATAGCTTTTTTTCTGACTTATGTTGTGGTTGCACCCCTCACAAATGGCAGAAC AGTATGTAAAGCTGGTAACACCTCGGTTTCAGTGCACCATGTGTTTGCTTTGTGAAGGTGAAGAATATGTTGGTTTAGAGAAAGAAATTG GATGTAATTTTATGCAATTTACTTTTAAAGACAAACATAACTATTTAGCAGAGAATATTTTAATAAATGCAAAACAACAGCTGGACTGCT GTACATCAAGGACAGATTAACTGGAAAACATATGTTCCTTATGTGTGATCGAGAGCCATTCAGAAAAGACTTCCTTTGTGTTCAGCCTAT ACTTTTCCATATGGTATACCTTGAAAAAAATTAGCACACCATGGTTATTTTTCTACCTTTTATAAAAGACAGAGCCTGTTTACTCATTTA GAAGATAGAGAAAATTGGTCTAAAATTGAACATCCTAGATTCACACTCCCAAGTCACTTAAGGTGATTTGATGGTGAGGAAAATGATTGA CAAAGCCCAACAATGATCTCAGGAATTACATTTTCCAACAGACCAAAAAATGTTTTCATGTAGCAGCAATGCAGATTTGGTGAATATTTA >68132_68132_4_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000507176_SLC39A8_chr4_103184350_ENST00000394833_length(amino acids)=199AA_BP= MEMSEPKAIDPKLSTTDRVVKVCGDIHGQFFDLMKLFEVGGSPANTRYLFLGDYVDRGYFSIECVLYLWALKILYPKTLFLLRGNHECRH LTEYFTFKQECKIKYSERVYDACMDAFDCLPLAALMNQQFLCVHGGLSPEINTLDDIRKLDRFKEPPAYGPMCDILWSDPLEDFGNEKTQ -------------------------------------------------------------- >68132_68132_5_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000523694_SLC39A8_chr4_103184350_ENST00000394833_length(transcript)=2109nt_BP=581nt ATGTCCGAGCCCAAGGCAATTGATCCCAAGTTGTCGACGACCGACAGGGTGGTGAAAGTTTGTGGGGACATTCATGGACAATTCTTTGAT TTGATGAAGCTCTTTGAAGTCGGGGGATCTCCTGCCAACACTCGCTACCTCTTCTTAGGGGACTATGTTGACAGAGGGTACTTCAGTATT GAATGTGTGCTGTATTTGTGGGCCTTGAAAATTCTCTACCCCAAAACACTGTTTTTACTTCGTGGAAATCATGAATGTAGACATCTAACA GAGTATTTCACATTTAAACAAGAATGTAAAATAAAGTATTCAGAACGCGTATATGATGCCTGTATGGATGCCTTTGACTGCCTTCCCCTG GCTGCCCTGATGAACCAACAGTTCCTGTGTGTGCATGGTGGTTTGTCTCCAGAGATTAACACTTTAGATGATATCAGAAAATTAGACCGA TTCAAAGAACCACCTGCATATGGACCTATGTGTGATATCCTGTGGTCAGACCCCCTGGAAGATTTTGGAAATGAGAAGACTCAGGAACAT TTCACTCACAACACAGTCAGGGGGTGTTCATACTTCTACAGTTTCCAGAGATGAATGATATGCTGAGAGAAAAGGTAACTGGAAGAAAAA CCGATTTCACCTTCTTCATGATTCAGAATGCTGGAATGTTAACTGGATTCACAGCCATTCTACTCATTACCTTGTATGCAGGAGAAATCG AATTGGAGTAATAGAAAATGGAAGATGGTGTTGTTAATAAAGGCATTTAATAGATAAAAACATCTCCAAAAAGGATTTTGAAGCTGATCC TATTTAGTTAAAAAGATAATTTTGCTTTCAACTGTAGGTCCAGAAAACTAATTATTGGCATCAGTCTGTGAAATAGTCCATTATTTGTTG TTAAAAATGCTTCAAAAGGTTTTCAGTGTCAGTCTGAGATGCCTGGTATATAGGAGCCTTTGGGAAATACCTATTTTTCAGTATTCCATG CATATTAGATATCACCATGAAGCAAGAGACATGCATTCTATAATCATGTAGACACTCAGACTCAGGGGAAAATACAAGTTATATCCTGAA AGCCTTTAAAACTCTATGGTAGGATCAAAGATTCAAATGGTTTCAGAGAGGTTTTATTTCAATTAATTTGTTCTAGTGCTTTCAAGAGCA AGTACATCAAAATGTAGAAGGTAAAATGTATGCAACACTAATATAAATTATTCCAAGTCTTTAAGGAGCCAAAGAAAAAAAAGATTTCTC ACAGCTTTTTGTTCTGTTTTGTATTTCAATTAGGAACTTGCAGTATTATTTTGAAAACCATTCTAAAATAATAGGAGTTAGGAAATAAAT AAAGTTTTGCTAGCCCTGCTAAGTTCAGGCTTAGAGGCTTATCGCTAAGTATAAACTTCACCAGATTCCACGAAAAGCTGGATAGCTTTT TTTCTGACTTATGTTGTGGTTGCACCCCTCACAAATGGCAGAACAGTATGTAAAGCTGGTAACACCTCGGTTTCAGTGCACCATGTGTTT GCTTTGTGAAGGTGAAGAATATGTTGGTTTAGAGAAAGAAATTGGATGTAATTTTATGCAATTTACTTTTAAAGACAAACATAACTATTT AGCAGAGAATATTTTAATAAATGCAAAACAACAGCTGGACTGCTGTACATCAAGGACAGATTAACTGGAAAACATATGTTCCTTATGTGT GATCGAGAGCCATTCAGAAAAGACTTCCTTTGTGTTCAGCCTATACTTTTCCATATGGTATACCTTGAAAAAAATTAGCACACCATGGTT ATTTTTCTACCTTTTATAAAAGACAGAGCCTGTTTACTCATTTAGAAGATAGAGAAAATTGGTCTAAAATTGAACATCCTAGATTCACAC TCCCAAGTCACTTAAGGTGATTTGATGGTGAGGAAAATGATTGACAAAGCCCAACAATGATCTCAGGAATTACATTTTCCAACAGACCAA AAAATGTTTTCATGTAGCAGCAATGCAGATTTGGTGAATATTTAATATATATTTTAGTATGTATTTCACTTTATGACTGACAATTAAAAA >68132_68132_5_PPP3CA-SLC39A8_PPP3CA_chr4_102014932_ENST00000523694_SLC39A8_chr4_103184350_ENST00000394833_length(amino acids)=197AA_BP= MSEPKAIDPKLSTTDRVVKVCGDIHGQFFDLMKLFEVGGSPANTRYLFLGDYVDRGYFSIECVLYLWALKILYPKTLFLLRGNHECRHLT EYFTFKQECKIKYSERVYDACMDAFDCLPLAALMNQQFLCVHGGLSPEINTLDDIRKLDRFKEPPAYGPMCDILWSDPLEDFGNEKTQEH -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for PPP3CA-SLC39A8 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000323055 | - | 6 | 12 | 327_336 | 260.6666666666667 | 470.0 | PxIxIF motif in substrate |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394853 | - | 6 | 13 | 327_336 | 260.6666666666667 | 512.0 | PxIxIF motif in substrate |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000394854 | - | 6 | 14 | 327_336 | 260.6666666666667 | 522.0 | PxIxIF motif in substrate |
| Hgene | PPP3CA | chr4:102014932 | chr4:103184350 | ENST00000512215 | - | 1 | 8 | 327_336 | 0 | 290.0 | PxIxIF motif in substrate |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for PPP3CA-SLC39A8 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for PPP3CA-SLC39A8 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
