
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:RAPGEF1-SEC16A (FusionGDB2 ID:72217) |
Fusion Gene Summary for RAPGEF1-SEC16A |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: RAPGEF1-SEC16A | Fusion gene ID: 72217 | Hgene | Tgene | Gene symbol | RAPGEF1 | SEC16A | Gene ID | 2889 | 9919 |
| Gene name | Rap guanine nucleotide exchange factor 1 | SEC16 homolog A, endoplasmic reticulum export factor | |
| Synonyms | C3G|GRF2 | KIAA0310|SEC16L|p250 | |
| Cytomap | 9q34.13 | 9q34.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | rap guanine nucleotide exchange factor 1CRK SH3-binding GNRPRap guanine nucleotide exchange factor (GEF) 1guanine nucleotide-releasing factor 2 (specific for crk proto-oncogene) | protein transport protein Sec16Aprotein SEC16 homolog A | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000372195, ENST00000372189, ENST00000372190, ENST00000481260, | ENST00000290037, ENST00000313084, ENST00000371706, ENST00000398335, ENST00000431893, ENST00000467838, ENST00000313050, | |
| Fusion gene scores | * DoF score | 20 X 16 X 11=3520 | 10 X 8 X 7=560 |
| # samples | 23 | 11 | |
| ** MAII score | log2(23/3520*10)=-3.93586966258028 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/560*10)=-2.34792330342031 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: RAPGEF1 [Title/Abstract] AND SEC16A [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | RAPGEF1(134615157)-SEC16A(139372136), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | RAPGEF1 | GO:0038180 | nerve growth factor signaling pathway | 17724123 |
| Hgene | RAPGEF1 | GO:0043547 | positive regulation of GTPase activity | 10548487 |
| Hgene | RAPGEF1 | GO:0071320 | cellular response to cAMP | 21840392 |
| Hgene | RAPGEF1 | GO:1990090 | cellular response to nerve growth factor stimulus | 17724123 |
| Tgene | SEC16A | GO:0007029 | endoplasmic reticulum organization | 21768384 |
| Tgene | SEC16A | GO:0034976 | response to endoplasmic reticulum stress | 28067262 |
 Fusion gene breakpoints across RAPGEF1 (5'-gene) Fusion gene breakpoints across RAPGEF1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
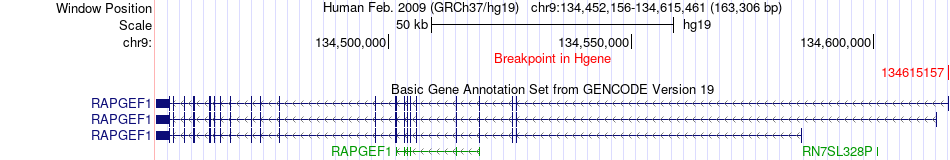 |
 Fusion gene breakpoints across SEC16A (3'-gene) Fusion gene breakpoints across SEC16A (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
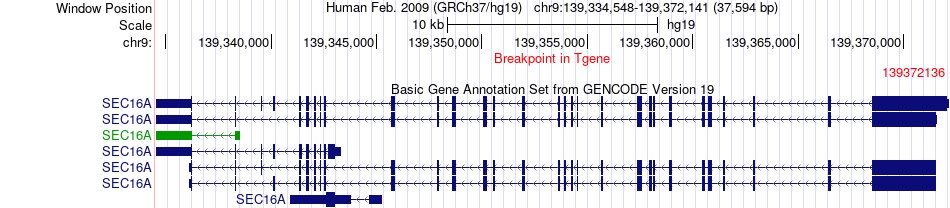 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LGG | TCGA-HT-7483-01A | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
Top |
Fusion Gene ORF analysis for RAPGEF1-SEC16A |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000372195 | ENST00000290037 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| 5CDS-intron | ENST00000372195 | ENST00000313084 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| 5CDS-intron | ENST00000372195 | ENST00000371706 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| 5CDS-intron | ENST00000372195 | ENST00000398335 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| 5CDS-intron | ENST00000372195 | ENST00000431893 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| 5CDS-intron | ENST00000372195 | ENST00000467838 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| In-frame | ENST00000372195 | ENST00000313050 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-3CDS | ENST00000372189 | ENST00000313050 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-3CDS | ENST00000372190 | ENST00000313050 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-3CDS | ENST00000481260 | ENST00000313050 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372189 | ENST00000290037 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372189 | ENST00000313084 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372189 | ENST00000371706 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372189 | ENST00000398335 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372189 | ENST00000431893 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372189 | ENST00000467838 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372190 | ENST00000290037 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372190 | ENST00000313084 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372190 | ENST00000371706 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372190 | ENST00000398335 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372190 | ENST00000431893 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000372190 | ENST00000467838 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000481260 | ENST00000290037 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000481260 | ENST00000313084 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000481260 | ENST00000371706 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000481260 | ENST00000398335 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000481260 | ENST00000431893 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
| intron-intron | ENST00000481260 | ENST00000467838 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000372195 | RAPGEF1 | chr9 | 134615157 | - | ENST00000313050 | SEC16A | chr9 | 139372136 | - | 9111 | 305 | 7 | 7452 | 2481 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000372195 | ENST00000313050 | RAPGEF1 | chr9 | 134615157 | - | SEC16A | chr9 | 139372136 | - | 0.00177575 | 0.9982242 |
Top |
Fusion Genomic Features for RAPGEF1-SEC16A |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
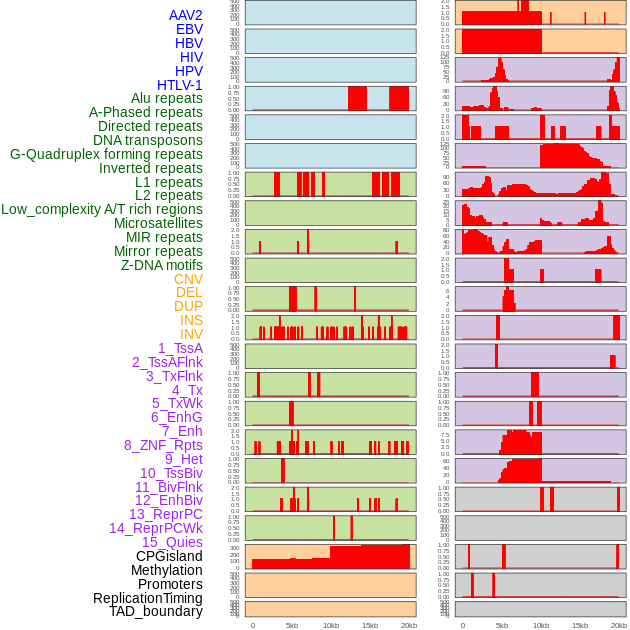 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for RAPGEF1-SEC16A |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr9:134615157/chr9:139372136) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1113_1181 | 0 | 2160.0 | Compositional bias | Note=Pro-rich | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1159_1162 | 0 | 2160.0 | Compositional bias | Note=Poly-Tyr | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1591_1594 | 0 | 2160.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1931_2285 | 0 | 2160.0 | Compositional bias | Note=Pro-rich | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1113_1181 | 0 | 2135.0 | Compositional bias | Note=Pro-rich | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1159_1162 | 0 | 2135.0 | Compositional bias | Note=Poly-Tyr | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1591_1594 | 0 | 2135.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1931_2285 | 0 | 2135.0 | Compositional bias | Note=Pro-rich | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1113_1181 | 0 | 2155.0 | Compositional bias | Note=Pro-rich | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1159_1162 | 0 | 2155.0 | Compositional bias | Note=Poly-Tyr | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1591_1594 | 0 | 2155.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1931_2285 | 0 | 2155.0 | Compositional bias | Note=Pro-rich | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1019_1890 | 0 | 2160.0 | Region | Required for localization to endoplasmic reticulum exit sites | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1102_1405 | 0 | 2160.0 | Region | Required for endoplasmic reticulum localization | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1019_1890 | 0 | 2135.0 | Region | Required for localization to endoplasmic reticulum exit sites | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1102_1405 | 0 | 2135.0 | Region | Required for endoplasmic reticulum localization | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1019_1890 | 0 | 2155.0 | Region | Required for localization to endoplasmic reticulum exit sites | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1102_1405 | 0 | 2155.0 | Region | Required for endoplasmic reticulum localization |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 963_966 | 0 | 1078.0 | Compositional bias | Note=Poly-Ser |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 963_966 | 0 | 1096.0 | Compositional bias | Note=Poly-Ser |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 688_810 | 0 | 1078.0 | Domain | N-terminal Ras-GEF |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 840_1064 | 0 | 1078.0 | Domain | Ras-GEF |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 688_810 | 0 | 1096.0 | Domain | N-terminal Ras-GEF |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 840_1064 | 0 | 1096.0 | Domain | Ras-GEF |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 281_292 | 0 | 1078.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 451_462 | 0 | 1078.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 538_549 | 0 | 1078.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372189 | - | 1 | 24 | 606_617 | 0 | 1078.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 281_292 | 0 | 1096.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 451_462 | 0 | 1096.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 538_549 | 0 | 1096.0 | Motif | Note=SH3-binding |
| Hgene | RAPGEF1 | chr9:134615157 | chr9:139372136 | ENST00000372190 | - | 1 | 24 | 606_617 | 0 | 1096.0 | Motif | Note=SH3-binding |
Top |
Fusion Gene Sequence for RAPGEF1-SEC16A |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >72217_72217_1_RAPGEF1-SEC16A_RAPGEF1_chr9_134615157_ENST00000372195_SEC16A_chr9_139372136_ENST00000313050_length(transcript)=9111nt_BP=305nt CGCTGCGCTGCGGCCGGCTCGCGGGCCGGGGGAGCGGCGGCGGGCGCGCGGGAGGCGGAGCGGCGGCGGCGGGCGCGGCGGGAGCGGCTG GGCCCGGGCGGGCTGGAGCGCGGTCGCCGAGCGCAGGGCCGCGGCCGCGCGGAGCCCGCCATGCCGGCGCGGGTCTGAGGCGTGGCGAGC CAGGGCCGAGGCGAGCGGCGGGCGCGCCGCCCCTGTCGCCCCGCGGCGGCCCGGCCCGGGCCCGATGAGCGGCGGCCTCGGCCTCCGGCG CAGCCCGGAAATGTCCGGCAAGATCGAGAAAGCAGTCCAGCTCCAATTAAGGAACAGCTATATCCTGCTTCCTTCTGTAAGCAGCATCGA CTTGTGCAAGGGTTCAGTCATGCAGCCACCGCCCCAGACGGTCCCGTCTGGCATGGCTGGGCCACCTCCAGCCGGGAATCCTCGGAGCGT GTTCTGGGCTAGCAGCCCTTACAGGAGACGGGCTAATAATAATGCAGCAGTGGCTCCGACAACTTGCCCGTTGCAGCCGGTCACGGATCC ATTTGCTTTTAGTAGACAGGCGCTCCAAAGTACACCACTGGGCAGTTCGTCCAAAAGCAGTCCACCTGTCTTGCAAGGCCCAGCCCCCGC AGGGTTTTCTCAGCACCCCGGTTTGCTTGTTCCTCACACACATGCCAGAGATAGCTCTCAGGGACCCTGTGAGCCCCTGCCTGGACCTCT GACACAGCCCAGAGCACATGCCAGTCCGTTTTCTGGTGCATTGACACCTTCAGCACCTCCTGGGCCTGAGATGAACAGGAGTGCAGAGGT CGGTCCCAGTTCAGAGCCTGAAGTTCAGACTCTGCCATATCTTCCTCACTACATTCCAGGAGTGGATCCTGAAACGTCTCATGGGGGCCA CCCTCATGGGAACATGCCTGGGCTCGACCGACCCCTGAGCAGGCAAAACCCACATGACGGTGTGGTCACCCCAGCAGCATCCCCTTCCCT CCCTCAGCCTGGTCTGCAGATGCCAGGACAGTGGGGGCCAGTGCAGGGAGGCCCACAGCCCTCGGGGCAACATCGTTCACCCTGCCCTGA AGGACCTGTTCCCAGCGGGGTGCCCTGTGCCACCAGCGTTCCTCATTTCCCCACCCCGTCCATCCTACATCAGGGCCCTGGTCATGAGCA ACACAGCCCTCTGGTGGCTCCCCCAGCAGCCTTGCCCAGTGACGGAAGAGACGAGGTGAGCCACTTGCAAAGTGGAAGCCACCTGGCCAA TAACTCTGATCCTGAAAGTACATTCAGGCAAAATCCCAGAATTGTGAATCACTGGGCAAGCCCAGAGCTCAGGCAGAATCCAGGAGTGAA GAATGAGCACCGGCCCGCCTCTGCTCTTGTGAACCCCCTCGCCCGGGGAGATAGCCCAGAAAACCGTACGCACCACCCACTGGGGGCTGG GGCCGGGTCTGGCTGTGCCCCGCTAGAAGCAGACTCAGGAGCTTCAGGAGCTCTGGCGATGTTTTTCCAAGGGGGAGAGACAGAAAATGA GGAGAATCTCTCATCTGAAAAAGCAGGCTTATCTGGTCAAGCGGACTTTGACGATTTCTGCTCCAGCCCTGGGCTAGGCCGTCCGCCCGC ACCTACACACGTGGGGGCAGGCAGCCTCTGCCAGGCCCTTCTCCCAGGCCCCAGCAATGAGGCTGCTGGTGATGTGTGGGGTGACACAGC GAGCACAGGGGTGCCGGATGCCAGCGGCTCGCAGTATGAGAATGTTGAGAACTTAGAATTTGTTCAGAATCAAGAAGTTCTGCCAAGTGA GCCCCTCAATTTGGACCCTTCCTCCCCGAGTGACCAGTTCAGATATGGGCCCCTTCCTGGGCCAGCTGTGCCCAGGCATGGTGCTGTGTG CCACACCGGAGCCCCTGATGCCACACTGCATACAGTGCACCCTGACAGCGTGTCATCCAGCTATAGCAGCAGAAGCCACGGAAGGCTCTC AGGCTCAGCCAGGCCCCAGGAGCTGGTTGGCACATTCATTCAGCAAGAAGTTGGAAAACCCGAGGATGAAGCTTCAGGTAGTTTTTTTAA GCAAATCGATTCTTCTCCCGTAGGAGGTGAAACAGACGAGACCACTGTGAGCCAGAATTACCGTGGCAGCGTGTCCCAGCCCTCAACCCC GAGCCCCCCGAAACCTACAGGAATATTTCAGACAAGTGCAAATAGTTCTTTCGAACCGGTAAAATCTCACTTAGTTGGGGTAAAACCATT TGAGGCAGATCGCGCCAACGTGGTTGGTGAAGTAAGGGAGACCTGTGTCCGCCAGAAGCAGTGCAGACCAGCTGCCGCCCTGCCCGATGC TTCCCCTGGCAACCTGGAGCAGCCACCAGACAACATGGAGACCCTCTGTGCACCCCAGGTCTGTCCCCTGCCTCTTAACTCCACCACGGA AGCTGTGCACATGCTTCCGCACGCAGGGGCACCGCCCTTGGATACTGTGTATCCAGCACCCGAGAAGAGGCCTTCAGCCAGGACCCAGGG GCCCGTGAAGTGTGAGAGCCCAGCAACGACTCTGTGGGCGCAAAGTGAGCTGCCAGATTTTGGAGGCAACGTCCTTCTGGCCCCTGCAGC CCCGGCGCTTTATGTGTGTGCAAAACCTCAGCCACCTGTTGTTCAGCCTCCAGAAGAGGCGATGTCCGGGCAGCAGTCACGGAACCCAAG CTCGGCGGCCCCGGTGCAGAGCCGAGGTGGCATTGGTGCTTCTGAGAACCTTGAGAATCCTCCCAAAATGGGAGAGGAGGAGGCCCTTCA GTCCCAGGCGAGTTCTGGTTATGCAAGTTTATTATCCTCACCGCCCACTGAGTCTCTGCAGAATCCTCCAGTCTTGATTGCTCAGCCTGA TCACAGCTATAATCTGGCTCAGCCCATTAACTTTTCTGTGTCCTTATCGAACTCTCATGAGAAGAATCAGTCCTGGAGAGAGGCTTTGGT GGGGGATAGACCTGCAGTCAGCAGTTGGGCTCTCGGTGGTGATTCTGGGGAGAACACTTCTTTGTCTGGGATTCCAACCAGCTCTGTCCT TAGCTTGTCTCTGCCTAGCAGTGTTGCCCAAAGTAATTTTCCACAAGGTTCTGGTGCTTCCGAAATGGTTTCTAATCAGCCTGCTAATTT GCTGGTTCAACCACCATCCCAGCCAGTTCCAGAGAACTTGGTTCCAGAAAGTCAAAAGGATCGTAAGGCAGGAAGTGCTCTTCCCGGATT TGCTAATAGCCCTGCTGGAAGCACAAGTGTGGTGTTAGTTCCACCTGCACACGGCACCCTGGTGCCTGATGGTAATAAGGCAAACCATTC CAGTCATCAGGAAGACACTTACGGAGCCCTAGACTTTACCTTAAGCAGGACTTTGGAAAATCCTGTAAACGTGTACAACCCGTCCCATTC TGACAGCCTCGCTTCTCAGCAAAGTGTTGCCAGTCATCCCAGACAATCTGGGCCTGGGGCGCCTAACCTTGACCGTTTTTATCAGCAGGT CACGAAAGATGCCCAGGGCCAGCCTGGCCTCGAAAGAGCCCAGCAGGAGCTGGTGCCACCCCAGCAACAGGCTTCTCCCCCACAACTACC CAAAGCCATGTTTTCGGAGCTGTCAAATCCAGAAAGTCTGCCCGCACAGGGACAGGCCCAGAACTCAGCACAGTCACCAGCAAGTCTGGT TCTGGTCGACGCGGGTCAGCAGCTGCCCCCTCGGCCTCCTCAGTCCTCTAGCGTGTCTCTGGTGTCCAGTGGCTCCGGCCAGGCAGCTGT GCCGTCAGAGCAGCCGTGGCCACAGCCAGTGCCTGCACTTGCCCCCGGCCCACCGCCTCAGGACCTGGCCGCCTACTACTACTACCGGCC TTTGTACGATGCCTACCAGCCTCAGTACTCTTTGCCGTACCCACCGGAGCCTGGCGCAGCCTCCCTCTATTACCAGGATGTCTACAGCCT CTATGAGCCTCGATACAGGCCCTATGATGGTGCTGCGTCTGCTTACGCCCAGAACTACCGCTATCCCGAGCCCGAGCGGCCCAGCTCCCG AGCCAGCCACTCCTCGGAACGGCCACCTCCCAGGCAAGGATATCCTGAAGGATACTATAGTTCCAAAAGTGGATGGAGCAGTCAGAGCGA TTACTATGCAAGCTATTACTCCAGCCAGTACGATTATGGAGATCCAGGTCACTGGGATCGTTACCACTACAGTGCTAGAGTCAGGGACCC CCGCACCTATGACCGGAGGTATTGGTGTGATGCAGAGTATGACGCATACAGGAGAGAGCACTCTGCCTTCGGGGACAGGCCCGAGAAACG TGACAACAACTGGAGGTACGATCCTCGCTTCACGGGGAGTTTTGACGATGACCCCGATCCGCACAGAGACCCTTATGGGGAAGAGGTGGA CCGGCGCAGCGTCCACAGCGAGCACTCGGCACGGAGCCTGCACAGCGCACACAGCCTGGCCAGCCGCCGCAGCAGCCTCAGCTCCCACTC GCACCAGAGTCAGATTTACAGAAGCCACAATGTGGCTGCCGGTTCCTACGAGGCCCCGCTTCCTCCAGGCTCCTTTCACGGCGATTTTGC CTACGGCACCTACCGCAGCAATTTCAGCAGTGGCCCCGGCTTCCCAGAGTATGGCTACCCTGCCGACACCGTCTGGCCTGCCATGGAGCA AGTTTCATCAAGACCAACTTCTCCTGAAAAATTTTCAGTGCCTCATGTCTGTGCCAGGTTTGGCCCTGGCGGTCAGCTTATCAAAGTGAT TCCCAATCTGCCTTCAGAAGGACAGCCGGCCTTGGTGGAGGTCCACAGCATGGAGGCCTTGCTGCAGCACACGTCTGAGCAGGAGGAGAT GCGGGCGTTCCCGGGACCCCTGGCCAAAGACGACACCCATAAGGTGGATGTCATTAATTTTGCACAGAACAAAGCTATGAAATGTTTGCA GAATGAAAACTTAATTGACAAAGAGTCTGCAAGTCTTCTTTGGAATTTTATTGTTCTCTTATGCAGACAAAATGGGACCGTGGTAGGGAC CGACATTGCGGAGCTTCTGTTACGAGACCACAGAACAGTGTGGCTTCCTGGGAAGTCGCCCAATGAAGCAAACCTGATTGATTTCACGAA TGAGGCAGTGGAGCAGGTGGAAGAGGAGGAGTCTGGTGAGGCCCAGCTCTCTTTCCTCACTGGTGGTCCGGCGGCTGCCGCCAGCTCGCT CGAGAGAGAGACCGAGAGGTTCAGGGAGCTGTTGCTGTATGGCCGTAAGAAGGATGCTTTGGAGTCTGCAATGAAGAATGGCCTGTGGGG TCACGCTCTGCTACTTGCAAGTAAGATGGACAGCCGGACACACGCCCGAGTCATGACCAGGTTTGCTAACAGCCTCCCAATCAACGACCC TCTGCAGACAGTCTACCAGCTCATGTCCGGACGGATGCCTGCCGCGTCCACGTGCTGTGGAGACGAGAAATGGGGAGATTGGAGGCCGCA CCTCGCCATGGTCTTGTCCAACTTGAACAACAACATGGACGTCGAGTCCAGGACGATGGCTACCATGGGCGACACTCTGGCTTCAAGGGG CCTCTTGGATGCGGCCCACTTCTGCTACCTCATGGCCCAGGCGGGATTTGGTGTTTACACGAAGAAAACTACAAAGCTTGTCTTAATCGG ATCCAATCACAGTTTGCCATTCTTAAAGTTCGCAACCAACGAAGCAATCCAGAGGACGGAAGCCTATGAGTACGCCCAGTCCCTGGGTGC CGAGACCTGCCCCCTGCCTAGTTTCCAGGTGTTTAAGTTCATCTACTCCTGCCGCCTGGCGGAAATGGGGCTGGCCACGCAAGCCTTCCA CTACTGTGAGGCCATCGCGAAGAGCATCCTGACGCAGCCGCACCTGTATTCCCCGGTGTTGATCAGCCAGCTTGTGCAGATGGCTTCCCA GTTACGACTCTTCGATCCCCAGCTGAAAGAGAAGCCAGAAGAGGAGTCCTTGGCCGCACCCACGTGGCTGGTTCACCTGCAGCAGGTGGA GCGGCAGATTAAGGAGGGGGCTGGAGTATGGCATCAGGATGGAGCCCTCCCGCAGCAGTGTCCTGGCACTCCGAGTTCCGAGATGGAGCA GTTGGACAGGCCAGGACTCAGTCAGCCAGGAGCCCTGGGGATCGCCAACCCTCTGCTGGCGGTGCCTGCACCGAGCCCTGAGCACTCGAG CCCGAGCGTGCGGCTGCTGCCCTCAGCTCCGCAGACGCTCCCTGACGGCCCATTGGCCAGTCCTGCCAGAGTGCCGATGTTCCCAGTGCC ACTGCCCCCGGGGCCCCTGGAGCCGGGTCCTGGCTGTGTGACCCCAGGGCCTGCACTTGGCTTCCTGGAGCCCTCCGGGCCTGGCCTCCC ACCTGGTGTGCCACCTCTGCAGGAAAGGAGACACTTGCTCCAGGAAGCCAGGAGCCCAGACCCAGGGATAGTGCCGCAGGAGGCGCCTGT TGGAAACTCACTTTCCGAGCTAAGCGAAGAAAATTTTGATGGAAAATTTGCTAATCTGACCCCCTCGAGGACGGTGCCAGACTCGGAGGC CCCCCCAGGGTGGGATCGTGCCGACTCGGGTCCCACGCAGCCACCTCTGTCTCTCTCACCCGCTCCCGAAACAAAGAGACCCGGACAGGC AGCCAAGAAAGAAACGAAGGAACCTAAGAAGGGTGAATCCTGGTTCTTTCGTTGGCTACCTGGAAAGAAAAAGACAGAAGCTTATTTGCC AGATGACAAGAACAAATCGATTGTTTGGGATGAAAAGAAAAACCAGTGGGTGAATTTAAATGAGCCAGAAGAGGAGAAGAAAGCCCCGCC CCCACCTCCAACCTCGATGCCCAAGACTGTGCAAGCTGCCCCGCCTGCCCTCCCAGGGCCTCCTGGAGCCCCCGTGAACATGTACTCTAG AAGAGCAGCAGGAACCAGAGCTCGCTACGTTGACGTCCTGAACCCAAGCGGGACCCAGCGGAGCGAGCCGGCTCTCGCTCCTGCGGACTT TGTCGCTCCACTCGCGCCACTCCCAATTCCTTCTAACTTGTTCGTGCCAACCCCAGATGCAGAAGAACCACAGCTTCCAGACGGGACTGG CAGGGAAGGGCCTGCAGCAGCTAGGGGCCTGGCCAATCCAGAGCCTGCCCCAGAGCCCAAGGTTTTAAGCTCTGCAGCGTCACTCCCTGG CTCTGAACTCCCCTCCTCCAGGCCTGAGGGTTCCCAGGGAGGAGAGCTTTCGCGCTGTAGTTCAATGAGTTCATTATCACGTGAAGTGAG CCAGCATTTTAATCAGGCTCCTGGCGACCTCCCTGCTGCAGGGGGCCCTCCCAGCGGGGCCATGCCCTTCTACAACCCTGCTCAGCTGGC ACAGGCCTGCGCCACCTCCGGGAGCTCAAGGCTAGGGAGGATTGGCCAGAGGAAGCACCTGGTGCTGAACTAGGCTTGCCCTGCTGTGAA CTTGCACTTGGAGCCCTGACGCTGCTGTTCTCCCCGAAGAACCCGACCGACCTCCGCGATCTCCGTCCCGCCCCCAGGGAGACACAGCAG TGACTCAGAGCTGGTCGCACACTGTGCCTCCCTCCTCACCGCCCATCGTAATGAATTATTTTGAAAATTAATTCCACCATCCTTTCAGAT TCTGGATGGAAAGACTGAATCTTTGACTCAGAATTGTTTGCCGAAAAGAATGATGTGACTTTCTTAGTCATTTAGGATGATTTAAGGATA TAGTATTCCTGGTCATTTAAGAATGTTCATTCATTGAAGCCGGAGCTGTCTCTGCCACAGGAGAGCCACATGGTCGGTAGTAACCAGGGC CTCTCCAAGCCCAGCTGTGAGTCACTGCCCAGTGAGTCCCGCGCTTCCTTTAAGGTGCTGGGAGCAAAGAGAGGGTGACTGAGGCAGACC CCAACCCCTGCTCTGCACCATCTGGGCCCTCGCCGTGTTTGAACCTGGCTGAATGAGTGGAGGGCGCTGTGTTCTCAATCAGCGCCTCCG AGGAGCCGTGGGGTTCCTTCGGCATTAGTTCACGGTTTTTGAGAGAGGCCTTAGTTACTGCAGTGAATTTCTTTCCTGTTGCAAAGACGC TTCCAGCCTCACTTTACTTTCTGTGGCCTGATGAGGACCATGGGTGATTTTGTGTACCCAAAGCGCTGGGGACTGCCCACCGTGTGGCCC AGTCACTGGGAAGGAGCCCCAGAGAGCCGGCTGTCTGACATGATGGCTCAGGGTGGTCATCCAGGTTGAAAACTGACCGTGTGATGTTTG ATTTGGGCTTCATTTCGTGTGTAGGAGCACGGTTAGACTCACTGTTAAGGAAGCTGGATGCACTTCTCTAAAAGGCTGCACTTTCCGTGA GCACTTTTCGTGGTACAATCCACATGACCCACTTTCTCCCCTGGGGGACGTTGGTTCAGAGGTTGGTAGCACTTGGGGAGAGTATCTTAA CACAGTTTCTTGACAGCAGCTCTGGAACTTAGTATTTCTGCCCCGAGTTTTGCCACACTGAGACTTTGAGTAGCTCCTGGTGGACTCAAC CCTGTTCAACTCAGAGACGGGCCTCCTCTCACTGATGCAAAGCTTTAAGGCTTCTCTGACTGTTCTGAAACTCTTCGTATTCTTGTCAAG TCTAAAGAGACTGAAGAAAAGATTTAAATACTAATAAAAATCAGTAGATAATTTCTGTAGGTTCTGCTGGAGGAATACAAACTGTTTGGT GTTTTAAATTTAAGTGTAGAAATTGTAGAATGTGGAATTAGCACAGATCCTTCCTGGCTTTCTGTTTCACTTGATCATTTAGCCCAGACC ACCCAGGATGTTTTCCAAAATGTTCCACAGGCGTGTCCCGCTGGATCCATTTGTCCTTGTCACTTGGAGAAAGGCCAGTCCCTGTGACGG GGCAGCCCTCTCTGTCCCTCGGTCAGCTCGTGTGAATCCTGGGACCTCTTCCGGTCGGCTCTGCCCGCTGTTCTGGGGTCGACTGCCACG ACTTTTGATTCAAGAAGCTTCCTCCAGGCGGGAGCGGCTATTTTTCCTAAATGAGAATTGTTACATTGCAAATTGTTGAATAAAATATTT >72217_72217_1_RAPGEF1-SEC16A_RAPGEF1_chr9_134615157_ENST00000372195_SEC16A_chr9_139372136_ENST00000313050_length(amino acids)=2481AA_BP=2 MRPARGPGERRRARGRRSGGGGRGGSGWARAGWSAVAERRAAAARSPPCRRGSEAWRARAEASGGRAAPVAPRRPGPGPMSGGLGLRRSP EMSGKIEKAVQLQLRNSYILLPSVSSIDLCKGSVMQPPPQTVPSGMAGPPPAGNPRSVFWASSPYRRRANNNAAVAPTTCPLQPVTDPFA FSRQALQSTPLGSSSKSSPPVLQGPAPAGFSQHPGLLVPHTHARDSSQGPCEPLPGPLTQPRAHASPFSGALTPSAPPGPEMNRSAEVGP SSEPEVQTLPYLPHYIPGVDPETSHGGHPHGNMPGLDRPLSRQNPHDGVVTPAASPSLPQPGLQMPGQWGPVQGGPQPSGQHRSPCPEGP VPSGVPCATSVPHFPTPSILHQGPGHEQHSPLVAPPAALPSDGRDEVSHLQSGSHLANNSDPESTFRQNPRIVNHWASPELRQNPGVKNE HRPASALVNPLARGDSPENRTHHPLGAGAGSGCAPLEADSGASGALAMFFQGGETENEENLSSEKAGLSGQADFDDFCSSPGLGRPPAPT HVGAGSLCQALLPGPSNEAAGDVWGDTASTGVPDASGSQYENVENLEFVQNQEVLPSEPLNLDPSSPSDQFRYGPLPGPAVPRHGAVCHT GAPDATLHTVHPDSVSSSYSSRSHGRLSGSARPQELVGTFIQQEVGKPEDEASGSFFKQIDSSPVGGETDETTVSQNYRGSVSQPSTPSP PKPTGIFQTSANSSFEPVKSHLVGVKPFEADRANVVGEVRETCVRQKQCRPAAALPDASPGNLEQPPDNMETLCAPQVCPLPLNSTTEAV HMLPHAGAPPLDTVYPAPEKRPSARTQGPVKCESPATTLWAQSELPDFGGNVLLAPAAPALYVCAKPQPPVVQPPEEAMSGQQSRNPSSA APVQSRGGIGASENLENPPKMGEEEALQSQASSGYASLLSSPPTESLQNPPVLIAQPDHSYNLAQPINFSVSLSNSHEKNQSWREALVGD RPAVSSWALGGDSGENTSLSGIPTSSVLSLSLPSSVAQSNFPQGSGASEMVSNQPANLLVQPPSQPVPENLVPESQKDRKAGSALPGFAN SPAGSTSVVLVPPAHGTLVPDGNKANHSSHQEDTYGALDFTLSRTLENPVNVYNPSHSDSLASQQSVASHPRQSGPGAPNLDRFYQQVTK DAQGQPGLERAQQELVPPQQQASPPQLPKAMFSELSNPESLPAQGQAQNSAQSPASLVLVDAGQQLPPRPPQSSSVSLVSSGSGQAAVPS EQPWPQPVPALAPGPPPQDLAAYYYYRPLYDAYQPQYSLPYPPEPGAASLYYQDVYSLYEPRYRPYDGAASAYAQNYRYPEPERPSSRAS HSSERPPPRQGYPEGYYSSKSGWSSQSDYYASYYSSQYDYGDPGHWDRYHYSARVRDPRTYDRRYWCDAEYDAYRREHSAFGDRPEKRDN NWRYDPRFTGSFDDDPDPHRDPYGEEVDRRSVHSEHSARSLHSAHSLASRRSSLSSHSHQSQIYRSHNVAAGSYEAPLPPGSFHGDFAYG TYRSNFSSGPGFPEYGYPADTVWPAMEQVSSRPTSPEKFSVPHVCARFGPGGQLIKVIPNLPSEGQPALVEVHSMEALLQHTSEQEEMRA FPGPLAKDDTHKVDVINFAQNKAMKCLQNENLIDKESASLLWNFIVLLCRQNGTVVGTDIAELLLRDHRTVWLPGKSPNEANLIDFTNEA VEQVEEEESGEAQLSFLTGGPAAAASSLERETERFRELLLYGRKKDALESAMKNGLWGHALLLASKMDSRTHARVMTRFANSLPINDPLQ TVYQLMSGRMPAASTCCGDEKWGDWRPHLAMVLSNLNNNMDVESRTMATMGDTLASRGLLDAAHFCYLMAQAGFGVYTKKTTKLVLIGSN HSLPFLKFATNEAIQRTEAYEYAQSLGAETCPLPSFQVFKFIYSCRLAEMGLATQAFHYCEAIAKSILTQPHLYSPVLISQLVQMASQLR LFDPQLKEKPEEESLAAPTWLVHLQQVERQIKEGAGVWHQDGALPQQCPGTPSSEMEQLDRPGLSQPGALGIANPLLAVPAPSPEHSSPS VRLLPSAPQTLPDGPLASPARVPMFPVPLPPGPLEPGPGCVTPGPALGFLEPSGPGLPPGVPPLQERRHLLQEARSPDPGIVPQEAPVGN SLSELSEENFDGKFANLTPSRTVPDSEAPPGWDRADSGPTQPPLSLSPAPETKRPGQAAKKETKEPKKGESWFFRWLPGKKKTEAYLPDD KNKSIVWDEKKNQWVNLNEPEEEKKAPPPPPTSMPKTVQAAPPALPGPPGAPVNMYSRRAAGTRARYVDVLNPSGTQRSEPALAPADFVA PLAPLPIPSNLFVPTPDAEEPQLPDGTGREGPAAARGLANPEPAPEPKVLSSAASLPGSELPSSRPEGSQGGELSRCSSMSSLSREVSQH -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for RAPGEF1-SEC16A |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000290037 | 0 | 29 | 1101_1400 | 0.0 | 2160.0 | MIA3 | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000371706 | 0 | 28 | 1101_1400 | 0 | 2135.0 | MIA3 | |
| Tgene | SEC16A | chr9:134615157 | chr9:139372136 | ENST00000431893 | 0 | 29 | 1101_1400 | 0.0 | 2155.0 | MIA3 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for RAPGEF1-SEC16A |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for RAPGEF1-SEC16A |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
