
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:RBPMS-RILPL1 (FusionGDB2 ID:73096) |
Fusion Gene Summary for RBPMS-RILPL1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: RBPMS-RILPL1 | Fusion gene ID: 73096 | Hgene | Tgene | Gene symbol | RBPMS | RILPL1 | Gene ID | 11030 | 353116 |
| Gene name | RNA binding protein, mRNA processing factor | Rab interacting lysosomal protein like 1 | |
| Synonyms | HERMES | GOSPEL|RLP1 | |
| Cytomap | 8p12 | 12q24.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | RNA-binding protein with multiple splicingRNA binding protein with multiple splicingheart and RRM expressed sequence | RILP-like protein 1GAPDH competitor of SIAH protein enhances life | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000287771, ENST00000320203, ENST00000339877, ENST00000397323, ENST00000517860, ENST00000538486, ENST00000519647, ENST00000520161, ENST00000520191, ENST00000520916, | ENST00000544468, ENST00000340724, ENST00000376874, | |
| Fusion gene scores | * DoF score | 8 X 6 X 5=240 | 8 X 6 X 4=192 |
| # samples | 8 | 8 | |
| ** MAII score | log2(8/240*10)=-1.58496250072116 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(8/192*10)=-1.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: RBPMS [Title/Abstract] AND RILPL1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | RBPMS(30242674)-RILPL1(123970352), # samples:4 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | RBPMS | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation | 17099224 |
| Hgene | RBPMS | GO:0060391 | positive regulation of SMAD protein signal transduction | 17099224 |
 Fusion gene breakpoints across RBPMS (5'-gene) Fusion gene breakpoints across RBPMS (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
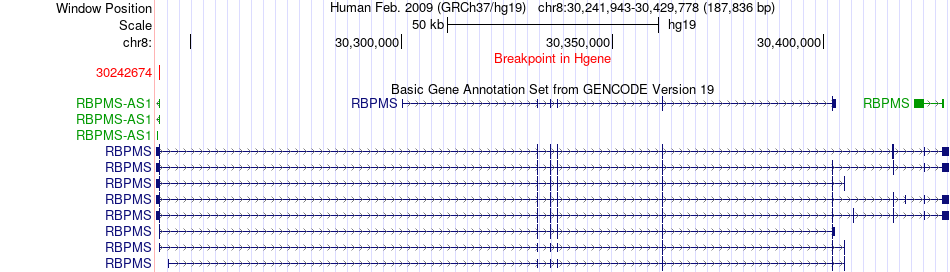 |
 Fusion gene breakpoints across RILPL1 (3'-gene) Fusion gene breakpoints across RILPL1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
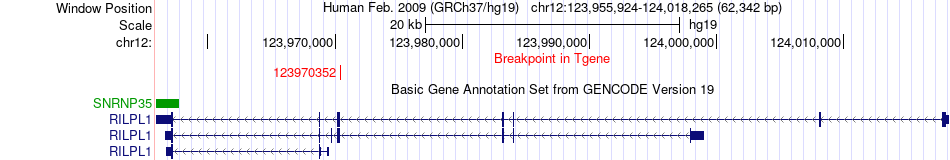 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-D8-A1JM-01A | RBPMS | chr8 | 30242674 | - | RILPL1 | chr12 | 123970352 | - |
| ChimerDB4 | BRCA | TCGA-D8-A1JM-01A | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| ChimerDB4 | BRCA | TCGA-D8-A1JM | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
Top |
Fusion Gene ORF analysis for RBPMS-RILPL1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000287771 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| 5CDS-intron | ENST00000320203 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| 5CDS-intron | ENST00000339877 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| 5CDS-intron | ENST00000397323 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| 5CDS-intron | ENST00000517860 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| 5CDS-intron | ENST00000538486 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000287771 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000287771 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000320203 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000320203 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000339877 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000339877 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000397323 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000397323 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000517860 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000517860 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000538486 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| In-frame | ENST00000538486 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000519647 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000519647 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000520161 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000520161 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000520191 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000520191 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000520916 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-3CDS | ENST00000520916 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-intron | ENST00000519647 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-intron | ENST00000520161 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-intron | ENST00000520191 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
| intron-intron | ENST00000520916 | ENST00000544468 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000538486 | RBPMS | chr8 | 30242674 | + | ENST00000376874 | RILPL1 | chr12 | 123970352 | - | 2302 | 731 | 350 | 1141 | 263 |
| ENST00000538486 | RBPMS | chr8 | 30242674 | + | ENST00000340724 | RILPL1 | chr12 | 123970352 | - | 1708 | 731 | 350 | 1237 | 295 |
| ENST00000397323 | RBPMS | chr8 | 30242674 | + | ENST00000376874 | RILPL1 | chr12 | 123970352 | - | 2251 | 680 | 299 | 1090 | 263 |
| ENST00000397323 | RBPMS | chr8 | 30242674 | + | ENST00000340724 | RILPL1 | chr12 | 123970352 | - | 1657 | 680 | 299 | 1186 | 295 |
| ENST00000320203 | RBPMS | chr8 | 30242674 | + | ENST00000376874 | RILPL1 | chr12 | 123970352 | - | 2219 | 648 | 267 | 1058 | 263 |
| ENST00000320203 | RBPMS | chr8 | 30242674 | + | ENST00000340724 | RILPL1 | chr12 | 123970352 | - | 1625 | 648 | 267 | 1154 | 295 |
| ENST00000287771 | RBPMS | chr8 | 30242674 | + | ENST00000376874 | RILPL1 | chr12 | 123970352 | - | 2203 | 632 | 251 | 1042 | 263 |
| ENST00000287771 | RBPMS | chr8 | 30242674 | + | ENST00000340724 | RILPL1 | chr12 | 123970352 | - | 1609 | 632 | 251 | 1138 | 295 |
| ENST00000339877 | RBPMS | chr8 | 30242674 | + | ENST00000376874 | RILPL1 | chr12 | 123970352 | - | 2219 | 648 | 267 | 1058 | 263 |
| ENST00000339877 | RBPMS | chr8 | 30242674 | + | ENST00000340724 | RILPL1 | chr12 | 123970352 | - | 1625 | 648 | 267 | 1154 | 295 |
| ENST00000517860 | RBPMS | chr8 | 30242674 | + | ENST00000376874 | RILPL1 | chr12 | 123970352 | - | 1645 | 74 | 8 | 484 | 158 |
| ENST00000517860 | RBPMS | chr8 | 30242674 | + | ENST00000340724 | RILPL1 | chr12 | 123970352 | - | 1051 | 74 | 8 | 580 | 190 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000538486 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.004968929 | 0.995031 |
| ENST00000538486 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.003086172 | 0.9969138 |
| ENST00000397323 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.005513236 | 0.99448675 |
| ENST00000397323 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.003049239 | 0.99695075 |
| ENST00000320203 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.003644451 | 0.99635553 |
| ENST00000320203 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.002227796 | 0.99777216 |
| ENST00000287771 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.004572833 | 0.99542713 |
| ENST00000287771 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.00286569 | 0.9971343 |
| ENST00000339877 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.003644451 | 0.99635553 |
| ENST00000339877 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.002227796 | 0.99777216 |
| ENST00000517860 | ENST00000376874 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.001730377 | 0.99826956 |
| ENST00000517860 | ENST00000340724 | RBPMS | chr8 | 30242674 | + | RILPL1 | chr12 | 123970352 | - | 0.001713835 | 0.9982862 |
Top |
Fusion Genomic Features for RBPMS-RILPL1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
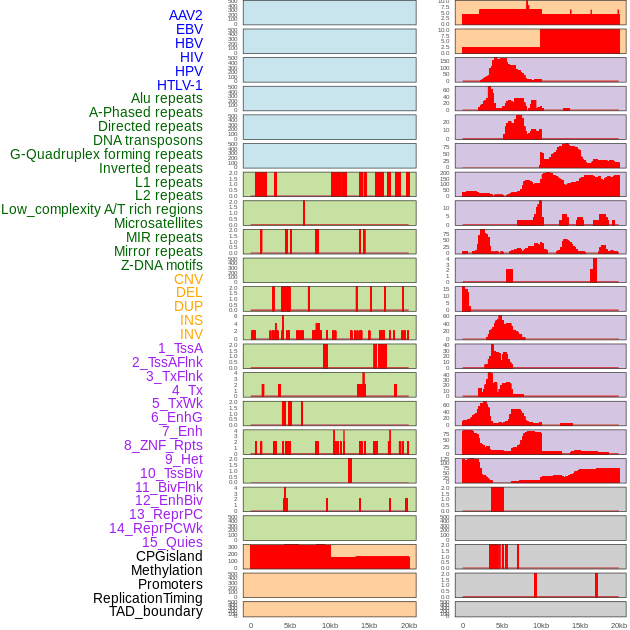 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for RBPMS-RILPL1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr8:30242674/chr12:123970352) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | RILPL1 | chr8:30242674 | chr12:123970352 | ENST00000376874 | 3 | 7 | 291_356 | 267 | 404.0 | Domain | RH2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000287771 | + | 1 | 10 | 24_101 | 22 | 543.0 | Domain | RRM |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000320203 | + | 1 | 10 | 24_101 | 22 | 585.3333333333334 | Domain | RRM |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000339877 | + | 1 | 7 | 24_101 | 22 | 220.0 | Domain | RRM |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000397323 | + | 1 | 9 | 24_101 | 22 | 520.6666666666666 | Domain | RRM |
| Tgene | RILPL1 | chr8:30242674 | chr12:123970352 | ENST00000376874 | 3 | 7 | 76_258 | 267 | 404.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | RILPL1 | chr8:30242674 | chr12:123970352 | ENST00000376874 | 3 | 7 | 10_97 | 267 | 404.0 | Domain | RH1 |
Top |
Fusion Gene Sequence for RBPMS-RILPL1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >73096_73096_1_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000287771_RILPL1_chr12_123970352_ENST00000340724_length(transcript)=1609nt_BP=632nt ATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCCGCGCTCCTCCGCCCCG CTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCCCAGCCCCAACTCTCCG CGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTGGCTCCCGCCCTCCCTC TCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTTGCACTTCCTGAGTCGC CCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCAGCTCCAGCCCCACAGC CCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCCGCGCCCCAGCCCTGCC CGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTCAGGAGGAGG AGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGTTCACCCTGC AGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATAAGAGGTTTC CACATGAGACCACAAGCAACATGAGACCAGAAGTCTTATTTATCTTGTCCATGGCTAAGTTCCCAGCACTTAGAATGGAGCTTGGCACAC ATGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCAAGC GACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGGCAA ACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTGGAC TGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTGACC CCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATCACT GTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAACACA AGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTATCA >73096_73096_1_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000287771_RILPL1_chr12_123970352_ENST00000340724_length(amino acids)=295AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL AYYKRFPHETTSNMRPEVLFILSMAKFPALRMELGTHEEMEEENRIPQPPPIAHPRTSPQPESGIKRLFSFFSRDKKRLANTQRNVHIQE -------------------------------------------------------------- >73096_73096_2_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000287771_RILPL1_chr12_123970352_ENST00000376874_length(transcript)=2203nt_BP=632nt ATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCCGCGCTCCTCCGCCCCG CTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCCCAGCCCCAACTCTCCG CGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTGGCTCCCGCCCTCCCTC TCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTTGCACTTCCTGAGTCGC CCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCAGCTCCAGCCCCACAGC CCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCCGCGCCCCAGCCCTGCC CGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTCAGGAGGAGG AGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGTTCACCCTGC AGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATAAGAGTGAAG AAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCAAGCGACTGT TTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGGCAAACACCC ACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTGGACTGCCCG CCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTGACCCCTCAA ACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATCACTGTTTCT AATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAACACAAGCTTC TTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTATCACAGGTG ACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTAACGTATTATACAGAATGTATCC ATCAGGCAGACAAGGGGTCTGAAGTCACAGGCTCAGTAGCCCAACTCAGACACAAAGCCACAGACAATATGGATGATGGTCCTCTTACCC AGAAACTGGAGACAACTTTGCAGGCACCCTGGCTGCGTTCCTGATAATGCCTAGAGCATGTAGCAATGTTCAAGGCAGGTGCCTTGGAAT CTGCTGTGAGTTATGCAGTACTAACCAAACAAAGTTGCTAAGGATGAGATGTTGCAACATTTCTTGTGTGCACTTTTTTTTCCTGACATC TTTTTTTCCTTTTAGGATTCATTAAGTCATATACTTAGTCCCCTGCAAAGAAGAAGACAGATGTCCCAGAGCAGACTGAAAGCGGGGCGT GGGATCCTAGGCAATGCAGGAGGGCAGGTGGGGGAGCTGTAGCAGAACAAGGGTGCTAGGTGGGAAGCTGGCTTGGGGACCACACGTCTG GCTGACTCACAGGGCTATAGCAGCAGACAGAGAGCCAGATGCTCTACTGCAACTGGTGAGTTGGAGGCTTGCCCACATAACACTGTCTTT CTCTCCACCCGCCAAGCATCCAGGGCACAGGGCTGGTTGCTCACAGATTGAGCGAGGTAGCACCTCCTACCAGCACTCAAGTTTATCAGG >73096_73096_2_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000287771_RILPL1_chr12_123970352_ENST00000376874_length(amino acids)=263AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL -------------------------------------------------------------- >73096_73096_3_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000320203_RILPL1_chr12_123970352_ENST00000340724_length(transcript)=1625nt_BP=648nt AAAGGGAGCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCC GCGCTCCTCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCC CAGCCCCAACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTG GCTCCCGCCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTT GCACTTCCTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCA GCTCCAGCCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCC GCGCCCCAGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCC AACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGC CCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCT TACTATAAGAGGTTTCCACATGAGACCACAAGCAACATGAGACCAGAAGTCTTATTTATCTTGTCCATGGCTAAGTTCCCAGCACTTAGA ATGGAGCTTGGCACACATGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCG GAGTCGGGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCC TTTGGACAGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCC ACCCTCCAACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGC ACCCTGAAAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTAT ACCCAGGGCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGT TAACTTCAGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGT AGAGGTTTTATTATCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTA >73096_73096_3_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000320203_RILPL1_chr12_123970352_ENST00000340724_length(amino acids)=295AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL AYYKRFPHETTSNMRPEVLFILSMAKFPALRMELGTHEEMEEENRIPQPPPIAHPRTSPQPESGIKRLFSFFSRDKKRLANTQRNVHIQE -------------------------------------------------------------- >73096_73096_4_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000320203_RILPL1_chr12_123970352_ENST00000376874_length(transcript)=2219nt_BP=648nt AAAGGGAGCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCC GCGCTCCTCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCC CAGCCCCAACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTG GCTCCCGCCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTT GCACTTCCTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCA GCTCCAGCCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCC GCGCCCCAGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCC AACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGC CCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCT TACTATAAGAGTGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCG GGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGA CAGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTC CAACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTG AAAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAG GGCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTT CAGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGT TTTATTATCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTAACGTAT TATACAGAATGTATCCATCAGGCAGACAAGGGGTCTGAAGTCACAGGCTCAGTAGCCCAACTCAGACACAAAGCCACAGACAATATGGAT GATGGTCCTCTTACCCAGAAACTGGAGACAACTTTGCAGGCACCCTGGCTGCGTTCCTGATAATGCCTAGAGCATGTAGCAATGTTCAAG GCAGGTGCCTTGGAATCTGCTGTGAGTTATGCAGTACTAACCAAACAAAGTTGCTAAGGATGAGATGTTGCAACATTTCTTGTGTGCACT TTTTTTTCCTGACATCTTTTTTTCCTTTTAGGATTCATTAAGTCATATACTTAGTCCCCTGCAAAGAAGAAGACAGATGTCCCAGAGCAG ACTGAAAGCGGGGCGTGGGATCCTAGGCAATGCAGGAGGGCAGGTGGGGGAGCTGTAGCAGAACAAGGGTGCTAGGTGGGAAGCTGGCTT GGGGACCACACGTCTGGCTGACTCACAGGGCTATAGCAGCAGACAGAGAGCCAGATGCTCTACTGCAACTGGTGAGTTGGAGGCTTGCCC ACATAACACTGTCTTTCTCTCCACCCGCCAAGCATCCAGGGCACAGGGCTGGTTGCTCACAGATTGAGCGAGGTAGCACCTCCTACCAGC >73096_73096_4_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000320203_RILPL1_chr12_123970352_ENST00000376874_length(amino acids)=263AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL -------------------------------------------------------------- >73096_73096_5_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000339877_RILPL1_chr12_123970352_ENST00000340724_length(transcript)=1625nt_BP=648nt AAAGGGAGCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCC GCGCTCCTCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCC CAGCCCCAACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTG GCTCCCGCCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTT GCACTTCCTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCA GCTCCAGCCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCC GCGCCCCAGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCC AACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGC CCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCT TACTATAAGAGGTTTCCACATGAGACCACAAGCAACATGAGACCAGAAGTCTTATTTATCTTGTCCATGGCTAAGTTCCCAGCACTTAGA ATGGAGCTTGGCACACATGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCG GAGTCGGGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCC TTTGGACAGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCC ACCCTCCAACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGC ACCCTGAAAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTAT ACCCAGGGCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGT TAACTTCAGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGT AGAGGTTTTATTATCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTA >73096_73096_5_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000339877_RILPL1_chr12_123970352_ENST00000340724_length(amino acids)=295AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL AYYKRFPHETTSNMRPEVLFILSMAKFPALRMELGTHEEMEEENRIPQPPPIAHPRTSPQPESGIKRLFSFFSRDKKRLANTQRNVHIQE -------------------------------------------------------------- >73096_73096_6_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000339877_RILPL1_chr12_123970352_ENST00000376874_length(transcript)=2219nt_BP=648nt AAAGGGAGCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCC GCGCTCCTCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCC CAGCCCCAACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTG GCTCCCGCCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTT GCACTTCCTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCA GCTCCAGCCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCC GCGCCCCAGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCC AACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGC CCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCT TACTATAAGAGTGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCG GGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGA CAGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTC CAACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTG AAAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAG GGCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTT CAGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGT TTTATTATCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTAACGTAT TATACAGAATGTATCCATCAGGCAGACAAGGGGTCTGAAGTCACAGGCTCAGTAGCCCAACTCAGACACAAAGCCACAGACAATATGGAT GATGGTCCTCTTACCCAGAAACTGGAGACAACTTTGCAGGCACCCTGGCTGCGTTCCTGATAATGCCTAGAGCATGTAGCAATGTTCAAG GCAGGTGCCTTGGAATCTGCTGTGAGTTATGCAGTACTAACCAAACAAAGTTGCTAAGGATGAGATGTTGCAACATTTCTTGTGTGCACT TTTTTTTCCTGACATCTTTTTTTCCTTTTAGGATTCATTAAGTCATATACTTAGTCCCCTGCAAAGAAGAAGACAGATGTCCCAGAGCAG ACTGAAAGCGGGGCGTGGGATCCTAGGCAATGCAGGAGGGCAGGTGGGGGAGCTGTAGCAGAACAAGGGTGCTAGGTGGGAAGCTGGCTT GGGGACCACACGTCTGGCTGACTCACAGGGCTATAGCAGCAGACAGAGAGCCAGATGCTCTACTGCAACTGGTGAGTTGGAGGCTTGCCC ACATAACACTGTCTTTCTCTCCACCCGCCAAGCATCCAGGGCACAGGGCTGGTTGCTCACAGATTGAGCGAGGTAGCACCTCCTACCAGC >73096_73096_6_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000339877_RILPL1_chr12_123970352_ENST00000376874_length(amino acids)=263AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL -------------------------------------------------------------- >73096_73096_7_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000397323_RILPL1_chr12_123970352_ENST00000340724_length(transcript)=1657nt_BP=680nt GGGAGGGGAGAGGGGAGGGAGGAAGGCGAGAGAAAGGGAGCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCG GCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCCGCGCTCCTCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCAC CCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCCCAGCCCCAACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTT CCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTGGCTCCCGCCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGC TCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTTGCACTTCCTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCA ACCCGAGCCCGACAGCCACTGCCCCGGCTCCAGCTCCAGCCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGC TCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCCGCGCCCCAGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCA AAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGA AGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGT CCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATAAGAGGTTTCCACATGAGACCACAAGCAACATGAGACCAGAAGTCTTATTTA TCTTGTCCATGGCTAAGTTCCCAGCACTTAGAATGGAGCTTGGCACACATGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCAC CCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCA ACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAG CCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGT CATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAAC TGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAA TAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCT GTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTATCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCT >73096_73096_7_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000397323_RILPL1_chr12_123970352_ENST00000340724_length(amino acids)=295AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL AYYKRFPHETTSNMRPEVLFILSMAKFPALRMELGTHEEMEEENRIPQPPPIAHPRTSPQPESGIKRLFSFFSRDKKRLANTQRNVHIQE -------------------------------------------------------------- >73096_73096_8_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000397323_RILPL1_chr12_123970352_ENST00000376874_length(transcript)=2251nt_BP=680nt GGGAGGGGAGAGGGGAGGGAGGAAGGCGAGAGAAAGGGAGCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCG GCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCCGCGCTCCTCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCAC CCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCCCAGCCCCAACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTT CCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTGGCTCCCGCCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGC TCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTTGCACTTCCTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCA ACCCGAGCCCGACAGCCACTGCCCCGGCTCCAGCTCCAGCCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGC TCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCCGCGCCCCAGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCA AAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGA AGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGT CCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATAAGAGTGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCG CCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACAC AGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGC AGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGC TGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGC CAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCC CACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGA CTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTATCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTA GTTACATTCATGAAATTGTGATGGTAACGTATTATACAGAATGTATCCATCAGGCAGACAAGGGGTCTGAAGTCACAGGCTCAGTAGCCC AACTCAGACACAAAGCCACAGACAATATGGATGATGGTCCTCTTACCCAGAAACTGGAGACAACTTTGCAGGCACCCTGGCTGCGTTCCT GATAATGCCTAGAGCATGTAGCAATGTTCAAGGCAGGTGCCTTGGAATCTGCTGTGAGTTATGCAGTACTAACCAAACAAAGTTGCTAAG GATGAGATGTTGCAACATTTCTTGTGTGCACTTTTTTTTCCTGACATCTTTTTTTCCTTTTAGGATTCATTAAGTCATATACTTAGTCCC CTGCAAAGAAGAAGACAGATGTCCCAGAGCAGACTGAAAGCGGGGCGTGGGATCCTAGGCAATGCAGGAGGGCAGGTGGGGGAGCTGTAG CAGAACAAGGGTGCTAGGTGGGAAGCTGGCTTGGGGACCACACGTCTGGCTGACTCACAGGGCTATAGCAGCAGACAGAGAGCCAGATGC TCTACTGCAACTGGTGAGTTGGAGGCTTGCCCACATAACACTGTCTTTCTCTCCACCCGCCAAGCATCCAGGGCACAGGGCTGGTTGCTC ACAGATTGAGCGAGGTAGCACCTCCTACCAGCACTCAAGTTTATCAGGGCCGCGTTCTAAACATTCTACTGAAGTTTGCTAAGAAACCAT >73096_73096_8_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000397323_RILPL1_chr12_123970352_ENST00000376874_length(amino acids)=263AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL -------------------------------------------------------------- >73096_73096_9_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000517860_RILPL1_chr12_123970352_ENST00000340724_length(transcript)=1051nt_BP=74nt CCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAG AGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGC TGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATAAGAGGTTTCCACATGAGACCACAAGCA ACATGAGACCAGAAGTCTTATTTATCTTGTCCATGGCTAAGTTCCCAGCACTTAGAATGGAGCTTGGCACACATGAAGAAATGGAAGAGG AAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCAAGCGACTGTTTAGCTTCTTCT CCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGGCAAACACCCACCGCGATGACG GTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTGGACTGCCCGCCACCAGCGCCT GCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTGACCCCTCAAACAGACTGTCTG ATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATCACTGTTTCTAATAGATGACTC TGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAACACAAGCTTCTTTAAGCCAAAT ACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTATCACAGGTGACATAGTTCAGC >73096_73096_9_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000517860_RILPL1_chr12_123970352_ENST00000340724_length(amino acids)=190AA_BP=22 MNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEELAYYKRFPHETTSNMR PEVLFILSMAKFPALRMELGTHEEMEEENRIPQPPPIAHPRTSPQPESGIKRLFSFFSRDKKRLANTQRNVHIQESFGQWANTHRDDGYT -------------------------------------------------------------- >73096_73096_10_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000517860_RILPL1_chr12_123970352_ENST00000376874_length(transcript)=1645nt_BP=74nt CCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTCAGGAGGAGGAGACGGAGCCGGTGGGAG AGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGTTCACCCTGCAGGAGCTGCGGGACGTGC TGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATAAGAGTGAAGAAATGGAAGAGGAAAACC GAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCAAGCGACTGTTTAGCTTCTTCTCCCGAG ATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGGCAAACACCCACCGCGATGACGGTTACA CAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTGGACTGCCCGCCACCAGCGCCTGCAACC GAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTGACCCCTCAAACAGACTGTCTGATTTGA GGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATCACTGTTTCTAATAGATGACTCTGATCC CGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAACACAAGCTTCTTTAAGCCAAATACATCA CTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTATCACAGGTGACATAGTTCAGCAGGAGG CATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTAACGTATTATACAGAATGTATCCATCAGGCAGACAAGGGGT CTGAAGTCACAGGCTCAGTAGCCCAACTCAGACACAAAGCCACAGACAATATGGATGATGGTCCTCTTACCCAGAAACTGGAGACAACTT TGCAGGCACCCTGGCTGCGTTCCTGATAATGCCTAGAGCATGTAGCAATGTTCAAGGCAGGTGCCTTGGAATCTGCTGTGAGTTATGCAG TACTAACCAAACAAAGTTGCTAAGGATGAGATGTTGCAACATTTCTTGTGTGCACTTTTTTTTCCTGACATCTTTTTTTCCTTTTAGGAT TCATTAAGTCATATACTTAGTCCCCTGCAAAGAAGAAGACAGATGTCCCAGAGCAGACTGAAAGCGGGGCGTGGGATCCTAGGCAATGCA GGAGGGCAGGTGGGGGAGCTGTAGCAGAACAAGGGTGCTAGGTGGGAAGCTGGCTTGGGGACCACACGTCTGGCTGACTCACAGGGCTAT AGCAGCAGACAGAGAGCCAGATGCTCTACTGCAACTGGTGAGTTGGAGGCTTGCCCACATAACACTGTCTTTCTCTCCACCCGCCAAGCA TCCAGGGCACAGGGCTGGTTGCTCACAGATTGAGCGAGGTAGCACCTCCTACCAGCACTCAAGTTTATCAGGGCCGCGTTCTAAACATTC >73096_73096_10_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000517860_RILPL1_chr12_123970352_ENST00000376874_length(amino acids)=158AA_BP=22 MNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEELAYYKSEEMEEENRIP -------------------------------------------------------------- >73096_73096_11_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000538486_RILPL1_chr12_123970352_ENST00000340724_length(transcript)=1708nt_BP=731nt GGCAGCCCACGTCCAGGGAGGGCGGAGGGAGGAGGAGAGGCGCGGAGAGGAGGGAGGGGAGAGGGGAGGGAGGAAGGCGAGAGAAAGGGA GCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCCGCGCTCC TCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCCCAGCCCC AACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTGGCTCCCG CCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTTGCACTTC CTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCAGCTCCAG CCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCCGCGCCCC AGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTC AGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGT TCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATA AGAGGTTTCCACATGAGACCACAAGCAACATGAGACCAGAAGTCTTATTTATCTTGTCCATGGCTAAGTTCCCAGCACTTAGAATGGAGC TTGGCACACATGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGG GCATCAAGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGAC AGTGGGCAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCC AACCTGGACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGA AAACTGACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGG GCTATCACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTC AGTAACACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTT >73096_73096_11_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000538486_RILPL1_chr12_123970352_ENST00000340724_length(amino acids)=295AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL AYYKRFPHETTSNMRPEVLFILSMAKFPALRMELGTHEEMEEENRIPQPPPIAHPRTSPQPESGIKRLFSFFSRDKKRLANTQRNVHIQE -------------------------------------------------------------- >73096_73096_12_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000538486_RILPL1_chr12_123970352_ENST00000376874_length(transcript)=2302nt_BP=731nt GGCAGCCCACGTCCAGGGAGGGCGGAGGGAGGAGGAGAGGCGCGGAGAGGAGGGAGGGGAGAGGGGAGGGAGGAAGGCGAGAGAAAGGGA GCTGCTTCCATCCCGGACTTCCCAGAGCCTGCCTGGAGCGCGTACTCAGCGGCTCTCGGGTCCCAGCGTCCCAGCCGCGGCCCGCGCTCC TCCGCCCCGCTCCTCCTCCTCCTCTTCCTCCTCCTCCTCCTCTCTAGGCACCCCCGTCCCCTCCTTCCAGCGGCTGCAGCCCCCAGCCCC AACTCTCCGCGCTTACTCCTGGGACGCGCGTCCTCGCCCCATCCTTTGCTTCCTTCCTTCCTTCCTTCTTCCTTCCTCCCCTGGCTCCCG CCCTCCCTCTCCAGGTCGCCCTCCCGGGGCCCGATTGTCTCGGTGCCCCGCTCCCGGCCCGCGCCCTGCCCCGTCTCTCCCTTGCACTTC CTGAGTCGCCCGCCGCCGCCGTCGCAGACTCGCCGCGGGAGCCCCAGCCCAACCCGAGCCCGACAGCCACTGCCCCGGCTCCAGCTCCAG CCCCACAGCCCGCGGCGCCCGCCCGAGGGAGCCCCGGCGCCCGGGGAAGGCTCCAGTGGGCTAGCGCGCCCTCGCCCAGCCCCGCGCCCC AGCCCTGCCCGGCCCGGCGAGGAAGGACCGGGAAGATGAACAACGGCGGCAAAGCCGAGAAGGAGAACACCCCGAGCGAGGCCAACCTTC AGGAGGAGGAGACGGAGCCGGTGGGAGAGGAGAGCATCTCCGACGCAGAGAAGGTGGCCATGGATCTCAAGGACCCCAACCGCCCCCGGT TCACCCTGCAGGAGCTGCGGGACGTGCTGCACGAGAGGAACGAGCTCAAGTCCAAGGTGTTCTTGCTGCAGGAGGAGCTGGCTTACTATA AGAGTGAAGAAATGGAAGAGGAAAACCGAATACCCCAACCCCCACCCATCGCCCACCCGAGGACGTCCCCCCAGCCGGAGTCGGGCATCA AGCGACTGTTTAGCTTCTTCTCCCGAGATAAGAAGCGCCTGGCCAACACACAGAGAAACGTGCACATCCAGGAGTCCTTTGGACAGTGGG CAAACACCCACCGCGATGACGGTTACACAGAGCAAGGACAGGAAGCCCTGCAGCATCTGTGACCTTGGCCCATCTCCACCCTCCAACCTG GACTGCCCGCCACCAGCGCCTGCAACCGAACTGCAGCCCAGGGGTCATTGCTGCCTCAAGCCTCTCGGTGCAGATGCACCCTGAAAACTG ACCCCTCAAACAGACTGTCTGATTTGAGGATGGACATTGAAAAACTGACGCCAAACTCTAAAGAAATGTTTATTTATACCCAGGGCTATC ACTGTTTCTAATAGATGACTCTGATCCCGTAGGATATATATTTAATAATCCCACAAACGGAGGCCAGACTTCTGCGTTAACTTCAGTAAC ACAAGCTTCTTTAAGCCAAATACATCACTTGCCACTATCATTGCTGTTTGACTTGCTTTGTATAAAATGCTATGTGTAGAGGTTTTATTA TCACAGGTGACATAGTTCAGCAGGAGGCATGGAAGGGCTAGATCTTCATTAGTTACATTCATGAAATTGTGATGGTAACGTATTATACAG AATGTATCCATCAGGCAGACAAGGGGTCTGAAGTCACAGGCTCAGTAGCCCAACTCAGACACAAAGCCACAGACAATATGGATGATGGTC CTCTTACCCAGAAACTGGAGACAACTTTGCAGGCACCCTGGCTGCGTTCCTGATAATGCCTAGAGCATGTAGCAATGTTCAAGGCAGGTG CCTTGGAATCTGCTGTGAGTTATGCAGTACTAACCAAACAAAGTTGCTAAGGATGAGATGTTGCAACATTTCTTGTGTGCACTTTTTTTT CCTGACATCTTTTTTTCCTTTTAGGATTCATTAAGTCATATACTTAGTCCCCTGCAAAGAAGAAGACAGATGTCCCAGAGCAGACTGAAA GCGGGGCGTGGGATCCTAGGCAATGCAGGAGGGCAGGTGGGGGAGCTGTAGCAGAACAAGGGTGCTAGGTGGGAAGCTGGCTTGGGGACC ACACGTCTGGCTGACTCACAGGGCTATAGCAGCAGACAGAGAGCCAGATGCTCTACTGCAACTGGTGAGTTGGAGGCTTGCCCACATAAC ACTGTCTTTCTCTCCACCCGCCAAGCATCCAGGGCACAGGGCTGGTTGCTCACAGATTGAGCGAGGTAGCACCTCCTACCAGCACTCAAG >73096_73096_12_RBPMS-RILPL1_RBPMS_chr8_30242674_ENST00000538486_RILPL1_chr12_123970352_ENST00000376874_length(amino acids)=263AA_BP=127 MAPALPLQVALPGPDCLGAPLPARALPRLSLALPESPAAAVADSPREPQPNPSPTATAPAPAPAPQPAAPARGSPGARGRLQWASAPSPS PAPQPCPARRGRTGKMNNGGKAEKENTPSEANLQEEETEPVGEESISDAEKVAMDLKDPNRPRFTLQELRDVLHERNELKSKVFLLQEEL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for RBPMS-RILPL1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000287771 | + | 1 | 10 | 98_105 | 22.0 | 543.0 | RNA |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000320203 | + | 1 | 10 | 98_105 | 22.0 | 585.3333333333334 | RNA |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000339877 | + | 1 | 7 | 98_105 | 22.0 | 220.0 | RNA |
| Hgene | RBPMS | chr8:30242674 | chr12:123970352 | ENST00000397323 | + | 1 | 9 | 98_105 | 22.0 | 520.6666666666666 | RNA |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for RBPMS-RILPL1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for RBPMS-RILPL1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
