
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:REPS2-GDPD4 (FusionGDB2 ID:73475) |
Fusion Gene Summary for REPS2-GDPD4 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: REPS2-GDPD4 | Fusion gene ID: 73475 | Hgene | Tgene | Gene symbol | REPS2 | GDPD4 | Gene ID | 9185 | 220032 |
| Gene name | RALBP1 associated Eps domain containing 2 | glycerophosphodiester phosphodiesterase domain containing 4 | |
| Synonyms | POB1 | GDE6 | |
| Cytomap | Xp22.2 | 11q13.5 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ralBP1-associated Eps domain-containing protein 2RALBP1-interacting protein 2partner of Ral-binding protein 1partner of RalBP1 | glycerophosphodiester phosphodiesterase domain-containing protein 4GDPD domain containing proteinglycerophosphodiester phosphodiesterase 6ugpQ | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000303843, ENST00000357277, ENST00000380064, ENST00000469714, | ENST00000527489, ENST00000315938, ENST00000376217, | |
| Fusion gene scores | * DoF score | 13 X 9 X 10=1170 | 10 X 5 X 3=150 |
| # samples | 19 | 11 | |
| ** MAII score | log2(19/1170*10)=-2.62243720613982 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/150*10)=-0.447458976971221 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: REPS2 [Title/Abstract] AND GDPD4 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | REPS2(16965257)-GDPD4(76998465), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across REPS2 (5'-gene) Fusion gene breakpoints across REPS2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
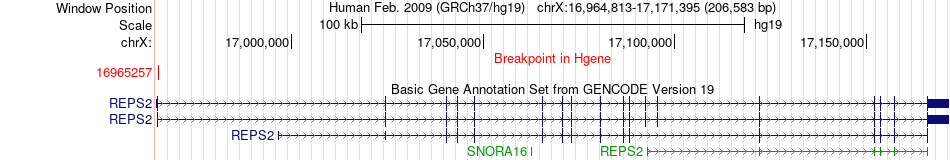 |
 Fusion gene breakpoints across GDPD4 (3'-gene) Fusion gene breakpoints across GDPD4 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
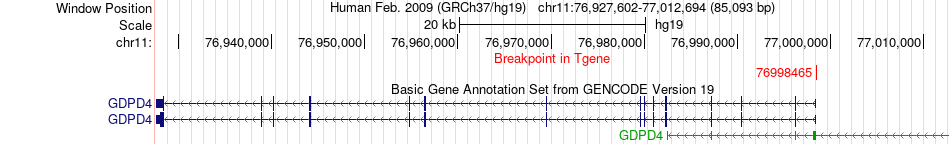 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | UCEC | TCGA-BK-A139-02A | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
Top |
Fusion Gene ORF analysis for REPS2-GDPD4 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000303843 | ENST00000527489 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| 5CDS-5UTR | ENST00000357277 | ENST00000527489 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| In-frame | ENST00000303843 | ENST00000315938 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| In-frame | ENST00000303843 | ENST00000376217 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| In-frame | ENST00000357277 | ENST00000315938 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| In-frame | ENST00000357277 | ENST00000376217 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| intron-3CDS | ENST00000380064 | ENST00000315938 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| intron-3CDS | ENST00000380064 | ENST00000376217 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| intron-3CDS | ENST00000469714 | ENST00000315938 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| intron-3CDS | ENST00000469714 | ENST00000376217 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| intron-5UTR | ENST00000380064 | ENST00000527489 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
| intron-5UTR | ENST00000469714 | ENST00000527489 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000357277 | REPS2 | chrX | 16965257 | + | ENST00000376217 | GDPD4 | chr11 | 76998465 | - | 2946 | 444 | 629 | 2566 | 645 |
| ENST00000357277 | REPS2 | chrX | 16965257 | + | ENST00000315938 | GDPD4 | chr11 | 76998465 | - | 2977 | 444 | 629 | 2257 | 542 |
| ENST00000303843 | REPS2 | chrX | 16965257 | + | ENST00000376217 | GDPD4 | chr11 | 76998465 | - | 2775 | 273 | 458 | 2395 | 645 |
| ENST00000303843 | REPS2 | chrX | 16965257 | + | ENST00000315938 | GDPD4 | chr11 | 76998465 | - | 2806 | 273 | 458 | 2086 | 542 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000357277 | ENST00000376217 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - | 0.008422268 | 0.99157774 |
| ENST00000357277 | ENST00000315938 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - | 0.013250249 | 0.9867497 |
| ENST00000303843 | ENST00000376217 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - | 0.006890988 | 0.993109 |
| ENST00000303843 | ENST00000315938 | REPS2 | chrX | 16965257 | + | GDPD4 | chr11 | 76998465 | - | 0.010914608 | 0.9890854 |
Top |
Fusion Genomic Features for REPS2-GDPD4 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
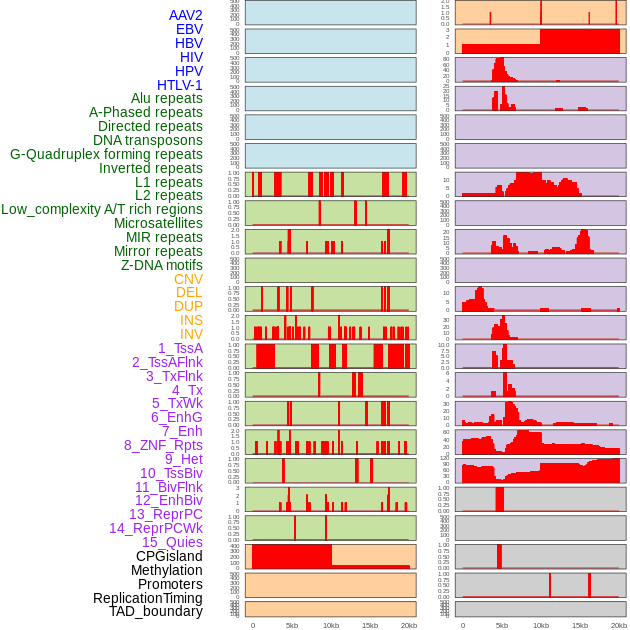 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for REPS2-GDPD4 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chrX:16965257/chr11:76998465) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 3_76 | 91 | 660.0 | Compositional bias | Note=Ala-rich |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 3_76 | 91 | 661.0 | Compositional bias | Note=Ala-rich |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 198_457 | 0 | 521.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 198_457 | 0 | 624.0 | Domain | Note=GP-PDE | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 131_162 | 0 | 521.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 184_468 | 0 | 521.0 | Topological domain | Extracellular | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 1_17 | 0 | 521.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 39_39 | 0 | 521.0 | Topological domain | Extracellular | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 490_623 | 0 | 521.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 61_69 | 0 | 521.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 91_109 | 0 | 521.0 | Topological domain | Extracellular | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 131_162 | 0 | 624.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 184_468 | 0 | 624.0 | Topological domain | Extracellular | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 1_17 | 0 | 624.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 39_39 | 0 | 624.0 | Topological domain | Extracellular | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 490_623 | 0 | 624.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 61_69 | 0 | 624.0 | Topological domain | Cytoplasmic | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 91_109 | 0 | 624.0 | Topological domain | Extracellular | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 110_130 | 0 | 521.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 163_183 | 0 | 521.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 18_38 | 0 | 521.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 40_60 | 0 | 521.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 469_489 | 0 | 521.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000315938 | -1 | 16 | 70_90 | 0 | 521.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 110_130 | 0 | 624.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 163_183 | 0 | 624.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 18_38 | 0 | 624.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 40_60 | 0 | 624.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 469_489 | 0 | 624.0 | Transmembrane | Helical | |
| Tgene | GDPD4 | chrX:16965257 | chr11:76998465 | ENST00000376217 | -1 | 17 | 70_90 | 0 | 624.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 328_339 | 91 | 660.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 328_339 | 91 | 661.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 601_657 | 91 | 660.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 601_657 | 91 | 661.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 477_524 | 91 | 660.0 | Compositional bias | Note=Pro-rich |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 477_524 | 91 | 661.0 | Compositional bias | Note=Pro-rich |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 282_373 | 91 | 660.0 | Domain | EH 2 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 315_350 | 91 | 660.0 | Domain | EF-hand |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 34_147 | 91 | 660.0 | Domain | EH 1 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 282_373 | 91 | 661.0 | Domain | EH 2 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 315_350 | 91 | 661.0 | Domain | EF-hand |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 34_147 | 91 | 661.0 | Domain | EH 1 |
Top |
Fusion Gene Sequence for REPS2-GDPD4 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >73475_73475_1_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000303843_GDPD4_chr11_76998465_ENST00000315938_length(transcript)=2806nt_BP=273nt ATGGAGGCGGCAGCGGCGGCGGCGGCGGCGGCAGCGGCAGCGGCAGCGGCGGGCGGGGGCTGTGGCTCCGGGCCGCCGCCGCTGCTGCTG AGCGAGGGCGAGCAGCAGTGCTACTCCGAGCTCTTCGCGCGCTGTGCCGGCGCCGCGGGCGGGGGCCCCGGGTCTGGGCCCCCCGAGGCC GCCAGAGTCGCCCCCGGCACGGCCACTGCGGCCGCCGGCCCCGTGGCTGACCTGTTTCGGGCATCGCAGCTGCCCGCCGAGACGCTGCAC CAGAAAGGGTGAATCATGTTTAAGAAGGTCAACAGGATTGAGCACAAGAGGAATAAGACAGAAAAACCAAAAAAGCTTAATCCATGGATT CCACGCATCTTCAACCACAAATGCTTTGTCACCTTCCTTACTGGCTGCTATAGTTGCCTCTGGAAATTCAGACAATGGGAAAAAACTAAA GTTGGATTTTGTTGCTGCACTTGGAAAGAGCAATTATTTTGCCGTGCCTGGTAATTTCATTTGTCTTGTCTCTGATGTTGCTGTTCCTGT GGATAGAAACATCCAGTGAATACTTTAACTTTGACTGGGTCACTTTTCTAGGAACAGGATACTGGTTTTTCTGGTCTATTTTTATTCTGA GTTTAGCTAGGATCCTGACTGCCTACTCTTCATTGCTGTTGTTACTTGGATTTTTGTTGCTTTGGGAAAGGATTGAACTATACCTGCACT TGTGTCATAAGATCCTGATTCTGCTAGTGATTCTTCTGTGTGTCATTTTAATGTTCATTATATGCAAATTCTGGAAAGAAAGGTGGCTGG TAGCTGGGCTGTCAATGCAGATCTTTGCTCCCTACGTGCACCTGGTCAGCATAACTGTGATGGTTATCCTTTTCTGGCCTGTGGCCTTCT ACGTGGCATGTTTGGAAAGAGAAGTTAGAATGAGAAGATACAGGATGACACATTCTGAGAAAAAAAGACTCAAGCAATGTAATGTAATCA CAAGGTTAAGAGGTCTACAAGTGCCTGTTGGATTACCCTTTCTTCTTATCCTTTTAGGTCTCTATTTGATGCCACTGGGGATTTATTCTC CCTGCATTCAGGAGAAGGAGAATTTGGGGCCCAAGCCAACCATCTTTGGACATAGAGGTGCACCCATGTTGGGGCCTGAGAATACCATGA TGTCCTTTGAGAAAGCTGTTGAACATGGAGCCCATGGATTGGAGACTGATATACACTTAAGTTATGATCATGTGCCTTTCCTCATGCATG ACTTTGACCTGAAAAGAACAACCAATATTGGGGAAGTTCAGCCAGAATCTGCCTGCGAGAACCCTGCCTTCTTCAACTGGGATTTCCTAT CGACTCTGAATGCAGGCAAATGGTTTGTAAAACCAGAGCTCAGGCCATTTTACAATATGAAACCTCTATCAGAGGCAGATAAAGAAAGAG CAAGAAATCAGTCAATTCCAACACTAGCTGATTTATTGACACTTGCAGAGAAGGAAAGAAAATTTGTGATATTTGATCTTCATCGCCCTC CACCAAAACATCCTCTCAGACACACATTTGTCCGCCAAGTAGTAAGCGTGATCCTTGCCTCTAAAATCGAGCAACATCTGATTTTTTGGT TGCCAGCTCATGATAGGCAATACGTCAGGTCCGTGGCTCCTGGTTTTCAGCATGTGGGCCGTTTAGTATCCATTGAAACCCTTGCTAAAA ACAATATCAGTATAATAAATGTTGACTACAAGAAGTTGTTCCCTAATGGGTTAAGAGATTATAAAGCAGCTAACATCCATATCAACGTAT ACACCGTCAATGAGCCTTGGCTTTTCTCACTGGCCTGGTGCTCCAGGATTAACTCAGTGACCACAGACAACATTGGGCTCCTGAGTCAGC TTGATCACCCTCACTTCTTCATGACACCAAAGTTCTATGTGTTCATGTGGCTCCTTGCAGATATCATTTCTGTGCTTTTTATTGTTGCCA TATTTTGTTTTCACTGGCGGAGAGAGACTGAAAAAGAAAAATTGTTTGAAACCTCCAGCACTCGCACAGATACTCAGAGTGGAAGTAAGA ATGAGGAAGATAGATAGATAGATAGATTTGCACATAGCCATGAAACCTCCTGTCCGAGTTGTGGAAGGTCCTTGGACTCTAGCAGCCCTC TACCCGGCTCTACCCAAGAGTGGAAAGGAACACCAAGGCCATTTCAATTTTGCAGCACCATCCAAGAAACTACTACCTATCAAGAATGCT GTCACACCTTTGAAGCCTGGCAAGCATGAGATTCAACCACCGATGCCCACCGTGGTCTTTGAACTCACCCAAGCACCCACCAGGCAGGCT ACAAGTGAGGCTACCTTTCAAACTACATTGCCCACCCTGAAGGTGGATAAGCCAACCATGCCCTCCATAGAGGTCCCCTATCCTTAAATA AGAGACCTCTAAGAGATCCCCATGGCTGAATACTCCAAATAATCCTATGTCACAATATATTCCATTGCATCTGACATAGCCTTCTGTGTT TCCAAAGAAGAATTAGCCACATCTCCCCTGCTTCTTCCTCTCTCTGCCTAAGGGTGGTGATGGTGGGGAGTGGTTGATAAAGGCACACAG AAGGGCCACGGAGGCTCTCAGGGGAACCATTGTGCTCTCGCTATCAAGAATGCATCTCAAGGGGCTAGGCAAGGGGAAAACTGCTGTGTG TGGTCTTGGAACTGGTTGCTGTCAGGTTTATCTCATCCTAACACCTCCATAATATTTTCTGCTAAAGGAGTGACTGTGCATATAAGTAAA >73475_73475_1_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000303843_GDPD4_chr11_76998465_ENST00000315938_length(amino acids)=542AA_BP= MLLHLERAIILPCLVISFVLSLMLLFLWIETSSEYFNFDWVTFLGTGYWFFWSIFILSLARILTAYSSLLLLLGFLLLWERIELYLHLCH KILILLVILLCVILMFIICKFWKERWLVAGLSMQIFAPYVHLVSITVMVILFWPVAFYVACLEREVRMRRYRMTHSEKKRLKQCNVITRL RGLQVPVGLPFLLILLGLYLMPLGIYSPCIQEKENLGPKPTIFGHRGAPMLGPENTMMSFEKAVEHGAHGLETDIHLSYDHVPFLMHDFD LKRTTNIGEVQPESACENPAFFNWDFLSTLNAGKWFVKPELRPFYNMKPLSEADKERARNQSIPTLADLLTLAEKERKFVIFDLHRPPPK HPLRHTFVRQVVSVILASKIEQHLIFWLPAHDRQYVRSVAPGFQHVGRLVSIETLAKNNISIINVDYKKLFPNGLRDYKAANIHINVYTV NEPWLFSLAWCSRINSVTTDNIGLLSQLDHPHFFMTPKFYVFMWLLADIISVLFIVAIFCFHWRRETEKEKLFETSSTRTDTQSGSKNEE -------------------------------------------------------------- >73475_73475_2_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000303843_GDPD4_chr11_76998465_ENST00000376217_length(transcript)=2775nt_BP=273nt ATGGAGGCGGCAGCGGCGGCGGCGGCGGCGGCAGCGGCAGCGGCAGCGGCGGGCGGGGGCTGTGGCTCCGGGCCGCCGCCGCTGCTGCTG AGCGAGGGCGAGCAGCAGTGCTACTCCGAGCTCTTCGCGCGCTGTGCCGGCGCCGCGGGCGGGGGCCCCGGGTCTGGGCCCCCCGAGGCC GCCAGAGTCGCCCCCGGCACGGCCACTGCGGCCGCCGGCCCCGTGGCTGACCTGTTTCGGGCATCGCAGCTGCCCGCCGAGACGCTGCAC CAGAAAGGGTGAATCATGTTTAAGAAGGTCAACAGGATTGAGCACAAGAGGAATAAGACAGAAAAACCAAAAAAGCTTAATCCATGGATT CCACGCATCTTCAACCACAAATGCTTTGTCACCTTCCTTACTGGCTGCTATAGTTGCCTCTGGAAATTCAGACAATGGGAAAAAACTAAA GTTGGATTTTGTTGCTGCACTTGGAAAGAGCAATTATTTTGCCGTGCCTGGTAATTTCATTTGTCTTGTCTCTGATGTTGCTGTTCCTGT GGATAGAAACATCCAGTGAATACTTTAACTTTGACTGGGTCACTTTTCTAGGAACAGGATACTGGTTTTTCTGGTCTATTTTTATTCTGA GTTTAGCTAGGATCCTGACTGCCTACTCTTCATTGCTGTTGTTACTTGGATTTTTGTTGCTTTGGGAAAGGATTGAACTATACCTGCACT TGTGTCATAAGATCCTGATTCTGCTAGTGATTCTTCTGTGTGTCATTTTAATGTTCATTATATGCAAATTCTGGAAAGAAAGGTGGCTGG TAGCTGGGCTGTCAATGCAGATCTTTGCTCCCTACGTGCACCTGGTCAGCATAACTGTGATGGTTATCCTTTTCTGGCCTGTGGCCTTCT ACGTGGCATGTTTGGAAAGAGAAGTTAGAATGAGAAGATACAGGATGACACATTCTGAGAAAAAAAGACTCAAGCAATGTAATGTAATCA CAAGGTTAAGAGGTCTACAAGTGCCTGTTGGATTACCCTTTCTTCTTATCCTTTTAGGTCTCTATTTGATGCCACTGGGGATTTATTCTC CCTGCATTCAGGAGAAGGAGAATTTGGGGCCCAAGCCAACCATCTTTGGACATAGAGGTGCACCCATGTTGGGGCCTGAGAATACCATGA TGTCCTTTGAGAAAGCTGTTGAACATGGAGCCCATGGATTGGAGACTGATATACACTTAAGTTATGATCATGTGCCTTTCCTCATGCATG ACTTTGACCTGAAAAGAACAACCAATATTGGGGAAGTTCAGCCAGAATCTGCCTGCGAGAACCCTGCCTTCTTCAACTGGGATTTCCTAT CGACTCTGAATGCAGGCAAATGGTTTGTAAAACCAGAGCTCAGGCCATTTTACAATATGAAACCTCTATCAGAGGCAGATAAAGAAAGAG CAAGAAATCAGTCAATTCCAACACTAGCTGATTTATTGACACTTGCAGAGAAGGAAAGAAAATTTGTGATATTTGATCTTCATCGCCCTC CACCAAAACATCCTCTCAGACACACATTTGTCCGCCAAGTAGTAAGCGTGATCCTTGCCTCTAAAATCGAGCAACATCTGATTTTTTGGT TGCCAGCTCATGATAGGCAATACGTCAGGTCCGTGGCTCCTGGTTTTCAGCATGTGGGCCGTTTAGTATCCATTGAAACCCTTGCTAAAA ACAATATCAGTATAATAAATGTTGACTACAAGAAGTTGTTCCCTAATGGGTTAAGAGATTATAAAGCAGCTAACATCCATATCAACGTAT ACACCGTCAATGAGCCTTGGCTTTTCTCACTGGCCTGGTGCTCCAGGATTAACTCAGTGACCACAGACAACATTGGGCTCCTGAGTCAGC TTGATCACCCTCACTTCTTCATGACACCAAAGTTCTATGTGTTCATGTGGCTCCTTGCAGATATCATTTCTGTGCTTTTTATTGTTGCCA TATTTTGTTTTCACTGGCGGAGAGAGACTGAAAAAGAAAAATTGTTTGAAACCTCCAGCACTCGCACAGATACTCAGAGTGGAAATTTGC ACATAGCCATGAAACCTCCTGTCCGAGTTGTGGAAGGTCCTTGGACTCTAGCAGCCCTCTACCCGGCTCTACCCAAGAGTGGAAAGGAAC ACCAAGGCCATTTCAATTTTGCAGCACCATCCAAGAAACTACTACCTATCAAGAATGCTGTCACACCTTTGAAGCCTGGCAAGCATGAGA TTCAACCACCGATGCCCACCGTGGTCTTTGAACTCACCCAAGCACCCACCAGGCAGGCTACAAGTGAGGCTACCTTTCAAACTACATTGC CCACCCTGAAGGTGGATAAGCCAACCATGCCCTCCATAGAGGTCCCCTATCCTTAAATAAGAGACCTCTAAGAGATCCCCATGGCTGAAT ACTCCAAATAATCCTATGTCACAATATATTCCATTGCATCTGACATAGCCTTCTGTGTTTCCAAAGAAGAATTAGCCACATCTCCCCTGC TTCTTCCTCTCTCTGCCTAAGGGTGGTGATGGTGGGGAGTGGTTGATAAAGGCACACAGAAGGGCCACGGAGGCTCTCAGGGGAACCATT GTGCTCTCGCTATCAAGAATGCATCTCAAGGGGCTAGGCAAGGGGAAAACTGCTGTGTGTGGTCTTGGAACTGGTTGCTGTCAGGTTTAT >73475_73475_2_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000303843_GDPD4_chr11_76998465_ENST00000376217_length(amino acids)=645AA_BP= MLLHLERAIILPCLVISFVLSLMLLFLWIETSSEYFNFDWVTFLGTGYWFFWSIFILSLARILTAYSSLLLLLGFLLLWERIELYLHLCH KILILLVILLCVILMFIICKFWKERWLVAGLSMQIFAPYVHLVSITVMVILFWPVAFYVACLEREVRMRRYRMTHSEKKRLKQCNVITRL RGLQVPVGLPFLLILLGLYLMPLGIYSPCIQEKENLGPKPTIFGHRGAPMLGPENTMMSFEKAVEHGAHGLETDIHLSYDHVPFLMHDFD LKRTTNIGEVQPESACENPAFFNWDFLSTLNAGKWFVKPELRPFYNMKPLSEADKERARNQSIPTLADLLTLAEKERKFVIFDLHRPPPK HPLRHTFVRQVVSVILASKIEQHLIFWLPAHDRQYVRSVAPGFQHVGRLVSIETLAKNNISIINVDYKKLFPNGLRDYKAANIHINVYTV NEPWLFSLAWCSRINSVTTDNIGLLSQLDHPHFFMTPKFYVFMWLLADIISVLFIVAIFCFHWRRETEKEKLFETSSTRTDTQSGNLHIA MKPPVRVVEGPWTLAALYPALPKSGKEHQGHFNFAAPSKKLLPIKNAVTPLKPGKHEIQPPMPTVVFELTQAPTRQATSEATFQTTLPTL -------------------------------------------------------------- >73475_73475_3_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000357277_GDPD4_chr11_76998465_ENST00000315938_length(transcript)=2977nt_BP=444nt CCGGGAGGAAGCGGCCGCGCGGCAGCTGCGGGGCGTGGGGGTGGTGGTGGCGGCGGCGGTGGTGGCGGCGGCGGCGGCGGCGGCAGCTGA GGCCGAGGAGGCGGTGGCTGTGGCGGACGCAGCAGCACCCCAGCTAGGGACAGGGCTCCGCCGCGCCCCCTTGCTGGCCCCATGGAGGCG GCAGCGGCGGCGGCGGCGGCGGCAGCGGCAGCGGCAGCGGCGGGCGGGGGCTGTGGCTCCGGGCCGCCGCCGCTGCTGCTGAGCGAGGGC GAGCAGCAGTGCTACTCCGAGCTCTTCGCGCGCTGTGCCGGCGCCGCGGGCGGGGGCCCCGGGTCTGGGCCCCCCGAGGCCGCCAGAGTC GCCCCCGGCACGGCCACTGCGGCCGCCGGCCCCGTGGCTGACCTGTTTCGGGCATCGCAGCTGCCCGCCGAGACGCTGCACCAGAAAGGG TGAATCATGTTTAAGAAGGTCAACAGGATTGAGCACAAGAGGAATAAGACAGAAAAACCAAAAAAGCTTAATCCATGGATTCCACGCATC TTCAACCACAAATGCTTTGTCACCTTCCTTACTGGCTGCTATAGTTGCCTCTGGAAATTCAGACAATGGGAAAAAACTAAAGTTGGATTT TGTTGCTGCACTTGGAAAGAGCAATTATTTTGCCGTGCCTGGTAATTTCATTTGTCTTGTCTCTGATGTTGCTGTTCCTGTGGATAGAAA CATCCAGTGAATACTTTAACTTTGACTGGGTCACTTTTCTAGGAACAGGATACTGGTTTTTCTGGTCTATTTTTATTCTGAGTTTAGCTA GGATCCTGACTGCCTACTCTTCATTGCTGTTGTTACTTGGATTTTTGTTGCTTTGGGAAAGGATTGAACTATACCTGCACTTGTGTCATA AGATCCTGATTCTGCTAGTGATTCTTCTGTGTGTCATTTTAATGTTCATTATATGCAAATTCTGGAAAGAAAGGTGGCTGGTAGCTGGGC TGTCAATGCAGATCTTTGCTCCCTACGTGCACCTGGTCAGCATAACTGTGATGGTTATCCTTTTCTGGCCTGTGGCCTTCTACGTGGCAT GTTTGGAAAGAGAAGTTAGAATGAGAAGATACAGGATGACACATTCTGAGAAAAAAAGACTCAAGCAATGTAATGTAATCACAAGGTTAA GAGGTCTACAAGTGCCTGTTGGATTACCCTTTCTTCTTATCCTTTTAGGTCTCTATTTGATGCCACTGGGGATTTATTCTCCCTGCATTC AGGAGAAGGAGAATTTGGGGCCCAAGCCAACCATCTTTGGACATAGAGGTGCACCCATGTTGGGGCCTGAGAATACCATGATGTCCTTTG AGAAAGCTGTTGAACATGGAGCCCATGGATTGGAGACTGATATACACTTAAGTTATGATCATGTGCCTTTCCTCATGCATGACTTTGACC TGAAAAGAACAACCAATATTGGGGAAGTTCAGCCAGAATCTGCCTGCGAGAACCCTGCCTTCTTCAACTGGGATTTCCTATCGACTCTGA ATGCAGGCAAATGGTTTGTAAAACCAGAGCTCAGGCCATTTTACAATATGAAACCTCTATCAGAGGCAGATAAAGAAAGAGCAAGAAATC AGTCAATTCCAACACTAGCTGATTTATTGACACTTGCAGAGAAGGAAAGAAAATTTGTGATATTTGATCTTCATCGCCCTCCACCAAAAC ATCCTCTCAGACACACATTTGTCCGCCAAGTAGTAAGCGTGATCCTTGCCTCTAAAATCGAGCAACATCTGATTTTTTGGTTGCCAGCTC ATGATAGGCAATACGTCAGGTCCGTGGCTCCTGGTTTTCAGCATGTGGGCCGTTTAGTATCCATTGAAACCCTTGCTAAAAACAATATCA GTATAATAAATGTTGACTACAAGAAGTTGTTCCCTAATGGGTTAAGAGATTATAAAGCAGCTAACATCCATATCAACGTATACACCGTCA ATGAGCCTTGGCTTTTCTCACTGGCCTGGTGCTCCAGGATTAACTCAGTGACCACAGACAACATTGGGCTCCTGAGTCAGCTTGATCACC CTCACTTCTTCATGACACCAAAGTTCTATGTGTTCATGTGGCTCCTTGCAGATATCATTTCTGTGCTTTTTATTGTTGCCATATTTTGTT TTCACTGGCGGAGAGAGACTGAAAAAGAAAAATTGTTTGAAACCTCCAGCACTCGCACAGATACTCAGAGTGGAAGTAAGAATGAGGAAG ATAGATAGATAGATAGATTTGCACATAGCCATGAAACCTCCTGTCCGAGTTGTGGAAGGTCCTTGGACTCTAGCAGCCCTCTACCCGGCT CTACCCAAGAGTGGAAAGGAACACCAAGGCCATTTCAATTTTGCAGCACCATCCAAGAAACTACTACCTATCAAGAATGCTGTCACACCT TTGAAGCCTGGCAAGCATGAGATTCAACCACCGATGCCCACCGTGGTCTTTGAACTCACCCAAGCACCCACCAGGCAGGCTACAAGTGAG GCTACCTTTCAAACTACATTGCCCACCCTGAAGGTGGATAAGCCAACCATGCCCTCCATAGAGGTCCCCTATCCTTAAATAAGAGACCTC TAAGAGATCCCCATGGCTGAATACTCCAAATAATCCTATGTCACAATATATTCCATTGCATCTGACATAGCCTTCTGTGTTTCCAAAGAA GAATTAGCCACATCTCCCCTGCTTCTTCCTCTCTCTGCCTAAGGGTGGTGATGGTGGGGAGTGGTTGATAAAGGCACACAGAAGGGCCAC GGAGGCTCTCAGGGGAACCATTGTGCTCTCGCTATCAAGAATGCATCTCAAGGGGCTAGGCAAGGGGAAAACTGCTGTGTGTGGTCTTGG AACTGGTTGCTGTCAGGTTTATCTCATCCTAACACCTCCATAATATTTTCTGCTAAAGGAGTGACTGTGCATATAAGTAAAGGCAACCTC >73475_73475_3_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000357277_GDPD4_chr11_76998465_ENST00000315938_length(amino acids)=542AA_BP= MLLHLERAIILPCLVISFVLSLMLLFLWIETSSEYFNFDWVTFLGTGYWFFWSIFILSLARILTAYSSLLLLLGFLLLWERIELYLHLCH KILILLVILLCVILMFIICKFWKERWLVAGLSMQIFAPYVHLVSITVMVILFWPVAFYVACLEREVRMRRYRMTHSEKKRLKQCNVITRL RGLQVPVGLPFLLILLGLYLMPLGIYSPCIQEKENLGPKPTIFGHRGAPMLGPENTMMSFEKAVEHGAHGLETDIHLSYDHVPFLMHDFD LKRTTNIGEVQPESACENPAFFNWDFLSTLNAGKWFVKPELRPFYNMKPLSEADKERARNQSIPTLADLLTLAEKERKFVIFDLHRPPPK HPLRHTFVRQVVSVILASKIEQHLIFWLPAHDRQYVRSVAPGFQHVGRLVSIETLAKNNISIINVDYKKLFPNGLRDYKAANIHINVYTV NEPWLFSLAWCSRINSVTTDNIGLLSQLDHPHFFMTPKFYVFMWLLADIISVLFIVAIFCFHWRRETEKEKLFETSSTRTDTQSGSKNEE -------------------------------------------------------------- >73475_73475_4_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000357277_GDPD4_chr11_76998465_ENST00000376217_length(transcript)=2946nt_BP=444nt CCGGGAGGAAGCGGCCGCGCGGCAGCTGCGGGGCGTGGGGGTGGTGGTGGCGGCGGCGGTGGTGGCGGCGGCGGCGGCGGCGGCAGCTGA GGCCGAGGAGGCGGTGGCTGTGGCGGACGCAGCAGCACCCCAGCTAGGGACAGGGCTCCGCCGCGCCCCCTTGCTGGCCCCATGGAGGCG GCAGCGGCGGCGGCGGCGGCGGCAGCGGCAGCGGCAGCGGCGGGCGGGGGCTGTGGCTCCGGGCCGCCGCCGCTGCTGCTGAGCGAGGGC GAGCAGCAGTGCTACTCCGAGCTCTTCGCGCGCTGTGCCGGCGCCGCGGGCGGGGGCCCCGGGTCTGGGCCCCCCGAGGCCGCCAGAGTC GCCCCCGGCACGGCCACTGCGGCCGCCGGCCCCGTGGCTGACCTGTTTCGGGCATCGCAGCTGCCCGCCGAGACGCTGCACCAGAAAGGG TGAATCATGTTTAAGAAGGTCAACAGGATTGAGCACAAGAGGAATAAGACAGAAAAACCAAAAAAGCTTAATCCATGGATTCCACGCATC TTCAACCACAAATGCTTTGTCACCTTCCTTACTGGCTGCTATAGTTGCCTCTGGAAATTCAGACAATGGGAAAAAACTAAAGTTGGATTT TGTTGCTGCACTTGGAAAGAGCAATTATTTTGCCGTGCCTGGTAATTTCATTTGTCTTGTCTCTGATGTTGCTGTTCCTGTGGATAGAAA CATCCAGTGAATACTTTAACTTTGACTGGGTCACTTTTCTAGGAACAGGATACTGGTTTTTCTGGTCTATTTTTATTCTGAGTTTAGCTA GGATCCTGACTGCCTACTCTTCATTGCTGTTGTTACTTGGATTTTTGTTGCTTTGGGAAAGGATTGAACTATACCTGCACTTGTGTCATA AGATCCTGATTCTGCTAGTGATTCTTCTGTGTGTCATTTTAATGTTCATTATATGCAAATTCTGGAAAGAAAGGTGGCTGGTAGCTGGGC TGTCAATGCAGATCTTTGCTCCCTACGTGCACCTGGTCAGCATAACTGTGATGGTTATCCTTTTCTGGCCTGTGGCCTTCTACGTGGCAT GTTTGGAAAGAGAAGTTAGAATGAGAAGATACAGGATGACACATTCTGAGAAAAAAAGACTCAAGCAATGTAATGTAATCACAAGGTTAA GAGGTCTACAAGTGCCTGTTGGATTACCCTTTCTTCTTATCCTTTTAGGTCTCTATTTGATGCCACTGGGGATTTATTCTCCCTGCATTC AGGAGAAGGAGAATTTGGGGCCCAAGCCAACCATCTTTGGACATAGAGGTGCACCCATGTTGGGGCCTGAGAATACCATGATGTCCTTTG AGAAAGCTGTTGAACATGGAGCCCATGGATTGGAGACTGATATACACTTAAGTTATGATCATGTGCCTTTCCTCATGCATGACTTTGACC TGAAAAGAACAACCAATATTGGGGAAGTTCAGCCAGAATCTGCCTGCGAGAACCCTGCCTTCTTCAACTGGGATTTCCTATCGACTCTGA ATGCAGGCAAATGGTTTGTAAAACCAGAGCTCAGGCCATTTTACAATATGAAACCTCTATCAGAGGCAGATAAAGAAAGAGCAAGAAATC AGTCAATTCCAACACTAGCTGATTTATTGACACTTGCAGAGAAGGAAAGAAAATTTGTGATATTTGATCTTCATCGCCCTCCACCAAAAC ATCCTCTCAGACACACATTTGTCCGCCAAGTAGTAAGCGTGATCCTTGCCTCTAAAATCGAGCAACATCTGATTTTTTGGTTGCCAGCTC ATGATAGGCAATACGTCAGGTCCGTGGCTCCTGGTTTTCAGCATGTGGGCCGTTTAGTATCCATTGAAACCCTTGCTAAAAACAATATCA GTATAATAAATGTTGACTACAAGAAGTTGTTCCCTAATGGGTTAAGAGATTATAAAGCAGCTAACATCCATATCAACGTATACACCGTCA ATGAGCCTTGGCTTTTCTCACTGGCCTGGTGCTCCAGGATTAACTCAGTGACCACAGACAACATTGGGCTCCTGAGTCAGCTTGATCACC CTCACTTCTTCATGACACCAAAGTTCTATGTGTTCATGTGGCTCCTTGCAGATATCATTTCTGTGCTTTTTATTGTTGCCATATTTTGTT TTCACTGGCGGAGAGAGACTGAAAAAGAAAAATTGTTTGAAACCTCCAGCACTCGCACAGATACTCAGAGTGGAAATTTGCACATAGCCA TGAAACCTCCTGTCCGAGTTGTGGAAGGTCCTTGGACTCTAGCAGCCCTCTACCCGGCTCTACCCAAGAGTGGAAAGGAACACCAAGGCC ATTTCAATTTTGCAGCACCATCCAAGAAACTACTACCTATCAAGAATGCTGTCACACCTTTGAAGCCTGGCAAGCATGAGATTCAACCAC CGATGCCCACCGTGGTCTTTGAACTCACCCAAGCACCCACCAGGCAGGCTACAAGTGAGGCTACCTTTCAAACTACATTGCCCACCCTGA AGGTGGATAAGCCAACCATGCCCTCCATAGAGGTCCCCTATCCTTAAATAAGAGACCTCTAAGAGATCCCCATGGCTGAATACTCCAAAT AATCCTATGTCACAATATATTCCATTGCATCTGACATAGCCTTCTGTGTTTCCAAAGAAGAATTAGCCACATCTCCCCTGCTTCTTCCTC TCTCTGCCTAAGGGTGGTGATGGTGGGGAGTGGTTGATAAAGGCACACAGAAGGGCCACGGAGGCTCTCAGGGGAACCATTGTGCTCTCG CTATCAAGAATGCATCTCAAGGGGCTAGGCAAGGGGAAAACTGCTGTGTGTGGTCTTGGAACTGGTTGCTGTCAGGTTTATCTCATCCTA >73475_73475_4_REPS2-GDPD4_REPS2_chrX_16965257_ENST00000357277_GDPD4_chr11_76998465_ENST00000376217_length(amino acids)=645AA_BP= MLLHLERAIILPCLVISFVLSLMLLFLWIETSSEYFNFDWVTFLGTGYWFFWSIFILSLARILTAYSSLLLLLGFLLLWERIELYLHLCH KILILLVILLCVILMFIICKFWKERWLVAGLSMQIFAPYVHLVSITVMVILFWPVAFYVACLEREVRMRRYRMTHSEKKRLKQCNVITRL RGLQVPVGLPFLLILLGLYLMPLGIYSPCIQEKENLGPKPTIFGHRGAPMLGPENTMMSFEKAVEHGAHGLETDIHLSYDHVPFLMHDFD LKRTTNIGEVQPESACENPAFFNWDFLSTLNAGKWFVKPELRPFYNMKPLSEADKERARNQSIPTLADLLTLAEKERKFVIFDLHRPPPK HPLRHTFVRQVVSVILASKIEQHLIFWLPAHDRQYVRSVAPGFQHVGRLVSIETLAKNNISIINVDYKKLFPNGLRDYKAANIHINVYTV NEPWLFSLAWCSRINSVTTDNIGLLSQLDHPHFFMTPKFYVFMWLLADIISVLFIVAIFCFHWRRETEKEKLFETSSTRTDTQSGNLHIA MKPPVRVVEGPWTLAALYPALPKSGKEHQGHFNFAAPSKKLLPIKNAVTPLKPGKHEIQPPMPTVVFELTQAPTRQATSEATFQTTLPTL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for REPS2-GDPD4 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 561_660 | 91.0 | 660.0 | ASAP1 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 561_660 | 91.0 | 661.0 | ASAP1 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000303843 | + | 1 | 18 | 514_660 | 91.0 | 660.0 | RALBP1 |
| Hgene | REPS2 | chrX:16965257 | chr11:76998465 | ENST00000357277 | + | 1 | 18 | 514_660 | 91.0 | 661.0 | RALBP1 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for REPS2-GDPD4 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for REPS2-GDPD4 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
