
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:RERE-SLC2A5 (FusionGDB2 ID:73530) |
Fusion Gene Summary for RERE-SLC2A5 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: RERE-SLC2A5 | Fusion gene ID: 73530 | Hgene | Tgene | Gene symbol | RERE | SLC2A5 | Gene ID | 473 | 6518 |
| Gene name | arginine-glutamic acid dipeptide repeats | solute carrier family 2 member 5 | |
| Synonyms | ARG|ARP|ATN1L|DNB1|NEDBEH | GLUT-5|GLUT5 | |
| Cytomap | 1p36.23 | 1p36.23 | |
| Type of gene | protein-coding | protein-coding | |
| Description | arginine-glutamic acid dipeptide repeats proteinarginine-glutamic acid dipeptide (RE) repeatsatrophin 2atrophin-1 like proteinatrophin-1 related protein | solute carrier family 2, facilitated glucose transporter member 5glucose transporter type 5, small intestineglucose transporter-like protein 5solute carrier family 2 (facilitated glucose/fructose transporter), member 5testicular tissue protein Li 81 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000337907, ENST00000377464, ENST00000400907, ENST00000400908, ENST00000460659, ENST00000476556, | ENST00000377414, ENST00000377424, ENST00000535586, ENST00000536305, | |
| Fusion gene scores | * DoF score | 27 X 28 X 11=8316 | 10 X 12 X 6=720 |
| # samples | 43 | 15 | |
| ** MAII score | log2(43/8316*10)=-4.27348119326891 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(15/720*10)=-2.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: RERE [Title/Abstract] AND SLC2A5 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | RERE(8555123)-SLC2A5(9118309), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | RERE-SLC2A5 seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. RERE-SLC2A5 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | SLC2A5 | GO:0015755 | fructose transmembrane transport | 28083649 |
| Tgene | SLC2A5 | GO:1990539 | fructose import across plasma membrane | 8333543 |
 Fusion gene breakpoints across RERE (5'-gene) Fusion gene breakpoints across RERE (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
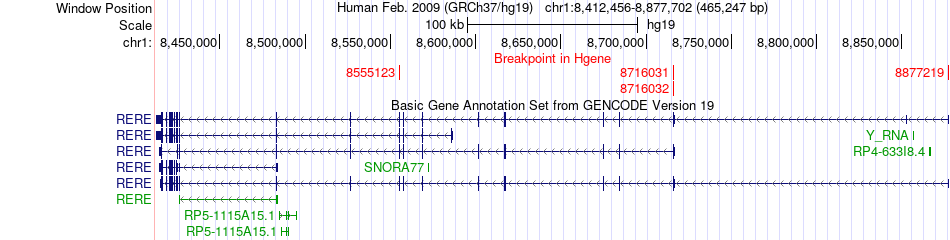 |
 Fusion gene breakpoints across SLC2A5 (3'-gene) Fusion gene breakpoints across SLC2A5 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
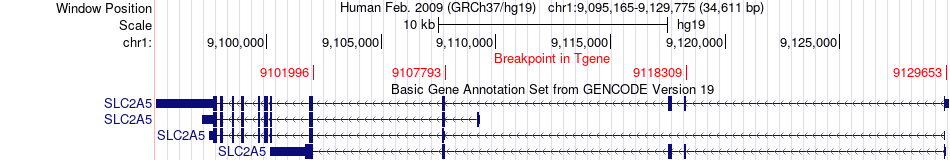 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LIHC | TCGA-DD-A73F-01A | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| ChimerDB4 | LUAD | TCGA-55-7576-01A | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| ChimerDB4 | UCEC | TCGA-AJ-A3BF-01A | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| ChimerDB4 | UCEC | TCGA-AJ-A3BF-01A | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| ChimerDB4 | UCEC | TCGA-FI-A2D2-01A | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| ChimerDB4 | UCEC | TCGA-FI-A2D2-01A | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| ChimerDB4 | UCEC | TCGA-FI-A2D2 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
Top |
Fusion Gene ORF analysis for RERE-SLC2A5 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| 5CDS-5UTR | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| 5CDS-5UTR | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| 5CDS-5UTR | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| 5CDS-5UTR | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| 5CDS-5UTR | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-5UTR | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| 5CDS-intron | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5CDS-intron | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| 5CDS-intron | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-3CDS | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-3CDS | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-3CDS | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-3CDS | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-5UTR | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-5UTR | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-5UTR | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-5UTR | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-5UTR | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-5UTR | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-5UTR | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-5UTR | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| 5UTR-intron | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-intron | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-intron | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| 5UTR-intron | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| Frame-shift | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000337907 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| Frame-shift | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| Frame-shift | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| Frame-shift | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| Frame-shift | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| Frame-shift | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| Frame-shift | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| In-frame | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000337907 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000337907 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000337907 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| In-frame | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| In-frame | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| In-frame | ENST00000400908 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| In-frame | ENST00000400908 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-3CDS | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-3CDS | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-3CDS | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-3CDS | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - |
| intron-5UTR | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000377464 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000377464 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000400907 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000400907 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000460659 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000476556 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-5UTR | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-5UTR | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - |
| intron-intron | ENST00000377464 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000377464 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000400907 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000400907 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000460659 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-intron | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-intron | ENST00000460659 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-intron | ENST00000460659 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000476556 | ENST00000377414 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-intron | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-intron | ENST00000476556 | ENST00000535586 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8877219 | - | SLC2A5 | chr1 | 9129653 | - |
| intron-intron | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - |
| intron-intron | ENST00000476556 | ENST00000536305 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9129653 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000337907 | RERE | chr1 | 8716031 | - | ENST00000377424 | SLC2A5 | chr1 | 9101996 | - | 4527 | 960 | 533 | 2047 | 504 |
| ENST00000337907 | RERE | chr1 | 8716031 | - | ENST00000536305 | SLC2A5 | chr1 | 9101996 | - | 2520 | 960 | 533 | 2047 | 504 |
| ENST00000337907 | RERE | chr1 | 8716031 | - | ENST00000535586 | SLC2A5 | chr1 | 9101996 | - | 2236 | 960 | 533 | 2047 | 504 |
| ENST00000400907 | RERE | chr1 | 8716031 | - | ENST00000377414 | SLC2A5 | chr1 | 9101996 | - | 2257 | 399 | 74 | 715 | 213 |
| ENST00000337907 | RERE | chr1 | 8716032 | - | ENST00000377424 | SLC2A5 | chr1 | 9101996 | - | 4527 | 960 | 533 | 2047 | 504 |
| ENST00000337907 | RERE | chr1 | 8716032 | - | ENST00000536305 | SLC2A5 | chr1 | 9101996 | - | 2520 | 960 | 533 | 2047 | 504 |
| ENST00000337907 | RERE | chr1 | 8716032 | - | ENST00000535586 | SLC2A5 | chr1 | 9101996 | - | 2236 | 960 | 533 | 2047 | 504 |
| ENST00000400907 | RERE | chr1 | 8716032 | - | ENST00000377414 | SLC2A5 | chr1 | 9101996 | - | 2257 | 399 | 74 | 715 | 213 |
| ENST00000400908 | RERE | chr1 | 8716032 | - | ENST00000377424 | SLC2A5 | chr1 | 9107793 | - | 4645 | 953 | 963 | 2165 | 400 |
| ENST00000400908 | RERE | chr1 | 8716032 | - | ENST00000536305 | SLC2A5 | chr1 | 9107793 | - | 2638 | 953 | 963 | 2165 | 400 |
| ENST00000337907 | RERE | chr1 | 8555123 | - | ENST00000377424 | SLC2A5 | chr1 | 9118309 | - | 5691 | 1739 | 533 | 3211 | 892 |
| ENST00000377464 | RERE | chr1 | 8555123 | - | ENST00000377424 | SLC2A5 | chr1 | 9118309 | - | 4407 | 455 | 155 | 1927 | 590 |
| ENST00000400907 | RERE | chr1 | 8555123 | - | ENST00000377424 | SLC2A5 | chr1 | 9118309 | - | 5130 | 1178 | 74 | 2650 | 858 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000337907 | ENST00000377424 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - | 0.00066843 | 0.9993316 |
| ENST00000337907 | ENST00000536305 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - | 0.001675154 | 0.9983248 |
| ENST00000337907 | ENST00000535586 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - | 0.002300386 | 0.99769956 |
| ENST00000400907 | ENST00000377414 | RERE | chr1 | 8716031 | - | SLC2A5 | chr1 | 9101996 | - | 0.008315794 | 0.9916842 |
| ENST00000337907 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - | 0.00066843 | 0.9993316 |
| ENST00000337907 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - | 0.001675154 | 0.9983248 |
| ENST00000337907 | ENST00000535586 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - | 0.002300386 | 0.99769956 |
| ENST00000400907 | ENST00000377414 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9101996 | - | 0.008315794 | 0.9916842 |
| ENST00000400908 | ENST00000377424 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - | 0.001540019 | 0.99845994 |
| ENST00000400908 | ENST00000536305 | RERE | chr1 | 8716032 | - | SLC2A5 | chr1 | 9107793 | - | 0.003851183 | 0.9961488 |
| ENST00000337907 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - | 0.00041347 | 0.9995865 |
| ENST00000377464 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - | 0.000668975 | 0.999331 |
| ENST00000400907 | ENST00000377424 | RERE | chr1 | 8555123 | - | SLC2A5 | chr1 | 9118309 | - | 0.000409569 | 0.99959046 |
Top |
Fusion Genomic Features for RERE-SLC2A5 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
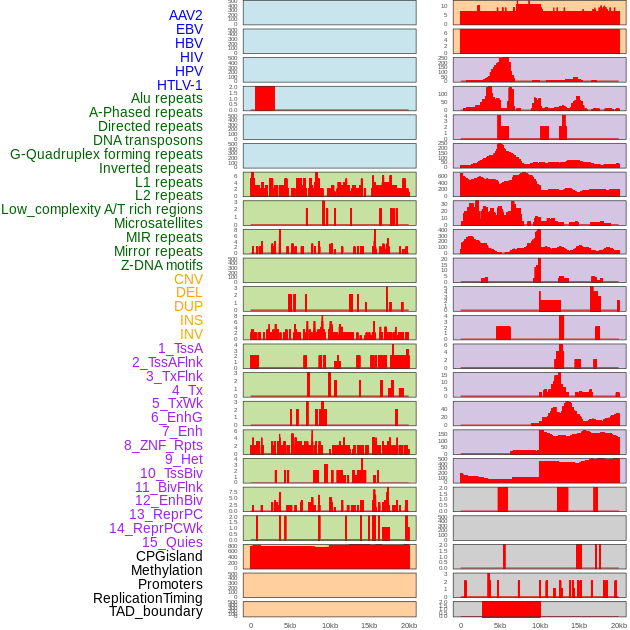 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for RERE-SLC2A5 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr1:8555123/chr1:9118309) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 103_283 | 368 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 103_283 | 368 | 1567.0 | Domain | BAH |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 296_298 | 11 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 419_420 | 11 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 296_298 | 11 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 419_420 | 11 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 296_298 | 139 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 419_420 | 139 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 296_298 | 139 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 419_420 | 139 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 296_298 | 139 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 419_420 | 139 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 296_298 | 139 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 419_420 | 139 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 296_298 | 97 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 419_420 | 97 | 245.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 296_298 | 97 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 419_420 | 97 | 502.0 | Region | Fructose binding | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 120_126 | 11 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 150_161 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 183_192 | 11 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 214_277 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 299_313 | 11 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 335_342 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 364_371 | 11 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 395_412 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 40_68 | 11 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 434_439 | 11 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 461_501 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 92_98 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 120_126 | 11 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 150_161 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 183_192 | 11 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 214_277 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 299_313 | 11 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 335_342 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 364_371 | 11 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 395_412 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 40_68 | 11 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 434_439 | 11 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 461_501 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 92_98 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 150_161 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 183_192 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 214_277 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 299_313 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 335_342 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 364_371 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 395_412 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 434_439 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 461_501 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 150_161 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 183_192 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 214_277 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 299_313 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 335_342 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 364_371 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 395_412 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 434_439 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 461_501 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 150_161 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 183_192 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 214_277 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 299_313 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 335_342 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 364_371 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 395_412 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 434_439 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 461_501 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 150_161 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 183_192 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 214_277 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 299_313 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 335_342 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 364_371 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 395_412 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 434_439 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 461_501 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 120_126 | 97 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 150_161 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 183_192 | 97 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 214_277 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 299_313 | 97 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 335_342 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 364_371 | 97 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 395_412 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 434_439 | 97 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 461_501 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 120_126 | 97 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 150_161 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 183_192 | 97 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 214_277 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 299_313 | 97 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 335_342 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 364_371 | 97 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 395_412 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 434_439 | 97 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 461_501 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 127_149 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 162_182 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 193_213 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 19_39 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 278_298 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 314_334 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 343_363 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 372_394 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 413_433 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 440_460 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 69_91 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 99_119 | 11 | 245.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 127_149 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 162_182 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 193_213 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 19_39 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 278_298 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 314_334 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 343_363 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 372_394 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 413_433 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 440_460 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 69_91 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 99_119 | 11 | 502.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 162_182 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 193_213 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 278_298 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 314_334 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 343_363 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 372_394 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 413_433 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 440_460 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 162_182 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 193_213 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 278_298 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 314_334 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 343_363 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 372_394 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 413_433 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 440_460 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 162_182 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 193_213 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 278_298 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 314_334 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 343_363 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 372_394 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 413_433 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 440_460 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 162_182 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 193_213 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 278_298 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 314_334 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 343_363 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 372_394 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 413_433 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 440_460 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 127_149 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 162_182 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 193_213 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 278_298 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 314_334 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 343_363 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 372_394 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 413_433 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 440_460 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 99_119 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 127_149 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 162_182 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 193_213 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 278_298 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 314_334 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 343_363 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 372_394 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 413_433 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D11 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 440_460 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D12 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 99_119 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D3 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 1156_1211 | 368 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 1156_1211 | 368 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 1156_1211 | 0 | 1013.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1156_1211 | 108 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1156_1211 | 108 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1156_1211 | 0 | 1013.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1156_1211 | 108 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1156_1211 | 108 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1156_1211 | 0 | 1013.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 1156_1211 | 108 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 1156_1211 | 108 | 1567.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 1156_1211 | 0 | 1013.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 1179_1204 | 368 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 1300_1322 | 368 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 1425_1445 | 368 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 1451_1510 | 368 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 738_1118 | 368 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 1179_1204 | 368 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 1300_1322 | 368 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 1425_1445 | 368 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 1451_1510 | 368 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 738_1118 | 368 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 1179_1204 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 1300_1322 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 1425_1445 | 0 | 1013.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 1451_1510 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 738_1118 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1179_1204 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1300_1322 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1425_1445 | 108 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1451_1510 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 738_1118 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1179_1204 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1300_1322 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1425_1445 | 108 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1451_1510 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 738_1118 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1179_1204 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1300_1322 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1425_1445 | 0 | 1013.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1451_1510 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 738_1118 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1179_1204 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1300_1322 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1425_1445 | 108 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 1451_1510 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 738_1118 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1179_1204 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1300_1322 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1425_1445 | 108 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 1451_1510 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 738_1118 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1179_1204 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1300_1322 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1425_1445 | 0 | 1013.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 1451_1510 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 738_1118 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 1179_1204 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 1300_1322 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 1425_1445 | 108 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 1451_1510 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 738_1118 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 1179_1204 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 1300_1322 | 108 | 1567.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 1425_1445 | 108 | 1567.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 1451_1510 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 738_1118 | 108 | 1567.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 1179_1204 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 1300_1322 | 0 | 1013.0 | Compositional bias | Note=Arg/Glu-rich (mixed charge) |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 1425_1445 | 0 | 1013.0 | Compositional bias | Note=His-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 1451_1510 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 738_1118 | 0 | 1013.0 | Compositional bias | Note=Pro-rich |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 284_387 | 368 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 391_443 | 368 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 284_387 | 368 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 391_443 | 368 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 103_283 | 0 | 1013.0 | Domain | BAH |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 284_387 | 0 | 1013.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 391_443 | 0 | 1013.0 | Domain | SANT |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 103_283 | 108 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 284_387 | 108 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 391_443 | 108 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 103_283 | 108 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 284_387 | 108 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 391_443 | 108 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 103_283 | 0 | 1013.0 | Domain | BAH |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 284_387 | 0 | 1013.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 391_443 | 0 | 1013.0 | Domain | SANT |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 103_283 | 108 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 284_387 | 108 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 391_443 | 108 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 103_283 | 108 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 284_387 | 108 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 391_443 | 108 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 103_283 | 0 | 1013.0 | Domain | BAH |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 284_387 | 0 | 1013.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 391_443 | 0 | 1013.0 | Domain | SANT |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 103_283 | 108 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 284_387 | 108 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 391_443 | 108 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 103_283 | 108 | 1567.0 | Domain | BAH |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 284_387 | 108 | 1567.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 391_443 | 108 | 1567.0 | Domain | SANT |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 103_283 | 0 | 1013.0 | Domain | BAH |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 284_387 | 0 | 1013.0 | Domain | ELM2 |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 391_443 | 0 | 1013.0 | Domain | SANT |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000337907 | - | 11 | 24 | 507_532 | 368 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000400908 | - | 10 | 23 | 507_532 | 368 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8555123 | chr1:9118309 | ENST00000476556 | - | 1 | 13 | 507_532 | 0 | 1013.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 507_532 | 108 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 507_532 | 108 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716031 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 507_532 | 0 | 1013.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000337907 | - | 3 | 24 | 507_532 | 108 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000400908 | - | 2 | 23 | 507_532 | 108 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716032 | chr1:9101996 | ENST00000476556 | - | 1 | 13 | 507_532 | 0 | 1013.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000337907 | - | 3 | 24 | 507_532 | 108 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000400908 | - | 2 | 23 | 507_532 | 108 | 1567.0 | Zinc finger | Note=GATA-type |
| Hgene | RERE | chr1:8716032 | chr1:9107793 | ENST00000476556 | - | 1 | 13 | 507_532 | 0 | 1013.0 | Zinc finger | Note=GATA-type |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377414 | 0 | 5 | 1_18 | 11 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8555123 | chr1:9118309 | ENST00000377424 | 0 | 12 | 1_18 | 11 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 120_126 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 1_18 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 40_68 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 92_98 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 120_126 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 1_18 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 40_68 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 92_98 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 120_126 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 1_18 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 40_68 | 139 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 92_98 | 139 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 120_126 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 1_18 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 40_68 | 139 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 92_98 | 139 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 1_18 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 40_68 | 97 | 245.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 92_98 | 97 | 245.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 1_18 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 40_68 | 97 | 502.0 | Topological domain | Extracellular | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 92_98 | 97 | 502.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 127_149 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 19_39 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 69_91 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377414 | 3 | 5 | 99_119 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 127_149 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 19_39 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 69_91 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8716031 | chr1:9101996 | ENST00000377424 | 3 | 12 | 99_119 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 127_149 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 19_39 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 69_91 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377414 | 3 | 5 | 99_119 | 139 | 245.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 127_149 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 19_39 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 69_91 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9101996 | ENST00000377424 | 3 | 12 | 99_119 | 139 | 502.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 19_39 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377414 | 2 | 5 | 69_91 | 97 | 245.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 19_39 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | SLC2A5 | chr1:8716032 | chr1:9107793 | ENST00000377424 | 2 | 12 | 69_91 | 97 | 502.0 | Transmembrane | Helical%3B Name%3D2 |
Top |
Fusion Gene Sequence for RERE-SLC2A5 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >73530_73530_1_RERE-SLC2A5_RERE_chr1_8555123_ENST00000337907_SLC2A5_chr1_9118309_ENST00000377424_length(transcript)=5691nt_BP=1739nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGACTGTGTGTATATCGAGAGTCGGAGGCCAA ACACACCGTATTTCATCTGTAGCATTCAAGACTTCAAACTGGTCCACAACTCCCAGGCCTGTTGCAGATCTCCAACTCCTGCTTTGTGTG ACCCCCCAGCATGCTCTCTGCCGGTGGCATCACAGCCACCACAGCATCTTTCTGAAGCCGGGAGAGGGCCTGTAGGGAGTAAGAGGGACC ATCTCCTCATGAACGTCAAATGGTACTACCGTCAATCTGAGGTTCCAGATTCTGTGTATCAGCATTTGGTTCAGGATCGACATAATGAAA ATGACTCTGGAAGAGAACTTGTCATTACAGACCCAGTTATCAAGAACCGAGAGCTCTTCATTTCTGATTACGTTGACACTTACCATGCTG CTGCCCTTAGAGGGAAGTGTAACATCTCCCATTTTTCTGACATATTTGCTGCTAGAGAGTTTAAAGCCCGAGTGGATTCATTTTTCTACA TATTAGGATATAACCCTGAGACAAGGAGGCTGAACAGTACCCAGGGGGAGATTCGTGTCGGTCCTAGTCATCAGGCCAAACTTCCAGATC TGCAACCATTTCCTTCTCCAGATGGTGATACAGTGACCCAACATGAGGAACTGGTCTGGATGCCTGGAGTTAACGACTGTGACCTCCTTA TGTACTTGAGGGCAGCAAGGAGCATGGCGGCATTTGCAGGAATGTGTGATGGAGGCTCTACAGAGGACGGCTGTGTCGCAGCCTCTCGGG ATGACACCACTCTGAATGCACTGAACACAAGGCTGACGCTTGTGCTTGCCCTGGCAACCCTGATAGCTGCCTTTGGGTCATCCTTCCAGT ATGGGTACAACGTGGCTGCTGTCAACTCCCCAGCACTGCTCATGCAACAATTTTACAATGAGACTTACTATGGTAGGACCGGTGAATTCA TGGAAGACTTCCCCTTGACGTTGCTGTGGTCTGTAACCGTGTCCATGTTTCCATTTGGAGGGTTTATCGGATCCCTCCTGGTCGGCCCCT TGGTGAATAAATTTGGCAGAAAAGGGGCCTTGCTGTTCAACAACATATTTTCTATCGTGCCTGCGATCTTAATGGGATGCAGCAGAGTCG CCACATCATTTGAGCTTATCATTATTTCCAGACTTTTGGTGGGAATATGTGCAGGTGTATCTTCCAACGTGGTCCCCATGTACTTAGGGG AGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTC TTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCT TCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCTGGGACT CTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCCGGATGC GCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACGCGGACC AGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCTTCTGCG CCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCACTGCAG CTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCGGGCCCA GTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACTGGCTCT CCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCCTCCTCA CCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATAAGGTGT CTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGAGCTGGT CTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGAATGCAG TCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCATGTTGGC CTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCCACCTTG ATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAGAGAAAC TAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAGAAATAC CTGTCTAGGAAGGTGTGATGTCAGAAACAATGACATCCAGAAAGCTGAGGAACAGGTTCCTGTGGAGACACTGAGTCAGAATTCTTCATC CTAAATTATTTTGTTAGTGGAAAATGGAATTGCTTCTGTGTAGTCAATAAAATGAACCTGATCACTTTTCAACAAGACGTGCTGCTGTTC TTTGAGCAAGTAGAAGGCAGGAGGAAGTTCCCTGCTCATTCAGGCAGTGTTTTTTTTGTTTTTATTTTTGCTTTTTTTTTTTTTTTTAAG ACGGAGTCTCGCTCTGTCACCCAGGCTGGAGTGCAGTGGTGCGATCTCAGCTCACTGCAAACTCCACCTCCCGGGTTCACACCATTCTCC TGCCTCAGCCTCCTGAGTAGCTGGGACTACAGGCGCCTGCCACCACGCCCGGCTAACTTTTTTGTATTTTTAGTAGAGACTGGGTTTCAC TGTGTTGGCCAGGATAGTCTCAATCTCCTGACCTCATGATCCGCCCACCTAGGCCCCTCAAAGTGCTGGGATTACAGGTGTGAGCCACTG CGCCCGGCCACTTCAGGCAGTTTGACTGCACGAGTGTCAGTTCCTTGAGGGCAGGGGCTTGGTCAGTTTTGTTTATGCCTGAATCCCCAG CATCTCGAAGAGGGCCTCTGGAGGCTGAGGTGGGAGGATTGCTTAAGCCTGGGAAGTAGAGGTTGCAGTGAGCTGAAATTGCACCCTATA CTCCAGCCTGGGCAACACAGTGAGATCCCATCTCAAAAAAAATTTTTTTAATTGGGCATGGTGACACGCCTTTGGGGTCCCAGCTATTTG GGAGGCTGAAATGGGAGGATCGCTTGAGCCTGGGAGGTCAAGGCTGCCCTGAGTCATGATTGCATCACTGCACACCAGCCTGCATGACAG AATGAGACCTTGTCTCAAATAAATAAATACATGTAAGATTGAGGAAGCTTCCAAATGTCCTCACATATGTAATGAAATCTTGAGGAGTCT TAAAATCTTGAGACTGTGGTGGGGGAGGAAGGGAAAAGGAACAGGGTCTTGATATGGTTTGGCTGTGTCCCCACCCAAATCTCATCTTGA ATTGTAGCTCCCATAATTCCCATGTGTTCTGGGAGGGACCCTGTGGGAGATAATTGAATCATGGGGGCAATTTCCCCCATACTGTTCTCA TGGTAGTGAATCAGCCTCATGAGATCTGATGGTTCTGTAAGGGGTAAAACCTTTTGCTTGGTTCTCATTCTCTCTTGCCTGCCACCAGGT AAGACGTGCCTTTTGCTTTCTGGCATGATTGTGAGGCCTCCCCAGCCATGTGGAATGGCGAGTCCATTAAATCTCTTTTTCTTTGCAATT TACCCAGTCTCTGATACATCTTTATCAGCAGCATGAAAATGGTCTAACACAGGTCTCTCCCCAAAAATATAGAGAGCTGGAGGTTTTTCT TAGAGATATGAATAAAGGGTGTGAGTGTGAGTGTATGTGCAAAAATATGGATAAAACCTTGAAAGTGCTGAATGGCCTGTAGCAAGAAGT GATGTCTTCTCATAGATGGGTGGCAGAGGCTGGTGGCCGGGAGAGAGAGCAGTGGGACTTGGGAGCACTTTTTGTGGGTATGTAATGGAG ATTCCACCAGCAACAAGCTAAACCACTCACAAGCGGGCCCAGAGGAAGTAAACAGAACACTGTGGTTGAACCATCCTTGGCCTTCTGCAG CAAGGATCTGCAGCCTGGAAGGGGCACAGAAACAACCAAAGATGAAGGCTCTCAGAGCTGAGGAGAGGCCAGGTCAGAATATCAGAAGTC AAGAAACTGCAGCACCTCTAACACAGGGGAGGCTATGAAGTTCATGGTTCTGGATGTCTGCTCCCTGGGTTCAAACCTTGGTCCTGCCAC ATTAGTTGTGCAACTTGGAAAGTTACTTGGCTTTTTTGTATCTCAGTCTTCTCTTCTGTTAAATGGCACTAATAGTAAATGTCTCACATG >73530_73530_1_RERE-SLC2A5_RERE_chr1_8555123_ENST00000337907_SLC2A5_chr1_9118309_ENST00000377424_length(amino acids)=892AA_BP=402 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGDCVYIESRRPNTPYFICSIQDFKLVHNSQACCRSPTPA LCDPPACSLPVASQPPQHLSEAGRGPVGSKRDHLLMNVKWYYRQSEVPDSVYQHLVQDRHNENDSGRELVITDPVIKNRELFISDYVDTY HAAALRGKCNISHFSDIFAAREFKARVDSFFYILGYNPETRRLNSTQGEIRVGPSHQAKLPDLQPFPSPDGDTVTQHEELVWMPGVNDCD LLMYLRAARSMAAFAGMCDGGSTEDGCVAASRDDTTLNALNTRLTLVLALATLIAAFGSSFQYGYNVAAVNSPALLMQQFYNETYYGRTG EFMEDFPLTLLWSVTVSMFPFGGFIGSLLVGPLVNKFGRKGALLFNNIFSIVPAILMGCSRVATSFELIIISRLLVGICAGVSSNVVPMY LGELAPKNLRGALGVVPQLFITVGILVAQIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRG WDSVDREVAEIRQEDEAEKAAGFISVLKLFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMT FCAVFVVELLGRRLLLLLGFSICLIACCVLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVH -------------------------------------------------------------- >73530_73530_2_RERE-SLC2A5_RERE_chr1_8555123_ENST00000377464_SLC2A5_chr1_9118309_ENST00000377424_length(transcript)=4407nt_BP=455nt GGCTTAGGTCATAGCACAAGCTCAGTGAAAGCTCTATGAGGTCTCTGAGACTGCATGTGCCTCCATTTTGAGTTGTTAGTGTAGAAAAGG GCAGAGTTTTAGCAGAAAGTAAAGGTTCAGGAATCCAGCTAGACTCACCAGAGCCCGTGTTAGTCATGGATGATCCATTCAGCCCCTGCA GGAGGCTGAACAGTACCCAGGGGGAGATTCGTGTCGGTCCTAGTCATCAGGCCAAACTTCCAGATCTGCAACCATTTCCTTCTCCAGATG GTGATACAGTGACCCAACATGAGGAACTGGTCTGGATGCCTGGAGTTAACGACTGTGACCTCCTTATGTACTTGAGGGCAGCAAGGAGCA TGGCGGCATTTGCAGGAATGTGTGATGGAGGCTCTACAGAGGACGGCTGTGTCGCAGCCTCTCGGGATGACACCACTCTGAATGCACTGA ACACAAGGCTGACGCTTGTGCTTGCCCTGGCAACCCTGATAGCTGCCTTTGGGTCATCCTTCCAGTATGGGTACAACGTGGCTGCTGTCA ACTCCCCAGCACTGCTCATGCAACAATTTTACAATGAGACTTACTATGGTAGGACCGGTGAATTCATGGAAGACTTCCCCTTGACGTTGC TGTGGTCTGTAACCGTGTCCATGTTTCCATTTGGAGGGTTTATCGGATCCCTCCTGGTCGGCCCCTTGGTGAATAAATTTGGCAGAAAAG GGGCCTTGCTGTTCAACAACATATTTTCTATCGTGCCTGCGATCTTAATGGGATGCAGCAGAGTCGCCACATCATTTGAGCTTATCATTA TTTCCAGACTTTTGGTGGGAATATGTGCAGGTGTATCTTCCAACGTGGTCCCCATGTACTTAGGGGAGCTGGCCCCTAAAAACCTGCGGG GGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTCTTCGGAATCTCCTTGCAAACGTAG ATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCTTCTTCCCCGAGAGCCCCAGGTACC TGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCTGGGACTCTGTGGACAGGGAGGTGGCCGAGA TCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCCGGATGCGCTCGCTGCGCTGGCAGCTGCTGT CCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACGCGGACCAGATCTACCTGAGCGCCGGCGTGC CGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCTTCTGCGCCGTGTTCGTGGTGGAGCTCCTGG GTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCACTGCAGCTCTGGCACTGCAGGACACAGTGT CCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCGGGCCCAGTCCCATACCCGCGCTGCTCATCA CTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACTGGCTCTCCAACTTCACCGTGGGCTTGATCT TCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCCTCCTCACCACCATCTACATCTTCTTGATTG TCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATAAGGTGTCTGAAGTGTACCCGGAAAAGGAGG AACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGAGCTGGTCTGCCAGGGGCTTCCCACTTTGGC TTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGAATGCAGTCCTCATCTCCAGCCTCCCCACCC CAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCATGTTGGCCTGTCACCAGACCCGAGTCAATTA AACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCCACCTTGATGGGTCAACCTTTGTGTGGCTCC TGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAGAGAAACTAACTTGGTGGCCCCGACAGGTCT TCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAGAAATACCTGTCTAGGAAGGTGTGATGTCAG AAACAATGACATCCAGAAAGCTGAGGAACAGGTTCCTGTGGAGACACTGAGTCAGAATTCTTCATCCTAAATTATTTTGTTAGTGGAAAA TGGAATTGCTTCTGTGTAGTCAATAAAATGAACCTGATCACTTTTCAACAAGACGTGCTGCTGTTCTTTGAGCAAGTAGAAGGCAGGAGG AAGTTCCCTGCTCATTCAGGCAGTGTTTTTTTTGTTTTTATTTTTGCTTTTTTTTTTTTTTTTAAGACGGAGTCTCGCTCTGTCACCCAG GCTGGAGTGCAGTGGTGCGATCTCAGCTCACTGCAAACTCCACCTCCCGGGTTCACACCATTCTCCTGCCTCAGCCTCCTGAGTAGCTGG GACTACAGGCGCCTGCCACCACGCCCGGCTAACTTTTTTGTATTTTTAGTAGAGACTGGGTTTCACTGTGTTGGCCAGGATAGTCTCAAT CTCCTGACCTCATGATCCGCCCACCTAGGCCCCTCAAAGTGCTGGGATTACAGGTGTGAGCCACTGCGCCCGGCCACTTCAGGCAGTTTG ACTGCACGAGTGTCAGTTCCTTGAGGGCAGGGGCTTGGTCAGTTTTGTTTATGCCTGAATCCCCAGCATCTCGAAGAGGGCCTCTGGAGG CTGAGGTGGGAGGATTGCTTAAGCCTGGGAAGTAGAGGTTGCAGTGAGCTGAAATTGCACCCTATACTCCAGCCTGGGCAACACAGTGAG ATCCCATCTCAAAAAAAATTTTTTTAATTGGGCATGGTGACACGCCTTTGGGGTCCCAGCTATTTGGGAGGCTGAAATGGGAGGATCGCT TGAGCCTGGGAGGTCAAGGCTGCCCTGAGTCATGATTGCATCACTGCACACCAGCCTGCATGACAGAATGAGACCTTGTCTCAAATAAAT AAATACATGTAAGATTGAGGAAGCTTCCAAATGTCCTCACATATGTAATGAAATCTTGAGGAGTCTTAAAATCTTGAGACTGTGGTGGGG GAGGAAGGGAAAAGGAACAGGGTCTTGATATGGTTTGGCTGTGTCCCCACCCAAATCTCATCTTGAATTGTAGCTCCCATAATTCCCATG TGTTCTGGGAGGGACCCTGTGGGAGATAATTGAATCATGGGGGCAATTTCCCCCATACTGTTCTCATGGTAGTGAATCAGCCTCATGAGA TCTGATGGTTCTGTAAGGGGTAAAACCTTTTGCTTGGTTCTCATTCTCTCTTGCCTGCCACCAGGTAAGACGTGCCTTTTGCTTTCTGGC ATGATTGTGAGGCCTCCCCAGCCATGTGGAATGGCGAGTCCATTAAATCTCTTTTTCTTTGCAATTTACCCAGTCTCTGATACATCTTTA TCAGCAGCATGAAAATGGTCTAACACAGGTCTCTCCCCAAAAATATAGAGAGCTGGAGGTTTTTCTTAGAGATATGAATAAAGGGTGTGA GTGTGAGTGTATGTGCAAAAATATGGATAAAACCTTGAAAGTGCTGAATGGCCTGTAGCAAGAAGTGATGTCTTCTCATAGATGGGTGGC AGAGGCTGGTGGCCGGGAGAGAGAGCAGTGGGACTTGGGAGCACTTTTTGTGGGTATGTAATGGAGATTCCACCAGCAACAAGCTAAACC ACTCACAAGCGGGCCCAGAGGAAGTAAACAGAACACTGTGGTTGAACCATCCTTGGCCTTCTGCAGCAAGGATCTGCAGCCTGGAAGGGG CACAGAAACAACCAAAGATGAAGGCTCTCAGAGCTGAGGAGAGGCCAGGTCAGAATATCAGAAGTCAAGAAACTGCAGCACCTCTAACAC AGGGGAGGCTATGAAGTTCATGGTTCTGGATGTCTGCTCCCTGGGTTCAAACCTTGGTCCTGCCACATTAGTTGTGCAACTTGGAAAGTT >73530_73530_2_RERE-SLC2A5_RERE_chr1_8555123_ENST00000377464_SLC2A5_chr1_9118309_ENST00000377424_length(amino acids)=590AA_BP=100 MDDPFSPCRRLNSTQGEIRVGPSHQAKLPDLQPFPSPDGDTVTQHEELVWMPGVNDCDLLMYLRAARSMAAFAGMCDGGSTEDGCVAASR DDTTLNALNTRLTLVLALATLIAAFGSSFQYGYNVAAVNSPALLMQQFYNETYYGRTGEFMEDFPLTLLWSVTVSMFPFGGFIGSLLVGP LVNKFGRKGALLFNNIFSIVPAILMGCSRVATSFELIIISRLLVGICAGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVAQIFG LRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLKLFRM RSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACCVLTA ALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAVICLL -------------------------------------------------------------- >73530_73530_3_RERE-SLC2A5_RERE_chr1_8555123_ENST00000400907_SLC2A5_chr1_9118309_ENST00000377424_length(transcript)=5130nt_BP=1178nt TGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGCGAATCATGACAGCGGACAAAG ACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAGCAAGAGAGAGTGAGAATTCAA GGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACGAGGACAATGACAACAATAGTG CCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGACAGATACCGGTGAGATAACAT CCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGACTGTGTGTATATCGAGAGTCGGAGGCCAAACACACCGTATTTCATCTGTA GCATTCAAGACTTCAAACTGGTCCACAACTCCCAGGCCTGTTGCAGATCTCCAACTCCTGCTTTGTGTGACCCCCCAGCATGCTCTCTGC CGGTGGCATCACAGCCACCACAGCATCTTTCTGAAGCCGGGAGAGGGCCTGTAGGGAGTAAGAGGGACCATCTCCTCATGAACGTCAAAT GGTACTACCGTCAATCTGAGGTTCCAGATTCTGTGTATCAGCATTTGGTTCAGGATCGACATAATGAAAATGACTCTGGAAGAGAACTTG TCATTACAGACCCAGTTATCAAGAACCGAGAGCTCTTCATTTCTGATTACGTTGACACTTACCATGCTGCTGCCCTTAGAGGGAAGTGTA ACATCTCCCATTTTTCTGACATATTTGCTGCTAGAGAGTTTAAAGCCCGAGTGGATTCATTTTTCTACATATTAGGATATAACCCTGAGA CAAGGAGGCTGAACAGTACCCAGGGGGAGATTCGTGTCGGTCCTAGTCATCAGGCCAAACTTCCAGATCTGCAACCATTTCCTTCTCCAG ATGGTGATACAGTGACCCAACATGAGGAACTGGTCTGGATGCCTGGAGTTAACGACTGTGACCTCCTTATGTACTTGAGGGCAGCAAGGA GCATGGCGGCATTTGCAGGAATGTGTGATGGAGGCTCTACAGAGGACGGCTGTGTCGCAGCCTCTCGGGATGACACCACTCTGAATGCAC TGAACACAAGGCTGACGCTTGTGCTTGCCCTGGCAACCCTGATAGCTGCCTTTGGGTCATCCTTCCAGTATGGGTACAACGTGGCTGCTG TCAACTCCCCAGCACTGCTCATGCAACAATTTTACAATGAGACTTACTATGGTAGGACCGGTGAATTCATGGAAGACTTCCCCTTGACGT TGCTGTGGTCTGTAACCGTGTCCATGTTTCCATTTGGAGGGTTTATCGGATCCCTCCTGGTCGGCCCCTTGGTGAATAAATTTGGCAGAA AAGGGGCCTTGCTGTTCAACAACATATTTTCTATCGTGCCTGCGATCTTAATGGGATGCAGCAGAGTCGCCACATCATTTGAGCTTATCA TTATTTCCAGACTTTTGGTGGGAATATGTGCAGGTGTATCTTCCAACGTGGTCCCCATGTACTTAGGGGAGCTGGCCCCTAAAAACCTGC GGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTCTTCGGAATCTCCTTGCAAACG TAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCTTCTTCCCCGAGAGCCCCAGGT ACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCTGGGACTCTGTGGACAGGGAGGTGGCCG AGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCCGGATGCGCTCGCTGCGCTGGCAGCTGC TGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACGCGGACCAGATCTACCTGAGCGCCGGCG TGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCTTCTGCGCCGTGTTCGTGGTGGAGCTCC TGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCACTGCAGCTCTGGCACTGCAGGACACAG TGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCGGGCCCAGTCCCATACCCGCGCTGCTCA TCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACTGGCTCTCCAACTTCACCGTGGGCTTGA TCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCCTCCTCACCACCATCTACATCTTCTTGA TTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATAAGGTGTCTGAAGTGTACCCGGAAAAGG AGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGAGCTGGTCTGCCAGGGGCTTCCCACTTT GGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGAATGCAGTCCTCATCTCCAGCCTCCCCA CCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCATGTTGGCCTGTCACCAGACCCGAGTCAA TTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCCACCTTGATGGGTCAACCTTTGTGTGGC TCCTGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAGAGAAACTAACTTGGTGGCCCCGACAGG TCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAGAAATACCTGTCTAGGAAGGTGTGATGT CAGAAACAATGACATCCAGAAAGCTGAGGAACAGGTTCCTGTGGAGACACTGAGTCAGAATTCTTCATCCTAAATTATTTTGTTAGTGGA AAATGGAATTGCTTCTGTGTAGTCAATAAAATGAACCTGATCACTTTTCAACAAGACGTGCTGCTGTTCTTTGAGCAAGTAGAAGGCAGG AGGAAGTTCCCTGCTCATTCAGGCAGTGTTTTTTTTGTTTTTATTTTTGCTTTTTTTTTTTTTTTTAAGACGGAGTCTCGCTCTGTCACC CAGGCTGGAGTGCAGTGGTGCGATCTCAGCTCACTGCAAACTCCACCTCCCGGGTTCACACCATTCTCCTGCCTCAGCCTCCTGAGTAGC TGGGACTACAGGCGCCTGCCACCACGCCCGGCTAACTTTTTTGTATTTTTAGTAGAGACTGGGTTTCACTGTGTTGGCCAGGATAGTCTC AATCTCCTGACCTCATGATCCGCCCACCTAGGCCCCTCAAAGTGCTGGGATTACAGGTGTGAGCCACTGCGCCCGGCCACTTCAGGCAGT TTGACTGCACGAGTGTCAGTTCCTTGAGGGCAGGGGCTTGGTCAGTTTTGTTTATGCCTGAATCCCCAGCATCTCGAAGAGGGCCTCTGG AGGCTGAGGTGGGAGGATTGCTTAAGCCTGGGAAGTAGAGGTTGCAGTGAGCTGAAATTGCACCCTATACTCCAGCCTGGGCAACACAGT GAGATCCCATCTCAAAAAAAATTTTTTTAATTGGGCATGGTGACACGCCTTTGGGGTCCCAGCTATTTGGGAGGCTGAAATGGGAGGATC GCTTGAGCCTGGGAGGTCAAGGCTGCCCTGAGTCATGATTGCATCACTGCACACCAGCCTGCATGACAGAATGAGACCTTGTCTCAAATA AATAAATACATGTAAGATTGAGGAAGCTTCCAAATGTCCTCACATATGTAATGAAATCTTGAGGAGTCTTAAAATCTTGAGACTGTGGTG GGGGAGGAAGGGAAAAGGAACAGGGTCTTGATATGGTTTGGCTGTGTCCCCACCCAAATCTCATCTTGAATTGTAGCTCCCATAATTCCC ATGTGTTCTGGGAGGGACCCTGTGGGAGATAATTGAATCATGGGGGCAATTTCCCCCATACTGTTCTCATGGTAGTGAATCAGCCTCATG AGATCTGATGGTTCTGTAAGGGGTAAAACCTTTTGCTTGGTTCTCATTCTCTCTTGCCTGCCACCAGGTAAGACGTGCCTTTTGCTTTCT GGCATGATTGTGAGGCCTCCCCAGCCATGTGGAATGGCGAGTCCATTAAATCTCTTTTTCTTTGCAATTTACCCAGTCTCTGATACATCT TTATCAGCAGCATGAAAATGGTCTAACACAGGTCTCTCCCCAAAAATATAGAGAGCTGGAGGTTTTTCTTAGAGATATGAATAAAGGGTG TGAGTGTGAGTGTATGTGCAAAAATATGGATAAAACCTTGAAAGTGCTGAATGGCCTGTAGCAAGAAGTGATGTCTTCTCATAGATGGGT GGCAGAGGCTGGTGGCCGGGAGAGAGAGCAGTGGGACTTGGGAGCACTTTTTGTGGGTATGTAATGGAGATTCCACCAGCAACAAGCTAA ACCACTCACAAGCGGGCCCAGAGGAAGTAAACAGAACACTGTGGTTGAACCATCCTTGGCCTTCTGCAGCAAGGATCTGCAGCCTGGAAG GGGCACAGAAACAACCAAAGATGAAGGCTCTCAGAGCTGAGGAGAGGCCAGGTCAGAATATCAGAAGTCAAGAAACTGCAGCACCTCTAA CACAGGGGAGGCTATGAAGTTCATGGTTCTGGATGTCTGCTCCCTGGGTTCAAACCTTGGTCCTGCCACATTAGTTGTGCAACTTGGAAA GTTACTTGGCTTTTTTGTATCTCAGTCTTCTCTTCTGTTAAATGGCACTAATAGTAAATGTCTCACATGTCCATTGTGAGAATTAAATGA >73530_73530_3_RERE-SLC2A5_RERE_chr1_8555123_ENST00000400907_SLC2A5_chr1_9118309_ENST00000377424_length(amino acids)=858AA_BP=368 MTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHSEDEDNDNNSATAEESTKKNKKKPPKKKSRYERTD TGEITSYITEDDVVYRPGDCVYIESRRPNTPYFICSIQDFKLVHNSQACCRSPTPALCDPPACSLPVASQPPQHLSEAGRGPVGSKRDHL LMNVKWYYRQSEVPDSVYQHLVQDRHNENDSGRELVITDPVIKNRELFISDYVDTYHAAALRGKCNISHFSDIFAAREFKARVDSFFYIL GYNPETRRLNSTQGEIRVGPSHQAKLPDLQPFPSPDGDTVTQHEELVWMPGVNDCDLLMYLRAARSMAAFAGMCDGGSTEDGCVAASRDD TTLNALNTRLTLVLALATLIAAFGSSFQYGYNVAAVNSPALLMQQFYNETYYGRTGEFMEDFPLTLLWSVTVSMFPFGGFIGSLLVGPLV NKFGRKGALLFNNIFSIVPAILMGCSRVATSFELIIISRLLVGICAGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVAQIFGLR NLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLKLFRMRS LRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACCVLTAAL ALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAVICLLTT -------------------------------------------------------------- >73530_73530_4_RERE-SLC2A5_RERE_chr1_8716031_ENST00000337907_SLC2A5_chr1_9101996_ENST00000377424_length(transcript)=4527nt_BP=960nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACT TAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCT TTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGC TGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCT GGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCC GGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACG CGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCT TCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCA CTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCG GGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACT GGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCC TCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATA AGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGA GCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGA ATGCAGTCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCAT GTTGGCCTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCC ACCTTGATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAG AGAAACTAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAG AAATACCTGTCTAGGAAGGTGTGATGTCAGAAACAATGACATCCAGAAAGCTGAGGAACAGGTTCCTGTGGAGACACTGAGTCAGAATTC TTCATCCTAAATTATTTTGTTAGTGGAAAATGGAATTGCTTCTGTGTAGTCAATAAAATGAACCTGATCACTTTTCAACAAGACGTGCTG CTGTTCTTTGAGCAAGTAGAAGGCAGGAGGAAGTTCCCTGCTCATTCAGGCAGTGTTTTTTTTGTTTTTATTTTTGCTTTTTTTTTTTTT TTTAAGACGGAGTCTCGCTCTGTCACCCAGGCTGGAGTGCAGTGGTGCGATCTCAGCTCACTGCAAACTCCACCTCCCGGGTTCACACCA TTCTCCTGCCTCAGCCTCCTGAGTAGCTGGGACTACAGGCGCCTGCCACCACGCCCGGCTAACTTTTTTGTATTTTTAGTAGAGACTGGG TTTCACTGTGTTGGCCAGGATAGTCTCAATCTCCTGACCTCATGATCCGCCCACCTAGGCCCCTCAAAGTGCTGGGATTACAGGTGTGAG CCACTGCGCCCGGCCACTTCAGGCAGTTTGACTGCACGAGTGTCAGTTCCTTGAGGGCAGGGGCTTGGTCAGTTTTGTTTATGCCTGAAT CCCCAGCATCTCGAAGAGGGCCTCTGGAGGCTGAGGTGGGAGGATTGCTTAAGCCTGGGAAGTAGAGGTTGCAGTGAGCTGAAATTGCAC CCTATACTCCAGCCTGGGCAACACAGTGAGATCCCATCTCAAAAAAAATTTTTTTAATTGGGCATGGTGACACGCCTTTGGGGTCCCAGC TATTTGGGAGGCTGAAATGGGAGGATCGCTTGAGCCTGGGAGGTCAAGGCTGCCCTGAGTCATGATTGCATCACTGCACACCAGCCTGCA TGACAGAATGAGACCTTGTCTCAAATAAATAAATACATGTAAGATTGAGGAAGCTTCCAAATGTCCTCACATATGTAATGAAATCTTGAG GAGTCTTAAAATCTTGAGACTGTGGTGGGGGAGGAAGGGAAAAGGAACAGGGTCTTGATATGGTTTGGCTGTGTCCCCACCCAAATCTCA TCTTGAATTGTAGCTCCCATAATTCCCATGTGTTCTGGGAGGGACCCTGTGGGAGATAATTGAATCATGGGGGCAATTTCCCCCATACTG TTCTCATGGTAGTGAATCAGCCTCATGAGATCTGATGGTTCTGTAAGGGGTAAAACCTTTTGCTTGGTTCTCATTCTCTCTTGCCTGCCA CCAGGTAAGACGTGCCTTTTGCTTTCTGGCATGATTGTGAGGCCTCCCCAGCCATGTGGAATGGCGAGTCCATTAAATCTCTTTTTCTTT GCAATTTACCCAGTCTCTGATACATCTTTATCAGCAGCATGAAAATGGTCTAACACAGGTCTCTCCCCAAAAATATAGAGAGCTGGAGGT TTTTCTTAGAGATATGAATAAAGGGTGTGAGTGTGAGTGTATGTGCAAAAATATGGATAAAACCTTGAAAGTGCTGAATGGCCTGTAGCA AGAAGTGATGTCTTCTCATAGATGGGTGGCAGAGGCTGGTGGCCGGGAGAGAGAGCAGTGGGACTTGGGAGCACTTTTTGTGGGTATGTA ATGGAGATTCCACCAGCAACAAGCTAAACCACTCACAAGCGGGCCCAGAGGAAGTAAACAGAACACTGTGGTTGAACCATCCTTGGCCTT CTGCAGCAAGGATCTGCAGCCTGGAAGGGGCACAGAAACAACCAAAGATGAAGGCTCTCAGAGCTGAGGAGAGGCCAGGTCAGAATATCA GAAGTCAAGAAACTGCAGCACCTCTAACACAGGGGAGGCTATGAAGTTCATGGTTCTGGATGTCTGCTCCCTGGGTTCAAACCTTGGTCC TGCCACATTAGTTGTGCAACTTGGAAAGTTACTTGGCTTTTTTGTATCTCAGTCTTCTCTTCTGTTAAATGGCACTAATAGTAAATGTCT >73530_73530_4_RERE-SLC2A5_RERE_chr1_8716031_ENST00000337907_SLC2A5_chr1_9101996_ENST00000377424_length(amino acids)=504AA_BP=142 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVA QIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLK LFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACC VLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAV -------------------------------------------------------------- >73530_73530_5_RERE-SLC2A5_RERE_chr1_8716031_ENST00000337907_SLC2A5_chr1_9101996_ENST00000535586_length(transcript)=2236nt_BP=960nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACT TAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCT TTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGC TGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCT GGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCC GGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACG CGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCT TCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCA CTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCG GGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACT GGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCC TCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATA AGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGA GCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGA >73530_73530_5_RERE-SLC2A5_RERE_chr1_8716031_ENST00000337907_SLC2A5_chr1_9101996_ENST00000535586_length(amino acids)=504AA_BP=142 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVA QIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLK LFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACC VLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAV -------------------------------------------------------------- >73530_73530_6_RERE-SLC2A5_RERE_chr1_8716031_ENST00000337907_SLC2A5_chr1_9101996_ENST00000536305_length(transcript)=2520nt_BP=960nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACT TAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCT TTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGC TGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCT GGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCC GGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACG CGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCT TCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCA CTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCG GGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACT GGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCC TCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATA AGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGA GCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGA ATGCAGTCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCAT GTTGGCCTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCC ACCTTGATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAG AGAAACTAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAG >73530_73530_6_RERE-SLC2A5_RERE_chr1_8716031_ENST00000337907_SLC2A5_chr1_9101996_ENST00000536305_length(amino acids)=504AA_BP=142 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVA QIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLK LFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACC VLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAV -------------------------------------------------------------- >73530_73530_7_RERE-SLC2A5_RERE_chr1_8716031_ENST00000400907_SLC2A5_chr1_9101996_ENST00000377414_length(transcript)=2257nt_BP=399nt TGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGCGAATCATGACAGCGGACAAAG ACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAGCAAGAGAGAGTGAGAATTCAA GGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACGAGGACAATGACAACAATAGTG CCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGACAGATACCGGTGAGATAACAT CCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACTTAGGGGAGCTGGCCCCTAAAA ACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTCTTCGGAATCTCCTTG CAAACGTAGATGGTGAGTTCAGGACATCTCGGGAGCACCCCCACCCCTTCACCACTACCCTTGGCCCCCTCCTTGTGTTCCAAAGCCACC ACCACAGGACAGGACTTTCTGCAGACTGGTCTCTTCTAACAGGCTGGATGTCCTTGGGGGGCCCATCCTGTCCCGAGCCAACATAGCAGG CCATGGACAAAGGCGCGATAAGGGACTGGCCAGCGTTTACTATCTAGTCCCTCTTAGGAGATGGATGAGGTGGCTTTGAGTGATGGGCTT CCCTGCCTGCTCAGCAGCTGTCCATATGGAGGCCCGTGGCATAGCAATGACAGCCACCAGTGTGCCAGGCTCTGTGCTTGGGGCGGACCC AGCATCAGTTCATAGGACCCTCCCAGCAACGCTGCTGCCTGGGACTCAGTGATTATCCCATTTTGCAGACCAGAAAGAGAAGTACCAGGA GTGAGGGTGGCCTGGCCCCATGGGGGAGGTGGTGAGGACATCCCAGACCCTGCCTCCACCTTCCCCAAGACAGCCTGGGGCCTGGCATGC GGCAGGTGCACAGACACCTGTAACCGGGGTTGTTTTTGCACGACTTTCCCTTTAAACTATCATCAGTCATCCCGTCAGCCAAAACATAAG CCAGTTCCCGCCAGGGTTCTACACACCGTCTGCAAGTCAGTCAAGTGGATCATCAAGCTCTGTGCTGCTTAGACACTCTCTGTAGAAAGT GCTTACCGTGAGCCAAGTGCCAAGTCTTTTTTAGTAACTCATGTATTCCTCAGAACAGCCCTATTAAGCCTGTACCAGTATCATACCCAT TTTGCAGGTGCATCACAGGTACAGAGAGATTGGAAGACTGGTCCAAGGTCTCATGAGGATGCAGACCCAGCGGCACAGACCTAGGTCCCT GTTTAAGTACCACATGCTCCTGCCTCTCCTTCACTCCTGCATGTGTTGGGGACACTTACGCCAAGGCGCCGCGTTCTCATTAGGAGCTGG GACCAGAAGTGAATAAGCCAGGTTCCTGTCTCAGGGAGCTCCATAGCAGGACTCAGAACCACACACGGCCCTCTAGGCATTTGTGAAGCT CTGTGCTTCATTTTTTTGCTTTGCCTCTAGTTTTGCCTTTGCAGTACCAATGCAGCCAGCCCATGTGTCCCCTCTATGTGGAATGTTAAC GATATTCCCACTGTTTCTGGTGTCCTTTCTGTAATCAGAGCTGCCGTGACCATTCCAGTTCAGGCATCCTGGTGGCCTGGCTTCTCTGGG CATAGAGCTGGGAGTGGGCAGTTGGGTCAGGCCGGGTATGCCCAGCTCCTCCAGCCCCTGCAAGACTGTCCTCTGAAGGACACGGGGACG TTCTCACCAGCACATGGTATTACCGGCCTCGTCGGTGTGGTCTGTCTGATGGGCGCAGGGGTGCGTCGCTGCTGATCCCTGCTGCAGGTG TTTCCCGAGTGTCCACCGTCGGGAGTTGCTGTTCTTAGGGCTGGGAAAGCTGTGCCCTCCTGGAGCCTGCACCCCACTCAGGATGACGGG GCTGGGGCCGGCTCTGCAGAGGGCGCCTTCGGGGTGGTGGGGAGGCCGCTGGTGCCAGCTCGCTGCCTTCCTCCCAGGCTGGCCGATCCT GCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAA >73530_73530_7_RERE-SLC2A5_RERE_chr1_8716031_ENST00000400907_SLC2A5_chr1_9101996_ENST00000377414_length(amino acids)=213AA_BP=108 MTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHSEDEDNDNNSATAEESTKKNKKKPPKKKSRYERTD TGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVAQIFGLRNLLANVDGEFRTSREHPHPFTTTLGPLL -------------------------------------------------------------- >73530_73530_8_RERE-SLC2A5_RERE_chr1_8716032_ENST00000337907_SLC2A5_chr1_9101996_ENST00000377424_length(transcript)=4527nt_BP=960nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACT TAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCT TTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGC TGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCT GGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCC GGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACG CGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCT TCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCA CTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCG GGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACT GGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCC TCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATA AGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGA GCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGA ATGCAGTCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCAT GTTGGCCTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCC ACCTTGATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAG AGAAACTAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAG AAATACCTGTCTAGGAAGGTGTGATGTCAGAAACAATGACATCCAGAAAGCTGAGGAACAGGTTCCTGTGGAGACACTGAGTCAGAATTC TTCATCCTAAATTATTTTGTTAGTGGAAAATGGAATTGCTTCTGTGTAGTCAATAAAATGAACCTGATCACTTTTCAACAAGACGTGCTG CTGTTCTTTGAGCAAGTAGAAGGCAGGAGGAAGTTCCCTGCTCATTCAGGCAGTGTTTTTTTTGTTTTTATTTTTGCTTTTTTTTTTTTT TTTAAGACGGAGTCTCGCTCTGTCACCCAGGCTGGAGTGCAGTGGTGCGATCTCAGCTCACTGCAAACTCCACCTCCCGGGTTCACACCA TTCTCCTGCCTCAGCCTCCTGAGTAGCTGGGACTACAGGCGCCTGCCACCACGCCCGGCTAACTTTTTTGTATTTTTAGTAGAGACTGGG TTTCACTGTGTTGGCCAGGATAGTCTCAATCTCCTGACCTCATGATCCGCCCACCTAGGCCCCTCAAAGTGCTGGGATTACAGGTGTGAG CCACTGCGCCCGGCCACTTCAGGCAGTTTGACTGCACGAGTGTCAGTTCCTTGAGGGCAGGGGCTTGGTCAGTTTTGTTTATGCCTGAAT CCCCAGCATCTCGAAGAGGGCCTCTGGAGGCTGAGGTGGGAGGATTGCTTAAGCCTGGGAAGTAGAGGTTGCAGTGAGCTGAAATTGCAC CCTATACTCCAGCCTGGGCAACACAGTGAGATCCCATCTCAAAAAAAATTTTTTTAATTGGGCATGGTGACACGCCTTTGGGGTCCCAGC TATTTGGGAGGCTGAAATGGGAGGATCGCTTGAGCCTGGGAGGTCAAGGCTGCCCTGAGTCATGATTGCATCACTGCACACCAGCCTGCA TGACAGAATGAGACCTTGTCTCAAATAAATAAATACATGTAAGATTGAGGAAGCTTCCAAATGTCCTCACATATGTAATGAAATCTTGAG GAGTCTTAAAATCTTGAGACTGTGGTGGGGGAGGAAGGGAAAAGGAACAGGGTCTTGATATGGTTTGGCTGTGTCCCCACCCAAATCTCA TCTTGAATTGTAGCTCCCATAATTCCCATGTGTTCTGGGAGGGACCCTGTGGGAGATAATTGAATCATGGGGGCAATTTCCCCCATACTG TTCTCATGGTAGTGAATCAGCCTCATGAGATCTGATGGTTCTGTAAGGGGTAAAACCTTTTGCTTGGTTCTCATTCTCTCTTGCCTGCCA CCAGGTAAGACGTGCCTTTTGCTTTCTGGCATGATTGTGAGGCCTCCCCAGCCATGTGGAATGGCGAGTCCATTAAATCTCTTTTTCTTT GCAATTTACCCAGTCTCTGATACATCTTTATCAGCAGCATGAAAATGGTCTAACACAGGTCTCTCCCCAAAAATATAGAGAGCTGGAGGT TTTTCTTAGAGATATGAATAAAGGGTGTGAGTGTGAGTGTATGTGCAAAAATATGGATAAAACCTTGAAAGTGCTGAATGGCCTGTAGCA AGAAGTGATGTCTTCTCATAGATGGGTGGCAGAGGCTGGTGGCCGGGAGAGAGAGCAGTGGGACTTGGGAGCACTTTTTGTGGGTATGTA ATGGAGATTCCACCAGCAACAAGCTAAACCACTCACAAGCGGGCCCAGAGGAAGTAAACAGAACACTGTGGTTGAACCATCCTTGGCCTT CTGCAGCAAGGATCTGCAGCCTGGAAGGGGCACAGAAACAACCAAAGATGAAGGCTCTCAGAGCTGAGGAGAGGCCAGGTCAGAATATCA GAAGTCAAGAAACTGCAGCACCTCTAACACAGGGGAGGCTATGAAGTTCATGGTTCTGGATGTCTGCTCCCTGGGTTCAAACCTTGGTCC TGCCACATTAGTTGTGCAACTTGGAAAGTTACTTGGCTTTTTTGTATCTCAGTCTTCTCTTCTGTTAAATGGCACTAATAGTAAATGTCT >73530_73530_8_RERE-SLC2A5_RERE_chr1_8716032_ENST00000337907_SLC2A5_chr1_9101996_ENST00000377424_length(amino acids)=504AA_BP=142 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVA QIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLK LFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACC VLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAV -------------------------------------------------------------- >73530_73530_9_RERE-SLC2A5_RERE_chr1_8716032_ENST00000337907_SLC2A5_chr1_9101996_ENST00000535586_length(transcript)=2236nt_BP=960nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACT TAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCT TTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGC TGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCT GGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCC GGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACG CGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCT TCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCA CTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCG GGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACT GGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCC TCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATA AGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGA GCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGA >73530_73530_9_RERE-SLC2A5_RERE_chr1_8716032_ENST00000337907_SLC2A5_chr1_9101996_ENST00000535586_length(amino acids)=504AA_BP=142 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVA QIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLK LFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACC VLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAV -------------------------------------------------------------- >73530_73530_10_RERE-SLC2A5_RERE_chr1_8716032_ENST00000337907_SLC2A5_chr1_9101996_ENST00000536305_length(transcript)=2520nt_BP=960nt AAAATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCG ACCGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCC GCCCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAA CACCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGGAGGTGATTGTCTAGGAGTAACTGTATGTGCGCTCAGCACCGGGTCACAACCAC CGTGGGATCCACTTACAGAGAAGATTTGTGGAGAGCTTCAGCCTGTTTATGGTGTTACTGAGAAAGGAATTGTGGCCATTTGATATAATT CTGGATGGAAGACCTCTTATTTGTATTACATTGAATTAGAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAAT TGAGTGAGAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGC GAATCATGACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAG CAAGAGAGAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACG AGGACAATGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGA CAGATACCGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACT TAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCT TTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGC TGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCCAAGAAAGCCCTACAGACGCTGCGCGGCT GGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCGGGCTTCATCTCCGTGCTGAAGCTGTTCC GGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTGTCGGGCGTCAACGCTATCTACTACTACG CGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGCACCGGGGCCGTGAACGTGGTCATGACCT TCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCCATCTGCCTCATAGCCTGCTGCGTGCTCA CTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTCATCTCCTACGTCATAGGACATGCCCTCG GGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCTGCCTTCATGGTGGGGGGCAGTGTGCACT GGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTACAGCTTCATTGTCTTCGCCGTGATCTGCC TCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAGATCAACCAGATTTTCACCAAGATGAATA AGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAACAGTGACTCTGGAGAGGAAGCCAGTGGA GCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATATCCAGAAATAAAACAACTCTGATGTGGA ATGCAGTCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTGTTCTTGAAGCTGGGCTGTCTCTCTCCAT GTTGGCCTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCCTTCGTGTGGCTCCTGGTAACGTGGCTCC ACCTTGATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGGTGAGATGGAAGGAATCAAATTTTGCCAG AGAAACTAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAATTCATCCTCTTACCAGATCCTTTTCCAG >73530_73530_10_RERE-SLC2A5_RERE_chr1_8716032_ENST00000337907_SLC2A5_chr1_9101996_ENST00000536305_length(amino acids)=504AA_BP=142 MELSEKFRRLKPRLTVYLSRRPSRERTEEGTWRIMTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHS EDEDNDNNSATAEESTKKNKKKPPKKKSRYERTDTGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVA QIFGLRNLLANVDGWPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLK LFRMRSLRWQLLSIIVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACC VLTAALALQDTVSWMPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAV -------------------------------------------------------------- >73530_73530_11_RERE-SLC2A5_RERE_chr1_8716032_ENST00000400907_SLC2A5_chr1_9101996_ENST00000377414_length(transcript)=2257nt_BP=399nt TGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGCGAATCATGACAGCGGACAAAG ACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAGCAAGAGAGAGTGAGAATTCAA GGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACGAGGACAATGACAACAATAGTG CCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGACAGATACCGGTGAGATAACAT CCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGGTGTATCTTCCAACGTGGTCCCCATGTACTTAGGGGAGCTGGCCCCTAAAA ACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATCACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTCTTCGGAATCTCCTTG CAAACGTAGATGGTGAGTTCAGGACATCTCGGGAGCACCCCCACCCCTTCACCACTACCCTTGGCCCCCTCCTTGTGTTCCAAAGCCACC ACCACAGGACAGGACTTTCTGCAGACTGGTCTCTTCTAACAGGCTGGATGTCCTTGGGGGGCCCATCCTGTCCCGAGCCAACATAGCAGG CCATGGACAAAGGCGCGATAAGGGACTGGCCAGCGTTTACTATCTAGTCCCTCTTAGGAGATGGATGAGGTGGCTTTGAGTGATGGGCTT CCCTGCCTGCTCAGCAGCTGTCCATATGGAGGCCCGTGGCATAGCAATGACAGCCACCAGTGTGCCAGGCTCTGTGCTTGGGGCGGACCC AGCATCAGTTCATAGGACCCTCCCAGCAACGCTGCTGCCTGGGACTCAGTGATTATCCCATTTTGCAGACCAGAAAGAGAAGTACCAGGA GTGAGGGTGGCCTGGCCCCATGGGGGAGGTGGTGAGGACATCCCAGACCCTGCCTCCACCTTCCCCAAGACAGCCTGGGGCCTGGCATGC GGCAGGTGCACAGACACCTGTAACCGGGGTTGTTTTTGCACGACTTTCCCTTTAAACTATCATCAGTCATCCCGTCAGCCAAAACATAAG CCAGTTCCCGCCAGGGTTCTACACACCGTCTGCAAGTCAGTCAAGTGGATCATCAAGCTCTGTGCTGCTTAGACACTCTCTGTAGAAAGT GCTTACCGTGAGCCAAGTGCCAAGTCTTTTTTAGTAACTCATGTATTCCTCAGAACAGCCCTATTAAGCCTGTACCAGTATCATACCCAT TTTGCAGGTGCATCACAGGTACAGAGAGATTGGAAGACTGGTCCAAGGTCTCATGAGGATGCAGACCCAGCGGCACAGACCTAGGTCCCT GTTTAAGTACCACATGCTCCTGCCTCTCCTTCACTCCTGCATGTGTTGGGGACACTTACGCCAAGGCGCCGCGTTCTCATTAGGAGCTGG GACCAGAAGTGAATAAGCCAGGTTCCTGTCTCAGGGAGCTCCATAGCAGGACTCAGAACCACACACGGCCCTCTAGGCATTTGTGAAGCT CTGTGCTTCATTTTTTTGCTTTGCCTCTAGTTTTGCCTTTGCAGTACCAATGCAGCCAGCCCATGTGTCCCCTCTATGTGGAATGTTAAC GATATTCCCACTGTTTCTGGTGTCCTTTCTGTAATCAGAGCTGCCGTGACCATTCCAGTTCAGGCATCCTGGTGGCCTGGCTTCTCTGGG CATAGAGCTGGGAGTGGGCAGTTGGGTCAGGCCGGGTATGCCCAGCTCCTCCAGCCCCTGCAAGACTGTCCTCTGAAGGACACGGGGACG TTCTCACCAGCACATGGTATTACCGGCCTCGTCGGTGTGGTCTGTCTGATGGGCGCAGGGGTGCGTCGCTGCTGATCCCTGCTGCAGGTG TTTCCCGAGTGTCCACCGTCGGGAGTTGCTGTTCTTAGGGCTGGGAAAGCTGTGCCCTCCTGGAGCCTGCACCCCACTCAGGATGACGGG GCTGGGGCCGGCTCTGCAGAGGGCGCCTTCGGGGTGGTGGGGAGGCCGCTGGTGCCAGCTCGCTGCCTTCCTCCCAGGCTGGCCGATCCT GCTGGGGCTGACCGGGGTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAA >73530_73530_11_RERE-SLC2A5_RERE_chr1_8716032_ENST00000400907_SLC2A5_chr1_9101996_ENST00000377414_length(amino acids)=213AA_BP=108 MTADKDKDKDKEKDRDRDRDREREKRDKARESENSRPRRSCTLEGGAKNYAESDHSEDEDNDNNSATAEESTKKNKKKPPKKKSRYERTD TGEITSYITEDDVVYRPGGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVAQIFGLRNLLANVDGEFRTSREHPHPFTTTLGPLL -------------------------------------------------------------- >73530_73530_12_RERE-SLC2A5_RERE_chr1_8716032_ENST00000400908_SLC2A5_chr1_9107793_ENST00000377424_length(transcript)=4645nt_BP=953nt CACAGACGCTCGCGAGCAACCCTCCGCGCTCCGGAGCCTCCTCAGCCTCCCGCACACCCCGAGACGCTCGCCCCGCTGGGCCTCGGTCCC CGCCTCCTCCCCCGTCCTTTTCCCCTCTCTCGCTCCCCACATTGTCTCTGCCTCATTTTCTCTCCTAGTTGAGGATTAAAAAAAAAAAAA AATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCGAC CGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCCGC CCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAACA CCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAATTGAGTGA GAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGCGAATCAT GACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAGCAAGAGA GAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACGAGGACAA TGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGACAGATAC CGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGAAAAGGGGCCTTGCTGTTCAACAACATATTTTCTATC GTGCCTGCGATCTTAATGGGATGCAGCAGAGTCGCCACATCATTTGAGCTTATCATTATTTCCAGACTTTTGGTGGGAATATGTGCAGGT GTATCTTCCAACGTGGTCCCCATGTACTTAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATC ACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGG GTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCC AAGAAAGCCCTACAGACGCTGCGCGGCTGGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCG GGCTTCATCTCCGTGCTGAAGCTGTTCCGGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTG TCGGGCGTCAACGCTATCTACTACTACGCGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGC ACCGGGGCCGTGAACGTGGTCATGACCTTCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCC ATCTGCCTCATAGCCTGCTGCGTGCTCACTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTC ATCTCCTACGTCATAGGACATGCCCTCGGGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCT GCCTTCATGGTGGGGGGCAGTGTGCACTGGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTAC AGCTTCATTGTCTTCGCCGTGATCTGCCTCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAG ATCAACCAGATTTTCACCAAGATGAATAAGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAA CAGTGACTCTGGAGAGGAAGCCAGTGGAGCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATA TCCAGAAATAAAACAACTCTGATGTGGAATGCAGTCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTG TTCTTGAAGCTGGGCTGTCTCTCTCCATGTTGGCCTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCC TTCGTGTGGCTCCTGGTAACGTGGCTCCACCTTGATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGG TGAGATGGAAGGAATCAAATTTTGCCAGAGAAACTAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAA TTCATCCTCTTACCAGATCCTTTTCCAGAAATACCTGTCTAGGAAGGTGTGATGTCAGAAACAATGACATCCAGAAAGCTGAGGAACAGG TTCCTGTGGAGACACTGAGTCAGAATTCTTCATCCTAAATTATTTTGTTAGTGGAAAATGGAATTGCTTCTGTGTAGTCAATAAAATGAA CCTGATCACTTTTCAACAAGACGTGCTGCTGTTCTTTGAGCAAGTAGAAGGCAGGAGGAAGTTCCCTGCTCATTCAGGCAGTGTTTTTTT TGTTTTTATTTTTGCTTTTTTTTTTTTTTTTAAGACGGAGTCTCGCTCTGTCACCCAGGCTGGAGTGCAGTGGTGCGATCTCAGCTCACT GCAAACTCCACCTCCCGGGTTCACACCATTCTCCTGCCTCAGCCTCCTGAGTAGCTGGGACTACAGGCGCCTGCCACCACGCCCGGCTAA CTTTTTTGTATTTTTAGTAGAGACTGGGTTTCACTGTGTTGGCCAGGATAGTCTCAATCTCCTGACCTCATGATCCGCCCACCTAGGCCC CTCAAAGTGCTGGGATTACAGGTGTGAGCCACTGCGCCCGGCCACTTCAGGCAGTTTGACTGCACGAGTGTCAGTTCCTTGAGGGCAGGG GCTTGGTCAGTTTTGTTTATGCCTGAATCCCCAGCATCTCGAAGAGGGCCTCTGGAGGCTGAGGTGGGAGGATTGCTTAAGCCTGGGAAG TAGAGGTTGCAGTGAGCTGAAATTGCACCCTATACTCCAGCCTGGGCAACACAGTGAGATCCCATCTCAAAAAAAATTTTTTTAATTGGG CATGGTGACACGCCTTTGGGGTCCCAGCTATTTGGGAGGCTGAAATGGGAGGATCGCTTGAGCCTGGGAGGTCAAGGCTGCCCTGAGTCA TGATTGCATCACTGCACACCAGCCTGCATGACAGAATGAGACCTTGTCTCAAATAAATAAATACATGTAAGATTGAGGAAGCTTCCAAAT GTCCTCACATATGTAATGAAATCTTGAGGAGTCTTAAAATCTTGAGACTGTGGTGGGGGAGGAAGGGAAAAGGAACAGGGTCTTGATATG GTTTGGCTGTGTCCCCACCCAAATCTCATCTTGAATTGTAGCTCCCATAATTCCCATGTGTTCTGGGAGGGACCCTGTGGGAGATAATTG AATCATGGGGGCAATTTCCCCCATACTGTTCTCATGGTAGTGAATCAGCCTCATGAGATCTGATGGTTCTGTAAGGGGTAAAACCTTTTG CTTGGTTCTCATTCTCTCTTGCCTGCCACCAGGTAAGACGTGCCTTTTGCTTTCTGGCATGATTGTGAGGCCTCCCCAGCCATGTGGAAT GGCGAGTCCATTAAATCTCTTTTTCTTTGCAATTTACCCAGTCTCTGATACATCTTTATCAGCAGCATGAAAATGGTCTAACACAGGTCT CTCCCCAAAAATATAGAGAGCTGGAGGTTTTTCTTAGAGATATGAATAAAGGGTGTGAGTGTGAGTGTATGTGCAAAAATATGGATAAAA CCTTGAAAGTGCTGAATGGCCTGTAGCAAGAAGTGATGTCTTCTCATAGATGGGTGGCAGAGGCTGGTGGCCGGGAGAGAGAGCAGTGGG ACTTGGGAGCACTTTTTGTGGGTATGTAATGGAGATTCCACCAGCAACAAGCTAAACCACTCACAAGCGGGCCCAGAGGAAGTAAACAGA ACACTGTGGTTGAACCATCCTTGGCCTTCTGCAGCAAGGATCTGCAGCCTGGAAGGGGCACAGAAACAACCAAAGATGAAGGCTCTCAGA GCTGAGGAGAGGCCAGGTCAGAATATCAGAAGTCAAGAAACTGCAGCACCTCTAACACAGGGGAGGCTATGAAGTTCATGGTTCTGGATG TCTGCTCCCTGGGTTCAAACCTTGGTCCTGCCACATTAGTTGTGCAACTTGGAAAGTTACTTGGCTTTTTTGTATCTCAGTCTTCTCTTC >73530_73530_12_RERE-SLC2A5_RERE_chr1_8716032_ENST00000400908_SLC2A5_chr1_9107793_ENST00000377424_length(amino acids)=400AA_BP= MLFNNIFSIVPAILMGCSRVATSFELIIISRLLVGICAGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVAQIFGLRNLLANVDG WPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLKLFRMRSLRWQLLSI IVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACCVLTAALALQDTVSW MPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAVICLLTTIYIFLIVP -------------------------------------------------------------- >73530_73530_13_RERE-SLC2A5_RERE_chr1_8716032_ENST00000400908_SLC2A5_chr1_9107793_ENST00000536305_length(transcript)=2638nt_BP=953nt CACAGACGCTCGCGAGCAACCCTCCGCGCTCCGGAGCCTCCTCAGCCTCCCGCACACCCCGAGACGCTCGCCCCGCTGGGCCTCGGTCCC CGCCTCCTCCCCCGTCCTTTTCCCCTCTCTCGCTCCCCACATTGTCTCTGCCTCATTTTCTCTCCTAGTTGAGGATTAAAAAAAAAAAAA AATCACCATCATCCTCAGGGAGCCCCCTCGCCCTCGGAAAACAAAACCAGACATGATCCTGGGCTCCTGCTTCTGAAAGATCTGCCCGAC CGAGGAGGAGGGAGCCTAGGTTCAAGTCGCAGGAAAGGTCACGGCAGGGAAGCCCCCTCAGCCCCTCCGCGTCGCCTCCCCGCGGGCCGC CCCTCTCGGCCCTGCGCCCCGCGGCCCCGCAGCCTCCGCGCGGCCTCCCGCCCCATCCCCGCCACTTTCCCCGGAGCTGGGCCGCGAACA CCTCACTATTGATGGTGTAGCGCTTTAGGGGAAGCATTTTTGAAACGTTGGGGTTGTTGGAGTGGTTGGATTTTCCCTGGAATTGAGTGA GAAATTCAGAAGACTGAAGCCCAGGCTTACTGTCTACCTTTCACGGAGGCCTAGCCGTGAGAGGACAGAAGAAGGCACGTGGCGAATCAT GACAGCGGACAAAGACAAAGACAAAGACAAAGAGAAGGACCGGGACCGAGACCGGGACCGAGAGAGAGAGAAAAGAGACAAAGCAAGAGA GAGTGAGAATTCAAGGCCACGCCGGAGCTGTACCTTGGAAGGAGGAGCCAAAAATTATGCTGAGAGTGATCACAGTGAAGACGAGGACAA TGACAACAATAGTGCCACCGCAGAGGAGTCCACGAAGAAGAATAAGAAGAAACCACCGAAAAAAAAGTCTCGTTATGAAAGGACAGATAC CGGTGAGATAACATCCTACATCACTGAAGATGATGTGGTCTACAGACCAGGAGAAAAGGGGCCTTGCTGTTCAACAACATATTTTCTATC GTGCCTGCGATCTTAATGGGATGCAGCAGAGTCGCCACATCATTTGAGCTTATCATTATTTCCAGACTTTTGGTGGGAATATGTGCAGGT GTATCTTCCAACGTGGTCCCCATGTACTTAGGGGAGCTGGCCCCTAAAAACCTGCGGGGGGCTCTCGGGGTGGTGCCCCAGCTCTTCATC ACTGTTGGCATCCTTGTGGCCCAGATCTTTGGTCTTCGGAATCTCCTTGCAAACGTAGATGGCTGGCCGATCCTGCTGGGGCTGACCGGG GTCCCCGCGGCGCTGCAGCTCCTTCTGCTGCCCTTCTTCCCCGAGAGCCCCAGGTACCTGCTGATTCAGAAGAAAGACGAAGCGGCCGCC AAGAAAGCCCTACAGACGCTGCGCGGCTGGGACTCTGTGGACAGGGAGGTGGCCGAGATCCGGCAGGAGGATGAGGCAGAGAAGGCCGCG GGCTTCATCTCCGTGCTGAAGCTGTTCCGGATGCGCTCGCTGCGCTGGCAGCTGCTGTCCATCATCGTCCTCATGGGCGGCCAGCAGCTG TCGGGCGTCAACGCTATCTACTACTACGCGGACCAGATCTACCTGAGCGCCGGCGTGCCGGAGGAGCACGTGCAGTACGTGACGGCCGGC ACCGGGGCCGTGAACGTGGTCATGACCTTCTGCGCCGTGTTCGTGGTGGAGCTCCTGGGTCGGAGGCTGCTGCTGCTGCTGGGCTTCTCC ATCTGCCTCATAGCCTGCTGCGTGCTCACTGCAGCTCTGGCACTGCAGGACACAGTGTCCTGGATGCCATACATCAGCATCGTCTGTGTC ATCTCCTACGTCATAGGACATGCCCTCGGGCCCAGTCCCATACCCGCGCTGCTCATCACTGAGATCTTCCTGCAGTCCTCTCGGCCATCT GCCTTCATGGTGGGGGGCAGTGTGCACTGGCTCTCCAACTTCACCGTGGGCTTGATCTTCCCGTTCATCCAGGAGGGCCTCGGCCCGTAC AGCTTCATTGTCTTCGCCGTGATCTGCCTCCTCACCACCATCTACATCTTCTTGATTGTCCCGGAGACCAAGGCCAAGACGTTCATAGAG ATCAACCAGATTTTCACCAAGATGAATAAGGTGTCTGAAGTGTACCCGGAAAAGGAGGAACTGAAAGAGCTTCCACCTGTCACTTCGGAA CAGTGACTCTGGAGAGGAAGCCAGTGGAGCTGGTCTGCCAGGGGCTTCCCACTTTGGCTTATTTTTCTGACTTCTAGCTGTCTGTGAATA TCCAGAAATAAAACAACTCTGATGTGGAATGCAGTCCTCATCTCCAGCCTCCCCACCCCAGTGGGAACTGTGCAAAGGGCTGCCTTGCTG TTCTTGAAGCTGGGCTGTCTCTCTCCATGTTGGCCTGTCACCAGACCCGAGTCAATTAAACAGCTGGTCCTCCACTTTGCTGGTTCAGCC TTCGTGTGGCTCCTGGTAACGTGGCTCCACCTTGATGGGTCAACCTTTGTGTGGCTCCTGGTAACATAACAACAACAGTTACTATAGTGG TGAGATGGAAGGAATCAAATTTTGCCAGAGAAACTAACTTGGTGGCCCCGACAGGTCTTCCGGGGCCATGGGCATTTGTTTAGAGCCAAA >73530_73530_13_RERE-SLC2A5_RERE_chr1_8716032_ENST00000400908_SLC2A5_chr1_9107793_ENST00000536305_length(amino acids)=400AA_BP= MLFNNIFSIVPAILMGCSRVATSFELIIISRLLVGICAGVSSNVVPMYLGELAPKNLRGALGVVPQLFITVGILVAQIFGLRNLLANVDG WPILLGLTGVPAALQLLLLPFFPESPRYLLIQKKDEAAAKKALQTLRGWDSVDREVAEIRQEDEAEKAAGFISVLKLFRMRSLRWQLLSI IVLMGGQQLSGVNAIYYYADQIYLSAGVPEEHVQYVTAGTGAVNVVMTFCAVFVVELLGRRLLLLLGFSICLIACCVLTAALALQDTVSW MPYISIVCVISYVIGHALGPSPIPALLITEIFLQSSRPSAFMVGGSVHWLSNFTVGLIFPFIQEGLGPYSFIVFAVICLLTTIYIFLIVP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for RERE-SLC2A5 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for RERE-SLC2A5 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for RERE-SLC2A5 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
