
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:RPRD1A-CEP192 (FusionGDB2 ID:77230) |
Fusion Gene Summary for RPRD1A-CEP192 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: RPRD1A-CEP192 | Fusion gene ID: 77230 | Hgene | Tgene | Gene symbol | RPRD1A | CEP192 | Gene ID | 55197 | 55125 |
| Gene name | regulation of nuclear pre-mRNA domain containing 1A | centrosomal protein 192 | |
| Synonyms | HsT3101|P15RS | PPP1R62 | |
| Cytomap | 18q12.2 | 18p11.21 | |
| Type of gene | protein-coding | protein-coding | |
| Description | regulation of nuclear pre-mRNA domain-containing protein 1ACDKN2B-related proteinCyclin-dependent kinase inhibitor 2B-related protein (p15INK4B-related protein)cyclin-dependent kinase 2B-inhibitor-related proteinp15INK4B-related protein | centrosomal protein of 192 kDa192 kDa centrosomal proteincentrosomal protein 192kDacep192/SPD-2protein phosphatase 1, regulatory subunit 62 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q8TEP8 | |
| Ensembl transtripts involved in fusion gene | ENST00000319040, ENST00000337059, ENST00000357384, ENST00000399022, ENST00000588737, ENST00000590898, ENST00000588459, | ENST00000540847, ENST00000325971, ENST00000506447, ENST00000430049, | |
| Fusion gene scores | * DoF score | 13 X 14 X 7=1274 | 4 X 4 X 4=64 |
| # samples | 17 | 4 | |
| ** MAII score | log2(17/1274*10)=-2.9057586261186 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/64*10)=-0.678071905112638 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: RPRD1A [Title/Abstract] AND CEP192 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | RPRD1A(33606863)-CEP192(13124631), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | RPRD1A-CEP192 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. RPRD1A-CEP192 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | CEP192 | GO:0010923 | negative regulation of phosphatase activity | 19389623 |
 Fusion gene breakpoints across RPRD1A (5'-gene) Fusion gene breakpoints across RPRD1A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
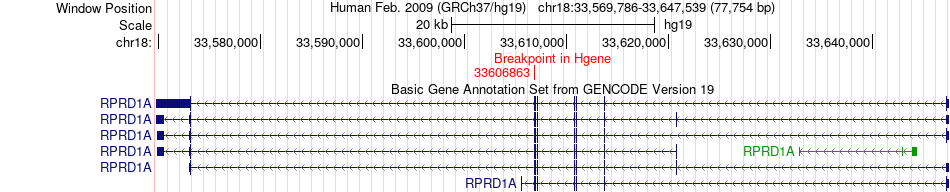 |
 Fusion gene breakpoints across CEP192 (3'-gene) Fusion gene breakpoints across CEP192 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
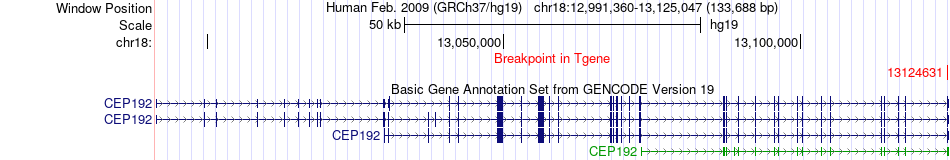 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | ERR315350 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
Top |
Fusion Gene ORF analysis for RPRD1A-CEP192 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000319040 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| 5CDS-3UTR | ENST00000337059 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| 5CDS-3UTR | ENST00000357384 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| 5CDS-3UTR | ENST00000399022 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| 5CDS-3UTR | ENST00000588737 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| 5CDS-3UTR | ENST00000590898 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000319040 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000319040 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000337059 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000357384 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000399022 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000399022 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000588737 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000588737 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| Frame-shift | ENST00000590898 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000319040 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000337059 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000337059 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000357384 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000357384 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000399022 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000588737 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000590898 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| In-frame | ENST00000590898 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| intron-3CDS | ENST00000588459 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| intron-3CDS | ENST00000588459 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| intron-3CDS | ENST00000588459 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
| intron-3UTR | ENST00000588459 | ENST00000540847 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000590898 | RPRD1A | chr18 | 33606863 | - | ENST00000506447 | CEP192 | chr18 | 13124631 | + | 1379 | 974 | 293 | 1033 | 246 |
| ENST00000590898 | RPRD1A | chr18 | 33606863 | - | ENST00000325971 | CEP192 | chr18 | 13124631 | + | 1391 | 974 | 293 | 1033 | 246 |
| ENST00000399022 | RPRD1A | chr18 | 33606863 | - | ENST00000430049 | CEP192 | chr18 | 13124631 | + | 1325 | 961 | 172 | 1020 | 282 |
| ENST00000337059 | RPRD1A | chr18 | 33606863 | - | ENST00000506447 | CEP192 | chr18 | 13124631 | + | 1086 | 681 | 0 | 740 | 246 |
| ENST00000337059 | RPRD1A | chr18 | 33606863 | - | ENST00000325971 | CEP192 | chr18 | 13124631 | + | 1098 | 681 | 0 | 740 | 246 |
| ENST00000357384 | RPRD1A | chr18 | 33606863 | - | ENST00000506447 | CEP192 | chr18 | 13124631 | + | 1346 | 941 | 152 | 1000 | 282 |
| ENST00000357384 | RPRD1A | chr18 | 33606863 | - | ENST00000325971 | CEP192 | chr18 | 13124631 | + | 1358 | 941 | 152 | 1000 | 282 |
| ENST00000588737 | RPRD1A | chr18 | 33606863 | - | ENST00000430049 | CEP192 | chr18 | 13124631 | + | 1367 | 1003 | 322 | 1062 | 246 |
| ENST00000319040 | RPRD1A | chr18 | 33606863 | - | ENST00000430049 | CEP192 | chr18 | 13124631 | + | 1305 | 941 | 152 | 1000 | 282 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000590898 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001409613 | 0.9985904 |
| ENST00000590898 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001337333 | 0.9986627 |
| ENST00000399022 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001835387 | 0.9981646 |
| ENST00000337059 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001343511 | 0.9986565 |
| ENST00000337059 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001291827 | 0.9987081 |
| ENST00000357384 | ENST00000506447 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001655046 | 0.9983449 |
| ENST00000357384 | ENST00000325971 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001548577 | 0.99845135 |
| ENST00000588737 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001538862 | 0.9984611 |
| ENST00000319040 | ENST00000430049 | RPRD1A | chr18 | 33606863 | - | CEP192 | chr18 | 13124631 | + | 0.001940006 | 0.9980599 |
Top |
Fusion Genomic Features for RPRD1A-CEP192 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| RPRD1A | chr18 | 33606862 | - | CEP192 | chr18 | 13124630 | + | 8.66E-06 | 0.9999913 |
| RPRD1A | chr18 | 33606862 | - | CEP192 | chr18 | 13124630 | + | 8.66E-06 | 0.9999913 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
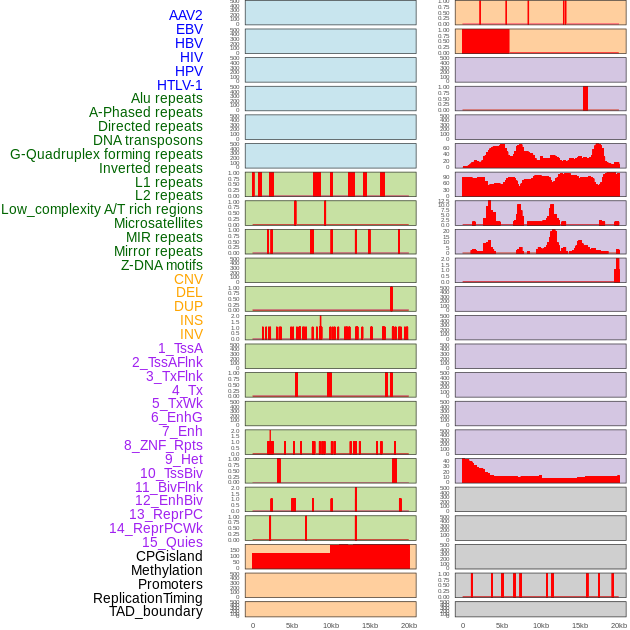 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
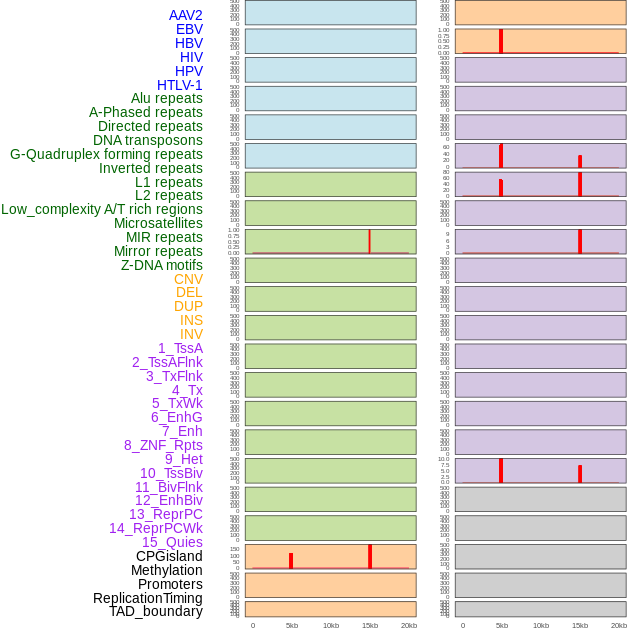 |
Top |
Fusion Protein Features for RPRD1A-CEP192 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr18:33606863/chr18:13124631) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CEP192 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Required for mitotic centrosome maturation and bipolar spindle assembly (PubMed:25042804, PubMed:17980596, PubMed:18207742). Appears to be a major regulator of pericentriolar material (PCM) recruitment, centrosome maturation, and centriole duplication (PubMed:25042804, PubMed:17980596, PubMed:18207742). Centrosome-specific activating scaffold for AURKA and PLK1 (PubMed:25042804). {ECO:0000269|PubMed:17980596, ECO:0000269|PubMed:18207742, ECO:0000269|PubMed:25042804}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000319040 | - | 6 | 7 | 2_133 | 263 | 290.0 | Domain | CID |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000337059 | - | 6 | 8 | 2_133 | 227 | 522.3333333333334 | Domain | CID |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000357384 | - | 6 | 8 | 2_133 | 263 | 471.6666666666667 | Domain | CID |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000399022 | - | 6 | 7 | 2_133 | 263 | 313.0 | Domain | CID |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000588737 | - | 7 | 8 | 2_133 | 227 | 277.0 | Domain | CID |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000590898 | - | 7 | 9 | 2_133 | 227 | 488.6666666666667 | Domain | CID |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000319040 | - | 6 | 7 | 244_286 | 263 | 290.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000337059 | - | 6 | 8 | 244_286 | 227 | 522.3333333333334 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000357384 | - | 6 | 8 | 244_286 | 263 | 471.6666666666667 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000399022 | - | 6 | 7 | 244_286 | 263 | 313.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000588737 | - | 7 | 8 | 244_286 | 227 | 277.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | RPRD1A | chr18:33606863 | chr18:13124631 | ENST00000590898 | - | 7 | 9 | 244_286 | 227 | 488.6666666666667 | Coiled coil | Ontology_term=ECO:0000255 |
Top |
Fusion Gene Sequence for RPRD1A-CEP192 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >77230_77230_1_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000319040_CEP192_chr18_13124631_ENST00000430049_length(transcript)=1305nt_BP=941nt TTGCTGGTTTGCGGCCGGTCTTGGATGAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGGGGGCGAGGCCGCAGCTAGGG CGGCGAAACTCTCCTCCCCTCGGCCCCACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGCCTTCTCTGAGGCGGCGCTGG AGAAGAAGCTGTCGGAGTTGAGCAACTCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCACCGTAAACACTCGCGTCCCA TCGTCACCGTGTGGGAGCGGGAGCTGCGGAAAGCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGCCAATGATGTCATACAGAACA GCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTCAAGTGAAACTGATGAAAGTT GTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGAACAACTTAAACAAGCTCTGT ATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTCTCTGGGATCTCCAAGTGAAC CACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGTTCATCAGAGGATAGCTTCTT TACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTCCAAAATGGTAGAGGATGCGT GTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAATGTTAGCAGATTTTCTTCGTT GTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGCAGTTCAAACCGAAGTCCGCA GGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTGAAGCTCTTGGAAAAAATTAA CTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACAATTATGCTTTTGTATATATA TTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAATATTATGTATCTAGCCCATAG >77230_77230_1_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000319040_CEP192_chr18_13124631_ENST00000430049_length(amino acids)=282AA_BP=263 MSAFSEAALEKKLSELSNSQQSVQTLSLWLIHHRKHSRPIVTVWERELRKAKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFK HVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLKQALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGD AAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMVEDACMLLADYNGRLAAEIDDRKQLTRMLADFLRCQKEALAEKEHKLESPALHQH -------------------------------------------------------------- >77230_77230_2_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000337059_CEP192_chr18_13124631_ENST00000325971_length(transcript)=1098nt_BP=681nt ATGAGAAATTCCAATTGGAGATTTTGTCAAACAGGAATTTACTCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGCCAATGATGTC ATACAGAACAGCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTCAAGTGAAACT GATGAAAGTTGTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGAACAACTTAAA CAAGCTCTGTATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTCTCTGGGATCT CCAAGTGAACCACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGTTCATCAGAGG ATAGCTTCTTTACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTCCAAAATGGTA GAGGATGCGTGTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAATGTTAGCAGAT TTTCTTCGTTGTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGCAGTTCAAACC GAAGTCCGCAGGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTGAAGCTCTTGG AAAAAATTAACTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACAATTATGCTTT TGTATATATATTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAATATTATGTATC TAGCCCATAGTATTGTACTTAACTTTTACAGGTGAGAAGAGAGTTCTGTGTTTGCATTGATTATGATATTCTGAATAAATATGGAATATA >77230_77230_2_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000337059_CEP192_chr18_13124631_ENST00000325971_length(amino acids)=246AA_BP=227 MRNSNWRFCQTGIYSKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFKHVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLK QALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGDAAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMV -------------------------------------------------------------- >77230_77230_3_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000337059_CEP192_chr18_13124631_ENST00000506447_length(transcript)=1086nt_BP=681nt ATGAGAAATTCCAATTGGAGATTTTGTCAAACAGGAATTTACTCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGCCAATGATGTC ATACAGAACAGCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTCAAGTGAAACT GATGAAAGTTGTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGAACAACTTAAA CAAGCTCTGTATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTCTCTGGGATCT CCAAGTGAACCACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGTTCATCAGAGG ATAGCTTCTTTACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTCCAAAATGGTA GAGGATGCGTGTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAATGTTAGCAGAT TTTCTTCGTTGTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGCAGTTCAAACC GAAGTCCGCAGGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTGAAGCTCTTGG AAAAAATTAACTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACAATTATGCTTT TGTATATATATTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAATATTATGTATC TAGCCCATAGTATTGTACTTAACTTTTACAGGTGAGAAGAGAGTTCTGTGTTTGCATTGATTATGATATTCTGAATAAATATGGAATATA >77230_77230_3_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000337059_CEP192_chr18_13124631_ENST00000506447_length(amino acids)=246AA_BP=227 MRNSNWRFCQTGIYSKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFKHVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLK QALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGDAAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMV -------------------------------------------------------------- >77230_77230_4_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000357384_CEP192_chr18_13124631_ENST00000325971_length(transcript)=1358nt_BP=941nt TTGCTGGTTTGCGGCCGGTCTTGGATGAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGGGGGCGAGGCCGCAGCTAGGG CGGCGAAACTCTCCTCCCCTCGGCCCCACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGCCTTCTCTGAGGCGGCGCTGG AGAAGAAGCTGTCGGAGTTGAGCAACTCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCACCGTAAACACTCGCGTCCCA TCGTCACCGTGTGGGAGCGGGAGCTGCGGAAAGCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGCCAATGATGTCATACAGAACA GCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTCAAGTGAAACTGATGAAAGTT GTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGAACAACTTAAACAAGCTCTGT ATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTCTCTGGGATCTCCAAGTGAAC CACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGTTCATCAGAGGATAGCTTCTT TACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTCCAAAATGGTAGAGGATGCGT GTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAATGTTAGCAGATTTTCTTCGTT GTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGCAGTTCAAACCGAAGTCCGCA GGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTGAAGCTCTTGGAAAAAATTAA CTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACAATTATGCTTTTGTATATATA TTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAATATTATGTATCTAGCCCATAG TATTGTACTTAACTTTTACAGGTGAGAAGAGAGTTCTGTGTTTGCATTGATTATGATATTCTGAATAAATATGGAATATATTTTAATGTG >77230_77230_4_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000357384_CEP192_chr18_13124631_ENST00000325971_length(amino acids)=282AA_BP=263 MSAFSEAALEKKLSELSNSQQSVQTLSLWLIHHRKHSRPIVTVWERELRKAKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFK HVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLKQALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGD AAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMVEDACMLLADYNGRLAAEIDDRKQLTRMLADFLRCQKEALAEKEHKLESPALHQH -------------------------------------------------------------- >77230_77230_5_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000357384_CEP192_chr18_13124631_ENST00000506447_length(transcript)=1346nt_BP=941nt TTGCTGGTTTGCGGCCGGTCTTGGATGAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGGGGGCGAGGCCGCAGCTAGGG CGGCGAAACTCTCCTCCCCTCGGCCCCACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGCCTTCTCTGAGGCGGCGCTGG AGAAGAAGCTGTCGGAGTTGAGCAACTCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCACCGTAAACACTCGCGTCCCA TCGTCACCGTGTGGGAGCGGGAGCTGCGGAAAGCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGCCAATGATGTCATACAGAACA GCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTCAAGTGAAACTGATGAAAGTT GTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGAACAACTTAAACAAGCTCTGT ATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTCTCTGGGATCTCCAAGTGAAC CACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGTTCATCAGAGGATAGCTTCTT TACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTCCAAAATGGTAGAGGATGCGT GTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAATGTTAGCAGATTTTCTTCGTT GTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGCAGTTCAAACCGAAGTCCGCA GGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTGAAGCTCTTGGAAAAAATTAA CTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACAATTATGCTTTTGTATATATA TTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAATATTATGTATCTAGCCCATAG >77230_77230_5_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000357384_CEP192_chr18_13124631_ENST00000506447_length(amino acids)=282AA_BP=263 MSAFSEAALEKKLSELSNSQQSVQTLSLWLIHHRKHSRPIVTVWERELRKAKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFK HVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLKQALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGD AAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMVEDACMLLADYNGRLAAEIDDRKQLTRMLADFLRCQKEALAEKEHKLESPALHQH -------------------------------------------------------------- >77230_77230_6_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000399022_CEP192_chr18_13124631_ENST00000430049_length(transcript)=1325nt_BP=961nt ACTGCCGGCCGGACGCCATCTTGCTGGTTTGCGGCCGGTCTTGGATGAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGG GGGCGAGGCCGCAGCTAGGGCGGCGAAACTCTCCTCCCCTCGGCCCCACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGC CTTCTCTGAGGCGGCGCTGGAGAAGAAGCTGTCGGAGTTGAGCAACTCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCA CCGTAAACACTCGCGTCCCATCGTCACCGTGTGGGAGCGGGAGCTGCGGAAAGCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGC CAATGATGTCATACAGAACAGCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTC AAGTGAAACTGATGAAAGTTGTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGA ACAACTTAAACAAGCTCTGTATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTC TCTGGGATCTCCAAGTGAACCACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGT TCATCAGAGGATAGCTTCTTTACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTC CAAAATGGTAGAGGATGCGTGTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAAT GTTAGCAGATTTTCTTCGTTGTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGC AGTTCAAACCGAAGTCCGCAGGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTG AAGCTCTTGGAAAAAATTAACTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACA ATTATGCTTTTGTATATATATTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAAT >77230_77230_6_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000399022_CEP192_chr18_13124631_ENST00000430049_length(amino acids)=282AA_BP=263 MSAFSEAALEKKLSELSNSQQSVQTLSLWLIHHRKHSRPIVTVWERELRKAKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFK HVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLKQALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGD AAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMVEDACMLLADYNGRLAAEIDDRKQLTRMLADFLRCQKEALAEKEHKLESPALHQH -------------------------------------------------------------- >77230_77230_7_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000588737_CEP192_chr18_13124631_ENST00000430049_length(transcript)=1367nt_BP=1003nt ATCTTGCTGGTTTGCGGCCGGTCTTGGATGAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGGGGGCGAGGCCGCAGCTA GGGCGGCGAAACTCTCCTCCCCTCGGCCCCACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGCCTTCTCTGAGGCGGCGC TGGAGAAGAAGCTGTCGGAGTTGAGCAACTCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCACCGTAAACACTCGCGTC CCATCGTCACCGTGTGGGAGCGGGAGCTGCGGAAAGAGTGGAGGTGCAACAAATGAGAAATTCCAATTGGAGATTTTGTCAAACAGGAAT TTACTCCAAACCAAACAGGAAGCTTACTTTTCTCTACCTAGCCAATGATGTCATACAGAACAGCAAGAGGAAGGGGCCAGAGTTTACAAA AGATTTTGCACCAGTTATAGTGGAGGCTTTTAAGCATGTTTCAAGTGAAACTGATGAAAGTTGTAAGAAGCACCTTGGAAGAGTGTTATC TATTTGGGAAGAAAGGTCTGTTTATGAAAATGATGTATTAGAACAACTTAAACAAGCTCTGTATGGTGATAAGAAGCCTAGGAAGCGAAC TTATGAACAGATAAAGGTGGATGAAAATGAAAACTGTTCCTCTCTGGGATCTCCAAGTGAACCACCACAGACTCTAGATCTCGTTAGAGC ATTACAAGATCTGGAAAATGCAGCCTCAGGTGATGCAGCAGTTCATCAGAGGATAGCTTCTTTACCTGTTGAAGTCCAAGAAGTATCTCT ATTAGATAAAATAACAGATAAAGAATCTGGAGAAAGGCTTTCCAAAATGGTAGAGGATGCGTGTATGTTGCTGGCAGATTACAATGGCAG ATTGGCGGCAGAAATAGATGATAGAAAGCAACTCACTCGAATGTTAGCAGATTTTCTTCGTTGTCAAAAGGAAGCCCTTGCAGAGAAAGA GCATAAATTGGAAAGCCCAGCATTACATCAACATGCCCGTGCAGTTCAAACCGAAGTCCGCAGGCAAATTTGAAGCTTTGCTTGTCATTC AAACAGATGAAGGCAAGAGTATTGCTATTCGACTAATTGGTGAAGCTCTTGGAAAAAATTAACTAGAATACATTTTTGTGTAAAGTAAAT TACATAAGTTGTATTTTGTTAACTTTATCTTTCTACACTACAATTATGCTTTTGTATATATATTTTGTATGATGGATATCTATAATTGTA GATTTTGTTTTTACAAGCTAATACTGAAGACTCGACTGAAATATTATGTATCTAGCCCATAGTATTGTACTTAACTTTTACAGGTGAGAA >77230_77230_7_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000588737_CEP192_chr18_13124631_ENST00000430049_length(amino acids)=246AA_BP=227 MRNSNWRFCQTGIYSKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFKHVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLK QALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGDAAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMV -------------------------------------------------------------- >77230_77230_8_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000590898_CEP192_chr18_13124631_ENST00000325971_length(transcript)=1391nt_BP=974nt GAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGGGGGCGAGGCCGCAGCTAGGGCGGCGAAACTCTCCTCCCCTCGGCCC CACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGCCTTCTCTGAGGCGGCGCTGGAGAAGAAGCTGTCGGAGTTGAGCAAC TCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCACCGTAAACACTCGCGTCCCATCGTCACCGTGTGGGAGCGGGAGCTG CGGAAAGAGTGGAGGTGCAACAAATGAGAAATTCCAATTGGAGATTTTGTCAAACAGGAATTTACTCCAAACCAAACAGGAAGCTTACTT TTCTCTACCTAGCCAATGATGTCATACAGAACAGCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTT TTAAGCATGTTTCAAGTGAAACTGATGAAAGTTGTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAA ATGATGTATTAGAACAACTTAAACAAGCTCTGTATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATG AAAACTGTTCCTCTCTGGGATCTCCAAGTGAACCACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAG GTGATGCAGCAGTTCATCAGAGGATAGCTTCTTTACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTG GAGAAAGGCTTTCCAAAATGGTAGAGGATGCGTGTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGC AACTCACTCGAATGTTAGCAGATTTTCTTCGTTGTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATC AACATGCCCGTGCAGTTCAAACCGAAGTCCGCAGGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATT CGACTAATTGGTGAAGCTCTTGGAAAAAATTAACTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATC TTTCTACACTACAATTATGCTTTTGTATATATATTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAG ACTCGACTGAAATATTATGTATCTAGCCCATAGTATTGTACTTAACTTTTACAGGTGAGAAGAGAGTTCTGTGTTTGCATTGATTATGAT >77230_77230_8_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000590898_CEP192_chr18_13124631_ENST00000325971_length(amino acids)=246AA_BP=227 MRNSNWRFCQTGIYSKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFKHVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLK QALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGDAAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMV -------------------------------------------------------------- >77230_77230_9_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000590898_CEP192_chr18_13124631_ENST00000506447_length(transcript)=1379nt_BP=974nt GAAGCGGCGGCCGTGGTGAGAGCGTGGGGAAGGGTGGGGTGAGGGGGCGAGGCCGCAGCTAGGGCGGCGAAACTCTCCTCCCCTCGGCCC CACCGCGTGGGACGGCGTGAACGTGGTGTCGGAGGGATGTCAGCCTTCTCTGAGGCGGCGCTGGAGAAGAAGCTGTCGGAGTTGAGCAAC TCGCAGCAGAGCGTGCAGACCTTGTCCCTGTGGCTCATTCACCACCGTAAACACTCGCGTCCCATCGTCACCGTGTGGGAGCGGGAGCTG CGGAAAGAGTGGAGGTGCAACAAATGAGAAATTCCAATTGGAGATTTTGTCAAACAGGAATTTACTCCAAACCAAACAGGAAGCTTACTT TTCTCTACCTAGCCAATGATGTCATACAGAACAGCAAGAGGAAGGGGCCAGAGTTTACAAAAGATTTTGCACCAGTTATAGTGGAGGCTT TTAAGCATGTTTCAAGTGAAACTGATGAAAGTTGTAAGAAGCACCTTGGAAGAGTGTTATCTATTTGGGAAGAAAGGTCTGTTTATGAAA ATGATGTATTAGAACAACTTAAACAAGCTCTGTATGGTGATAAGAAGCCTAGGAAGCGAACTTATGAACAGATAAAGGTGGATGAAAATG AAAACTGTTCCTCTCTGGGATCTCCAAGTGAACCACCACAGACTCTAGATCTCGTTAGAGCATTACAAGATCTGGAAAATGCAGCCTCAG GTGATGCAGCAGTTCATCAGAGGATAGCTTCTTTACCTGTTGAAGTCCAAGAAGTATCTCTATTAGATAAAATAACAGATAAAGAATCTG GAGAAAGGCTTTCCAAAATGGTAGAGGATGCGTGTATGTTGCTGGCAGATTACAATGGCAGATTGGCGGCAGAAATAGATGATAGAAAGC AACTCACTCGAATGTTAGCAGATTTTCTTCGTTGTCAAAAGGAAGCCCTTGCAGAGAAAGAGCATAAATTGGAAAGCCCAGCATTACATC AACATGCCCGTGCAGTTCAAACCGAAGTCCGCAGGCAAATTTGAAGCTTTGCTTGTCATTCAAACAGATGAAGGCAAGAGTATTGCTATT CGACTAATTGGTGAAGCTCTTGGAAAAAATTAACTAGAATACATTTTTGTGTAAAGTAAATTACATAAGTTGTATTTTGTTAACTTTATC TTTCTACACTACAATTATGCTTTTGTATATATATTTTGTATGATGGATATCTATAATTGTAGATTTTGTTTTTACAAGCTAATACTGAAG ACTCGACTGAAATATTATGTATCTAGCCCATAGTATTGTACTTAACTTTTACAGGTGAGAAGAGAGTTCTGTGTTTGCATTGATTATGAT >77230_77230_9_RPRD1A-CEP192_RPRD1A_chr18_33606863_ENST00000590898_CEP192_chr18_13124631_ENST00000506447_length(amino acids)=246AA_BP=227 MRNSNWRFCQTGIYSKPNRKLTFLYLANDVIQNSKRKGPEFTKDFAPVIVEAFKHVSSETDESCKKHLGRVLSIWEERSVYENDVLEQLK QALYGDKKPRKRTYEQIKVDENENCSSLGSPSEPPQTLDLVRALQDLENAASGDAAVHQRIASLPVEVQEVSLLDKITDKESGERLSKMV -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for RPRD1A-CEP192 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for RPRD1A-CEP192 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for RPRD1A-CEP192 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
