
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:RSU1-CUBN (FusionGDB2 ID:78485) |
Fusion Gene Summary for RSU1-CUBN |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: RSU1-CUBN | Fusion gene ID: 78485 | Hgene | Tgene | Gene symbol | RSU1 | CUBN | Gene ID | 6251 | 8029 |
| Gene name | Ras suppressor protein 1 | cubilin | |
| Synonyms | RSP-1 | IFCR|MGA1|gp280 | |
| Cytomap | 10p13 | 10p13 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ras suppressor protein 1ras suppressor protein 1 variant 1ras suppressor protein 1 variant 2ras suppressor protein 1 variant 3ras suppressor protein 1 variant 5rsu-1 | cubilin460 kDa receptorcubilin (intrinsic factor-cobalamin receptor)cubilin precursor variant 1cubilin precursor variant 2cubilin precursor variant 3intestinal intrinsic factor receptorintrinsic factor-vitamin B12 receptor | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | . | O60494 | |
| Ensembl transtripts involved in fusion gene | ENST00000345264, ENST00000377921, ENST00000602389, ENST00000464074, | ENST00000377823, ENST00000377833, | |
| Fusion gene scores | * DoF score | 13 X 8 X 11=1144 | 10 X 11 X 5=550 |
| # samples | 14 | 12 | |
| ** MAII score | log2(14/1144*10)=-3.03058831983342 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(12/550*10)=-2.1963972128035 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: RSU1 [Title/Abstract] AND CUBN [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | RSU1(16858972)-CUBN(16949678), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | RSU1-CUBN seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. RSU1-CUBN seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. RSU1-CUBN seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across RSU1 (5'-gene) Fusion gene breakpoints across RSU1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
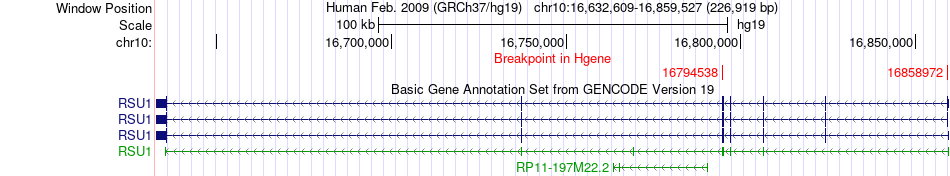 |
 Fusion gene breakpoints across CUBN (3'-gene) Fusion gene breakpoints across CUBN (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
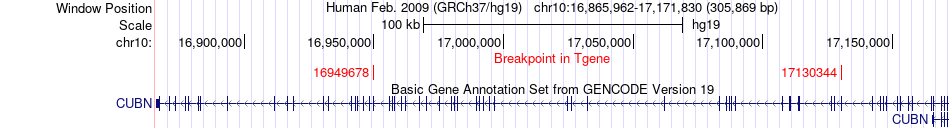 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-A2-A3XT-01A | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| ChimerDB4 | KIRC | TCGA-B0-4848-01A | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| ChimerDB4 | STAD | TCGA-CG-4449-01A | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
Top |
Fusion Gene ORF analysis for RSU1-CUBN |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000345264 | ENST00000377823 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| 5CDS-intron | ENST00000345264 | ENST00000377823 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| 5CDS-intron | ENST00000377921 | ENST00000377823 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| 5CDS-intron | ENST00000377921 | ENST00000377823 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| 5CDS-intron | ENST00000602389 | ENST00000377823 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| 5UTR-3CDS | ENST00000464074 | ENST00000377833 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| 5UTR-intron | ENST00000464074 | ENST00000377823 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| Frame-shift | ENST00000345264 | ENST00000377833 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| Frame-shift | ENST00000345264 | ENST00000377833 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| Frame-shift | ENST00000602389 | ENST00000377833 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| In-frame | ENST00000377921 | ENST00000377833 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| In-frame | ENST00000377921 | ENST00000377833 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - |
| intron-3CDS | ENST00000464074 | ENST00000377833 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| intron-3CDS | ENST00000602389 | ENST00000377833 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| intron-intron | ENST00000464074 | ENST00000377823 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
| intron-intron | ENST00000602389 | ENST00000377823 | RSU1 | chr10 | 16858972 | - | CUBN | chr10 | 16949678 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000377921 | RSU1 | chr10 | 16794538 | - | ENST00000377833 | CUBN | chr10 | 17130344 | - | 11018 | 900 | 107 | 10006 | 3299 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000377921 | ENST00000377833 | RSU1 | chr10 | 16794538 | - | CUBN | chr10 | 17130344 | - | 0.000595295 | 0.99940467 |
Top |
Fusion Genomic Features for RSU1-CUBN |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
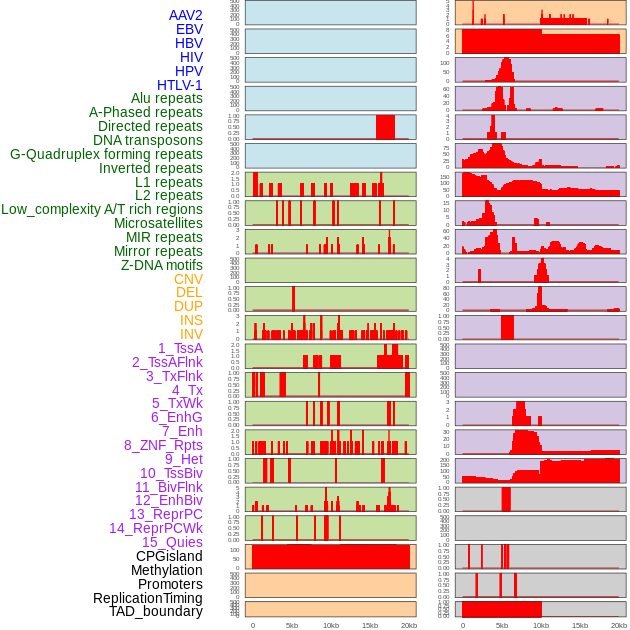 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for RSU1-CUBN |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr10:16858972/chr10:16949678) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CUBN |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Endocytic receptor which plays a role in lipoprotein, vitamin and iron metabolism by facilitating their uptake (PubMed:9572993, PubMed:10371504, PubMed:11717447, PubMed:11606717, PubMed:14576052). Acts together with LRP2 to mediate endocytosis of high-density lipoproteins, GC, hemoglobin, ALB, TF and SCGB1A1. Acts together with AMN to mediate endocytosis of the CBLIF-cobalamin complex (PubMed:9572993, PubMed:14576052). Binds to ALB, MB, Kappa and lambda-light chains, TF, hemoglobin, GC, SCGB1A1, APOA1, high density lipoprotein, and the CBLIF-cobalamin complex. Ligand binding requires calcium (PubMed:9572993). Serves as important transporter in several absorptive epithelia, including intestine, renal proximal tubules and embryonic yolk sac. May play an important role in the development of the peri-implantation embryo through internalization of APOA1 and cholesterol. Binds to LGALS3 at the maternal-fetal interface. {ECO:0000269|PubMed:10371504, ECO:0000269|PubMed:11606717, ECO:0000269|PubMed:11717447, ECO:0000269|PubMed:14576052, ECO:0000269|PubMed:9572993}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 110_133 | 199 | 278.0 | Repeat | Note=LRR 4 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 135_156 | 199 | 278.0 | Repeat | Note=LRR 5 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 158_179 | 199 | 278.0 | Repeat | Note=LRR 6 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 181_202 | 199 | 278.0 | Repeat | Note=LRR 7 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 41_63 | 199 | 278.0 | Repeat | Note=LRR 1 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 64_85 | 199 | 278.0 | Repeat | Note=LRR 2 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000345264 | - | 7 | 9 | 87_109 | 199 | 278.0 | Repeat | Note=LRR 3 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 110_133 | 199 | 278.0 | Repeat | Note=LRR 4 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 135_156 | 199 | 278.0 | Repeat | Note=LRR 5 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 158_179 | 199 | 278.0 | Repeat | Note=LRR 6 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 181_202 | 199 | 278.0 | Repeat | Note=LRR 7 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 41_63 | 199 | 278.0 | Repeat | Note=LRR 1 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 64_85 | 199 | 278.0 | Repeat | Note=LRR 2 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000377921 | - | 6 | 8 | 87_109 | 199 | 278.0 | Repeat | Note=LRR 3 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 110_133 | 146 | 225.0 | Repeat | Note=LRR 4 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 41_63 | 146 | 225.0 | Repeat | Note=LRR 1 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 64_85 | 146 | 225.0 | Repeat | Note=LRR 2 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 87_109 | 146 | 225.0 | Repeat | Note=LRR 3 |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1048_1161 | 588 | 3624.0 | Domain | CUB 6 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1165_1277 | 588 | 3624.0 | Domain | CUB 7 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1278_1389 | 588 | 3624.0 | Domain | CUB 8 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1391_1506 | 588 | 3624.0 | Domain | CUB 9 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1510_1619 | 588 | 3624.0 | Domain | CUB 10 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1620_1734 | 588 | 3624.0 | Domain | CUB 11 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1738_1850 | 588 | 3624.0 | Domain | CUB 12 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1852_1963 | 588 | 3624.0 | Domain | CUB 13 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 1978_2091 | 588 | 3624.0 | Domain | CUB 14 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2092_2213 | 588 | 3624.0 | Domain | CUB 15 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2217_2334 | 588 | 3624.0 | Domain | CUB 16 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2336_2448 | 588 | 3624.0 | Domain | CUB 17 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2452_2565 | 588 | 3624.0 | Domain | CUB 18 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2570_2687 | 588 | 3624.0 | Domain | CUB 19 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2689_2801 | 588 | 3624.0 | Domain | CUB 20 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2805_2919 | 588 | 3624.0 | Domain | CUB 21 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 2920_3035 | 588 | 3624.0 | Domain | CUB 22 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 3037_3150 | 588 | 3624.0 | Domain | CUB 23 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 3157_3274 | 588 | 3624.0 | Domain | CUB 24 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 3278_3393 | 588 | 3624.0 | Domain | CUB 25 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 3395_3507 | 588 | 3624.0 | Domain | CUB 26 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 3511_3623 | 588 | 3624.0 | Domain | CUB 27 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 590_702 | 588 | 3624.0 | Domain | CUB 2 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 708_816 | 588 | 3624.0 | Domain | CUB 3 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 816_928 | 588 | 3624.0 | Domain | CUB 4 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 932_1042 | 588 | 3624.0 | Domain | CUB 5 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2570_2687 | 2511 | 3624.0 | Domain | CUB 19 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2689_2801 | 2511 | 3624.0 | Domain | CUB 20 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2805_2919 | 2511 | 3624.0 | Domain | CUB 21 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2920_3035 | 2511 | 3624.0 | Domain | CUB 22 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 3037_3150 | 2511 | 3624.0 | Domain | CUB 23 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 3157_3274 | 2511 | 3624.0 | Domain | CUB 24 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 3278_3393 | 2511 | 3624.0 | Domain | CUB 25 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 3395_3507 | 2511 | 3624.0 | Domain | CUB 26 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 3511_3623 | 2511 | 3624.0 | Domain | CUB 27 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 135_156 | 146 | 225.0 | Repeat | Note=LRR 5 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 158_179 | 146 | 225.0 | Repeat | Note=LRR 6 |
| Hgene | RSU1 | chr10:16794538 | chr10:17130344 | ENST00000602389 | - | 6 | 8 | 181_202 | 146 | 225.0 | Repeat | Note=LRR 7 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 110_133 | 36 | 278.0 | Repeat | Note=LRR 4 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 135_156 | 36 | 278.0 | Repeat | Note=LRR 5 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 158_179 | 36 | 278.0 | Repeat | Note=LRR 6 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 181_202 | 36 | 278.0 | Repeat | Note=LRR 7 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 41_63 | 36 | 278.0 | Repeat | Note=LRR 1 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 64_85 | 36 | 278.0 | Repeat | Note=LRR 2 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000345264 | - | 2 | 9 | 87_109 | 36 | 278.0 | Repeat | Note=LRR 3 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 110_133 | 36 | 278.0 | Repeat | Note=LRR 4 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 135_156 | 36 | 278.0 | Repeat | Note=LRR 5 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 158_179 | 36 | 278.0 | Repeat | Note=LRR 6 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 181_202 | 36 | 278.0 | Repeat | Note=LRR 7 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 41_63 | 36 | 278.0 | Repeat | Note=LRR 1 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 64_85 | 36 | 278.0 | Repeat | Note=LRR 2 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000377921 | - | 1 | 8 | 87_109 | 36 | 278.0 | Repeat | Note=LRR 3 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 110_133 | 0 | 225.0 | Repeat | Note=LRR 4 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 135_156 | 0 | 225.0 | Repeat | Note=LRR 5 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 158_179 | 0 | 225.0 | Repeat | Note=LRR 6 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 181_202 | 0 | 225.0 | Repeat | Note=LRR 7 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 41_63 | 0 | 225.0 | Repeat | Note=LRR 1 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 64_85 | 0 | 225.0 | Repeat | Note=LRR 2 |
| Hgene | RSU1 | chr10:16858972 | chr10:16949678 | ENST00000602389 | - | 1 | 8 | 87_109 | 0 | 225.0 | Repeat | Note=LRR 3 |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 132_168 | 588 | 3624.0 | Domain | EGF-like 1 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 170_211 | 588 | 3624.0 | Domain | EGF-like 2%3B calcium-binding | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 263_304 | 588 | 3624.0 | Domain | EGF-like 3%3B calcium-binding | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 305_348 | 588 | 3624.0 | Domain | EGF-like 4%3B calcium-binding | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 349_385 | 588 | 3624.0 | Domain | EGF-like 5 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 395_430 | 588 | 3624.0 | Domain | EGF-like 6 | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 432_468 | 588 | 3624.0 | Domain | EGF-like 7%3B calcium-binding | |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 474_586 | 588 | 3624.0 | Domain | CUB 1 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1048_1161 | 2511 | 3624.0 | Domain | CUB 6 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1165_1277 | 2511 | 3624.0 | Domain | CUB 7 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1278_1389 | 2511 | 3624.0 | Domain | CUB 8 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 132_168 | 2511 | 3624.0 | Domain | EGF-like 1 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1391_1506 | 2511 | 3624.0 | Domain | CUB 9 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1510_1619 | 2511 | 3624.0 | Domain | CUB 10 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1620_1734 | 2511 | 3624.0 | Domain | CUB 11 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 170_211 | 2511 | 3624.0 | Domain | EGF-like 2%3B calcium-binding | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1738_1850 | 2511 | 3624.0 | Domain | CUB 12 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1852_1963 | 2511 | 3624.0 | Domain | CUB 13 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 1978_2091 | 2511 | 3624.0 | Domain | CUB 14 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2092_2213 | 2511 | 3624.0 | Domain | CUB 15 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2217_2334 | 2511 | 3624.0 | Domain | CUB 16 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2336_2448 | 2511 | 3624.0 | Domain | CUB 17 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 2452_2565 | 2511 | 3624.0 | Domain | CUB 18 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 263_304 | 2511 | 3624.0 | Domain | EGF-like 3%3B calcium-binding | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 305_348 | 2511 | 3624.0 | Domain | EGF-like 4%3B calcium-binding | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 349_385 | 2511 | 3624.0 | Domain | EGF-like 5 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 395_430 | 2511 | 3624.0 | Domain | EGF-like 6 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 432_468 | 2511 | 3624.0 | Domain | EGF-like 7%3B calcium-binding | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 474_586 | 2511 | 3624.0 | Domain | CUB 1 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 590_702 | 2511 | 3624.0 | Domain | CUB 2 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 708_816 | 2511 | 3624.0 | Domain | CUB 3 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 816_928 | 2511 | 3624.0 | Domain | CUB 4 | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 932_1042 | 2511 | 3624.0 | Domain | CUB 5 |
Top |
Fusion Gene Sequence for RSU1-CUBN |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >78485_78485_1_RSU1-CUBN_RSU1_chr10_16794538_ENST00000377921_CUBN_chr10_17130344_ENST00000377833_length(transcript)=11018nt_BP=900nt GGGCAGCTGGAGTGCGTTCTGCCGAAGCTTGTGGTTGCACGCCCATCGTCTTAGGGGCTACCTTCCGTGGTGAGTGTGTGCGGTGTTGCA CTTGGGGTTGCTTCTCGCTGCGTGGCCGCAGCGGGGCTGAACCCTTCCCACTCCATCCCCTCCCCGCACCCGGACCAGCCCCTGGGACGC CCTCGGACGCCCACCCGCTCCAACCCTGGGGAAGCCTCAGAGTGGCAGCGAAGGCCTCTTGCCTTTCCGCCTCTGCGCTTGCTGTCATCG CCCACGTGCTTTGTTGTTGTTCGGTGCAGACCATGTCCAAGTCTCTGAAGAAGTTGGTGGAGGAGAGCCGGGAGAAGAACCAGCCCGAGG TGGACATGAGTGACCGGGGCATCTCCAACATGCTGGATGTCAACGGCCTCTTTACCTTATCCCATATCACACAACTGGTCCTCAGCCATA ACAAGCTAACAATGGTGCCACCGAACATCGCAGAACTGAAGAATTTGGAGGTGCTCAACTTTTTTAATAACCAAATCGAGGAGCTGCCCA CACAGATCAGTAGCCTTCAGAAACTCAAACACCTGAACCTTGGCATGAACAGGCTGAACACTTTGCCACGAGGCTTCGGCTCCCTGCCAG CTCTTGAGGTTCTGGACTTGACGTACAACAACTTGAGCGAAAATTCTCTTCCTGGAAACTTCTTCTACCTGACCACCCTGCGTGCACTCT ATCTAAGTGACAACGATTTTGAAATCCTGCCGCCAGATATTGGGAAGCTCACAAAGTTGCAGATACTCAGCCTTAGGGATAACGACCTGA TCTCGCTGCCTAAGGAAATCGGGGAGCTTACCCAGCTTAAAGAGCTCCACATTCAGGGGAACCGCCTCACCGTTCTGCCCCCAGAACTAG AGTGTGGAGGTATCCTGACTGGTCCTTACGGTTCTATTAAGTCTCCGGGGTATCCTGGAAACTATCCCCCAGGAAGAGATTGTGTCTGGA TTGTTGTAACTAGTCCTGACCTCCTGGTAACATTTACTTTTGGGACCTTGAGCCTCGAGCACCATGATGACTGCAACAAAGATTACCTTG AGATTCGAGATGGTCCTTTGTATCAGGACCCCCTTCTTGGGAAGTTCTGCACCACTTTCTCTGTCCCACCGCTCCAGACTACTGGCCCCT TTGCCAGAATTCACTTCCATTCAGACTCCCAGATTAGTGACCAAGGCTTCCATATCACCTACTTAACATCACCTTCGGATCTGCGTTGTG GTGGGAACTACACGGACCCAGAGGGTGAACTCTTCTTGCCTGAGTTGTCTGGGCCTTTCACTCACACCAGGCAATGCGTCTATATGATGA AGCAGCCCCAGGGAGAACAAATACAAATCAACTTCACCCACGTGGAGCTGCAATGCCAGAGTGACAGTTCTCAGAATTACATTGAGGTTC GAGATGGTGAAACCTTACTTGGAAAAGTCTGTGGCAACGGAACCATCTCTCACATTAAATCCATTACTAATAGTGTCTGGATCAGGTTTA AAATAGATGCTTCTGTTGAAAAAGCTAGTTTCAGAGCTGTTTATCAAGTCGCTTGCGGGGATGAATTAACTGGAGAAGGGGTCATTCGCT CGCCTTTTTTTCCTAACGTGTATCCTGGAGAAAGAACCTGTAGGTGGACCATCCACCAGCCCCAAAGCCAAGTCATTCTCCTCAACTTCA CTGTCTTTGAAATTGGAAGTTCTGCCCACTGTGAAACAGATTATGTTGAGATTGGTAGCAGTTCCATTTTGGGTTCTCCTGAAAATAAAA AGTATTGCGGTACAGACATACCTTCATTTATAACATCTGTGTACAATTTTCTTTATGTCACATTCGTGAAAAGTTCTTCTACTGAAAACC ATGGTTTCATGGCTAAGTTCAGTGCTGAGGATTTGGCATGTGGAGAAATTCTTACAGAATCAACAGGGACCATTCAAAGTCCTGGCCATC CAAATGTCTACCCCCACGGTATCAACTGTACTTGGCATATATTAGTCCAACCTAATCACCTGATTCATTTAATGTTCGAAACATTTCATC TGGAGTTTCATTACAATTGCACAAACGACTACTTGGAAGTTTATGACACCGACTCTGAGACATCCCTTGGAAGATACTGTGGAAAGTCGA TCCCGCCATCTCTCACAAGCAGTGGTAACTCATTGATGCTGGTGTTTGTGACTGACTCCGACCTCGCTTATGAAGGCTTCTTAATAAACT ATGAAGCAATCAGTGCAGCAACAGCATGTTTGCAAGACTACACAGATGATTTGGGGACATTCACTTCTCCAAACTTCCCCAATAATTATC CCAACAACTGGGAATGCATTTATCGGATCACAGTGAGAACTGGCCAACTGATTGCAGTGCACTTCACAAACTTCTCCTTGGAGGAAGCCA TTGGAAACTATTATACAGATTTTCTGGAAATCAGAGATGGAGGCTATGAAAAATCACCATTGCTGGGAATATTCTATGGCTCAAATCTAC CCCCAACAATCATCTCTCATAGTAACAAACTATGGTTAAAATTTAAGAGTGACCAAATAGACACAAGGTCTGGATTCTCAGCTTACTGGG ATGGGTCATCAACAGGTTGCGGGGGTAATCTCACCACTTCAAGCGGCACGTTCATATCTCCCAACTACCCGATGCCCTATTACCACAGCT CTGAATGCTACTGGTGGTTGAAATCTAGCCACGGCAGCGCATTTGAACTGGAATTCAAAGACTTTCACTTGGAGCATCATCCAAACTGCA CTTTAGATTACCTGGCTGTATATGATGGCCCAAGTAGCAACTCTCATCTGCTAACTCAGCTTTGTGGGGATGAGAAACCCCCTCTTATTC GTTCTAGTGGAGACAGCATGTTTATAAAACTGAGGACAGATGAAGGTCAGCAAGGACGTGGCTTCAAGGCTGAATACCGGCAGACATGTG AGAATGTGGTAATAGTCAATCAAACCTATGGCATCTTAGAGAGTATAGGGTATCCGAATCCTTATTCTGAAAATCAGCATTGCAACTGGA CCATCCGGGCAACAACAGGCAACACTGTGAACTACACATTTTTAGCATTTGACTTGGAACATCACATAAACTGCTCCACAGATTATTTAG AGCTCTATGATGGACCACGGCAGATGGGACGCTACTGTGGAGTAGACCTGCCCCCTCCAGGGAGTACTACAAGCTCCAAGCTTCAAGTGC TGCTCCTTACAGATGGGGTTGGCCGCCGTGAGAAAGGATTTCAGATGCAGTGGTTTGTTTACGGTTGTGGTGGAGAGCTGTCTGGGGCCA CAGGCTCCTTCAGCAGCCCCGGGTTCCCCAACAGGTATCCACCAAACAAGGAGTGTATCTGGTACATTAGGACGGACCCCGGGAGTAGCA TTCAGCTCACCATCCATGACTTCGATGTGGAGTATCATTCAAGGTGCAACTTTGATGTCTTGGAGATCTATGGAGGCCCCGATTTCCACT CTCCCAGAATAGCCCAACTGTGTACCCAGAGATCACCTGAGAACCCCATGCAGGTCTCCAGCACTGGAAATGAGCTAGCAATTCGATTCA AGACCGACTTGTCCATAAATGGGAGAGGCTTCAATGCGTCATGGCAAGCAGTCACTGGAGGTTGTGGTGGGATTTTCCAGGCTCCCAGTG GAGAGATTCATTCTCCAAATTACCCCAGTCCTTATAGGAGCAACACAGACTGTTCTTGGGTCATTCGGGTTGACAGAAATCATCGTGTTC TCTTGAACTTCACTGACTTTGATCTTGAACCACAAGACTCTTGTATTATGGCATACGATGGCTTAAGCTCCACAATGTCCCGCCTTGCCA GGACGTGTGGAAGGGAGCAGCTGGCTAACCCCATCGTCTCCTCAGGAAACAGCCTCTTCTTGAGATTTCAGTCTGGCCCTTCCAGACAGA ACAGAGGCTTCCGAGCTCAATTCAGGCAAGCCTGCGGAGGCCACATCCTCACCAGCTCATTTGATACTGTTTCCTCTCCACGGTTCCCTG CCAATTATCCAAACAATCAGAACTGCAGCTGGATCATTCAAGCGCAACCTCCATTAAATCATATCACCCTCTCTTTTACCCACTTTGAAC TTGAAAGAAGCACAACGTGTGCACGTGACTTTGTAGAAATTTTGGATGGCGGCCACGAAGACGCGCCCCTCCGAGGCCGTTACTGTGGCA CCGACATGCCCCATCCTATCACATCCTTCAGCAGCGCCCTGACGCTGAGATTCGTCTCTGATTCTAGCATCAGTGCTGGGGGTTTCCACA CCACGGTCACCGCATCAGTGTCGGCTTGTGGTGGAACGTTCTACATGGCTGAAGGCATCTTCAACAGCCCTGGCTACCCAGACATTTATC CCCCTAATGTGGAATGTGTCTGGAACATCGTCAGTTCCCCTGGCAACCGGCTCCAGCTGTCTTTTATATCTTTCCAGTTGGAAGACTCTC AGGACTGCAGCAGAGATTTTGTGGAGATCCGTGAAGGAAATGCCACGGGTCACTTGGTGGGACGATACTGTGGAAACTCCTTCCCTCTCA ATTATTCTTCCATCGTTGGACATACCCTGTGGGTCAGATTTATCTCAGATGGTTCTGGCAGCGGCACGGGCTTCCAGGCCACATTTATGA AGATATTTGGCAATGATAATATTGTGGGAACTCATGGGAAAGTCGCCTCTCCTTTCTGGCCTGAAAACTACCCACATAACTCCAATTACC AATGGACAGTAAATGTGAATGCATCTCACGTTGTCCATGGTAGAATCTTGGAGATGGACATAGAAGAAATACAAAACTGCTATTATGACA AATTAAGGATCTATGATGGGCCTAGCATTCACGCCCGCCTAATTGGAGCTTACTGTGGTACCCAGACTGAATCTTTCAGCTCCACTGGAA ATTCTTTGACATTTCATTTTTACTCCGACTCTTCAATCTCAGGGAAGGGATTCCTTCTGGAGTGGTTTGCAGTGGATGCACCTGATGGTG TTTTACCTACCATTGCTCCAGGTGCTTGTGGTGGCTTCCTGAGGACGGGAGATGCACCCGTGTTTCTCTTCTCCCCGGGCTGGCCTGACA GTTACAGTAATAGAGTGGACTGTACGTGGCTCATCCAGGCTCCCGACTCTACCGTGGAACTCAACATTCTTTCCCTGGACATTGAATCTC ACCGAACGTGTGCCTATGATAGCCTTGTGATACGAGATGGAGATAATAACTTGGCCCAGCAGCTAGCAGTTCTCTGTGGCAGAGAGATCC CTGGGCCCATCCGGTCTACTGGAGAGTACATGTTCATCCGCTTCACCTCGGACTCCAGTGTAACCAGGGCAGGCTTCAATGCATCCTTTC ACAAGAGCTGCGGTGGATATTTGCATGCAGACAGAGGGATCATCACGTCCCCCAAGTATCCAGAGACTTACCCATCCAACCTCAACTGTT CTTGGCACGTCCTGGTCCAAAGTGGCCTGACCATTGCTGTCCATTTTGAACAGCCTTTCCAGATTCCAAATGGAGATTCTTCTTGCAACC AGGGGGATTACTTGGTGCTAAGAAATGGTCCTGATATCTGTTCTCCACCCTTGGGACCCCCTGGAGGAAATGGTCATTTTTGTGGCAGTC ATGCTTCATCAACTCTGTTCACCTCGGATAATCAAATGTTTGTTCAGTTTATTTCTGATCACAGTAATGAAGGGCAAGGATTTAAAATCA AATATGAGGCAAAGAGTTTAGCCTGTGGGGGCAACGTCTACATCCATGATGCTGATTCTGCTGGGTATGTGACCTCCCCCAACCACCCTC ATAATTATCCCCCGCACGCTGATTGCATTTGGATCTTAGCGGCTCCACCGGAAACACGCATACAGCTGCAATTTGAAGATCGATTCGATA TTGAAGTAACACCCAACTGTACTTCCAACTACCTTGAGTTGCGGGATGGAGTGGATTCGGATGCACCAATACTTTCCAAATTTTGTGGGA CATCTTTGCCCAGCAGTCAGTGGTCCTCAGGAGAGGTTATGTATTTGAGATTTCGATCTGACAACAGCCCCACACATGTGGGATTCAAGG CCAAGTATTCTATAGCTCAGTGTGGGGGAAGAGTACCAGGGCAAAGTGGTGTTGTTGAAAGCATTGGACATCCAACACTTCCATACAGAG ACAACTTATTCTGTGAGTGGCATCTCCAGGGGCTCTCTGGACACTATCTCACCATCTCTTTTGAAGACTTTAACCTTCAGAATTCTTCTG GCTGTGAAAAAGACTTCGTGGAGATCTGGGACAATCATACCTCTGGAAACATCTTGGGCAGATACTGTGGAAACACCATTCCTGACAGCA TAGACACTTCTAGCAATACTGCTGTGGTCAGGTTTGTCACAGACGGCTCTGTGACTGCCTCAGGATTCAGACTGCGATTTGAATCCAGTA TGGAAGAGTGTGGTGGGGATCTTCAGGGCTCTATTGGAACATTTACTTCTCCCAACTACCCGAACCCAAATCCTCATGGCCGGATCTGCG AGTGGAGAATCACTGCCCCGGAGGGAAGGCGGATCACCCTAATGTTTAACAACCTGAGGCTGGCCACGCATCCGTCCTGCAACAATGAGC ATGTGATAGTATTCAATGGCATTAGAAGTAACTCACCCCAGCTAGAGAAACTGTGTAGTAGTGTGAATGTAAGCAATGAGATTAAATCTT CAGGAAACACAATGAAAGTCATTTTTTTCACGGATGGATCCAGGCCATATGGCGGCTTCACTGCTTCCTATACCTCCAGTGAAGATGCAG TGTGTGGTGGGTCTCTTCCAAATACTCCTGAAGGAAACTTTACTTCTCCTGGCTATGACGGAGTCAGGAATTACTCAAGAAACCTGAACT GCGAATGGACTCTCAGCAATCCAAATCAGGGAAATTCATCCATTTCCATTCACTTTGAAGATTTTTACCTAGAAAGTCACCAAGACTGTC AATTTGATGTCCTCGAGTTTCGAGTGGGTGATGCTGATGGGCCCCTGATGTGGAGACTTTGTGGTCCTTCAAAGCCTACATTGCCATTGG TTATACCTTATTCTCAGGTATGGATTCACTTTGTCACCAACGAACGTGTAGAACACATTGGATTCCATGCAAAGTATTCCTTTACAGATT GTGGCGGAATACAGATAGGTGACAGTGGAGTGATCACAAGCCCCAACTATCCAAATGCTTATGACAGCCTGACCCACTGCTCTTCGCTGT TGGAGGCCCCACAAGGGCACACCATCACTCTCACATTTAGTGACTTTGATATTGAACCCCATACAACTTGTGCTTGGGACTCTGTCACTG TCAGGAATGGTGGGTCCCCTGAATCACCCATCATAGGACAATACTGTGGAAATTCAAACCCCAGGACAATACAGTCAGGTTCCAATCAGC TGGTCGTGACTTTTAACTCAGACCATTCATTGCAAGGTGGTGGATTTTATGCTACGTGGAACACACAAACTTTAGGTTGTGGTGGAATAT TTCATTCTGATAATGGTACAATCAGATCCCCTCACTGGCCTCAGAATTTTCCCGAAAACAGCAGATGTTCCTGGACGGCCATTACTCACA AAAGTAAACACTTGGAGATCAGCTTTGACAACAACTTCCTAATCCCCAGCGGTGATGGACAATGTCAGAATAGCTTCGTGAAGGTGTGGG CAGGAACTGAGGAGGTGGACAAAGCCCTGCTAGCCACTGGCTGTGGGAACGTGGCTCCGGGTCCCGTTATCACACCAAGTAACACATTCA CTGCCGTCTTCCAGTCTCAGGAGGCACCAGCTCAGGGCTTCTCCGCGTCCTTTGTTAGCCGATGTGGAAGTAATTTCACTGGCCCTTCAG GTTACATCATTTCTCCAAATTACCCAAAACAATATGACAACAACATGAATTGCACCTATGTCATAGAGGCTAATCCTCTGTCAGTGGTCC TCTTGACTTTTGTGTCCTTCCACTTAGAAGCTCGTTCCGCTGTGACGGGAAGCTGTGTCAACGATGGCGTGCACATTATCAGAGGTTACA GCGTCATGTCCACCCCATTTGCTACTGTGTGTGGGGATGAGATGCCAGCTCCCCTCACCATCGCTGGGCCGGTTCTGCTTAACTTCTACT CCAACGAGCAAATCACAGACTTCGGATTCAAGTTTTCCTATAGGATAATCTCCTGTGGTGGTGTGTTCAATTTCTCTTCTGGAATCATCA CAAGTCCTGCCTATTCATACGCAGACTACCCAAATGATATGCACTGTCTGTATACCATCACCGTTAGTGACGACAAGGTGATCGAGCTCA AGTTCAGTGATTTTGATGTGGTTCCCTCCACCTCCTGCTCCCATGACTACCTGGCAATTTACGATGGTGCCAATACCAGCGATCCCCTTC TTGGCAAATTCTGCGGTTCCAAGCGCCCACCAAATGTGAAGAGCAGCAATAATAGTATGCTCCTGGTGTTCAAGACAGATTCATTTCAGA CAGCAAAAGGCTGGAAGATGTCTTTCCGGCAGACATTGGGGCCTCAGCAAGGATGTGGTGGTTATCTGACAGGCTCGAATAATACCTTTG CCTCTCCTGATTCTGATTCGAATGGAATGTATGACAAGAATTTAAACTGTGTATGGATCATAATTGCACCTGTAAACAAAGTAATTCACC TCACCTTCAATACATTTGCTCTGGAGGCAGCAAGTACTAGGCAAAGATGCCTTTATGATTATGTAAAGTTATATGATGGGGATAGTGAAA ATGCGAACTTGGCTGGAACGTTTTGTGGTTCCACAGTACCTGCTCCTTTTATCTCTTCTGGTAACTTCCTTACGGTTCAATTCATCAGTG ACTTAACATTAGAGAGGGAAGGATTTAATGCTACATACACCATCATGGACATGCCTTGTGGTGGAACATACAATGCAACTTGGACCCCAC AAAATATTTCATCACCCAATTCATCAGACCCAGATGTCCCATTTTCCATCTGTACTTGGGTCATTGATTCCCCTCCGCATCAGCAGGTCA AGATAACTGTGTGGGCATTACAGCTGACCTCGCAAGACTGCACGCAGAATTACTTACAGCTTCAGGACTCACCGCAGGGTCACGGAAATT CAAGATTTCAGTTCTGTGGCAGAAATGCTTCGGCTGTGCCAGTGTTTTATTCTTCTATGAGTACTGCAATGGTCATTTTCAAATCTGGAG TTGTAAACAGAAACTCTAGAATGAGTTTCACCTATCAGATTGCAGATTGCAACAGAGACTATCACAAGGCATTTGGCAACCTGAGAAGCC CTGGATGGCCAGATAACTACGACAATGACAAGGATTGCACCGTTACTCTCACAGCCCCCCAGAACCACACCATTTCCCTCTTTTTTCATT CACTTGGCATCGAGAACTCAGTTGAATGCAGAAACGATTTCTTGGAGGTGAGAAATGGAAGTAACAGCAATTCACCATTACTGGGCAAGT ACTGTGGAACTCTGCTGCCAAACCCTGTCTTCTCTCAAAATAATGAACTATACCTACGATTTAAGAGTGATAGTGTAACTTCTGATCGTG GATATGAAATCATCTGGACTTCATCACCCTCTGGATGTGGTGGAACTCTTTATGGAGACAGAGGCTCATTCACCAGCCCCGGCTATCCAG GCACATACCCAAACAACACGTACTGCGAGTGGGTCCTTGTTGCTCCTGCTGGAAGGCTTGTCACCATCAACTTCTACTTCATCAGCATTG ACGATCCAGGAGACTGTGTCCAGAACTATCTCACACTCTATGATGGGCCCAACGCCAGCTCTCCATCCTCTGGACCATACTGCGGAGGCG ACACCAGCATAGCTCCCTTCGTGGCTTCCTCAAATCAGGTCTTCATAAAATTTCATGCTGATTATGCACGGCGTCCATCCGCATTCCGAT TAACTTGGGACAGCTAAGTGGGTAACAACTCGTGTTCACTCAGCACTTTCCCTCTGCAGCACGCTGGACAGCACTCTGCCATCCTGATAC ATGACCCCTGCTGATGCCACAGAGAATAAGCTGAACTTGTATGGTTTTTCACCAAACCATGGATAGAATCAATATTTGTAGGCCAGGCGT GGTGGCTCACCCCCCTGTATTCTCAGCACTTTGGGAGGCCGAGGCAGGTTGATCACCTGAGGTCAGGAGTTTGAGACTAGCCTGGCCAGC ATGGTGAAACCTCATCTCTCTAACAATATAATAATTAGCCAGGCGTGGTGGTGGGTGCCTGTAATTCCAGCCACTCGAGAGGCTGAGGCA GGAGAATTGCTTGAACCCAGGAGGCAGAGGTTGCAGTGAGCTAAGATCACACCACTACACTCCAGCCTGGGCGAGACGGCAAGACTCCAT CTCAAAAAAAAAAGAAACAAAAAAAACCAGAATCAATATTTGTACATTTTCTCGAACATAGAATATAGCTTCTTTAGTCTTGAGTGTGCA TTTCATTCTAATATTTTGAGCTGAAATTTAAAAAAACTTTGAAAGAGTTGGAAATGATTATGGCATATGTGACATACATTTTTAAAAGTT AATAATAATAGCCAGGGGCAGTGGCTCATACCCATAATCCCAGCACGCTGGGAGGCCATGATGGGAGGATTGCTTGAACCTAGGAGTTTG AGACCAGCCTGGGCAACAAAGTGAGACCTGATTTTTACAAAAAATCAAAAAATTAGCCAGGCATGGTGGCATGCACCCGTGGTTCCAGCT ACACAGGAGGTTGAAGCAGGAGGATCACTTGAGCCCAGTAGGTTAAGGCTGCAGTGAAACCCTGTGAATTAACCACTGTACTCCAGCCTG GGTGACAGACTGAGACCCTATCTCAAAAATGACAACAAGAACAACAAAAGTTAATGATAATATAGAAGCATAAATTTCCTGTGAATGTTC >78485_78485_1_RSU1-CUBN_RSU1_chr10_16794538_ENST00000377921_CUBN_chr10_17130344_ENST00000377833_length(amino acids)=3299AA_BP=263 MRGRSGAEPFPLHPLPAPGPAPGTPSDAHPLQPWGSLRVAAKASCLSASALAVIAHVLCCCSVQTMSKSLKKLVEESREKNQPEVDMSDR GISNMLDVNGLFTLSHITQLVLSHNKLTMVPPNIAELKNLEVLNFFNNQIEELPTQISSLQKLKHLNLGMNRLNTLPRGFGSLPALEVLD LTYNNLSENSLPGNFFYLTTLRALYLSDNDFEILPPDIGKLTKLQILSLRDNDLISLPKEIGELTQLKELHIQGNRLTVLPPELECGGIL TGPYGSIKSPGYPGNYPPGRDCVWIVVTSPDLLVTFTFGTLSLEHHDDCNKDYLEIRDGPLYQDPLLGKFCTTFSVPPLQTTGPFARIHF HSDSQISDQGFHITYLTSPSDLRCGGNYTDPEGELFLPELSGPFTHTRQCVYMMKQPQGEQIQINFTHVELQCQSDSSQNYIEVRDGETL LGKVCGNGTISHIKSITNSVWIRFKIDASVEKASFRAVYQVACGDELTGEGVIRSPFFPNVYPGERTCRWTIHQPQSQVILLNFTVFEIG SSAHCETDYVEIGSSSILGSPENKKYCGTDIPSFITSVYNFLYVTFVKSSSTENHGFMAKFSAEDLACGEILTESTGTIQSPGHPNVYPH GINCTWHILVQPNHLIHLMFETFHLEFHYNCTNDYLEVYDTDSETSLGRYCGKSIPPSLTSSGNSLMLVFVTDSDLAYEGFLINYEAISA ATACLQDYTDDLGTFTSPNFPNNYPNNWECIYRITVRTGQLIAVHFTNFSLEEAIGNYYTDFLEIRDGGYEKSPLLGIFYGSNLPPTIIS HSNKLWLKFKSDQIDTRSGFSAYWDGSSTGCGGNLTTSSGTFISPNYPMPYYHSSECYWWLKSSHGSAFELEFKDFHLEHHPNCTLDYLA VYDGPSSNSHLLTQLCGDEKPPLIRSSGDSMFIKLRTDEGQQGRGFKAEYRQTCENVVIVNQTYGILESIGYPNPYSENQHCNWTIRATT GNTVNYTFLAFDLEHHINCSTDYLELYDGPRQMGRYCGVDLPPPGSTTSSKLQVLLLTDGVGRREKGFQMQWFVYGCGGELSGATGSFSS PGFPNRYPPNKECIWYIRTDPGSSIQLTIHDFDVEYHSRCNFDVLEIYGGPDFHSPRIAQLCTQRSPENPMQVSSTGNELAIRFKTDLSI NGRGFNASWQAVTGGCGGIFQAPSGEIHSPNYPSPYRSNTDCSWVIRVDRNHRVLLNFTDFDLEPQDSCIMAYDGLSSTMSRLARTCGRE QLANPIVSSGNSLFLRFQSGPSRQNRGFRAQFRQACGGHILTSSFDTVSSPRFPANYPNNQNCSWIIQAQPPLNHITLSFTHFELERSTT CARDFVEILDGGHEDAPLRGRYCGTDMPHPITSFSSALTLRFVSDSSISAGGFHTTVTASVSACGGTFYMAEGIFNSPGYPDIYPPNVEC VWNIVSSPGNRLQLSFISFQLEDSQDCSRDFVEIREGNATGHLVGRYCGNSFPLNYSSIVGHTLWVRFISDGSGSGTGFQATFMKIFGND NIVGTHGKVASPFWPENYPHNSNYQWTVNVNASHVVHGRILEMDIEEIQNCYYDKLRIYDGPSIHARLIGAYCGTQTESFSSTGNSLTFH FYSDSSISGKGFLLEWFAVDAPDGVLPTIAPGACGGFLRTGDAPVFLFSPGWPDSYSNRVDCTWLIQAPDSTVELNILSLDIESHRTCAY DSLVIRDGDNNLAQQLAVLCGREIPGPIRSTGEYMFIRFTSDSSVTRAGFNASFHKSCGGYLHADRGIITSPKYPETYPSNLNCSWHVLV QSGLTIAVHFEQPFQIPNGDSSCNQGDYLVLRNGPDICSPPLGPPGGNGHFCGSHASSTLFTSDNQMFVQFISDHSNEGQGFKIKYEAKS LACGGNVYIHDADSAGYVTSPNHPHNYPPHADCIWILAAPPETRIQLQFEDRFDIEVTPNCTSNYLELRDGVDSDAPILSKFCGTSLPSS QWSSGEVMYLRFRSDNSPTHVGFKAKYSIAQCGGRVPGQSGVVESIGHPTLPYRDNLFCEWHLQGLSGHYLTISFEDFNLQNSSGCEKDF VEIWDNHTSGNILGRYCGNTIPDSIDTSSNTAVVRFVTDGSVTASGFRLRFESSMEECGGDLQGSIGTFTSPNYPNPNPHGRICEWRITA PEGRRITLMFNNLRLATHPSCNNEHVIVFNGIRSNSPQLEKLCSSVNVSNEIKSSGNTMKVIFFTDGSRPYGGFTASYTSSEDAVCGGSL PNTPEGNFTSPGYDGVRNYSRNLNCEWTLSNPNQGNSSISIHFEDFYLESHQDCQFDVLEFRVGDADGPLMWRLCGPSKPTLPLVIPYSQ VWIHFVTNERVEHIGFHAKYSFTDCGGIQIGDSGVITSPNYPNAYDSLTHCSSLLEAPQGHTITLTFSDFDIEPHTTCAWDSVTVRNGGS PESPIIGQYCGNSNPRTIQSGSNQLVVTFNSDHSLQGGGFYATWNTQTLGCGGIFHSDNGTIRSPHWPQNFPENSRCSWTAITHKSKHLE ISFDNNFLIPSGDGQCQNSFVKVWAGTEEVDKALLATGCGNVAPGPVITPSNTFTAVFQSQEAPAQGFSASFVSRCGSNFTGPSGYIISP NYPKQYDNNMNCTYVIEANPLSVVLLTFVSFHLEARSAVTGSCVNDGVHIIRGYSVMSTPFATVCGDEMPAPLTIAGPVLLNFYSNEQIT DFGFKFSYRIISCGGVFNFSSGIITSPAYSYADYPNDMHCLYTITVSDDKVIELKFSDFDVVPSTSCSHDYLAIYDGANTSDPLLGKFCG SKRPPNVKSSNNSMLLVFKTDSFQTAKGWKMSFRQTLGPQQGCGGYLTGSNNTFASPDSDSNGMYDKNLNCVWIIIAPVNKVIHLTFNTF ALEAASTRQRCLYDYVKLYDGDSENANLAGTFCGSTVPAPFISSGNFLTVQFISDLTLEREGFNATYTIMDMPCGGTYNATWTPQNISSP NSSDPDVPFSICTWVIDSPPHQQVKITVWALQLTSQDCTQNYLQLQDSPQGHGNSRFQFCGRNASAVPVFYSSMSTAMVIFKSGVVNRNS RMSFTYQIADCNRDYHKAFGNLRSPGWPDNYDNDKDCTVTLTAPQNHTISLFFHSLGIENSVECRNDFLEVRNGSNSNSPLLGKYCGTLL PNPVFSQNNELYLRFKSDSVTSDRGYEIIWTSSPSGCGGTLYGDRGSFTSPGYPGTYPNNTYCEWVLVAPAGRLVTINFYFISIDDPGDC -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for RSU1-CUBN |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | CUBN | chr10:16794538 | chr10:17130344 | ENST00000377833 | 13 | 67 | 42_49 | 588.3333333333334 | 3624.0 | AMN | |
| Tgene | CUBN | chr10:16858972 | chr10:16949678 | ENST00000377833 | 47 | 67 | 42_49 | 2511.0 | 3624.0 | AMN |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for RSU1-CUBN |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for RSU1-CUBN |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
