
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SAMD8-KAT6B (FusionGDB2 ID:79103) |
Fusion Gene Summary for SAMD8-KAT6B |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SAMD8-KAT6B | Fusion gene ID: 79103 | Hgene | Tgene | Gene symbol | SAMD8 | KAT6B | Gene ID | 142891 | 23522 |
| Gene name | sterile alpha motif domain containing 8 | lysine acetyltransferase 6B | |
| Synonyms | HEL-177|SMSr | GTPTS|MORF|MOZ2|MYST4|ZC2HC6B|qkf|querkopf | |
| Cytomap | 10q22.2 | 10q22.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | sphingomyelin synthase-related protein 1CPE synthaseHEL-S-181mPSAM domain-containing protein 8ceramide phosphoethanolamine synthaseepididymis luminal protein 177epididymis secretory sperm binding protein Li 181mPsphingomyelin synthase relatedsteri | histone acetyltransferase KAT6BK(lysine) acetyltransferase 6BMOZ, YBF2/SAS3, SAS2 and TIP60 protein 4MOZ-related factorMYST histone acetyltransferase (monocytic leukemia) 4MYST-4histone acetyltransferase MORFhistone acetyltransferase MOZ2histone a | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q8WYB5 | |
| Ensembl transtripts involved in fusion gene | ENST00000372687, ENST00000372690, ENST00000542569, | ENST00000490365, ENST00000287239, ENST00000372711, ENST00000372724, ENST00000372725, ENST00000372714, | |
| Fusion gene scores | * DoF score | 6 X 6 X 5=180 | 7 X 11 X 4=308 |
| # samples | 8 | 10 | |
| ** MAII score | log2(8/180*10)=-1.16992500144231 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(10/308*10)=-1.62293035092018 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SAMD8 [Title/Abstract] AND KAT6B [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SAMD8(76928416)-KAT6B(76788247), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | SAMD8-KAT6B seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. SAMD8-KAT6B seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. SAMD8-KAT6B seems lost the major protein functional domain in Tgene partner, which is a epigenetic factor due to the frame-shifted ORF. SAMD8-KAT6B seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. SAMD8-KAT6B seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SAMD8 | GO:0046513 | ceramide biosynthetic process | 19506037 |
| Hgene | SAMD8 | GO:2000303 | regulation of ceramide biosynthetic process | 19506037 |
| Tgene | KAT6B | GO:0016573 | histone acetylation | 11965546 |
| Tgene | KAT6B | GO:0043966 | histone H3 acetylation | 16387653 |
| Tgene | KAT6B | GO:0045892 | negative regulation of transcription, DNA-templated | 10497217 |
| Tgene | KAT6B | GO:0045893 | positive regulation of transcription, DNA-templated | 10497217|11965546 |
 Fusion gene breakpoints across SAMD8 (5'-gene) Fusion gene breakpoints across SAMD8 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
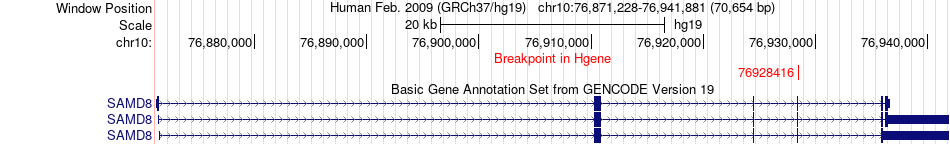 |
 Fusion gene breakpoints across KAT6B (3'-gene) Fusion gene breakpoints across KAT6B (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
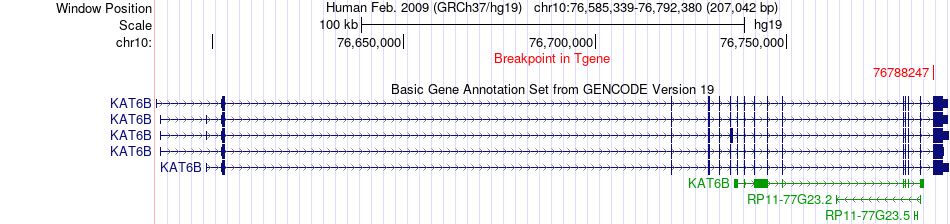 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUAD | TCGA-44-7660-01A | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
Top |
Fusion Gene ORF analysis for SAMD8-KAT6B |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000372687 | ENST00000490365 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| 5CDS-intron | ENST00000372690 | ENST00000490365 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| 5CDS-intron | ENST00000542569 | ENST00000490365 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372687 | ENST00000287239 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372687 | ENST00000372711 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372687 | ENST00000372724 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372687 | ENST00000372725 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372690 | ENST00000287239 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372690 | ENST00000372711 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372690 | ENST00000372724 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000372690 | ENST00000372725 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000542569 | ENST00000287239 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000542569 | ENST00000372711 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000542569 | ENST00000372724 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| Frame-shift | ENST00000542569 | ENST00000372725 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| In-frame | ENST00000372687 | ENST00000372714 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| In-frame | ENST00000372690 | ENST00000372714 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
| In-frame | ENST00000542569 | ENST00000372714 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000542569 | SAMD8 | chr10 | 76928416 | - | ENST00000372714 | KAT6B | chr10 | 76788247 | + | 3597 | 895 | 831 | 3452 | 873 |
| ENST00000372687 | SAMD8 | chr10 | 76928416 | - | ENST00000372714 | KAT6B | chr10 | 76788247 | + | 3579 | 877 | 813 | 3434 | 873 |
| ENST00000372690 | SAMD8 | chr10 | 76928416 | - | ENST00000372714 | KAT6B | chr10 | 76788247 | + | 3761 | 1059 | 995 | 3616 | 873 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000542569 | ENST00000372714 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + | 0.006066816 | 0.99393314 |
| ENST00000372687 | ENST00000372714 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + | 0.006415662 | 0.99358433 |
| ENST00000372690 | ENST00000372714 | SAMD8 | chr10 | 76928416 | - | KAT6B | chr10 | 76788247 | + | 0.006175128 | 0.99382484 |
Top |
Fusion Genomic Features for SAMD8-KAT6B |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
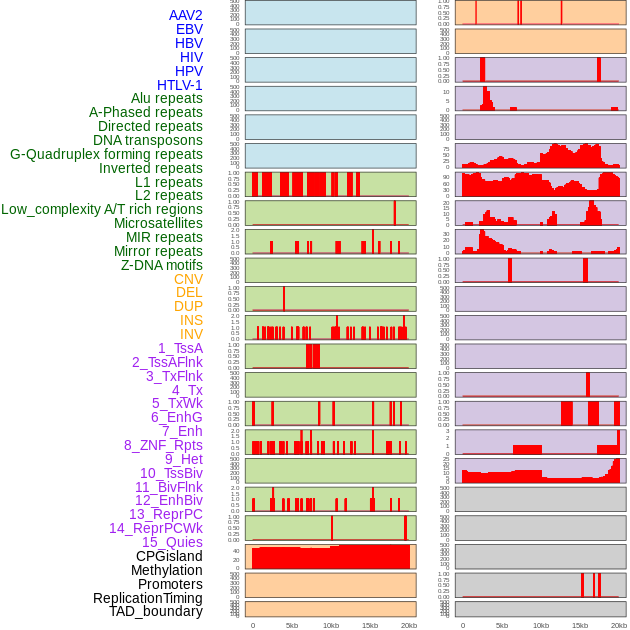 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for SAMD8-KAT6B |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr10:76928416/chr10:76788247) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | KAT6B |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Histone acetyltransferase which may be involved in both positive and negative regulation of transcription. Required for RUNX2-dependent transcriptional activation. May be involved in cerebral cortex development. Component of the MOZ/MORF complex which has a histone H3 acetyltransferase activity. {ECO:0000269|PubMed:10497217, ECO:0000269|PubMed:11965546, ECO:0000269|PubMed:16387653}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 12_78 | 264 | 327.0 | Domain | SAM |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 12_78 | 327 | 479.0 | Domain | SAM |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 12_78 | 264 | 416.0 | Domain | SAM |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 153_173 | 264 | 327.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 201_221 | 264 | 327.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 232_252 | 264 | 327.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 153_173 | 327 | 479.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 201_221 | 327 | 479.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 232_252 | 327 | 479.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 277_297 | 327 | 479.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 153_173 | 264 | 416.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 201_221 | 264 | 416.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 232_252 | 264 | 416.0 | Transmembrane | Helical |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1351_1373 | 1221 | 2074.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1409_1417 | 1221 | 2074.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1594_1763 | 1221 | 2074.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1961_2061 | 1221 | 2074.0 | Compositional bias | Note=Met-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1070_1104 | 1038 | 1891.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1204_1207 | 1038 | 1891.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1351_1373 | 1038 | 1891.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1409_1417 | 1038 | 1891.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1594_1763 | 1038 | 1891.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1961_2061 | 1038 | 1891.0 | Compositional bias | Note=Met-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1070_1104 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1204_1207 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1351_1373 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1409_1417 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1594_1763 | 929 | 1782.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1961_2061 | 929 | 1782.0 | Compositional bias | Note=Met-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1070_1104 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1204_1207 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1351_1373 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1409_1417 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1594_1763 | 929 | 1782.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1961_2061 | 929 | 1782.0 | Compositional bias | Note=Met-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1070_1104 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1204_1207 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1351_1373 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1409_1417 | 929 | 1782.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1594_1763 | 929 | 1782.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1961_2061 | 929 | 1782.0 | Compositional bias | Note=Met-rich |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 368_415 | 264 | 327.0 | Topological domain | Cytoplasmic |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 368_415 | 327 | 479.0 | Topological domain | Cytoplasmic |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 368_415 | 264 | 416.0 | Topological domain | Cytoplasmic |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 277_297 | 264 | 327.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 322_342 | 264 | 327.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372687 | - | 4 | 5 | 347_367 | 264 | 327.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 322_342 | 327 | 479.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000372690 | - | 4 | 6 | 347_367 | 327 | 479.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 277_297 | 264 | 416.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 322_342 | 264 | 416.0 | Transmembrane | Helical |
| Hgene | SAMD8 | chr10:76928416 | chr10:76788247 | ENST00000542569 | - | 4 | 6 | 347_367 | 264 | 416.0 | Transmembrane | Helical |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1070_1104 | 1221 | 2074.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1204_1207 | 1221 | 2074.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 492_533 | 1221 | 2074.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 521_524 | 1221 | 2074.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 599_605 | 1221 | 2074.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 492_533 | 1038 | 1891.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 521_524 | 1038 | 1891.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 599_605 | 1038 | 1891.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 492_533 | 929 | 1782.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 521_524 | 929 | 1782.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 599_605 | 929 | 1782.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 492_533 | 929 | 1782.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 521_524 | 929 | 1782.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 599_605 | 929 | 1782.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 492_533 | 929 | 1782.0 | Compositional bias | Note=Ser-rich | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 521_524 | 929 | 1782.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 599_605 | 929 | 1782.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 103_176 | 1221 | 2074.0 | Domain | H15 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 715_989 | 1221 | 2074.0 | Domain | MYST-type HAT | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 103_176 | 1038 | 1891.0 | Domain | H15 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 715_989 | 1038 | 1891.0 | Domain | MYST-type HAT | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 103_176 | 929 | 1782.0 | Domain | H15 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 715_989 | 929 | 1782.0 | Domain | MYST-type HAT | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 103_176 | 929 | 1782.0 | Domain | H15 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 715_989 | 929 | 1782.0 | Domain | MYST-type HAT | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 103_176 | 929 | 1782.0 | Domain | H15 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 715_989 | 929 | 1782.0 | Domain | MYST-type HAT | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 361_717 | 1221 | 2074.0 | Region | Note=Negatively regulates HAT activity | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 718_1008 | 1221 | 2074.0 | Region | Note=Catalytic | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 856_860 | 1221 | 2074.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 865_871 | 1221 | 2074.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 361_717 | 1038 | 1891.0 | Region | Note=Negatively regulates HAT activity | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 718_1008 | 1038 | 1891.0 | Region | Note=Catalytic | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 856_860 | 1038 | 1891.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 865_871 | 1038 | 1891.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 361_717 | 929 | 1782.0 | Region | Note=Negatively regulates HAT activity | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 718_1008 | 929 | 1782.0 | Region | Note=Catalytic | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 856_860 | 929 | 1782.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 865_871 | 929 | 1782.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 361_717 | 929 | 1782.0 | Region | Note=Negatively regulates HAT activity | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 718_1008 | 929 | 1782.0 | Region | Note=Catalytic | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 856_860 | 929 | 1782.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 865_871 | 929 | 1782.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 361_717 | 929 | 1782.0 | Region | Note=Negatively regulates HAT activity | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 718_1008 | 929 | 1782.0 | Region | Note=Catalytic | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 856_860 | 929 | 1782.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 865_871 | 929 | 1782.0 | Region | Acetyl-CoA binding | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 213_272 | 1221 | 2074.0 | Zinc finger | PHD-type 1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 269_320 | 1221 | 2074.0 | Zinc finger | PHD-type 2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 748_773 | 1221 | 2074.0 | Zinc finger | C2HC MYST-type | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 213_272 | 1038 | 1891.0 | Zinc finger | PHD-type 1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 269_320 | 1038 | 1891.0 | Zinc finger | PHD-type 2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 748_773 | 1038 | 1891.0 | Zinc finger | C2HC MYST-type | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 213_272 | 929 | 1782.0 | Zinc finger | PHD-type 1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 269_320 | 929 | 1782.0 | Zinc finger | PHD-type 2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 748_773 | 929 | 1782.0 | Zinc finger | C2HC MYST-type | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 213_272 | 929 | 1782.0 | Zinc finger | PHD-type 1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 269_320 | 929 | 1782.0 | Zinc finger | PHD-type 2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 748_773 | 929 | 1782.0 | Zinc finger | C2HC MYST-type | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 213_272 | 929 | 1782.0 | Zinc finger | PHD-type 1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 269_320 | 929 | 1782.0 | Zinc finger | PHD-type 2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 748_773 | 929 | 1782.0 | Zinc finger | C2HC MYST-type |
Top |
Fusion Gene Sequence for SAMD8-KAT6B |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >79103_79103_1_SAMD8-KAT6B_SAMD8_chr10_76928416_ENST00000372687_KAT6B_chr10_76788247_ENST00000372714_length(transcript)=3579nt_BP=877nt CTCGGACCGCGGAGGTGAGCGGGAGCTGAGGCTGAGGAGAGGGGAGCTTGGGGGGCGCCTGCTGCCAAGGGCAGCGGAGGAGGAAATGGC AGGTCCTAATCAACTCTGCATTCGCCGCTGGACTACCAAGCATGTAGCTGTGTGGCTGAAGGATGAAGGCTTTTTTGAATATGTGGACAT TTTATGCAATAAGCACCGACTTGATGGAATCACATTGCTAACATTGACTGAATATGATCTCCGGTCTCCTCCTCTGGAAATCAAAGTCTT AGGGGACATTAAAAGGTTAATGCTCTCAGTCCGAAAATTGCAGAAAATACATATTGATGTTTTAGAAGAGATGGGCTACAACAGTGACAG TCCCATGGGTTCCATGACCCCTTTCATCAGTGCTCTTCAGAGTACAGACTGGCTCTGTAATGGGGAGCTTTCCCATGACTGTGACGGACC CATAACTGACTTGAATTCTGATCAGTACCAGTACATGAATGGTAAAAACAAACATTCTGTTCGAAGATTGGACCCAGAATACTGGAAGAC TATACTGAGTTGTATATATGTTTTTATAGTATTTGGATTTACATCTTTCATTATGGTTATAGTCCATGAGCGAGTGCCTGACATGCAGAC CTATCCACCACTCCCAGATATATTCTTAGACAGCGTTCCTAGAATCCCATGGGCCTTTGCCATGACGGAAGTATGTGGCATGATTCTGTG CTATATTTGGCTCCTGGTTCTTCTTCTTCACAAGCACAGGTCAATACTTCTGCGAAGGCTCTGTAGTCTGATGGGAACTGTATTCTTGCT TCGCTGCTTTACCATGTTTGTGACCTCCCTCTCCGTGCCAGGACAACACCTGCAGTGTACTGGAAAGACAATATGAATGATGATTCAAGT AACTTGAAAGAAGGCAGTAAAGACAATCCCGAACCTCTAAAGTGCAAACAAGTGTGGCCAAAAGGAACAAAGCGCGGTCTATCTAAGTGG AGGCAAAACAAAGAGAGGAAGACCGGATTTAAACTGAATTTGTACACCCCGCCAGAAACACCCATGGAGCCTGACGAGCAGGTAACAGTG GAAGAACAGAAGGAGACTTCAGAAGGAAAAACCAGCCCCAGTCCCATCAGGATTGAGGAGGAGGTCAAGGAAACTGGGGAAGCCCTGTTG CCTCAAGAGGAAAACAGAAGGGAAGAAACATGTGCCCCTGTAAGTCCAAACACATCACCAGGTGAAAAACCAGAAGATGATCTCATCAAA CCTGAGGAAGAGGAAGAGGAGGAGGAGGAGGAAGAGGAAGAAGAGGAAGAAGAGGAAGGGGAAGAAGAAGAAGGAGGAGGAAATGTAGAA AAAGATCCAGATGGTGCTAAAAGCCAAGAAAAAGAGGAACCAGAAATCTCCACGGAAAAAGAAGACTCTGCACGTTTGGATGATCACGAA GAGGAGGAGGAAGAGGATGAAGAGCCATCCCACAACGAGGACCATGATGCCGATGACGAGGATGACAGCCACATGGAGTCTGCCGAAGTG GAGAAGGAAGAGCTGCCCAGAGAAAGCTTCAAAGAAGTACTGGAAAACCAGGAGACTTTTTTAGACCTTAATGTGCAGCCTGGTCACTCG AACCCAGAGGTCTTAATGGACTGTGGCGTCGACCTGACAGCTTCTTGTAACAGTGAGCCCAAGGAGCTTGCTGGGGACCCTGAAGCTGTA CCCGAATCTGACGAGGAGCCACCCCCAGGAGAACAGGCACAGAAGCAGGACCAAAAGAACAGCAAGGAAGTCGATACAGAGTTCAAAGAG GGAAACCCAGCAACCATGGAAATCGACTCTGAGACTGTCCAGGCCGTTCAGTCTTTGACCCAGGAGAGCAGCGAACAGGACGACACCTTT CAGGATTGTGCCGAGACTCAAGAGGCCTGTAGAAGCCTACAGAACTACACCCGTGCAGACCAAAGTCCACAGATTGCCACCACGCTCGAC GATTGCCAACAGTCGGACCACAGTAGCCCAGTTTCATCCGTCCACTCCCATCCTGGCCAGTCCGTACGTTCTGTCAACAGCCCAAGTGTC CCTGCTCTGGAAAACAGCTACGCCCAAATCAGCCCAGATCAAAGTGCCATCTCAGTGCCATCTCTGCAGAACATGGAAACCAGTCCCATG ATGGATGTCCCATCAGTTTCAGATCATTCACAGCAAGTCGTAGACAGTGGATTTAGTGACCTGGGCAGTATCGAGAGCACAACTGAGAAC TACGAAAACCCAAGCAGCTACGATTCTACTATGGGAGGCAGCATCTGTGGAAACGGCTCTTCACAGAACAGCTGCTCCTATAGCAACCTC ACCTCCAGCAGTCTGACACAGAGCAGCTGTGCTGTCACCCAGCAGATGTCCAACATCAGCGGGAGCTGCAGCATGCTGCAGCAAACCAGC ATCAGCTCCCCTCCGACCTGCAGCGTCAAGTCTCCTCAAGGCTGTGTGGTGGAGAGGCCTCCGAGCAGCAGCCAGCAGCTGGCTCAGTGC AGCATGGCTGCTAACTTCACCCCACCCATGCAGCTGGCTGAAATCCCCGAGACGAGCAACGCCAACATTGGCTTATACGAGCGAATGGGT CAGAGTGATTTTGGGGCTGGGCATTACCCGCAGCCGTCAGCCACCTTCAGCCTTGCCAAACTGCAGCAGTTAACTAATACACTTATTGAT CATTCATTGCCTTACAGCCATTCCGCTGCTGTGACTTCCTATGCAAACAGTGCCTCTTTGTCCACACCATTAAGTAACACAGGGCTTGTT CAACTTTCTCAGTCTCCACACTCCGTCCCTGGGGGACCCCAAGCACAAGCTACCATGACCCCACCCCCCAACCTGACTCCTCCTCCAATG AATCTGCCGCCGCCTCTTTTGCAACGGAACATGGCTGCATCAAATATTGGCATCTCTCACAGCCAAAGACTGCAAACCCAGATTGCCAGC AAGGGCCACATCTCCATGAGAACCAAGTCAGCGTCTCTGTCACCAGCCGCTGCCACCCATCAGTCACAAATCTATGGGCGCTCCCAGACT GTAGCCATGCAGGGTCCTGCACGGACTTTAACGATGCAAAGAGGCATGAACATGAGTGTGAACCTGATGCCAGCGCCAGCCTACAATGTC AACTCTGTGAACATGAACATGAACACTCTCAACGCCATGAATGGGTACAGCATGTCCCAGCCAATGATGAACAGTGGCTACCACAGCAAT CATGGCTATATGAATCAAACGCCCCAATACCCTATGCAGATGCAGATGGGCATGATGGGCACCCAGCCATATGCCCAGCAGCCAATGCAG ACCCCACCCCACGGTAACATGATGTACACGGCCCCCGGACATCACGGCTACATGAACACAGGCATGTCCAAACAGTCTCTCAATGGCTCC TACATGAGAAGGTAGACAACGTGGGCAGTCCACAAAACCTACGGGGCATCACTATTGGATTGATCTGCACAAATACCTTTGAAGAGTACG >79103_79103_1_SAMD8-KAT6B_SAMD8_chr10_76928416_ENST00000372687_KAT6B_chr10_76788247_ENST00000372714_length(amino acids)=873AA_BP=21 MLYHVCDLPLRARTTPAVYWKDNMNDDSSNLKEGSKDNPEPLKCKQVWPKGTKRGLSKWRQNKERKTGFKLNLYTPPETPMEPDEQVTVE EQKETSEGKTSPSPIRIEEEVKETGEALLPQEENRREETCAPVSPNTSPGEKPEDDLIKPEEEEEEEEEEEEEEEEEEGEEEEGGGNVEK DPDGAKSQEKEEPEISTEKEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKEELPRESFKEVLENQETFLDLNVQPGHSN PEVLMDCGVDLTASCNSEPKELAGDPEAVPESDEEPPPGEQAQKQDQKNSKEVDTEFKEGNPATMEIDSETVQAVQSLTQESSEQDDTFQ DCAETQEACRSLQNYTRADQSPQIATTLDDCQQSDHSSPVSSVHSHPGQSVRSVNSPSVPALENSYAQISPDQSAISVPSLQNMETSPMM DVPSVSDHSQQVVDSGFSDLGSIESTTENYENPSSYDSTMGGSICGNGSSQNSCSYSNLTSSSLTQSSCAVTQQMSNISGSCSMLQQTSI SSPPTCSVKSPQGCVVERPPSSSQQLAQCSMAANFTPPMQLAEIPETSNANIGLYERMGQSDFGAGHYPQPSATFSLAKLQQLTNTLIDH SLPYSHSAAVTSYANSASLSTPLSNTGLVQLSQSPHSVPGGPQAQATMTPPPNLTPPPMNLPPPLLQRNMAASNIGISHSQRLQTQIASK GHISMRTKSASLSPAAATHQSQIYGRSQTVAMQGPARTLTMQRGMNMSVNLMPAPAYNVNSVNMNMNTLNAMNGYSMSQPMMNSGYHSNH -------------------------------------------------------------- >79103_79103_2_SAMD8-KAT6B_SAMD8_chr10_76928416_ENST00000372690_KAT6B_chr10_76788247_ENST00000372714_length(transcript)=3761nt_BP=1059nt GGTAGGGCCCCGCCTCCCGAAGCAGTTCCGGTTCGGCTCCGAGCAGCTGCCGGCGCGGCGGTGACGGCGGCCGGGACGATGCCTGCGCGC AGTCGCCACCGCCCCCGCCTCCACTCCGGCTCCCCGCCCCGGGCTCCGCCCCCGCCGCTTGAGGCGCTTCACTCCGGCGAGGCGGGGAGG GCCCCGGACTCCGACGGCGGCTCGGACGCCGACTCGGAGGTGGGTCCGGGGAGCCCGACTCGGACCGCGGAGGCAGCGGAGGAGGAAATG GCAGGTCCTAATCAACTCTGCATTCGCCGCTGGACTACCAAGCATGTAGCTGTGTGGCTGAAGGATGAAGGCTTTTTTGAATATGTGGAC ATTTTATGCAATAAGCACCGACTTGATGGAATCACATTGCTAACATTGACTGAATATGATCTCCGGTCTCCTCCTCTGGAAATCAAAGTC TTAGGGGACATTAAAAGGTTAATGCTCTCAGTCCGAAAATTGCAGAAAATACATATTGATGTTTTAGAAGAGATGGGCTACAACAGTGAC AGTCCCATGGGTTCCATGACCCCTTTCATCAGTGCTCTTCAGAGTACAGACTGGCTCTGTAATGGGGAGCTTTCCCATGACTGTGACGGA CCCATAACTGACTTGAATTCTGATCAGTACCAGTACATGAATGGTAAAAACAAACATTCTGTTCGAAGATTGGACCCAGAATACTGGAAG ACTATACTGAGTTGTATATATGTTTTTATAGTATTTGGATTTACATCTTTCATTATGGTTATAGTCCATGAGCGAGTGCCTGACATGCAG ACCTATCCACCACTCCCAGATATATTCTTAGACAGCGTTCCTAGAATCCCATGGGCCTTTGCCATGACGGAAGTATGTGGCATGATTCTG TGCTATATTTGGCTCCTGGTTCTTCTTCTTCACAAGCACAGGTCAATACTTCTGCGAAGGCTCTGTAGTCTGATGGGAACTGTATTCTTG CTTCGCTGCTTTACCATGTTTGTGACCTCCCTCTCCGTGCCAGGACAACACCTGCAGTGTACTGGAAAGACAATATGAATGATGATTCAA GTAACTTGAAAGAAGGCAGTAAAGACAATCCCGAACCTCTAAAGTGCAAACAAGTGTGGCCAAAAGGAACAAAGCGCGGTCTATCTAAGT GGAGGCAAAACAAAGAGAGGAAGACCGGATTTAAACTGAATTTGTACACCCCGCCAGAAACACCCATGGAGCCTGACGAGCAGGTAACAG TGGAAGAACAGAAGGAGACTTCAGAAGGAAAAACCAGCCCCAGTCCCATCAGGATTGAGGAGGAGGTCAAGGAAACTGGGGAAGCCCTGT TGCCTCAAGAGGAAAACAGAAGGGAAGAAACATGTGCCCCTGTAAGTCCAAACACATCACCAGGTGAAAAACCAGAAGATGATCTCATCA AACCTGAGGAAGAGGAAGAGGAGGAGGAGGAGGAAGAGGAAGAAGAGGAAGAAGAGGAAGGGGAAGAAGAAGAAGGAGGAGGAAATGTAG AAAAAGATCCAGATGGTGCTAAAAGCCAAGAAAAAGAGGAACCAGAAATCTCCACGGAAAAAGAAGACTCTGCACGTTTGGATGATCACG AAGAGGAGGAGGAAGAGGATGAAGAGCCATCCCACAACGAGGACCATGATGCCGATGACGAGGATGACAGCCACATGGAGTCTGCCGAAG TGGAGAAGGAAGAGCTGCCCAGAGAAAGCTTCAAAGAAGTACTGGAAAACCAGGAGACTTTTTTAGACCTTAATGTGCAGCCTGGTCACT CGAACCCAGAGGTCTTAATGGACTGTGGCGTCGACCTGACAGCTTCTTGTAACAGTGAGCCCAAGGAGCTTGCTGGGGACCCTGAAGCTG TACCCGAATCTGACGAGGAGCCACCCCCAGGAGAACAGGCACAGAAGCAGGACCAAAAGAACAGCAAGGAAGTCGATACAGAGTTCAAAG AGGGAAACCCAGCAACCATGGAAATCGACTCTGAGACTGTCCAGGCCGTTCAGTCTTTGACCCAGGAGAGCAGCGAACAGGACGACACCT TTCAGGATTGTGCCGAGACTCAAGAGGCCTGTAGAAGCCTACAGAACTACACCCGTGCAGACCAAAGTCCACAGATTGCCACCACGCTCG ACGATTGCCAACAGTCGGACCACAGTAGCCCAGTTTCATCCGTCCACTCCCATCCTGGCCAGTCCGTACGTTCTGTCAACAGCCCAAGTG TCCCTGCTCTGGAAAACAGCTACGCCCAAATCAGCCCAGATCAAAGTGCCATCTCAGTGCCATCTCTGCAGAACATGGAAACCAGTCCCA TGATGGATGTCCCATCAGTTTCAGATCATTCACAGCAAGTCGTAGACAGTGGATTTAGTGACCTGGGCAGTATCGAGAGCACAACTGAGA ACTACGAAAACCCAAGCAGCTACGATTCTACTATGGGAGGCAGCATCTGTGGAAACGGCTCTTCACAGAACAGCTGCTCCTATAGCAACC TCACCTCCAGCAGTCTGACACAGAGCAGCTGTGCTGTCACCCAGCAGATGTCCAACATCAGCGGGAGCTGCAGCATGCTGCAGCAAACCA GCATCAGCTCCCCTCCGACCTGCAGCGTCAAGTCTCCTCAAGGCTGTGTGGTGGAGAGGCCTCCGAGCAGCAGCCAGCAGCTGGCTCAGT GCAGCATGGCTGCTAACTTCACCCCACCCATGCAGCTGGCTGAAATCCCCGAGACGAGCAACGCCAACATTGGCTTATACGAGCGAATGG GTCAGAGTGATTTTGGGGCTGGGCATTACCCGCAGCCGTCAGCCACCTTCAGCCTTGCCAAACTGCAGCAGTTAACTAATACACTTATTG ATCATTCATTGCCTTACAGCCATTCCGCTGCTGTGACTTCCTATGCAAACAGTGCCTCTTTGTCCACACCATTAAGTAACACAGGGCTTG TTCAACTTTCTCAGTCTCCACACTCCGTCCCTGGGGGACCCCAAGCACAAGCTACCATGACCCCACCCCCCAACCTGACTCCTCCTCCAA TGAATCTGCCGCCGCCTCTTTTGCAACGGAACATGGCTGCATCAAATATTGGCATCTCTCACAGCCAAAGACTGCAAACCCAGATTGCCA GCAAGGGCCACATCTCCATGAGAACCAAGTCAGCGTCTCTGTCACCAGCCGCTGCCACCCATCAGTCACAAATCTATGGGCGCTCCCAGA CTGTAGCCATGCAGGGTCCTGCACGGACTTTAACGATGCAAAGAGGCATGAACATGAGTGTGAACCTGATGCCAGCGCCAGCCTACAATG TCAACTCTGTGAACATGAACATGAACACTCTCAACGCCATGAATGGGTACAGCATGTCCCAGCCAATGATGAACAGTGGCTACCACAGCA ATCATGGCTATATGAATCAAACGCCCCAATACCCTATGCAGATGCAGATGGGCATGATGGGCACCCAGCCATATGCCCAGCAGCCAATGC AGACCCCACCCCACGGTAACATGATGTACACGGCCCCCGGACATCACGGCTACATGAACACAGGCATGTCCAAACAGTCTCTCAATGGCT CCTACATGAGAAGGTAGACAACGTGGGCAGTCCACAAAACCTACGGGGCATCACTATTGGATTGATCTGCACAAATACCTTTGAAGAGTA >79103_79103_2_SAMD8-KAT6B_SAMD8_chr10_76928416_ENST00000372690_KAT6B_chr10_76788247_ENST00000372714_length(amino acids)=873AA_BP=21 MLYHVCDLPLRARTTPAVYWKDNMNDDSSNLKEGSKDNPEPLKCKQVWPKGTKRGLSKWRQNKERKTGFKLNLYTPPETPMEPDEQVTVE EQKETSEGKTSPSPIRIEEEVKETGEALLPQEENRREETCAPVSPNTSPGEKPEDDLIKPEEEEEEEEEEEEEEEEEEGEEEEGGGNVEK DPDGAKSQEKEEPEISTEKEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKEELPRESFKEVLENQETFLDLNVQPGHSN PEVLMDCGVDLTASCNSEPKELAGDPEAVPESDEEPPPGEQAQKQDQKNSKEVDTEFKEGNPATMEIDSETVQAVQSLTQESSEQDDTFQ DCAETQEACRSLQNYTRADQSPQIATTLDDCQQSDHSSPVSSVHSHPGQSVRSVNSPSVPALENSYAQISPDQSAISVPSLQNMETSPMM DVPSVSDHSQQVVDSGFSDLGSIESTTENYENPSSYDSTMGGSICGNGSSQNSCSYSNLTSSSLTQSSCAVTQQMSNISGSCSMLQQTSI SSPPTCSVKSPQGCVVERPPSSSQQLAQCSMAANFTPPMQLAEIPETSNANIGLYERMGQSDFGAGHYPQPSATFSLAKLQQLTNTLIDH SLPYSHSAAVTSYANSASLSTPLSNTGLVQLSQSPHSVPGGPQAQATMTPPPNLTPPPMNLPPPLLQRNMAASNIGISHSQRLQTQIASK GHISMRTKSASLSPAAATHQSQIYGRSQTVAMQGPARTLTMQRGMNMSVNLMPAPAYNVNSVNMNMNTLNAMNGYSMSQPMMNSGYHSNH -------------------------------------------------------------- >79103_79103_3_SAMD8-KAT6B_SAMD8_chr10_76928416_ENST00000542569_KAT6B_chr10_76788247_ENST00000372714_length(transcript)=3597nt_BP=895nt CGGCGAGGCGGGGAGGGCCCCGGACTCCGACGGCGGCTCGGACGCCGACTCGGAGGTGGGTCCGGGGAGCCCGACTCGGACCGCGGAGGC AGCGGAGGAGGAAATGGCAGGTCCTAATCAACTCTGCATTCGCCGCTGGACTACCAAGCATGTAGCTGTGTGGCTGAAGGATGAAGGCTT TTTTGAATATGTGGACATTTTATGCAATAAGCACCGACTTGATGGAATCACATTGCTAACATTGACTGAATATGATCTCCGGTCTCCTCC TCTGGAAATCAAAGTCTTAGGGGACATTAAAAGGTTAATGCTCTCAGTCCGAAAATTGCAGAAAATACATATTGATGTTTTAGAAGAGAT GGGCTACAACAGTGACAGTCCCATGGGTTCCATGACCCCTTTCATCAGTGCTCTTCAGAGTACAGACTGGCTCTGTAATGGGGAGCTTTC CCATGACTGTGACGGACCCATAACTGACTTGAATTCTGATCAGTACCAGTACATGAATGGTAAAAACAAACATTCTGTTCGAAGATTGGA CCCAGAATACTGGAAGACTATACTGAGTTGTATATATGTTTTTATAGTATTTGGATTTACATCTTTCATTATGGTTATAGTCCATGAGCG AGTGCCTGACATGCAGACCTATCCACCACTCCCAGATATATTCTTAGACAGCGTTCCTAGAATCCCATGGGCCTTTGCCATGACGGAAGT ATGTGGCATGATTCTGTGCTATATTTGGCTCCTGGTTCTTCTTCTTCACAAGCACAGGTCAATACTTCTGCGAAGGCTCTGTAGTCTGAT GGGAACTGTATTCTTGCTTCGCTGCTTTACCATGTTTGTGACCTCCCTCTCCGTGCCAGGACAACACCTGCAGTGTACTGGAAAGACAAT ATGAATGATGATTCAAGTAACTTGAAAGAAGGCAGTAAAGACAATCCCGAACCTCTAAAGTGCAAACAAGTGTGGCCAAAAGGAACAAAG CGCGGTCTATCTAAGTGGAGGCAAAACAAAGAGAGGAAGACCGGATTTAAACTGAATTTGTACACCCCGCCAGAAACACCCATGGAGCCT GACGAGCAGGTAACAGTGGAAGAACAGAAGGAGACTTCAGAAGGAAAAACCAGCCCCAGTCCCATCAGGATTGAGGAGGAGGTCAAGGAA ACTGGGGAAGCCCTGTTGCCTCAAGAGGAAAACAGAAGGGAAGAAACATGTGCCCCTGTAAGTCCAAACACATCACCAGGTGAAAAACCA GAAGATGATCTCATCAAACCTGAGGAAGAGGAAGAGGAGGAGGAGGAGGAAGAGGAAGAAGAGGAAGAAGAGGAAGGGGAAGAAGAAGAA GGAGGAGGAAATGTAGAAAAAGATCCAGATGGTGCTAAAAGCCAAGAAAAAGAGGAACCAGAAATCTCCACGGAAAAAGAAGACTCTGCA CGTTTGGATGATCACGAAGAGGAGGAGGAAGAGGATGAAGAGCCATCCCACAACGAGGACCATGATGCCGATGACGAGGATGACAGCCAC ATGGAGTCTGCCGAAGTGGAGAAGGAAGAGCTGCCCAGAGAAAGCTTCAAAGAAGTACTGGAAAACCAGGAGACTTTTTTAGACCTTAAT GTGCAGCCTGGTCACTCGAACCCAGAGGTCTTAATGGACTGTGGCGTCGACCTGACAGCTTCTTGTAACAGTGAGCCCAAGGAGCTTGCT GGGGACCCTGAAGCTGTACCCGAATCTGACGAGGAGCCACCCCCAGGAGAACAGGCACAGAAGCAGGACCAAAAGAACAGCAAGGAAGTC GATACAGAGTTCAAAGAGGGAAACCCAGCAACCATGGAAATCGACTCTGAGACTGTCCAGGCCGTTCAGTCTTTGACCCAGGAGAGCAGC GAACAGGACGACACCTTTCAGGATTGTGCCGAGACTCAAGAGGCCTGTAGAAGCCTACAGAACTACACCCGTGCAGACCAAAGTCCACAG ATTGCCACCACGCTCGACGATTGCCAACAGTCGGACCACAGTAGCCCAGTTTCATCCGTCCACTCCCATCCTGGCCAGTCCGTACGTTCT GTCAACAGCCCAAGTGTCCCTGCTCTGGAAAACAGCTACGCCCAAATCAGCCCAGATCAAAGTGCCATCTCAGTGCCATCTCTGCAGAAC ATGGAAACCAGTCCCATGATGGATGTCCCATCAGTTTCAGATCATTCACAGCAAGTCGTAGACAGTGGATTTAGTGACCTGGGCAGTATC GAGAGCACAACTGAGAACTACGAAAACCCAAGCAGCTACGATTCTACTATGGGAGGCAGCATCTGTGGAAACGGCTCTTCACAGAACAGC TGCTCCTATAGCAACCTCACCTCCAGCAGTCTGACACAGAGCAGCTGTGCTGTCACCCAGCAGATGTCCAACATCAGCGGGAGCTGCAGC ATGCTGCAGCAAACCAGCATCAGCTCCCCTCCGACCTGCAGCGTCAAGTCTCCTCAAGGCTGTGTGGTGGAGAGGCCTCCGAGCAGCAGC CAGCAGCTGGCTCAGTGCAGCATGGCTGCTAACTTCACCCCACCCATGCAGCTGGCTGAAATCCCCGAGACGAGCAACGCCAACATTGGC TTATACGAGCGAATGGGTCAGAGTGATTTTGGGGCTGGGCATTACCCGCAGCCGTCAGCCACCTTCAGCCTTGCCAAACTGCAGCAGTTA ACTAATACACTTATTGATCATTCATTGCCTTACAGCCATTCCGCTGCTGTGACTTCCTATGCAAACAGTGCCTCTTTGTCCACACCATTA AGTAACACAGGGCTTGTTCAACTTTCTCAGTCTCCACACTCCGTCCCTGGGGGACCCCAAGCACAAGCTACCATGACCCCACCCCCCAAC CTGACTCCTCCTCCAATGAATCTGCCGCCGCCTCTTTTGCAACGGAACATGGCTGCATCAAATATTGGCATCTCTCACAGCCAAAGACTG CAAACCCAGATTGCCAGCAAGGGCCACATCTCCATGAGAACCAAGTCAGCGTCTCTGTCACCAGCCGCTGCCACCCATCAGTCACAAATC TATGGGCGCTCCCAGACTGTAGCCATGCAGGGTCCTGCACGGACTTTAACGATGCAAAGAGGCATGAACATGAGTGTGAACCTGATGCCA GCGCCAGCCTACAATGTCAACTCTGTGAACATGAACATGAACACTCTCAACGCCATGAATGGGTACAGCATGTCCCAGCCAATGATGAAC AGTGGCTACCACAGCAATCATGGCTATATGAATCAAACGCCCCAATACCCTATGCAGATGCAGATGGGCATGATGGGCACCCAGCCATAT GCCCAGCAGCCAATGCAGACCCCACCCCACGGTAACATGATGTACACGGCCCCCGGACATCACGGCTACATGAACACAGGCATGTCCAAA CAGTCTCTCAATGGCTCCTACATGAGAAGGTAGACAACGTGGGCAGTCCACAAAACCTACGGGGCATCACTATTGGATTGATCTGCACAA >79103_79103_3_SAMD8-KAT6B_SAMD8_chr10_76928416_ENST00000542569_KAT6B_chr10_76788247_ENST00000372714_length(amino acids)=873AA_BP=21 MLYHVCDLPLRARTTPAVYWKDNMNDDSSNLKEGSKDNPEPLKCKQVWPKGTKRGLSKWRQNKERKTGFKLNLYTPPETPMEPDEQVTVE EQKETSEGKTSPSPIRIEEEVKETGEALLPQEENRREETCAPVSPNTSPGEKPEDDLIKPEEEEEEEEEEEEEEEEEEGEEEEGGGNVEK DPDGAKSQEKEEPEISTEKEDSARLDDHEEEEEEDEEPSHNEDHDADDEDDSHMESAEVEKEELPRESFKEVLENQETFLDLNVQPGHSN PEVLMDCGVDLTASCNSEPKELAGDPEAVPESDEEPPPGEQAQKQDQKNSKEVDTEFKEGNPATMEIDSETVQAVQSLTQESSEQDDTFQ DCAETQEACRSLQNYTRADQSPQIATTLDDCQQSDHSSPVSSVHSHPGQSVRSVNSPSVPALENSYAQISPDQSAISVPSLQNMETSPMM DVPSVSDHSQQVVDSGFSDLGSIESTTENYENPSSYDSTMGGSICGNGSSQNSCSYSNLTSSSLTQSSCAVTQQMSNISGSCSMLQQTSI SSPPTCSVKSPQGCVVERPPSSSQQLAQCSMAANFTPPMQLAEIPETSNANIGLYERMGQSDFGAGHYPQPSATFSLAKLQQLTNTLIDH SLPYSHSAAVTSYANSASLSTPLSNTGLVQLSQSPHSVPGGPQAQATMTPPPNLTPPPMNLPPPLLQRNMAASNIGISHSQRLQTQIASK GHISMRTKSASLSPAAATHQSQIYGRSQTVAMQGPARTLTMQRGMNMSVNLMPAPAYNVNSVNMNMNTLNAMNGYSMSQPMMNSGYHSNH -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SAMD8-KAT6B |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 1560_2073 | 1221.3333333333333 | 2074.0 | RUNX1 and RUNX2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 1560_2073 | 1038.3333333333333 | 1891.0 | RUNX1 and RUNX2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 1560_2073 | 929.3333333333334 | 1782.0 | RUNX1 and RUNX2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 1560_2073 | 929.3333333333334 | 1782.0 | RUNX1 and RUNX2 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 1560_2073 | 929.3333333333334 | 1782.0 | RUNX1 and RUNX2 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000287239 | 16 | 18 | 752_1008 | 1221.3333333333333 | 2074.0 | BRPF1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372711 | 15 | 17 | 752_1008 | 1038.3333333333333 | 1891.0 | BRPF1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372714 | 15 | 17 | 752_1008 | 929.3333333333334 | 1782.0 | BRPF1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372724 | 16 | 18 | 752_1008 | 929.3333333333334 | 1782.0 | BRPF1 | |
| Tgene | KAT6B | chr10:76928416 | chr10:76788247 | ENST00000372725 | 15 | 17 | 752_1008 | 929.3333333333334 | 1782.0 | BRPF1 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SAMD8-KAT6B |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SAMD8-KAT6B |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
