
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SART3-CORO1C (FusionGDB2 ID:79219) |
Fusion Gene Summary for SART3-CORO1C |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SART3-CORO1C | Fusion gene ID: 79219 | Hgene | Tgene | Gene symbol | SART3 | CORO1C | Gene ID | 9733 | 23603 |
| Gene name | spliceosome associated factor 3, U4/U6 recycling protein | coronin 1C | |
| Synonyms | DSAP1|P100|RP11-13G14|TIP110|p110|p110(nrb) | HCRNN4 | |
| Cytomap | 12q23.3 | 12q24.11 | |
| Type of gene | protein-coding | protein-coding | |
| Description | squamous cell carcinoma antigen recognized by T-cells 3HIV-1 Tat-interacting protein of 110kDaPRP24 homologSART-3hSART-3p110 nuclear RNA-binding proteinsquamous cell carcinoma antigen recognized by T cells 3tat-interacting protein of 110 kDa | coronin-1Ccoronin, actin binding protein, 1Ccoronin-3 | |
| Modification date | 20200313 | 20200322 | |
| UniProtAcc | . | Q9ULV4 | |
| Ensembl transtripts involved in fusion gene | ENST00000228284, ENST00000431469, ENST00000552221, ENST00000546611, | ENST00000420959, ENST00000421578, ENST00000549384, ENST00000549772, ENST00000261401, ENST00000541050, | |
| Fusion gene scores | * DoF score | 9 X 11 X 6=594 | 16 X 9 X 8=1152 |
| # samples | 11 | 17 | |
| ** MAII score | log2(11/594*10)=-2.43295940727611 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(17/1152*10)=-2.76053406530461 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SART3 [Title/Abstract] AND CORO1C [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SART3(108954619)-CORO1C(109041298), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | SART3-CORO1C seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. SART3-CORO1C seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SART3 | GO:0000244 | spliceosomal tri-snRNP complex assembly | 12032085|20595234 |
| Hgene | SART3 | GO:0000387 | spliceosomal snRNP assembly | 14749385|15314151 |
| Hgene | SART3 | GO:0000398 | mRNA splicing, via spliceosome | 15314151 |
| Hgene | SART3 | GO:0006334 | nucleosome assembly | 24526689 |
| Hgene | SART3 | GO:0010468 | regulation of gene expression | 21447833 |
| Hgene | SART3 | GO:1903586 | positive regulation of histone deubiquitination | 24526689 |
| Tgene | CORO1C | GO:0016197 | endosomal transport | 30220460 |
| Tgene | CORO1C | GO:0090148 | membrane fission | 30220460 |
| Tgene | CORO1C | GO:0097750 | endosome membrane tubulation | 30220460 |
 Fusion gene breakpoints across SART3 (5'-gene) Fusion gene breakpoints across SART3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
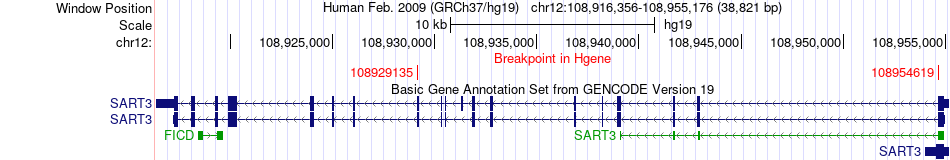 |
 Fusion gene breakpoints across CORO1C (3'-gene) Fusion gene breakpoints across CORO1C (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
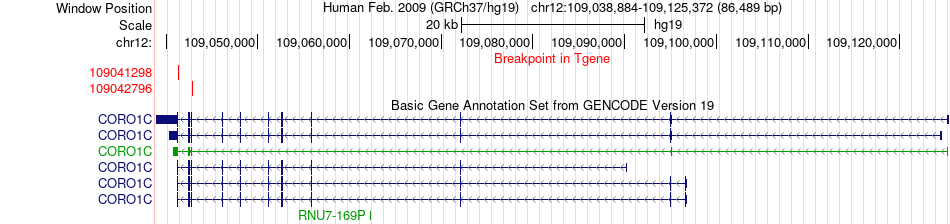 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUAD | TCGA-44-7669-01A | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| ChimerDB4 | LUAD | TCGA-MP-A4SW-01A | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
Top |
Fusion Gene ORF analysis for SART3-CORO1C |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000228284 | ENST00000420959 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000228284 | ENST00000421578 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000228284 | ENST00000549384 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000228284 | ENST00000549772 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000431469 | ENST00000420959 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000431469 | ENST00000421578 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000431469 | ENST00000549384 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-5UTR | ENST00000431469 | ENST00000549772 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5CDS-intron | ENST00000228284 | ENST00000420959 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000228284 | ENST00000421578 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000228284 | ENST00000549384 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000228284 | ENST00000549772 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000431469 | ENST00000420959 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000431469 | ENST00000421578 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000431469 | ENST00000549384 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5CDS-intron | ENST00000431469 | ENST00000549772 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| 5UTR-3CDS | ENST00000552221 | ENST00000261401 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5UTR-3CDS | ENST00000552221 | ENST00000541050 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5UTR-5UTR | ENST00000552221 | ENST00000420959 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5UTR-5UTR | ENST00000552221 | ENST00000421578 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5UTR-5UTR | ENST00000552221 | ENST00000549384 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| 5UTR-5UTR | ENST00000552221 | ENST00000549772 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| Frame-shift | ENST00000228284 | ENST00000261401 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| Frame-shift | ENST00000228284 | ENST00000541050 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| Frame-shift | ENST00000431469 | ENST00000261401 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| Frame-shift | ENST00000431469 | ENST00000541050 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| In-frame | ENST00000228284 | ENST00000261401 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| In-frame | ENST00000228284 | ENST00000541050 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| In-frame | ENST00000431469 | ENST00000261401 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| In-frame | ENST00000431469 | ENST00000541050 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-3CDS | ENST00000546611 | ENST00000261401 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-3CDS | ENST00000546611 | ENST00000261401 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| intron-3CDS | ENST00000546611 | ENST00000541050 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-3CDS | ENST00000546611 | ENST00000541050 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| intron-3CDS | ENST00000552221 | ENST00000261401 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-3CDS | ENST00000552221 | ENST00000541050 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-5UTR | ENST00000546611 | ENST00000420959 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| intron-5UTR | ENST00000546611 | ENST00000421578 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| intron-5UTR | ENST00000546611 | ENST00000549384 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| intron-5UTR | ENST00000546611 | ENST00000549772 | SART3 | chr12 | 108954619 | - | CORO1C | chr12 | 109041298 | - |
| intron-intron | ENST00000546611 | ENST00000420959 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000546611 | ENST00000421578 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000546611 | ENST00000549384 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000546611 | ENST00000549772 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000552221 | ENST00000420959 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000552221 | ENST00000421578 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000552221 | ENST00000549384 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
| intron-intron | ENST00000552221 | ENST00000549772 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000228284 | SART3 | chr12 | 108929135 | - | ENST00000261401 | CORO1C | chr12 | 109042796 | - | 4509 | 1791 | 235 | 2214 | 659 |
| ENST00000228284 | SART3 | chr12 | 108929135 | - | ENST00000541050 | CORO1C | chr12 | 109042796 | - | 3051 | 1791 | 235 | 2214 | 659 |
| ENST00000431469 | SART3 | chr12 | 108929135 | - | ENST00000261401 | CORO1C | chr12 | 109042796 | - | 4171 | 1453 | 5 | 1876 | 623 |
| ENST00000431469 | SART3 | chr12 | 108929135 | - | ENST00000541050 | CORO1C | chr12 | 109042796 | - | 2713 | 1453 | 5 | 1876 | 623 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000228284 | ENST00000261401 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - | 0.000401433 | 0.99959856 |
| ENST00000228284 | ENST00000541050 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - | 0.000879338 | 0.99912065 |
| ENST00000431469 | ENST00000261401 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - | 0.000527074 | 0.9994729 |
| ENST00000431469 | ENST00000541050 | SART3 | chr12 | 108929135 | - | CORO1C | chr12 | 109042796 | - | 0.001589011 | 0.99841094 |
Top |
Fusion Genomic Features for SART3-CORO1C |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
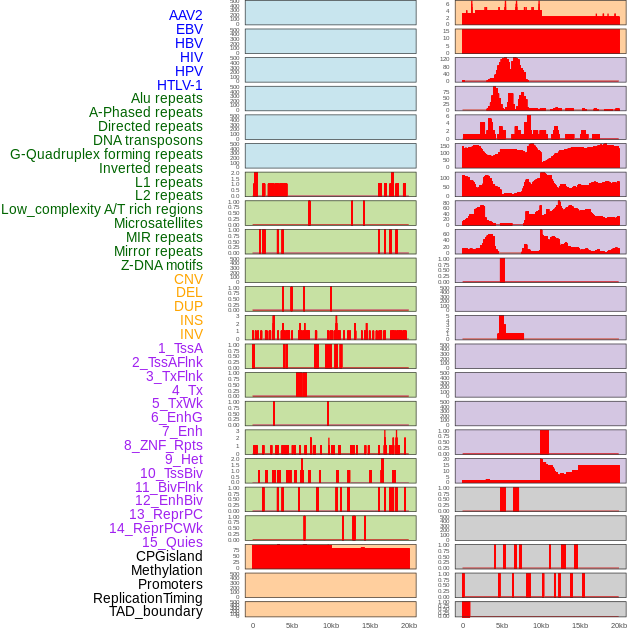 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for SART3-CORO1C |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:108954619/chr12:109041298) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CORO1C |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Plays a role in directed cell migration by regulating the activation and subcellular location of RAC1 (PubMed:25074804, PubMed:25925950). Increases the presence of activated RAC1 at the leading edge of migrating cells (PubMed:25074804, PubMed:25925950). Required for normal organization of the cytoskeleton, including the actin cytoskeleton, microtubules and the vimentin intermediate filaments (By similarity). Plays a role in endoplasmic reticulum-associated endosome fission: localizes to endosome membrane tubules and promotes recruitment of TMCC1, leading to recruitment of the endoplasmic reticulum to endosome tubules for fission (PubMed:30220460). Endosome membrane fission of early and late endosomes is essential to separate regions destined for lysosomal degradation from carriers to be recycled to the plasma membrane (PubMed:30220460). Required for normal cell proliferation, cell migration, and normal formation of lamellipodia (By similarity). Required for normal distribution of mitochondria within cells (By similarity). {ECO:0000250|UniProtKB:Q9WUM4, ECO:0000269|PubMed:25074804, ECO:0000269|PubMed:25925950, ECO:0000269|PubMed:30220460}.; FUNCTION: [Isoform 3]: Involved in myogenic differentiation. {ECO:0000269|PubMed:19651142}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 21_46 | 518 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 82_110 | 518 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 89_92 | 518 | 964.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 126_158 | 518 | 964.0 | Repeat | Note=HAT 1 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 164_195 | 518 | 964.0 | Repeat | Note=HAT 2 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 201_237 | 518 | 964.0 | Repeat | Note=HAT 3 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 242_275 | 518 | 964.0 | Repeat | Note=HAT 4 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 324_356 | 518 | 964.0 | Repeat | Note=HAT 5 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 359_391 | 518 | 964.0 | Repeat | Note=HAT 6 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 394_430 | 518 | 964.0 | Repeat | Note=HAT 7 |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 436_474 | 333 | 475.0 | Coiled coil | Ontology_term=ECO:0000269 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 436_474 | 386 | 528.0 | Coiled coil | Ontology_term=ECO:0000269 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 436_474 | 333 | 475.0 | Coiled coil | Ontology_term=ECO:0000269 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 436_474 | 339 | 481.0 | Coiled coil | Ontology_term=ECO:0000269 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 559_619 | 518 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 21_46 | 0 | 130.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 559_619 | 0 | 130.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 82_110 | 0 | 130.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 612_616 | 518 | 964.0 | Compositional bias | Note=Poly-Lys |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 612_616 | 0 | 130.0 | Compositional bias | Note=Poly-Lys |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 89_92 | 0 | 130.0 | Compositional bias | Note=Poly-Glu |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 704_782 | 518 | 964.0 | Domain | RRM 1 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 801_878 | 518 | 964.0 | Domain | RRM 2 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 704_782 | 0 | 130.0 | Domain | RRM 1 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 801_878 | 0 | 130.0 | Domain | RRM 2 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 601_617 | 518 | 964.0 | Motif | Nuclear localization signal |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 601_617 | 0 | 130.0 | Motif | Nuclear localization signal |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 537_953 | 518 | 964.0 | Region | Necessary and sufficient for U6 snRNA binding |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000228284 | - | 12 | 19 | 600_670 | 518 | 964.0 | Region | Required for nuclear localization |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 537_953 | 0 | 130.0 | Region | Necessary and sufficient for U6 snRNA binding |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 600_670 | 0 | 130.0 | Region | Required for nuclear localization |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 126_158 | 0 | 130.0 | Repeat | Note=HAT 1 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 164_195 | 0 | 130.0 | Repeat | Note=HAT 2 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 201_237 | 0 | 130.0 | Repeat | Note=HAT 3 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 242_275 | 0 | 130.0 | Repeat | Note=HAT 4 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 324_356 | 0 | 130.0 | Repeat | Note=HAT 5 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 359_391 | 0 | 130.0 | Repeat | Note=HAT 6 |
| Hgene | SART3 | chr12:108929135 | chr12:109042796 | ENST00000546611 | - | 1 | 1 | 394_430 | 0 | 130.0 | Repeat | Note=HAT 7 |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 128_168 | 333 | 475.0 | Repeat | WD 3 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 172_202 | 333 | 475.0 | Repeat | WD 4 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 215_249 | 333 | 475.0 | Repeat | WD 5 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 25_70 | 333 | 475.0 | Repeat | WD 1 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 263_303 | 333 | 475.0 | Repeat | WD 6 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000261401 | 7 | 11 | 78_118 | 333 | 475.0 | Repeat | WD 2 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 128_168 | 386 | 528.0 | Repeat | WD 3 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 172_202 | 386 | 528.0 | Repeat | WD 4 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 215_249 | 386 | 528.0 | Repeat | WD 5 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 25_70 | 386 | 528.0 | Repeat | WD 1 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 263_303 | 386 | 528.0 | Repeat | WD 6 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000420959 | 7 | 11 | 78_118 | 386 | 528.0 | Repeat | WD 2 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 128_168 | 333 | 475.0 | Repeat | WD 3 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 172_202 | 333 | 475.0 | Repeat | WD 4 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 215_249 | 333 | 475.0 | Repeat | WD 5 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 25_70 | 333 | 475.0 | Repeat | WD 1 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 263_303 | 333 | 475.0 | Repeat | WD 6 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000541050 | 7 | 11 | 78_118 | 333 | 475.0 | Repeat | WD 2 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 128_168 | 339 | 481.0 | Repeat | WD 3 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 172_202 | 339 | 481.0 | Repeat | WD 4 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 215_249 | 339 | 481.0 | Repeat | WD 5 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 25_70 | 339 | 481.0 | Repeat | WD 1 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 263_303 | 339 | 481.0 | Repeat | WD 6 | |
| Tgene | CORO1C | chr12:108929135 | chr12:109042796 | ENST00000549772 | 7 | 11 | 78_118 | 339 | 481.0 | Repeat | WD 2 |
Top |
Fusion Gene Sequence for SART3-CORO1C |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >79219_79219_1_SART3-CORO1C_SART3_chr12_108929135_ENST00000228284_CORO1C_chr12_109042796_ENST00000261401_length(transcript)=4509nt_BP=1791nt AGTTGTGACGGAACGCATGCCATTATAATGAGTTTTTATATCTGGTGGGAGCTTTTACAGGGTATAAAATAATGTGAATTTAGTGTATGT GGAGATTTTCCTGTTATCTAAGTTCCCAGTTTCTCTTTCAATGGCAAGAAAACGGGTTTTTGATAAGGCGCGCGCCGCGGGCAGCTTCTG CCTCAGCGGGTCTTCGGCGGCCGCGGGAAGACCCGAGAGTCATTAGAAGCGCAAGATGGCGACTGCGGCCGAAACCTCGGCTTCAGAACC CGAGGCTGAGTCCAAGGCTGGGCCCAAGGCTGACGGAGAGGAGGATGAGGTTAAGGCGGCTAGGACAAGGAGGAAGGTGTTATCGCGGGC TGTGGCCGCTGCGACATACAAGACCATGGGGCCAGCGTGGGATCAGCAGGAGGAAGGCGTGAGCGAGAGCGATGGGGATGAGTACGCCAT GGCTTCCTCCGCGGAGAGCTCCCCCGGGGAGTACGAGTGGGAATATGACGAAGAGGAGGAGAAAAACCAGCTGGAGATTGAGAGACTGGA GGAGCAGTTGTCTATCAACGTCTATGACTACAACTGCCATGTGGACTTGATCAGACTGCTCAGGCTGGAAGGGGAGCTTACCAAGGTGAG GATGGCCCGCCAGAAGATGAGTGAAATCTTTCCCTTGACTGAAGAGCTCTGGCTGGAGTGGCTGCATGACGAGATCAGCATGGCCCAGGA TGGCCTGGACAGAGAGCACGTGTATGACCTCTTTGAGAAAGCCGTGAAGGATTACATTTGTCCTAACATTTGGCTAGAGTATGGCCAGTA CTCAGTTGGTGGGATTGGTCAGAAAGGTGGCCTTGAGAAAGTTCGCTCCGTGTTTGAAAGGGCTCTCTCGTCTGTTGGTTTACATATGAC CAAAGGACTCGCCCTCTGGGAGGCTTACCGAGAGTTTGAAAGTGCGATTGTGGAAGCTGCTCGGCTTGAGAAAGTCCACAGTCTTTTCCG GCGACAGTTGGCGATCCCACTCTATGATATGGAGGCCACATTTGCAGAGTATGAAGAATGGTCAGAAGACCCAATACCAGAGTCAGTAAT TCAGAACTATAACAAAGCACTACAGCAGCTGGAGAAATATAAACCCTATGAAGAAGCACTGTTGCAGGCAGAGGCACCAAGGCTGGCAGA ATATCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCT TGTCCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTAT TAGAAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGTAAC CTTCGAGAAAGCTTTGAATGCCGGCTTCATCCAGGCCACTGATTATGTGGAGATTTGGCAGGCATACCTTGATTACCTGAGGAGAAGGGT TGATTTCAAACAAGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGA GCGTTTCAATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGC TCGGGAACTCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGATTCTTCAA ACTTCATGAGAGAAAGTGTGAACCTATTATTATGACTGTTCCCAGGAAGTCTGACCTTTTCCAAGATGACCTGTATCCTGACACAGCGGG GCCAGAGGCCGCGCTGGAGGCAGAAGAGTGGTTCGAAGGCAAGAATGCAGACCCAATCCTCATCTCCTTGAAGCACGGGTACATTCCAGG CAAAAACAGGGATCTCAAGGTGGTCAAGAAGAACATTCTGGATAGCAAGCCCACTGCAAACAAGAAGTGCGACCTGATCAGCATCCCCAA GAAAACCACAGACACGGCCAGTGTGCAAAATGAAGCCAAGTTGGATGAGATTTTAAAAGAGATCAAATCTATAAAAGACACAATCTGCAA TCAAGATGAGCGTATTTCCAAGTTAGAACAGCAGATGGCAAAGATAGCAGCCTGAAGGTCCCACCCCCACCCCTACAGAAAAAATGGGAG CAAGAACTTGTGCTTGGGAGCTGGTTATTGGTGTGGTCCTAGGGAGGGCGGAAAGGGAGGCACTGCCATTTGGAGACATTCCATTTCAGA TTTGTCAACCAGCGATAGGCCACATTCCAGTAAGAACTCAATTTGTCTCCCAAATTTGCAGAAACAAAACGTGATTTAAAAGCTGAGCTT TTTATCAGAAAGCTTTTTTGATGTTTTAAGTGTTATGTGACTTGTTGAACTTTTTAAAAAGTGCTACTTTTAAAATCCCAGATACTCTGA ATTTTAGAAAACAAACTAATTCTGATTGTGTCGTGCCCAAGTACCCTTTTTTTTTTAATGAATAGGGACCAATGCCACATTGCTTTTTAT ATTTCTTTCTTTTTTAATGTTGCCAAAACCAAAAGTAGCTTTGTTTTCCTTTGTATTTTGCTACTTTGCAGTATTTGTGTGTGTGGTTTT TTTTCCTTAATTTGAAAGGGACAGCACTGTGTATGTTTATAAACTAAATGAAGATAAGATATTATTTTGTATAAACATTCATCTGAGAAC AATCAAAGCAGTAGCCACATGGTGCTGGCTCCTTTGCAGCACAAACCTGGTCATTTTGATGACTGTACAACAGGAAGACTTGAAAAATCA CGTGGATTCATATTACCACCGCTCTCATTTCATGGAGTCTTCTGATCAAAAAAGCTCACGTCGTATTTCTTCTTTTCCTTTCTCTTTTCT AGAAATTGGGTGTTTGTACCAGAATGGAATTTTGCTTCTCGGTTATCCTGTGCTTCAGATGATTATAATCTAACCCAAACTAGCATGTGT TTCTGCAGTTTGTTACACACCTAGGATCATATTGCATTCATCACTTTAAACATCATGTTTCAGGTTTTGGTCAATACTTGACAAGGGTGC CCAGGACAGGAAGACGTGTACTGCTGAGTGTTCCTTCTTGCCCTTTTCAGCAGCTTGCCCAGCTCTTGAGTACAGTGGTGGGGACTAAAA ATGTGGGCATGTGGAGAGGGGTATTTGCCCTGGGTGATCCTGTTTCCCTGTGCTGTCCCCATGCTGTGTTGGAGGAGGAAGTGGCTCTCC TTCCACCAACAAAGCTCCTGCTCTACCCTCTTCCTCACATGTGCTGCGACCTCTCTCAGGGCTCCCCCAGCCATTCCTTCTTTCCTTCCT GCCTTTTAGCTCTAACCACATTAAGCTAAGACAAGGCCAGAGGGTGCGATTGAATGAGTATTGAGACTGAGGAGAATGATAGAGAGTGAA GCAGAAACAGGAGCACAGACCTCTGCTGTAGCTTTAATGCATACAAACATGTCCCTCCGCACAACTAACCTGCCCTGCCTCTCCATCTCT CACCAAGGCTGCGTCAAAGCACAGAGGCTCCCCGGACTCGGAGGGGGCCAGAGACTGAGCTCTGGTCACCTGTTCATTCCTCGGTTAGCT GGAACTTTGCCCCGTTTCCAGTTTCTTATAGTGCATGCTTGGGAAACAAGATTTAAGGAGCCTCTGTTTTGGAAGGGCTGTCTGTGATTG AACGTGAAATGTGTAGTGCCATTGGGACCACGAAGGGAATTCTTGCACATGCTCGTGCTGGTGTGGGCATGGGACTGGCTGGAAACGTCT GTATGCAGGGAGCCAGGGTGAGGGCAGAGTGTGGTGACAGCCGAACTTGGAGTAATGTCCGTGTAGAAAAAGGACCATGTTCTTATCCAG CCAATACTGGGAGTGCTGTCTCCACAATTTCAGGGCATCTGAATGTTTGATGTGGTTTTGTGTGTGTGTATGTATGTGTTTAATATTGAA GTGGATCATGAGATGTAAAGAAAACAATAATGGCAATGACTTATATTCAAATCTGTATTTGTTTCTTTATCAATGTAATCTGCTGAGGAC CTTTTGTCTAAGATTCAGTAGTGTTTTAAGGTTCTGATATCGAATTAATGAAGTAAAGTTGTTGATGGTGGTGAAACACCGTAGGGCATG TGGTTCAAAGAGAAGCAGGAGGGCAAGGGAAAGTTACCCTGATCTTAGTTTGTAGCTTATGACTTATTTAATGAATGGATGCCCAGCCAA GCTCAGAGTAGGCGCCCAAAGCATTGTGGATTATTTTCCTGTTTTGTCTTTTTTTTTTTTTTTTTTTAAGCCATGACATCCCAGAAGAGG ACAGTGAATTACTCCTAGGTCGGCTCTTATAGAGTGGCCATAGTGTTCTGTCAAAACACTTGCTTCCATTTTCAGAGATAAAAATCATTG >79219_79219_1_SART3-CORO1C_SART3_chr12_108929135_ENST00000228284_CORO1C_chr12_109042796_ENST00000261401_length(amino acids)=659AA_BP=516 MATAAETSASEPEAESKAGPKADGEEDEVKAARTRRKVLSRAVAAATYKTMGPAWDQQEEGVSESDGDEYAMASSAESSPGEYEWEYDEE EEKNQLEIERLEEQLSINVYDYNCHVDLIRLLRLEGELTKVRMARQKMSEIFPLTEELWLEWLHDEISMAQDGLDREHVYDLFEKAVKDY ICPNIWLEYGQYSVGGIGQKGGLEKVRSVFERALSSVGLHMTKGLALWEAYREFESAIVEAARLEKVHSLFRRQLAIPLYDMEATFAEYE EWSEDPIPESVIQNYNKALQQLEKYKPYEEALLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVPDLWIRYSQYLDRQLKV KDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISVTFEKALNAGFIQATDYVEIWQAYLDYLRRRVDFKQDSSKELEELRAAFT RALEYLKQEVEERFNESGDPSCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERFFKLHERKCEPIIMTVPRKSD LFQDDLYPDTAGPEAALEAEEWFEGKNADPILISLKHGYIPGKNRDLKVVKKNILDSKPTANKKCDLISIPKKTTDTASVQNEAKLDEIL -------------------------------------------------------------- >79219_79219_2_SART3-CORO1C_SART3_chr12_108929135_ENST00000228284_CORO1C_chr12_109042796_ENST00000541050_length(transcript)=3051nt_BP=1791nt AGTTGTGACGGAACGCATGCCATTATAATGAGTTTTTATATCTGGTGGGAGCTTTTACAGGGTATAAAATAATGTGAATTTAGTGTATGT GGAGATTTTCCTGTTATCTAAGTTCCCAGTTTCTCTTTCAATGGCAAGAAAACGGGTTTTTGATAAGGCGCGCGCCGCGGGCAGCTTCTG CCTCAGCGGGTCTTCGGCGGCCGCGGGAAGACCCGAGAGTCATTAGAAGCGCAAGATGGCGACTGCGGCCGAAACCTCGGCTTCAGAACC CGAGGCTGAGTCCAAGGCTGGGCCCAAGGCTGACGGAGAGGAGGATGAGGTTAAGGCGGCTAGGACAAGGAGGAAGGTGTTATCGCGGGC TGTGGCCGCTGCGACATACAAGACCATGGGGCCAGCGTGGGATCAGCAGGAGGAAGGCGTGAGCGAGAGCGATGGGGATGAGTACGCCAT GGCTTCCTCCGCGGAGAGCTCCCCCGGGGAGTACGAGTGGGAATATGACGAAGAGGAGGAGAAAAACCAGCTGGAGATTGAGAGACTGGA GGAGCAGTTGTCTATCAACGTCTATGACTACAACTGCCATGTGGACTTGATCAGACTGCTCAGGCTGGAAGGGGAGCTTACCAAGGTGAG GATGGCCCGCCAGAAGATGAGTGAAATCTTTCCCTTGACTGAAGAGCTCTGGCTGGAGTGGCTGCATGACGAGATCAGCATGGCCCAGGA TGGCCTGGACAGAGAGCACGTGTATGACCTCTTTGAGAAAGCCGTGAAGGATTACATTTGTCCTAACATTTGGCTAGAGTATGGCCAGTA CTCAGTTGGTGGGATTGGTCAGAAAGGTGGCCTTGAGAAAGTTCGCTCCGTGTTTGAAAGGGCTCTCTCGTCTGTTGGTTTACATATGAC CAAAGGACTCGCCCTCTGGGAGGCTTACCGAGAGTTTGAAAGTGCGATTGTGGAAGCTGCTCGGCTTGAGAAAGTCCACAGTCTTTTCCG GCGACAGTTGGCGATCCCACTCTATGATATGGAGGCCACATTTGCAGAGTATGAAGAATGGTCAGAAGACCCAATACCAGAGTCAGTAAT TCAGAACTATAACAAAGCACTACAGCAGCTGGAGAAATATAAACCCTATGAAGAAGCACTGTTGCAGGCAGAGGCACCAAGGCTGGCAGA ATATCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCT TGTCCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTAT TAGAAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGTAAC CTTCGAGAAAGCTTTGAATGCCGGCTTCATCCAGGCCACTGATTATGTGGAGATTTGGCAGGCATACCTTGATTACCTGAGGAGAAGGGT TGATTTCAAACAAGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGA GCGTTTCAATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGC TCGGGAACTCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGATTCTTCAA ACTTCATGAGAGAAAGTGTGAACCTATTATTATGACTGTTCCCAGGAAGTCTGACCTTTTCCAAGATGACCTGTATCCTGACACAGCGGG GCCAGAGGCCGCGCTGGAGGCAGAAGAGTGGTTCGAAGGCAAGAATGCAGACCCAATCCTCATCTCCTTGAAGCACGGGTACATTCCAGG CAAAAACAGGGATCTCAAGGTGGTCAAGAAGAACATTCTGGATAGCAAGCCCACTGCAAACAAGAAGTGCGACCTGATCAGCATCCCCAA GAAAACCACAGACACGGCCAGTGTGCAAAATGAAGCCAAGTTGGATGAGATTTTAAAAGAGATCAAATCTATAAAAGACACAATCTGCAA TCAAGATGAGCGTATTTCCAAGTTAGAACAGCAGATGGCAAAGATAGCAGCCTGAAGGTCCCACCCCCACCCCTACAGAAAAAATGGGAG CAAGAACTTGTGCTTGGGAGCTGGTTATTGGTGTGGTCCTAGGGAGGGCGGAAAGGGAGGCACTGCCATTTGGAGACATTCCATTTCAGA TTTGTCAACCAGCGATAGGCCACATTCCAGTAAGAACTCAATTTGTCTCCCAAATTTGCAGAAACAAAACGTGATTTAAAAGCTGAGCTT TTTATCAGAAAGCTTTTTTGATGTTTTAAGTGTTATGTGACTTGTTGAACTTTTTAAAAAGTGCTACTTTTAAAATCCCAGATACTCTGA ATTTTAGAAAACAAACTAATTCTGATTGTGTCGTGCCCAAGTACCCTTTTTTTTTTAATGAATAGGGACCAATGCCACATTGCTTTTTAT ATTTCTTTCTTTTTTAATGTTGCCAAAACCAAAAGTAGCTTTGTTTTCCTTTGTATTTTGCTACTTTGCAGTATTTGTGTGTGTGGTTTT TTTTCCTTAATTTGAAAGGGACAGCACTGTGTATGTTTATAAACTAAATGAAGATAAGATATTATTTTGTATAAACATTCATCTGAGAAC AATCAAAGCAGTAGCCACATGGTGCTGGCTCCTTTGCAGCACAAACCTGGTCATTTTGATGACTGTACAACAGGAAGACTTGAAAAATCA CGTGGATTCATATTACCACCGCTCTCATTTCATGGAGTCTTCTGATCAAAAAAGCTCACGTCGTATTTCTTCTTTTCCTTTCTCTTTTCT >79219_79219_2_SART3-CORO1C_SART3_chr12_108929135_ENST00000228284_CORO1C_chr12_109042796_ENST00000541050_length(amino acids)=659AA_BP=516 MATAAETSASEPEAESKAGPKADGEEDEVKAARTRRKVLSRAVAAATYKTMGPAWDQQEEGVSESDGDEYAMASSAESSPGEYEWEYDEE EEKNQLEIERLEEQLSINVYDYNCHVDLIRLLRLEGELTKVRMARQKMSEIFPLTEELWLEWLHDEISMAQDGLDREHVYDLFEKAVKDY ICPNIWLEYGQYSVGGIGQKGGLEKVRSVFERALSSVGLHMTKGLALWEAYREFESAIVEAARLEKVHSLFRRQLAIPLYDMEATFAEYE EWSEDPIPESVIQNYNKALQQLEKYKPYEEALLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVPDLWIRYSQYLDRQLKV KDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISVTFEKALNAGFIQATDYVEIWQAYLDYLRRRVDFKQDSSKELEELRAAFT RALEYLKQEVEERFNESGDPSCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERFFKLHERKCEPIIMTVPRKSD LFQDDLYPDTAGPEAALEAEEWFEGKNADPILISLKHGYIPGKNRDLKVVKKNILDSKPTANKKCDLISIPKKTTDTASVQNEAKLDEIL -------------------------------------------------------------- >79219_79219_3_SART3-CORO1C_SART3_chr12_108929135_ENST00000431469_CORO1C_chr12_109042796_ENST00000261401_length(transcript)=4171nt_BP=1453nt GCAAGATGGCGACTGCGGCCGAAACCTCGGCTTCAGAACCCGAGGCTGAGTCCAAGGCTGGGCCCAAGGCTGACGGAGAGGAGGATGAGG TTAAGGCGGCTAGGACAAGGAGGAAGGTGTTATCGCGGGCTGTGGCCGCTGCGACATACAAGACCATGGGGCCAGCGTGGGATCAGCAGG AGGAAGGCGTGAGCGAGAGCGATGGGGATGAGTACGCCATGGCTTCCTCCGCGGAGAGCTCCCCCGGGGAGTACGAGTGGGAATATGACG AAGAGGAGGAGAAAAACCAGCTGGAGATTGAGAGACTGGAGGAGCAGTTGTCTATCAACGTCTATGACTACAACTGCCATGTGGACTTGA TCAGACTGCTCAGGCTGGAAGGGGAGCTTACCAAGGTGAGGATGGCCCGCCAGAAGATGAGTGAAATCTTTCCCTTGACTGAAGAGCTCT GGCTGGAGTGGCTGCATGACGAGATCAGCATGGCCCAGGATGGCCTGGACAGAGAGCACGTGTATGACCTCTTTGAGAAAGCCGTGAAGG ATTACATTTGTCCTAACATTTGGCTAGAGTATGGCCAGTACTCAGTTGGTGGGATTGGTCAGAAAGGTGGCCTTGAGAAAGTTCGCTCCG TGTTTGAAAGGGCTCTCTCGTCTGTTGGTTTACATATGACCAAAGGACTCGCCCTCTGGGAGGCTTACCGAGAGTTTGAAAGTGCGATTG TGGAAGCTGCTCGGCTTGAGAAAGTCCACAGTCTTTTCCGGCGACAGTTGGCGATCCCACTCTATGATATGGAGGCCACATTTGCAGAGT ATGAAGAATGGTCAGAAGACCCAATACCAGAGTCAGTAATTCAGAACTATAACAAAGCACTACAGCAGCTGGAGAAATATAAACCCTATG AAGAAGCACTGTTGCAGGCAGAGGCACCAAGGCTGGCAGAATATCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCA TTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGTCCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGA AAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAGAAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCA TGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGT ATCTGAAGCAGGAGGTGGAAGAGCGTTTCAATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGAC TGTGCAATAACATGCAGAAAGCTCGGGAACTCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATT ACAACCTGGAAAGATTCTTCAAACTTCATGAGAGAAAGTGTGAACCTATTATTATGACTGTTCCCAGGAAGTCTGACCTTTTCCAAGATG ACCTGTATCCTGACACAGCGGGGCCAGAGGCCGCGCTGGAGGCAGAAGAGTGGTTCGAAGGCAAGAATGCAGACCCAATCCTCATCTCCT TGAAGCACGGGTACATTCCAGGCAAAAACAGGGATCTCAAGGTGGTCAAGAAGAACATTCTGGATAGCAAGCCCACTGCAAACAAGAAGT GCGACCTGATCAGCATCCCCAAGAAAACCACAGACACGGCCAGTGTGCAAAATGAAGCCAAGTTGGATGAGATTTTAAAAGAGATCAAAT CTATAAAAGACACAATCTGCAATCAAGATGAGCGTATTTCCAAGTTAGAACAGCAGATGGCAAAGATAGCAGCCTGAAGGTCCCACCCCC ACCCCTACAGAAAAAATGGGAGCAAGAACTTGTGCTTGGGAGCTGGTTATTGGTGTGGTCCTAGGGAGGGCGGAAAGGGAGGCACTGCCA TTTGGAGACATTCCATTTCAGATTTGTCAACCAGCGATAGGCCACATTCCAGTAAGAACTCAATTTGTCTCCCAAATTTGCAGAAACAAA ACGTGATTTAAAAGCTGAGCTTTTTATCAGAAAGCTTTTTTGATGTTTTAAGTGTTATGTGACTTGTTGAACTTTTTAAAAAGTGCTACT TTTAAAATCCCAGATACTCTGAATTTTAGAAAACAAACTAATTCTGATTGTGTCGTGCCCAAGTACCCTTTTTTTTTTAATGAATAGGGA CCAATGCCACATTGCTTTTTATATTTCTTTCTTTTTTAATGTTGCCAAAACCAAAAGTAGCTTTGTTTTCCTTTGTATTTTGCTACTTTG CAGTATTTGTGTGTGTGGTTTTTTTTCCTTAATTTGAAAGGGACAGCACTGTGTATGTTTATAAACTAAATGAAGATAAGATATTATTTT GTATAAACATTCATCTGAGAACAATCAAAGCAGTAGCCACATGGTGCTGGCTCCTTTGCAGCACAAACCTGGTCATTTTGATGACTGTAC AACAGGAAGACTTGAAAAATCACGTGGATTCATATTACCACCGCTCTCATTTCATGGAGTCTTCTGATCAAAAAAGCTCACGTCGTATTT CTTCTTTTCCTTTCTCTTTTCTAGAAATTGGGTGTTTGTACCAGAATGGAATTTTGCTTCTCGGTTATCCTGTGCTTCAGATGATTATAA TCTAACCCAAACTAGCATGTGTTTCTGCAGTTTGTTACACACCTAGGATCATATTGCATTCATCACTTTAAACATCATGTTTCAGGTTTT GGTCAATACTTGACAAGGGTGCCCAGGACAGGAAGACGTGTACTGCTGAGTGTTCCTTCTTGCCCTTTTCAGCAGCTTGCCCAGCTCTTG AGTACAGTGGTGGGGACTAAAAATGTGGGCATGTGGAGAGGGGTATTTGCCCTGGGTGATCCTGTTTCCCTGTGCTGTCCCCATGCTGTG TTGGAGGAGGAAGTGGCTCTCCTTCCACCAACAAAGCTCCTGCTCTACCCTCTTCCTCACATGTGCTGCGACCTCTCTCAGGGCTCCCCC AGCCATTCCTTCTTTCCTTCCTGCCTTTTAGCTCTAACCACATTAAGCTAAGACAAGGCCAGAGGGTGCGATTGAATGAGTATTGAGACT GAGGAGAATGATAGAGAGTGAAGCAGAAACAGGAGCACAGACCTCTGCTGTAGCTTTAATGCATACAAACATGTCCCTCCGCACAACTAA CCTGCCCTGCCTCTCCATCTCTCACCAAGGCTGCGTCAAAGCACAGAGGCTCCCCGGACTCGGAGGGGGCCAGAGACTGAGCTCTGGTCA CCTGTTCATTCCTCGGTTAGCTGGAACTTTGCCCCGTTTCCAGTTTCTTATAGTGCATGCTTGGGAAACAAGATTTAAGGAGCCTCTGTT TTGGAAGGGCTGTCTGTGATTGAACGTGAAATGTGTAGTGCCATTGGGACCACGAAGGGAATTCTTGCACATGCTCGTGCTGGTGTGGGC ATGGGACTGGCTGGAAACGTCTGTATGCAGGGAGCCAGGGTGAGGGCAGAGTGTGGTGACAGCCGAACTTGGAGTAATGTCCGTGTAGAA AAAGGACCATGTTCTTATCCAGCCAATACTGGGAGTGCTGTCTCCACAATTTCAGGGCATCTGAATGTTTGATGTGGTTTTGTGTGTGTG TATGTATGTGTTTAATATTGAAGTGGATCATGAGATGTAAAGAAAACAATAATGGCAATGACTTATATTCAAATCTGTATTTGTTTCTTT ATCAATGTAATCTGCTGAGGACCTTTTGTCTAAGATTCAGTAGTGTTTTAAGGTTCTGATATCGAATTAATGAAGTAAAGTTGTTGATGG TGGTGAAACACCGTAGGGCATGTGGTTCAAAGAGAAGCAGGAGGGCAAGGGAAAGTTACCCTGATCTTAGTTTGTAGCTTATGACTTATT TAATGAATGGATGCCCAGCCAAGCTCAGAGTAGGCGCCCAAAGCATTGTGGATTATTTTCCTGTTTTGTCTTTTTTTTTTTTTTTTTTTA AGCCATGACATCCCAGAAGAGGACAGTGAATTACTCCTAGGTCGGCTCTTATAGAGTGGCCATAGTGTTCTGTCAAAACACTTGCTTCCA >79219_79219_3_SART3-CORO1C_SART3_chr12_108929135_ENST00000431469_CORO1C_chr12_109042796_ENST00000261401_length(amino acids)=623AA_BP=480 MATAAETSASEPEAESKAGPKADGEEDEVKAARTRRKVLSRAVAAATYKTMGPAWDQQEEGVSESDGDEYAMASSAESSPGEYEWEYDEE EEKNQLEIERLEEQLSINVYDYNCHVDLIRLLRLEGELTKVRMARQKMSEIFPLTEELWLEWLHDEISMAQDGLDREHVYDLFEKAVKDY ICPNIWLEYGQYSVGGIGQKGGLEKVRSVFERALSSVGLHMTKGLALWEAYREFESAIVEAARLEKVHSLFRRQLAIPLYDMEATFAEYE EWSEDPIPESVIQNYNKALQQLEKYKPYEEALLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVPDLWIRYSQYLDRQLKV KDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISDSSKELEELRAAFTRALEYLKQEVEERFNESGDPSCVIMQNWARIEARLC NNMQKARELWDSIMTRGNAKYANMWLEYYNLERFFKLHERKCEPIIMTVPRKSDLFQDDLYPDTAGPEAALEAEEWFEGKNADPILISLK -------------------------------------------------------------- >79219_79219_4_SART3-CORO1C_SART3_chr12_108929135_ENST00000431469_CORO1C_chr12_109042796_ENST00000541050_length(transcript)=2713nt_BP=1453nt GCAAGATGGCGACTGCGGCCGAAACCTCGGCTTCAGAACCCGAGGCTGAGTCCAAGGCTGGGCCCAAGGCTGACGGAGAGGAGGATGAGG TTAAGGCGGCTAGGACAAGGAGGAAGGTGTTATCGCGGGCTGTGGCCGCTGCGACATACAAGACCATGGGGCCAGCGTGGGATCAGCAGG AGGAAGGCGTGAGCGAGAGCGATGGGGATGAGTACGCCATGGCTTCCTCCGCGGAGAGCTCCCCCGGGGAGTACGAGTGGGAATATGACG AAGAGGAGGAGAAAAACCAGCTGGAGATTGAGAGACTGGAGGAGCAGTTGTCTATCAACGTCTATGACTACAACTGCCATGTGGACTTGA TCAGACTGCTCAGGCTGGAAGGGGAGCTTACCAAGGTGAGGATGGCCCGCCAGAAGATGAGTGAAATCTTTCCCTTGACTGAAGAGCTCT GGCTGGAGTGGCTGCATGACGAGATCAGCATGGCCCAGGATGGCCTGGACAGAGAGCACGTGTATGACCTCTTTGAGAAAGCCGTGAAGG ATTACATTTGTCCTAACATTTGGCTAGAGTATGGCCAGTACTCAGTTGGTGGGATTGGTCAGAAAGGTGGCCTTGAGAAAGTTCGCTCCG TGTTTGAAAGGGCTCTCTCGTCTGTTGGTTTACATATGACCAAAGGACTCGCCCTCTGGGAGGCTTACCGAGAGTTTGAAAGTGCGATTG TGGAAGCTGCTCGGCTTGAGAAAGTCCACAGTCTTTTCCGGCGACAGTTGGCGATCCCACTCTATGATATGGAGGCCACATTTGCAGAGT ATGAAGAATGGTCAGAAGACCCAATACCAGAGTCAGTAATTCAGAACTATAACAAAGCACTACAGCAGCTGGAGAAATATAAACCCTATG AAGAAGCACTGTTGCAGGCAGAGGCACCAAGGCTGGCAGAATATCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCA TTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGTCCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGA AAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAGAAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCA TGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGT ATCTGAAGCAGGAGGTGGAAGAGCGTTTCAATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGAC TGTGCAATAACATGCAGAAAGCTCGGGAACTCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATT ACAACCTGGAAAGATTCTTCAAACTTCATGAGAGAAAGTGTGAACCTATTATTATGACTGTTCCCAGGAAGTCTGACCTTTTCCAAGATG ACCTGTATCCTGACACAGCGGGGCCAGAGGCCGCGCTGGAGGCAGAAGAGTGGTTCGAAGGCAAGAATGCAGACCCAATCCTCATCTCCT TGAAGCACGGGTACATTCCAGGCAAAAACAGGGATCTCAAGGTGGTCAAGAAGAACATTCTGGATAGCAAGCCCACTGCAAACAAGAAGT GCGACCTGATCAGCATCCCCAAGAAAACCACAGACACGGCCAGTGTGCAAAATGAAGCCAAGTTGGATGAGATTTTAAAAGAGATCAAAT CTATAAAAGACACAATCTGCAATCAAGATGAGCGTATTTCCAAGTTAGAACAGCAGATGGCAAAGATAGCAGCCTGAAGGTCCCACCCCC ACCCCTACAGAAAAAATGGGAGCAAGAACTTGTGCTTGGGAGCTGGTTATTGGTGTGGTCCTAGGGAGGGCGGAAAGGGAGGCACTGCCA TTTGGAGACATTCCATTTCAGATTTGTCAACCAGCGATAGGCCACATTCCAGTAAGAACTCAATTTGTCTCCCAAATTTGCAGAAACAAA ACGTGATTTAAAAGCTGAGCTTTTTATCAGAAAGCTTTTTTGATGTTTTAAGTGTTATGTGACTTGTTGAACTTTTTAAAAAGTGCTACT TTTAAAATCCCAGATACTCTGAATTTTAGAAAACAAACTAATTCTGATTGTGTCGTGCCCAAGTACCCTTTTTTTTTTAATGAATAGGGA CCAATGCCACATTGCTTTTTATATTTCTTTCTTTTTTAATGTTGCCAAAACCAAAAGTAGCTTTGTTTTCCTTTGTATTTTGCTACTTTG CAGTATTTGTGTGTGTGGTTTTTTTTCCTTAATTTGAAAGGGACAGCACTGTGTATGTTTATAAACTAAATGAAGATAAGATATTATTTT GTATAAACATTCATCTGAGAACAATCAAAGCAGTAGCCACATGGTGCTGGCTCCTTTGCAGCACAAACCTGGTCATTTTGATGACTGTAC AACAGGAAGACTTGAAAAATCACGTGGATTCATATTACCACCGCTCTCATTTCATGGAGTCTTCTGATCAAAAAAGCTCACGTCGTATTT CTTCTTTTCCTTTCTCTTTTCTAGAAATTGGGTGTTTGTACCAGAATGGAATTTTGCTTCTCGGTTATCCTGTGCTTCAGATGATTATAA >79219_79219_4_SART3-CORO1C_SART3_chr12_108929135_ENST00000431469_CORO1C_chr12_109042796_ENST00000541050_length(amino acids)=623AA_BP=480 MATAAETSASEPEAESKAGPKADGEEDEVKAARTRRKVLSRAVAAATYKTMGPAWDQQEEGVSESDGDEYAMASSAESSPGEYEWEYDEE EEKNQLEIERLEEQLSINVYDYNCHVDLIRLLRLEGELTKVRMARQKMSEIFPLTEELWLEWLHDEISMAQDGLDREHVYDLFEKAVKDY ICPNIWLEYGQYSVGGIGQKGGLEKVRSVFERALSSVGLHMTKGLALWEAYREFESAIVEAARLEKVHSLFRRQLAIPLYDMEATFAEYE EWSEDPIPESVIQNYNKALQQLEKYKPYEEALLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVPDLWIRYSQYLDRQLKV KDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISDSSKELEELRAAFTRALEYLKQEVEERFNESGDPSCVIMQNWARIEARLC NNMQKARELWDSIMTRGNAKYANMWLEYYNLERFFKLHERKCEPIIMTVPRKSDLFQDDLYPDTAGPEAALEAEEWFEGKNADPILISLK -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SART3-CORO1C |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SART3-CORO1C |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SART3-CORO1C |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
