
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SEC31A-AGMO (FusionGDB2 ID:80097) |
Fusion Gene Summary for SEC31A-AGMO |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SEC31A-AGMO | Fusion gene ID: 80097 | Hgene | Tgene | Gene symbol | SEC31A | AGMO | Gene ID | 22872 | 392636 |
| Gene name | SEC31 homolog A, COPII coat complex component | alkylglycerol monooxygenase | |
| Synonyms | ABP125|ABP130|HSPC275|HSPC334|NEDSOSB|SEC31L1 | TMEM195 | |
| Cytomap | 4q21.22 | 7p21.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein transport protein Sec31ASEC31 homolog A, COPII coating complex componentSEC31-like protein 1SEC31-related protein Aweb1-like proteinyeast Sec31p homolog | alkylglycerol monooxygenaseglyceryl-ether monooxygenasetransmembrane protein 195 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q6ZNB7 | |
| Ensembl transtripts involved in fusion gene | ENST00000311785, ENST00000326950, ENST00000348405, ENST00000355196, ENST00000395310, ENST00000432794, ENST00000443462, ENST00000448323, ENST00000500777, ENST00000505472, ENST00000505984, ENST00000508479, ENST00000508502, ENST00000509142, ENST00000513858, ENST00000436790, ENST00000264405, | ENST00000498264, ENST00000342526, | |
| Fusion gene scores | * DoF score | 22 X 23 X 10=5060 | 6 X 5 X 4=120 |
| # samples | 25 | 7 | |
| ** MAII score | log2(25/5060*10)=-4.33913738491959 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/120*10)=-0.777607578663552 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SEC31A [Title/Abstract] AND AGMO [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SEC31A(83801952)-AGMO(15405244), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | SEC31A-AGMO seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SEC31A | GO:0051592 | response to calcium ion | 17196169 |
| Hgene | SEC31A | GO:0090110 | cargo loading into COPII-coated vesicle | 17499046|18843296 |
| Tgene | AGMO | GO:0006643 | membrane lipid metabolic process | 20643956 |
| Tgene | AGMO | GO:0046485 | ether lipid metabolic process | 20643956 |
 Fusion gene breakpoints across SEC31A (5'-gene) Fusion gene breakpoints across SEC31A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
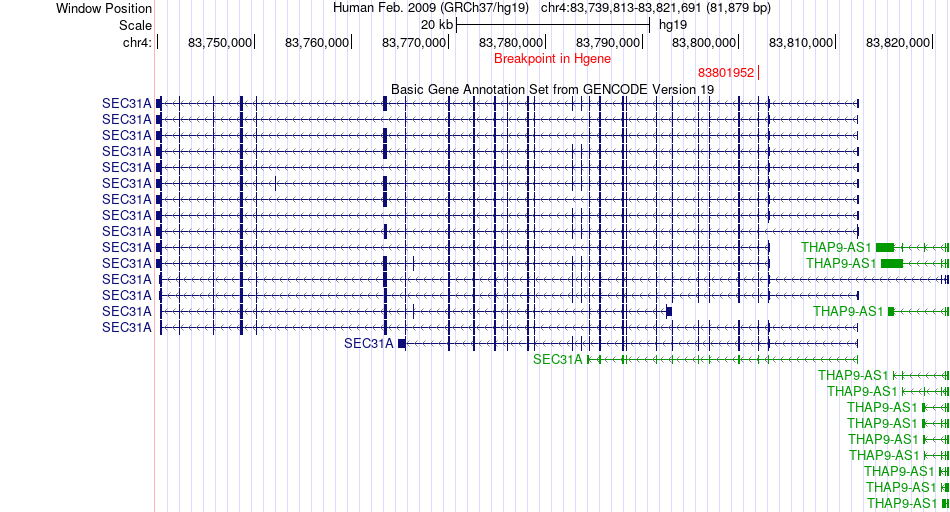 |
 Fusion gene breakpoints across AGMO (3'-gene) Fusion gene breakpoints across AGMO (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
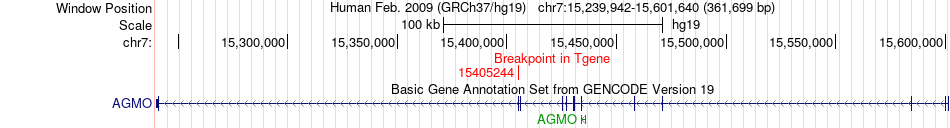 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUSC | TCGA-77-8131-01A | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
Top |
Fusion Gene ORF analysis for SEC31A-AGMO |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000311785 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000326950 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000348405 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000355196 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000395310 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000432794 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000443462 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000448323 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000500777 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000505472 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000505984 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000508479 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000508502 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000509142 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5CDS-intron | ENST00000513858 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5UTR-3CDS | ENST00000436790 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| 5UTR-intron | ENST00000436790 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| Frame-shift | ENST00000355196 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000311785 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000326950 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000348405 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000395310 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000432794 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000443462 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000448323 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000500777 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000505472 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000505984 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000508479 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000508502 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000509142 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| In-frame | ENST00000513858 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| intron-3CDS | ENST00000264405 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
| intron-intron | ENST00000264405 | ENST00000498264 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000348405 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1404 | 256 | 29 | 436 | 135 |
| ENST00000513858 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1421 | 273 | 46 | 453 | 135 |
| ENST00000395310 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1534 | 386 | 159 | 566 | 135 |
| ENST00000443462 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1363 | 215 | 0 | 395 | 131 |
| ENST00000509142 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1479 | 331 | 107 | 511 | 134 |
| ENST00000326950 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1512 | 364 | 140 | 544 | 134 |
| ENST00000432794 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1515 | 367 | 140 | 547 | 135 |
| ENST00000311785 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1515 | 367 | 140 | 547 | 135 |
| ENST00000448323 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1512 | 364 | 140 | 544 | 134 |
| ENST00000505472 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1352 | 204 | 1 | 384 | 127 |
| ENST00000500777 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1355 | 207 | 4 | 387 | 127 |
| ENST00000508502 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1471 | 323 | 96 | 503 | 135 |
| ENST00000505984 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1416 | 268 | 41 | 448 | 135 |
| ENST00000508479 | SEC31A | chr4 | 83801952 | - | ENST00000342526 | AGMO | chr7 | 15405244 | - | 1421 | 273 | 46 | 453 | 135 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000348405 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.28075916 | 0.71924084 |
| ENST00000513858 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.28664082 | 0.7133592 |
| ENST00000395310 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.19186321 | 0.8081368 |
| ENST00000443462 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.48554683 | 0.5144532 |
| ENST00000509142 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.22684449 | 0.7731555 |
| ENST00000326950 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.16872098 | 0.83127904 |
| ENST00000432794 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.18172514 | 0.8182749 |
| ENST00000311785 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.18172514 | 0.8182749 |
| ENST00000448323 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.16872098 | 0.83127904 |
| ENST00000505472 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.36260274 | 0.6373972 |
| ENST00000500777 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.32157603 | 0.678424 |
| ENST00000508502 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.35504875 | 0.6449512 |
| ENST00000505984 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.2743849 | 0.7256151 |
| ENST00000508479 | ENST00000342526 | SEC31A | chr4 | 83801952 | - | AGMO | chr7 | 15405244 | - | 0.28664082 | 0.7133592 |
Top |
Fusion Genomic Features for SEC31A-AGMO |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for SEC31A-AGMO |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr4:83801952/chr7:15405244) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | AGMO |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Glyceryl-ether monooxygenase that cleaves the O-alkyl bond of ether lipids. Ether lipids are essential components of brain membranes. {ECO:0000269|PubMed:20643956}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 4_47 | 67 | 1107.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 4_47 | 67 | 1182.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 4_47 | 67 | 1182.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 4_47 | 67 | 1221.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 4_47 | 67 | 1221.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 4_47 | 67 | 1234.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 4_47 | 62 | 1201.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 4_47 | 67 | 1221.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 4_47 | 67 | 1068.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 4_47 | 67 | 1206.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 4_47 | 67 | 1107.0 | Repeat | Note=WD 1 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 4_47 | 67 | 1068.0 | Repeat | Note=WD 1 |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 413_433 | 385 | 446.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 800_1091 | 67 | 1107.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 800_1091 | 67 | 1182.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 800_1091 | 67 | 1182.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 800_1091 | 67 | 1221.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 800_1091 | 67 | 1221.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 800_1091 | 67 | 1234.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 800_1091 | 62 | 1201.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 800_1091 | 67 | 1221.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 800_1091 | 67 | 1068.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 800_1091 | 67 | 1206.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 800_1091 | 67 | 1107.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 800_1091 | 67 | 1068.0 | Compositional bias | Note=Pro-rich |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 842_848 | 67 | 1107.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 842_848 | 67 | 1182.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 842_848 | 67 | 1182.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 842_848 | 67 | 1221.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 842_848 | 67 | 1221.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 842_848 | 67 | 1234.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 842_848 | 62 | 1201.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 842_848 | 67 | 1221.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 842_848 | 67 | 1068.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 842_848 | 67 | 1206.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 842_848 | 67 | 1107.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 842_848 | 67 | 1068.0 | Motif | Note=ALG-2-binding site motif-2 (ABS-2)%2C |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 120_160 | 67 | 1107.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 166_206 | 67 | 1107.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 209_254 | 67 | 1107.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 258_298 | 67 | 1107.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 301_342 | 67 | 1107.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 68_111 | 67 | 1107.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 120_160 | 67 | 1182.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 166_206 | 67 | 1182.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 209_254 | 67 | 1182.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 258_298 | 67 | 1182.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 301_342 | 67 | 1182.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 68_111 | 67 | 1182.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 120_160 | 67 | 1182.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 166_206 | 67 | 1182.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 209_254 | 67 | 1182.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 258_298 | 67 | 1182.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 301_342 | 67 | 1182.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 68_111 | 67 | 1182.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 120_160 | 67 | 1221.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 166_206 | 67 | 1221.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 209_254 | 67 | 1221.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 258_298 | 67 | 1221.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 301_342 | 67 | 1221.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 68_111 | 67 | 1221.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 120_160 | 67 | 1221.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 166_206 | 67 | 1221.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 209_254 | 67 | 1221.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 258_298 | 67 | 1221.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 301_342 | 67 | 1221.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 68_111 | 67 | 1221.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 120_160 | 67 | 1234.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 166_206 | 67 | 1234.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 209_254 | 67 | 1234.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 258_298 | 67 | 1234.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 301_342 | 67 | 1234.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 68_111 | 67 | 1234.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 120_160 | 62 | 1201.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 166_206 | 62 | 1201.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 209_254 | 62 | 1201.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 258_298 | 62 | 1201.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 301_342 | 62 | 1201.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 68_111 | 62 | 1201.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 120_160 | 67 | 1221.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 166_206 | 67 | 1221.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 209_254 | 67 | 1221.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 258_298 | 67 | 1221.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 301_342 | 67 | 1221.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 68_111 | 67 | 1221.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 120_160 | 67 | 1068.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 166_206 | 67 | 1068.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 209_254 | 67 | 1068.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 258_298 | 67 | 1068.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 301_342 | 67 | 1068.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 68_111 | 67 | 1068.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 120_160 | 67 | 1206.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 166_206 | 67 | 1206.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 209_254 | 67 | 1206.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 258_298 | 67 | 1206.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 301_342 | 67 | 1206.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 68_111 | 67 | 1206.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 120_160 | 67 | 1107.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 166_206 | 67 | 1107.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 209_254 | 67 | 1107.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 258_298 | 67 | 1107.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 301_342 | 67 | 1107.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 68_111 | 67 | 1107.0 | Repeat | Note=WD 2 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 120_160 | 67 | 1068.0 | Repeat | Note=WD 3 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 166_206 | 67 | 1068.0 | Repeat | Note=WD 4 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 209_254 | 67 | 1068.0 | Repeat | Note=WD 5 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 258_298 | 67 | 1068.0 | Repeat | Note=WD 6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 301_342 | 67 | 1068.0 | Repeat | Note=WD 7 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 68_111 | 67 | 1068.0 | Repeat | Note=WD 2 |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 120_249 | 385 | 446.0 | Domain | Fatty acid hydroxylase | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 132_136 | 385 | 446.0 | Motif | Note=Histidine box-1 | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 145_149 | 385 | 446.0 | Motif | Note=Histidine box-2 | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 221_225 | 385 | 446.0 | Motif | Note=Histidine box-3 | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 111_131 | 385 | 446.0 | Transmembrane | Helical | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 334_354 | 385 | 446.0 | Transmembrane | Helical | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 363_383 | 385 | 446.0 | Transmembrane | Helical | |
| Tgene | AGMO | chr4:83801952 | chr7:15405244 | ENST00000342526 | 10 | 13 | 43_63 | 385 | 446.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for SEC31A-AGMO |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >80097_80097_1_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000311785_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1515nt_BP=367nt CGGAAGTGTTGGGCGCAGGGCGTGGCTGCGGCTGCGGCTGCCACGTTGGAACGGAACGTGGAGGTGGCCCTGGCCGGGGAGGAGGGGCGG CGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAG TAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAA CATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTT CTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCC CTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTT GGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTA TGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAA AATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTAT ATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCC TTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTT TGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTT GTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAA ATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTT GGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAA TGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTG >80097_80097_1_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000311785_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_2_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000326950_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1512nt_BP=364nt CGGAAGTGTTGGGCGCAGGGCGTGGCTGCGGCTGCGGCTGCCACGTTGGAACGGAACGTGGAGGTGGCCCTGGCCGGGGAGGAGGGGCGG CGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGGATGAAGTTAAAGGAAGTAG ATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACAT TTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTC ACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTT CATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGA AATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGT AATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAAT TAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATG ACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTT GATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGC AATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTG GGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATT TAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGA GTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGA ACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGC >80097_80097_2_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000326950_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=134AA_BP=74 MPKVLARMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMFL -------------------------------------------------------------- >80097_80097_3_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000348405_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1404nt_BP=256nt GACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAGTAGATCGTACAGCCATGCAGG CATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCC TTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTA TTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTG AGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTAC ATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGTAATGCTTATATGAACTAT TTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTA CTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGG GATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTAT GCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGA AGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGAT CATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTG TGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACC ACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATT AGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGC >80097_80097_3_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000348405_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_4_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000395310_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1534nt_BP=386nt CGGGGAAAGATGTCGCTCCCGGAAGTGTTGGGCGCAGGGCGTGGCTGCGGCTGCGGCTGCCACGTTGGAACGGAACGTGGAGGTGGCCCT GGCCGGGGAGGAGGGGCGGCGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGC AGGATGAAGTTAAAGGAAGTAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCT GCTCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCT TGTGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGT CACCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAA CAACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATA TTAAAATGTAATTATTTTATGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACA TTTGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTA TATTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCA ATTTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTC CTAAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACC CCTGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGT CTCTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTG AACCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAA GACATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTC AATTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGCATTATTTAAAGTTATCAGAACAATGTTAATTCTCTAAATAAAATTACCCA >80097_80097_4_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000395310_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_5_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000432794_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1515nt_BP=367nt CGGAAGTGTTGGGCGCAGGGCGTGGCTGCGGCTGCGGCTGCCACGTTGGAACGGAACGTGGAGGTGGCCCTGGCCGGGGAGGAGGGGCGG CGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAG TAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAA CATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTT CTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCC CTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTT GGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTA TGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAA AATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTAT ATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCC TTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTT TGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTT GTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAA ATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTT GGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAA TGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTG >80097_80097_5_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000432794_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_6_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000443462_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1363nt_BP=215nt CTGGCCGGGGAGGAGGGGCGGCGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCA GGAACATCTGCTCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGAT ATGAAATCTTGTGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTAC CGATTTGGTCACCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGA AGCATGAAACAACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTAT GACTGCATATTAAAATGTAATTATTTTATGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTG CCTTTCACATTTGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATA ATGATGTTATATTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCC TTTACATCAATTTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTG GTTGTCTTCCTAAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGG TGGCTCACCCCTGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTG AAGTCCTGTCTCTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAG AATCGCTTGAACCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCA AAGAAAAAAGACATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATA TGCTTGGTCAATTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGCATTATTTAAAGTTATCAGAACAATGTTAATTCTCTAAATAA >80097_80097_6_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000443462_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=131AA_BP=71 LAGEEGRRRMLGESDERCTNAGSGCRRSSPGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMFLMLY -------------------------------------------------------------- >80097_80097_7_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000448323_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1512nt_BP=364nt CGGAAGTGTTGGGCGCAGGGCGTGGCTGCGGCTGCGGCTGCCACGTTGGAACGGAACGTGGAGGTGGCCCTGGCCGGGGAGGAGGGGCGG CGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGGATGAAGTTAAAGGAAGTAG ATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACAT TTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTC ACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTT CATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGA AATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGT AATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAAT TAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATG ACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTT GATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGC AATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTG GGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATT TAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGA GTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGA ACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGC >80097_80097_7_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000448323_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=134AA_BP=74 MPKVLARMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMFL -------------------------------------------------------------- >80097_80097_8_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000500777_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1355nt_BP=207nt CAGGATGAAGTTAAAGGAAGTAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATC TGCTCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATC TTGTGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGG TCACCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAA ACAACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCAT ATTAAAATGTAATTATTTTATGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCAC ATTTGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTT ATATTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATC AATTTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTT CCTAAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCAC CCCTGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTG TCTCTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTT GAACCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAA AGACATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGT CAATTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGCATTATTTAAAGTTATCAGAACAATGTTAATTCTCTAAATAAAATTACCC >80097_80097_8_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000500777_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=127AA_BP=67 MKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMFLMLYRFGH -------------------------------------------------------------- >80097_80097_9_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000505472_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1352nt_BP=204nt GATGAAGTTAAAGGAAGTAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGC TCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTG TGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCA CCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACA ACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATT AAAATGTAATTATTTTATGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATT TGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATA TTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAAT TTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCT AAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCC TGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCT CTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAA CCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGA CATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAA TTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGCATTATTTAAAGTTATCAGAACAATGTTAATTCTCTAAATAAAATTACCCAAG >80097_80097_9_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000505472_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=127AA_BP=67 MKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMFLMLYRFGH -------------------------------------------------------------- >80097_80097_10_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000505984_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1416nt_BP=268nt CTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAGTAGATCGTA CAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACATTTAGTA CGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTCACAGAC CCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTTCATTGT CATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGAAATAAC CTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGTAATGCT TATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAATTAAAGT CAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATGACAAGT ATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTTGATGTT TGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGCAATCCA TATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTGGGGCTG AGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATTTAGCTG GACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGAGTGAGT GGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGAACAGAA TTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGCAGCATG >80097_80097_10_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000505984_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_11_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000508479_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1421nt_BP=273nt GAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAGTAGA TCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACATT TAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTCA CAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTTC ATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGAA ATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGTA ATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAATT AAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATGA CAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTTG ATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGCA ATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTGG GGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATTT AGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGAG TGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGAA CAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGCA >80097_80097_11_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000508479_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_12_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000508502_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1471nt_BP=323nt GTTGGAACGGAACGTGGAGGTGGCCCTGGCCGGGGAGGAGGGGCGGCGGCGAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGG ATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAGTAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCAC CCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTC TCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTG ATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATT GCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGT TGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGTAATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTA CTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAA AAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAA ACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCA TAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACAT CACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACC ATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTC TGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACA GAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTG TCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGCATTATTTAAAGTTATCAGAACAA >80097_80097_12_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000508502_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- >80097_80097_13_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000509142_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1479nt_BP=331nt GCGGCTGCCACGTTGGAACGGAACGTGGAGGTGGCCCTGGCCGGGGAGGAGGGGCGGCGGCGAATGCTGGGAGAGTCCGACGAGCGCTGC ACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGGATGAAGTTAAAGGAAGTAGATCGTACAGCCATGCAGGCATGGAGCCCTGCCC AGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACATTTAGTACGAATGCTTCCCTTGAGATATTTGAAT TAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTCACAGACCCAAGGCAGCTATTATGGAAACTCTCC GTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTTCATTGTCATCTGCTTTTGAGATTGTTTTTTCCA TTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGAAATAACCTGAATTTGTACATAATTCTCTTCTTT TAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGTAATGCTTATATGAACTATTTCTTCAATGAAAAG TAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAATTAAAGTCAGTTTTGGTTACTTCCCCCCTTTACT ACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATGACAAGTATAAAAAGAAGGGATGAAACTTAAAAA CAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTTGATGTTTGCTAAAATTATGCATAGATACATGTT TGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGCAATCCATATAACCTAAGAAGCTGCATTCCAGAA AAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTGGGGCTGAGGTGGGCGGATCATGAGGTCCAGAGA TAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATTTAGCTGGACATGGTGGTGTGCGCCTGTAGTCCC AGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGAGTGAGTGGAGATTGCACCACTGCACTCCAGCCT GGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGAACAGAATTTGTCAGAATTAGTTTTTTCAACAGG TTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGCAGCATGCAGCAATTCTGCATTATTTAAAGTTAT >80097_80097_13_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000509142_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=134AA_BP=74 MPKVLARMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMFL -------------------------------------------------------------- >80097_80097_14_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000513858_AGMO_chr7_15405244_ENST00000342526_length(transcript)=1421nt_BP=273nt GAATGCTGGGAGAGTCCGACGAGCGCTGCACTAACGCAGGATCCGGCTGCCGAAGGTCCTCGCCAGCAGGATGAAGTTAAAGGAAGTAGA TCGTACAGCCATGCAGGCATGGAGCCCTGCCCAGAATCACCCCATTTACCTAGCAACAGGAACATCTGCTCAGCAATTGGATGCAACATT TAGTACGAATGCTTCCCTTGAGATATTTGAATTAGACCTCTCTGATCCATCCTTGGATATGAAATCTTGTGCCACATTCTCCTCTTCTCA CAGACCCAAGGCAGCTATTATGGAAACTCTCCGTTGCTTGATGTTCTTAATGCTGTACCGATTTGGTCACCTGAAGCCTCTTGTCCCTTC ATTGTCATCTGCTTTTGAGATTGTTTTTTCCATTTGCATTGCTTTCTGGGGAGTTAGAAGCATGAAACAACTCACCTCTCACCCTTGGAA ATAACCTGAATTTGTACATAATTCTCTTCTTTTAATGAGTTGTCCACACGCATATTATGACTGCATATTAAAATGTAATTATTTTATGTA ATGCTTATATGAACTATTTCTTCAATGAAAAGTAAAATTACTTATTTACTATTGTTTGCCTTTCACATTTGTTATTTTCTATTAAAAATT AAAGTCAGTTTTGGTTACTTCCCCCCTTTACTACAATTAAAAAAAGATTTCAAATATAATGATGTTATATTAACTGATAGCCTTATATGA CAAGTATAAAAAGAAGGGATGAAACTTAAAAACAGTAAAAACAAGAAGGAATATTGCCTTTACATCAATTTGAAAACAATGTTTCCTTTG ATGTTTGCTAAAATTATGCATAGATACATGTTTGTAGTCATAAAAATGTATTACATTGGTTGTCTTCCTAAGGCCACAGTTACCTTTGCA ATCCATATAACCTAAGAAGCTGCATTCCAGAAAAAGACATCACTGAGGCCAGGCGCGGTGGCTCACCCCTGTAATCCCAGCACTTTGTGG GGCTGAGGTGGGCGGATCATGAGGTCCAGAGATAGAGACCATCCTGGCCAACATGGTGAAGTCCTGTCTCTACTAAAAATATAAAAATTT AGCTGGACATGGTGGTGTGCGCCTGTAGTCCCAGCTACTCTGGAGGCTGAGGCAGGAGAATCGCTTGAACCTGGGAGGCAGAAGTTGGAG TGAGTGGAGATTGCACCACTGCACTCCAGCCTGGGCGACAGAGCGAGACTCCGTCTCAAAGAAAAAAGACATCACTGAAAGAAAAATGAA CAGAATTTGTCAGAATTAGTTTTTTCAACAGGTTACTTTGTCATACATTTCTCTAATATGCTTGGTCAATTTGTTTTGGCAGACTGGGCA >80097_80097_14_SEC31A-AGMO_SEC31A_chr4_83801952_ENST00000513858_AGMO_chr7_15405244_ENST00000342526_length(amino acids)=135AA_BP=75 MPKVLASRMKLKEVDRTAMQAWSPAQNHPIYLATGTSAQQLDATFSTNASLEIFELDLSDPSLDMKSCATFSSSHRPKAAIMETLRCLMF -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SEC31A-AGMO |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 800_1113 | 67.66666666666667 | 1107.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 800_1113 | 67.66666666666667 | 1182.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 800_1113 | 67.66666666666667 | 1182.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 800_1113 | 67.66666666666667 | 1221.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 800_1113 | 67.66666666666667 | 1221.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 800_1113 | 67.66666666666667 | 1234.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 800_1113 | 62.666666666666664 | 1201.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 800_1113 | 67.66666666666667 | 1221.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 800_1113 | 67.66666666666667 | 1068.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 800_1113 | 67.66666666666667 | 1206.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 800_1113 | 67.66666666666667 | 1107.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 800_1113 | 67.66666666666667 | 1068.0 | PDCD6 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000311785 | - | 3 | 26 | 161_471 | 67.66666666666667 | 1107.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000326950 | - | 3 | 25 | 161_471 | 67.66666666666667 | 1182.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000348405 | - | 3 | 25 | 161_471 | 67.66666666666667 | 1182.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000355196 | - | 5 | 29 | 161_471 | 67.66666666666667 | 1221.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000395310 | - | 3 | 27 | 161_471 | 67.66666666666667 | 1221.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000432794 | - | 3 | 28 | 161_471 | 67.66666666666667 | 1234.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000443462 | - | 2 | 26 | 161_471 | 62.666666666666664 | 1201.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000448323 | - | 3 | 27 | 161_471 | 67.66666666666667 | 1221.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000500777 | - | 2 | 23 | 161_471 | 67.66666666666667 | 1068.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000508502 | - | 3 | 27 | 161_471 | 67.66666666666667 | 1206.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000509142 | - | 3 | 26 | 161_471 | 67.66666666666667 | 1107.0 | SEC13 |
| Hgene | SEC31A | chr4:83801952 | chr7:15405244 | ENST00000513858 | - | 3 | 24 | 161_471 | 67.66666666666667 | 1068.0 | SEC13 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SEC31A-AGMO |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SEC31A-AGMO |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
