
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SH3GLB1-CLCA1 (FusionGDB2 ID:81479) |
Fusion Gene Summary for SH3GLB1-CLCA1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SH3GLB1-CLCA1 | Fusion gene ID: 81479 | Hgene | Tgene | Gene symbol | SH3GLB1 | CLCA1 | Gene ID | 51100 | 1179 |
| Gene name | SH3 domain containing GRB2 like, endophilin B1 | chloride channel accessory 1 | |
| Synonyms | Bif-1|CGI-61|PPP1R70|dJ612B15.2 | CACC|CACC1|CLCRG1|CaCC-1|GOB5|hCLCA1|hCaCC-1 | |
| Cytomap | 1p22.3 | 1p22.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | endophilin-B1Bax-interacting factor 1SH3 domain-containing GRB2-like protein B1SH3-containing protein SH3GLB1SH3-domain GRB2 like endophilin B1protein phosphatase 1, regulatory subunit 70testicular tissue protein Li 172 | calcium-activated chloride channel regulator 1CLCA family member 1, chloride channel regulatorcalcium-activated chloride channel family member 1calcium-activated chloride channel protein 1calcium-dependent chloride channel-1chloride channel regulator | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | A8K7I4 | |
| Ensembl transtripts involved in fusion gene | ENST00000535010, ENST00000370558, ENST00000482504, | ENST00000234701, ENST00000394711, | |
| Fusion gene scores | * DoF score | 4 X 5 X 2=40 | 3 X 3 X 3=27 |
| # samples | 4 | 3 | |
| ** MAII score | log2(4/40*10)=0 | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: SH3GLB1 [Title/Abstract] AND CLCA1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SH3GLB1(87181548)-CLCA1(86934704), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | SH3GLB1-CLCA1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. SH3GLB1-CLCA1 seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SH3GLB1 | GO:0031334 | positive regulation of protein complex assembly | 19074440 |
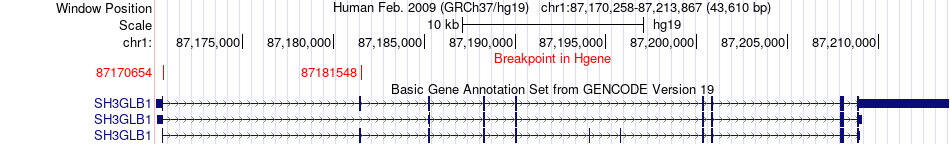
 Fusion gene breakpoints across SH3GLB1 (5'-gene) Fusion gene breakpoints across SH3GLB1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
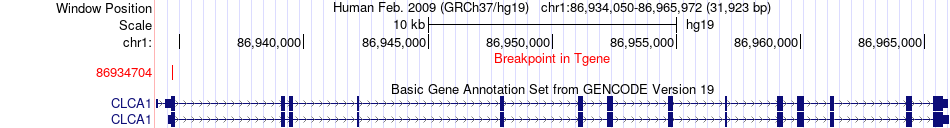
 Fusion gene breakpoints across CLCA1 (3'-gene) Fusion gene breakpoints across CLCA1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | CESC | TCGA-JX-A3PZ-01A | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| ChimerDB4 | CESC | TCGA-JX-A3PZ-01A | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
Top |
Fusion Gene ORF analysis for SH3GLB1-CLCA1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5UTR-3CDS | ENST00000535010 | ENST00000234701 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| 5UTR-3CDS | ENST00000535010 | ENST00000394711 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| Frame-shift | ENST00000370558 | ENST00000234701 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| Frame-shift | ENST00000370558 | ENST00000394711 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
| Frame-shift | ENST00000482504 | ENST00000234701 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| Frame-shift | ENST00000482504 | ENST00000394711 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
| In-frame | ENST00000370558 | ENST00000234701 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
| In-frame | ENST00000370558 | ENST00000394711 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| In-frame | ENST00000482504 | ENST00000234701 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
| In-frame | ENST00000482504 | ENST00000394711 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + |
| intron-3CDS | ENST00000535010 | ENST00000234701 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
| intron-3CDS | ENST00000535010 | ENST00000394711 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000370558 | SH3GLB1 | chr1 | 87181548 | + | ENST00000234701 | CLCA1 | chr1 | 86934704 | + | 3447 | 538 | 545 | 3232 | 895 |
| ENST00000482504 | SH3GLB1 | chr1 | 87181548 | + | ENST00000234701 | CLCA1 | chr1 | 86934704 | + | 3128 | 219 | 226 | 2913 | 895 |
| ENST00000370558 | SH3GLB1 | chr1 | 87170654 | + | ENST00000394711 | CLCA1 | chr1 | 86934704 | + | 3335 | 396 | 403 | 3090 | 895 |
| ENST00000482504 | SH3GLB1 | chr1 | 87170654 | + | ENST00000394711 | CLCA1 | chr1 | 86934704 | + | 3016 | 77 | 84 | 2771 | 895 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000370558 | ENST00000234701 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + | 0.001593454 | 0.9984066 |
| ENST00000482504 | ENST00000234701 | SH3GLB1 | chr1 | 87181548 | + | CLCA1 | chr1 | 86934704 | + | 0.001568078 | 0.99843186 |
| ENST00000370558 | ENST00000394711 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + | 0.001522921 | 0.99847704 |
| ENST00000482504 | ENST00000394711 | SH3GLB1 | chr1 | 87170654 | + | CLCA1 | chr1 | 86934704 | + | 0.001544991 | 0.99845505 |
Top |
Fusion Genomic Features for SH3GLB1-CLCA1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
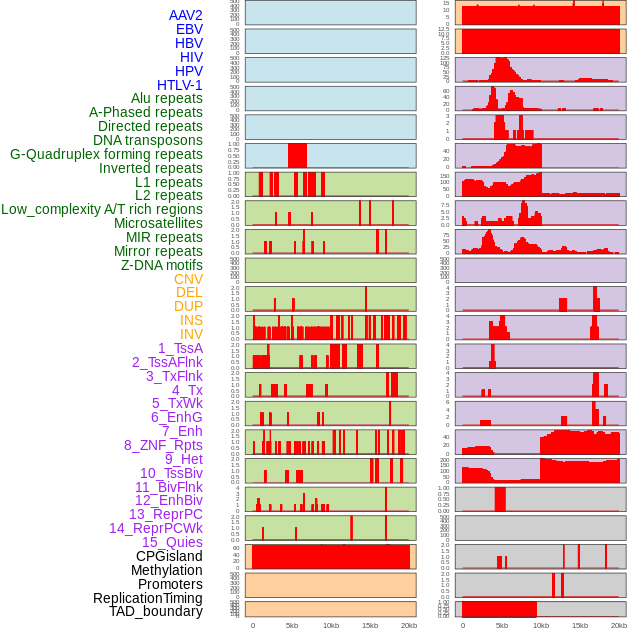 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for SH3GLB1-CLCA1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr1:87181548/chr1:86934704) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CLCA1 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: May be involved in mediating calcium-activated chloride conductance. May play critical roles in goblet cell metaplasia, mucus hypersecretion, cystic fibrosis and AHR. May be involved in the regulation of mucus production and/or secretion by goblet cells. Involved in the regulation of tissue inflammation in the innate immune response. May play a role as a tumor suppressor. Induces MUC5AC. {ECO:0000269|PubMed:11445004, ECO:0000269|PubMed:11842292, ECO:0000269|PubMed:11956057, ECO:0000269|PubMed:23112050, ECO:0000269|PubMed:9828122}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000370558 | + | 2 | 9 | 1_30 | 71 | 366.0 | Region | Note=Membrane-binding amphipathic helix |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000370558 | + | 2 | 9 | 1_37 | 71 | 366.0 | Region | Required for membrane binding |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000482504 | + | 2 | 11 | 1_30 | 71 | 387.0 | Region | Note=Membrane-binding amphipathic helix |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000482504 | + | 2 | 11 | 1_37 | 71 | 387.0 | Region | Required for membrane binding |
| Tgene | CLCA1 | chr1:87170654 | chr1:86934704 | ENST00000234701 | 0 | 15 | 306_475 | 0 | 915.0 | Domain | VWFA | |
| Tgene | CLCA1 | chr1:87170654 | chr1:86934704 | ENST00000394711 | 0 | 14 | 306_475 | 0 | 915.0 | Domain | VWFA | |
| Tgene | CLCA1 | chr1:87181548 | chr1:86934704 | ENST00000234701 | 0 | 15 | 306_475 | 0 | 915.0 | Domain | VWFA | |
| Tgene | CLCA1 | chr1:87181548 | chr1:86934704 | ENST00000394711 | 0 | 14 | 306_475 | 0 | 915.0 | Domain | VWFA | |
| Tgene | CLCA1 | chr1:87170654 | chr1:86934704 | ENST00000234701 | 0 | 15 | 46_199 | 0 | 915.0 | Region | Metalloprotease domain | |
| Tgene | CLCA1 | chr1:87170654 | chr1:86934704 | ENST00000394711 | 0 | 14 | 46_199 | 0 | 915.0 | Region | Metalloprotease domain | |
| Tgene | CLCA1 | chr1:87181548 | chr1:86934704 | ENST00000234701 | 0 | 15 | 46_199 | 0 | 915.0 | Region | Metalloprotease domain | |
| Tgene | CLCA1 | chr1:87181548 | chr1:86934704 | ENST00000394711 | 0 | 14 | 46_199 | 0 | 915.0 | Region | Metalloprotease domain |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000370558 | + | 1 | 9 | 155_195 | 24 | 366.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000482504 | + | 1 | 11 | 155_195 | 24 | 387.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 155_195 | 0 | 266.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000370558 | + | 2 | 9 | 155_195 | 71 | 366.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000482504 | + | 2 | 11 | 155_195 | 71 | 387.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 155_195 | 0 | 266.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000370558 | + | 1 | 9 | 27_261 | 24 | 366.0 | Domain | BAR |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000370558 | + | 1 | 9 | 305_365 | 24 | 366.0 | Domain | SH3 |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000482504 | + | 1 | 11 | 27_261 | 24 | 387.0 | Domain | BAR |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000482504 | + | 1 | 11 | 305_365 | 24 | 387.0 | Domain | SH3 |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 27_261 | 0 | 266.0 | Domain | BAR |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 305_365 | 0 | 266.0 | Domain | SH3 |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000370558 | + | 2 | 9 | 27_261 | 71 | 366.0 | Domain | BAR |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000370558 | + | 2 | 9 | 305_365 | 71 | 366.0 | Domain | SH3 |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000482504 | + | 2 | 11 | 27_261 | 71 | 387.0 | Domain | BAR |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000482504 | + | 2 | 11 | 305_365 | 71 | 387.0 | Domain | SH3 |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 27_261 | 0 | 266.0 | Domain | BAR |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 305_365 | 0 | 266.0 | Domain | SH3 |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000370558 | + | 1 | 9 | 1_30 | 24 | 366.0 | Region | Note=Membrane-binding amphipathic helix |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000370558 | + | 1 | 9 | 1_37 | 24 | 366.0 | Region | Required for membrane binding |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000482504 | + | 1 | 11 | 1_30 | 24 | 387.0 | Region | Note=Membrane-binding amphipathic helix |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000482504 | + | 1 | 11 | 1_37 | 24 | 387.0 | Region | Required for membrane binding |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 1_30 | 0 | 266.0 | Region | Note=Membrane-binding amphipathic helix |
| Hgene | SH3GLB1 | chr1:87170654 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 1_37 | 0 | 266.0 | Region | Required for membrane binding |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 1_30 | 0 | 266.0 | Region | Note=Membrane-binding amphipathic helix |
| Hgene | SH3GLB1 | chr1:87181548 | chr1:86934704 | ENST00000535010 | + | 1 | 8 | 1_37 | 0 | 266.0 | Region | Required for membrane binding |
Top |
Fusion Gene Sequence for SH3GLB1-CLCA1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >81479_81479_1_SH3GLB1-CLCA1_SH3GLB1_chr1_87170654_ENST00000370558_CLCA1_chr1_86934704_ENST00000394711_length(transcript)=3335nt_BP=396nt GTTTTTCCCTTGGGACCCGGGTCCACACGGCGGGGTCGCCCGTCCATCTCCGGCTCGCCCGCGGGGCCCATCGTCGACGTTAGCGGCCGT TCTCCGAGCCGACTGACCCATCCTTGGCGCTGCCGCCGCGCGCTTGTTCTCCTCCCTCGCCCCGCCTTCATCCTCCCCGTTCACGGAAAC GACAGCTGCGGCTGCGGGGCTGGCGCCGCCTCCCTCCACCTACCACGTCTGCCCTCGCCGCTCTAGCCCTGCGCCCCAGCCCGGCCGCGG CACCTCCGCCTCGCCGCCGCTAGGTCGGCCGGCTCCGCCCGGCTGCCGCCTAGGATGAATATCATGGACTTCAACGTGAAGAAGCTGGCG GCCGACGCAGGCACCTTCCTCAGTCGCGCCGTGCAGAGGGGCCCTGAGTAATTCACTCATTCAGCTGAACAACAATGGCTATGAAGGCAT TGTCGTTGCAATCGACCCCAATGTGCCAGAAGATGAAACACTCATTCAACAAATAAAGGACATGGTGACCCAGGCATCTCTGTATCTGCT TGAAGCTACAGGAAAGCGATTTTATTTCAAAAATGTTGCCATTTTGATTCCTGAAACATGGAAGACAAAGGCTGACTATGTGAGACCAAA ACTTGAGACCTACAAAAATGCTGATGTTCTGGTTGCTGAGTCTACTCCTCCAGGTAATGATGAACCCTACACTGAGCAGATGGGCAACTG TGGAGAGAAGGGTGAAAGGATCCACCTCACTCCTGATTTCATTGCAGGAAAAAAGTTAGCTGAATATGGACCACAAGGTAGGGCATTTGT CCATGAGTGGGCTCATCTACGATGGGGAGTATTTGACGAGTACAATAATGATGAGAAATTCTACTTATCCAATGGAAGAATACAAGCAGT AAGATGTTCAGCAGGTATTACTGGTACAAATGTAGTAAAGAAGTGTCAGGGAGGCAGCTGTTACACCAAAAGATGCACATTCAATAAAGT AACAGGACTCTATGAAAAAGGATGTGAGTTTGTTCTCCAATCCCGCCAGACGGAGAAGGCTTCTATAATGTTTGCACAACATGTTGATTC TATAGTTGAATTCTGTACAGAACAAAACCACAACAAAGAAGCTCCAAACAAGCAAAATCAAAAATGCAATCTCCGAAGCACATGGGAAGT GATCCGTGATTCTGAGGACTTTAAGAAAACCACTCCTATGACAACACAGCCACCAAATCCCACCTTCTCATTGCTGCAGATTGGACAAAG AATTGTGTGTTTAGTCCTTGACAAATCTGGAAGCATGGCGACTGGTAACCGCCTCAATCGACTGAATCAAGCAGGCCAGCTTTTCCTGCT GCAGACAGTTGAGCTGGGGTCCTGGGTTGGGATGGTGACATTTGACAGTGCTGCCCATGTACAAAATGAACTCATACAGATAAACAGTGG CAGTGACAGGGACACACTCGCCAAAAGATTACCTGCAGCAGCTTCAGGAGGGACGTCCATCTGCAGCGGGCTTCGATCGGCATTTACTGT GATTAGGAAGAAATATCCAACTGATGGATCTGAAATTGTGCTGCTGACGGATGGGGAAGACAACACTATAAGTGGGTGCTTTAACGAGGT CAAACAAAGTGGTGCCATCATCCACACAGTCGCTTTGGGGCCCTCTGCAGCTCAAGAACTAGAGGAGCTGTCCAAAATGACAGGAGGTTT ACAGACATATGCTTCAGATCAAGTTCAGAACAATGGCCTCATTGATGCTTTTGGGGCCCTTTCATCAGGAAATGGAGCTGTCTCTCAGCG CTCCATCCAGCTTGAGAGTAAGGGATTAACCCTCCAGAACAGCCAGTGGATGAATGGCACAGTGATCGTGGACAGCACCGTGGGAAAGGA CACTTTGTTTCTTATCACCTGGACAATGCAGCCTCCCCAAATCCTTCTCTGGGATCCCAGTGGACAGAAGCAAGGTGGCTTTGTAGTGGA CAAAAACACCAAAATGGCCTACCTCCAAATCCCAGGCATTGCTAAGGTTGGCACTTGGAAATACAGTCTGCAAGCAAGCTCACAAACCTT GACCCTGACTGTCACGTCCCGTGCGTCCAATGCTACCCTGCCTCCAATTACAGTGACTTCCAAAACGAACAAGGACACCAGCAAATTCCC CAGCCCTCTGGTAGTTTATGCAAATATTCGCCAAGGAGCCTCCCCAATTCTCAGGGCCAGTGTCACAGCCCTGATTGAATCAGTGAATGG AAAAACAGTTACCTTGGAACTACTGGATAATGGAGCAGGTGCTGATGCTACTAAGGATGACGGTGTCTACTCAAGGTATTTCACAACTTA TGACACGAATGGTAGATACAGTGTAAAAGTGCGGGCTCTGGGAGGAGTTAACGCAGCCAGACGGAGAGTGATACCCCAGCAGAGTGGAGC ACTGTACATACCTGGCTGGATTGAGAATGATGAAATACAATGGAATCCACCAAGACCTGAAATTAATAAGGATGATGTTCAACACAAGCA AGTGTGTTTCAGCAGAACATCCTCGGGAGGCTCATTTGTGGCTTCTGATGTCCCAAATGCTCCCATACCTGATCTCTTCCCACCTGGCCA AATCACCGACCTGAAGGCGGAAATTCACGGGGGCAGTCTCATTAATCTGACTTGGACAGCTCCTGGGGATGATTATGACCATGGAACAGC TCACAAGTATATCATTCGAATAAGTACAAGTATTCTTGATCTCAGAGACAAGTTCAATGAATCTCTTCAAGTGAATACTACTGCTCTCAT CCCAAAGGAAGCCAACTCTGAGGAAGTCTTTTTGTTTAAACCAGAAAACATTACTTTTGAAAATGGCACAGATCTTTTCATTGCTATTCA GGCTGTTGATAAGGTCGATCTGAAATCAGAAATATCCAACATTGCACGAGTATCTTTGTTTATTCCTCCACAGACTCCGCCAGAGACACC TAGTCCTGATGAAACGTCTGCTCCTTGTCCTAATATTCATATCAACAGCACCATTCCTGGCATTCACATTTTAAAAATTATGTGGAAGTG GATAGGAGAACTGCAGCTGTCAATAGCCTAGGGCTGAATTTTTGTCAGATAAATAAAATAAATCATTCATCCTTTTTTTTGATTATAAAA TTTTCTAAAATGTATTTTAGACTTCCTGTAGGGGGCGATATACTAAATGTATATAGTACATTTATACTAAATGTATTCCTGTAGGGGGCG ATATACTAAATGTATTTTAGACTTCCTGTAGGGGGCGATAAAATAAAATGCTAAACAACTGGGTATACATGCATAAAAACTATCCATTCA >81479_81479_1_SH3GLB1-CLCA1_SH3GLB1_chr1_87170654_ENST00000370558_CLCA1_chr1_86934704_ENST00000394711_length(amino acids)=895AA_BP= MSNSLIQLNNNGYEGIVVAIDPNVPEDETLIQQIKDMVTQASLYLLEATGKRFYFKNVAILIPETWKTKADYVRPKLETYKNADVLVAES TPPGNDEPYTEQMGNCGEKGERIHLTPDFIAGKKLAEYGPQGRAFVHEWAHLRWGVFDEYNNDEKFYLSNGRIQAVRCSAGITGTNVVKK CQGGSCYTKRCTFNKVTGLYEKGCEFVLQSRQTEKASIMFAQHVDSIVEFCTEQNHNKEAPNKQNQKCNLRSTWEVIRDSEDFKKTTPMT TQPPNPTFSLLQIGQRIVCLVLDKSGSMATGNRLNRLNQAGQLFLLQTVELGSWVGMVTFDSAAHVQNELIQINSGSDRDTLAKRLPAAA SGGTSICSGLRSAFTVIRKKYPTDGSEIVLLTDGEDNTISGCFNEVKQSGAIIHTVALGPSAAQELEELSKMTGGLQTYASDQVQNNGLI DAFGALSSGNGAVSQRSIQLESKGLTLQNSQWMNGTVIVDSTVGKDTLFLITWTMQPPQILLWDPSGQKQGGFVVDKNTKMAYLQIPGIA KVGTWKYSLQASSQTLTLTVTSRASNATLPPITVTSKTNKDTSKFPSPLVVYANIRQGASPILRASVTALIESVNGKTVTLELLDNGAGA DATKDDGVYSRYFTTYDTNGRYSVKVRALGGVNAARRRVIPQQSGALYIPGWIENDEIQWNPPRPEINKDDVQHKQVCFSRTSSGGSFVA SDVPNAPIPDLFPPGQITDLKAEIHGGSLINLTWTAPGDDYDHGTAHKYIIRISTSILDLRDKFNESLQVNTTALIPKEANSEEVFLFKP -------------------------------------------------------------- >81479_81479_2_SH3GLB1-CLCA1_SH3GLB1_chr1_87170654_ENST00000482504_CLCA1_chr1_86934704_ENST00000394711_length(transcript)=3016nt_BP=77nt CTAGGATGAATATCATGGACTTCAACGTGAAGAAGCTGGCGGCCGACGCAGGCACCTTCCTCAGTCGCGCCGTGCAGAGGGGCCCTGAGT AATTCACTCATTCAGCTGAACAACAATGGCTATGAAGGCATTGTCGTTGCAATCGACCCCAATGTGCCAGAAGATGAAACACTCATTCAA CAAATAAAGGACATGGTGACCCAGGCATCTCTGTATCTGCTTGAAGCTACAGGAAAGCGATTTTATTTCAAAAATGTTGCCATTTTGATT CCTGAAACATGGAAGACAAAGGCTGACTATGTGAGACCAAAACTTGAGACCTACAAAAATGCTGATGTTCTGGTTGCTGAGTCTACTCCT CCAGGTAATGATGAACCCTACACTGAGCAGATGGGCAACTGTGGAGAGAAGGGTGAAAGGATCCACCTCACTCCTGATTTCATTGCAGGA AAAAAGTTAGCTGAATATGGACCACAAGGTAGGGCATTTGTCCATGAGTGGGCTCATCTACGATGGGGAGTATTTGACGAGTACAATAAT GATGAGAAATTCTACTTATCCAATGGAAGAATACAAGCAGTAAGATGTTCAGCAGGTATTACTGGTACAAATGTAGTAAAGAAGTGTCAG GGAGGCAGCTGTTACACCAAAAGATGCACATTCAATAAAGTAACAGGACTCTATGAAAAAGGATGTGAGTTTGTTCTCCAATCCCGCCAG ACGGAGAAGGCTTCTATAATGTTTGCACAACATGTTGATTCTATAGTTGAATTCTGTACAGAACAAAACCACAACAAAGAAGCTCCAAAC AAGCAAAATCAAAAATGCAATCTCCGAAGCACATGGGAAGTGATCCGTGATTCTGAGGACTTTAAGAAAACCACTCCTATGACAACACAG CCACCAAATCCCACCTTCTCATTGCTGCAGATTGGACAAAGAATTGTGTGTTTAGTCCTTGACAAATCTGGAAGCATGGCGACTGGTAAC CGCCTCAATCGACTGAATCAAGCAGGCCAGCTTTTCCTGCTGCAGACAGTTGAGCTGGGGTCCTGGGTTGGGATGGTGACATTTGACAGT GCTGCCCATGTACAAAATGAACTCATACAGATAAACAGTGGCAGTGACAGGGACACACTCGCCAAAAGATTACCTGCAGCAGCTTCAGGA GGGACGTCCATCTGCAGCGGGCTTCGATCGGCATTTACTGTGATTAGGAAGAAATATCCAACTGATGGATCTGAAATTGTGCTGCTGACG GATGGGGAAGACAACACTATAAGTGGGTGCTTTAACGAGGTCAAACAAAGTGGTGCCATCATCCACACAGTCGCTTTGGGGCCCTCTGCA GCTCAAGAACTAGAGGAGCTGTCCAAAATGACAGGAGGTTTACAGACATATGCTTCAGATCAAGTTCAGAACAATGGCCTCATTGATGCT TTTGGGGCCCTTTCATCAGGAAATGGAGCTGTCTCTCAGCGCTCCATCCAGCTTGAGAGTAAGGGATTAACCCTCCAGAACAGCCAGTGG ATGAATGGCACAGTGATCGTGGACAGCACCGTGGGAAAGGACACTTTGTTTCTTATCACCTGGACAATGCAGCCTCCCCAAATCCTTCTC TGGGATCCCAGTGGACAGAAGCAAGGTGGCTTTGTAGTGGACAAAAACACCAAAATGGCCTACCTCCAAATCCCAGGCATTGCTAAGGTT GGCACTTGGAAATACAGTCTGCAAGCAAGCTCACAAACCTTGACCCTGACTGTCACGTCCCGTGCGTCCAATGCTACCCTGCCTCCAATT ACAGTGACTTCCAAAACGAACAAGGACACCAGCAAATTCCCCAGCCCTCTGGTAGTTTATGCAAATATTCGCCAAGGAGCCTCCCCAATT CTCAGGGCCAGTGTCACAGCCCTGATTGAATCAGTGAATGGAAAAACAGTTACCTTGGAACTACTGGATAATGGAGCAGGTGCTGATGCT ACTAAGGATGACGGTGTCTACTCAAGGTATTTCACAACTTATGACACGAATGGTAGATACAGTGTAAAAGTGCGGGCTCTGGGAGGAGTT AACGCAGCCAGACGGAGAGTGATACCCCAGCAGAGTGGAGCACTGTACATACCTGGCTGGATTGAGAATGATGAAATACAATGGAATCCA CCAAGACCTGAAATTAATAAGGATGATGTTCAACACAAGCAAGTGTGTTTCAGCAGAACATCCTCGGGAGGCTCATTTGTGGCTTCTGAT GTCCCAAATGCTCCCATACCTGATCTCTTCCCACCTGGCCAAATCACCGACCTGAAGGCGGAAATTCACGGGGGCAGTCTCATTAATCTG ACTTGGACAGCTCCTGGGGATGATTATGACCATGGAACAGCTCACAAGTATATCATTCGAATAAGTACAAGTATTCTTGATCTCAGAGAC AAGTTCAATGAATCTCTTCAAGTGAATACTACTGCTCTCATCCCAAAGGAAGCCAACTCTGAGGAAGTCTTTTTGTTTAAACCAGAAAAC ATTACTTTTGAAAATGGCACAGATCTTTTCATTGCTATTCAGGCTGTTGATAAGGTCGATCTGAAATCAGAAATATCCAACATTGCACGA GTATCTTTGTTTATTCCTCCACAGACTCCGCCAGAGACACCTAGTCCTGATGAAACGTCTGCTCCTTGTCCTAATATTCATATCAACAGC ACCATTCCTGGCATTCACATTTTAAAAATTATGTGGAAGTGGATAGGAGAACTGCAGCTGTCAATAGCCTAGGGCTGAATTTTTGTCAGA TAAATAAAATAAATCATTCATCCTTTTTTTTGATTATAAAATTTTCTAAAATGTATTTTAGACTTCCTGTAGGGGGCGATATACTAAATG TATATAGTACATTTATACTAAATGTATTCCTGTAGGGGGCGATATACTAAATGTATTTTAGACTTCCTGTAGGGGGCGATAAAATAAAAT >81479_81479_2_SH3GLB1-CLCA1_SH3GLB1_chr1_87170654_ENST00000482504_CLCA1_chr1_86934704_ENST00000394711_length(amino acids)=895AA_BP= MSNSLIQLNNNGYEGIVVAIDPNVPEDETLIQQIKDMVTQASLYLLEATGKRFYFKNVAILIPETWKTKADYVRPKLETYKNADVLVAES TPPGNDEPYTEQMGNCGEKGERIHLTPDFIAGKKLAEYGPQGRAFVHEWAHLRWGVFDEYNNDEKFYLSNGRIQAVRCSAGITGTNVVKK CQGGSCYTKRCTFNKVTGLYEKGCEFVLQSRQTEKASIMFAQHVDSIVEFCTEQNHNKEAPNKQNQKCNLRSTWEVIRDSEDFKKTTPMT TQPPNPTFSLLQIGQRIVCLVLDKSGSMATGNRLNRLNQAGQLFLLQTVELGSWVGMVTFDSAAHVQNELIQINSGSDRDTLAKRLPAAA SGGTSICSGLRSAFTVIRKKYPTDGSEIVLLTDGEDNTISGCFNEVKQSGAIIHTVALGPSAAQELEELSKMTGGLQTYASDQVQNNGLI DAFGALSSGNGAVSQRSIQLESKGLTLQNSQWMNGTVIVDSTVGKDTLFLITWTMQPPQILLWDPSGQKQGGFVVDKNTKMAYLQIPGIA KVGTWKYSLQASSQTLTLTVTSRASNATLPPITVTSKTNKDTSKFPSPLVVYANIRQGASPILRASVTALIESVNGKTVTLELLDNGAGA DATKDDGVYSRYFTTYDTNGRYSVKVRALGGVNAARRRVIPQQSGALYIPGWIENDEIQWNPPRPEINKDDVQHKQVCFSRTSSGGSFVA SDVPNAPIPDLFPPGQITDLKAEIHGGSLINLTWTAPGDDYDHGTAHKYIIRISTSILDLRDKFNESLQVNTTALIPKEANSEEVFLFKP -------------------------------------------------------------- >81479_81479_3_SH3GLB1-CLCA1_SH3GLB1_chr1_87181548_ENST00000370558_CLCA1_chr1_86934704_ENST00000234701_length(transcript)=3447nt_BP=538nt GTTTTTCCCTTGGGACCCGGGTCCACACGGCGGGGTCGCCCGTCCATCTCCGGCTCGCCCGCGGGGCCCATCGTCGACGTTAGCGGCCGT TCTCCGAGCCGACTGACCCATCCTTGGCGCTGCCGCCGCGCGCTTGTTCTCCTCCCTCGCCCCGCCTTCATCCTCCCCGTTCACGGAAAC GACAGCTGCGGCTGCGGGGCTGGCGCCGCCTCCCTCCACCTACCACGTCTGCCCTCGCCGCTCTAGCCCTGCGCCCCAGCCCGGCCGCGG CACCTCCGCCTCGCCGCCGCTAGGTCGGCCGGCTCCGCCCGGCTGCCGCCTAGGATGAATATCATGGACTTCAACGTGAAGAAGCTGGCG GCCGACGCAGGCACCTTCCTCAGTCGCGCCGTGCAGTTCACAGAAGAAAAGCTTGGCCAGGCTGAGAAGACAGAATTGGATGCTCACTTA GAGAACCTCCTTAGCAAAGCTGAATGTACCAAAATATGGACAGAAAAAATAATGAAACAAACTGAAGTGTTATTGCAGCCAAATCCAAAG GGGCCCTGAGTAATTCACTCATTCAGCTGAACAACAATGGCTATGAAGGCATTGTCGTTGCAATCGACCCCAATGTGCCAGAAGATGAAA CACTCATTCAACAAATAAAGGACATGGTGACCCAGGCATCTCTGTATCTGCTTGAAGCTACAGGAAAGCGATTTTATTTCAAAAATGTTG CCATTTTGATTCCTGAAACATGGAAGACAAAGGCTGACTATGTGAGACCAAAACTTGAGACCTACAAAAATGCTGATGTTCTGGTTGCTG AGTCTACTCCTCCAGGTAATGATGAACCCTACACTGAGCAGATGGGCAACTGTGGAGAGAAGGGTGAAAGGATCCACCTCACTCCTGATT TCATTGCAGGAAAAAAGTTAGCTGAATATGGACCACAAGGTAGGGCATTTGTCCATGAGTGGGCTCATCTACGATGGGGAGTATTTGACG AGTACAATAATGATGAGAAATTCTACTTATCCAATGGAAGAATACAAGCAGTAAGATGTTCAGCAGGTATTACTGGTACAAATGTAGTAA AGAAGTGTCAGGGAGGCAGCTGTTACACCAAAAGATGCACATTCAATAAAGTAACAGGACTCTATGAAAAAGGATGTGAGTTTGTTCTCC AATCCCGCCAGACGGAGAAGGCTTCTATAATGTTTGCACAACATGTTGATTCTATAGTTGAATTCTGTACAGAACAAAACCACAACAAAG AAGCTCCAAACAAGCAAAATCAAAAATGCAATCTCCGAAGCACATGGGAAGTGATCCGTGATTCTGAGGACTTTAAGAAAACCACTCCTA TGACAACACAGCCACCAAATCCCACCTTCTCATTGCTGCAGATTGGACAAAGAATTGTGTGTTTAGTCCTTGACAAATCTGGAAGCATGG CGACTGGTAACCGCCTCAATCGACTGAATCAAGCAGGCCAGCTTTTCCTGCTGCAGACAGTTGAGCTGGGGTCCTGGGTTGGGATGGTGA CATTTGACAGTGCTGCCCATGTACAAAATGAACTCATACAGATAAACAGTGGCAGTGACAGGGACACACTCGCCAAAAGATTACCTGCAG CAGCTTCAGGAGGGACGTCCATCTGCAGCGGGCTTCGATCGGCATTTACTGTGATTAGGAAGAAATATCCAACTGATGGATCTGAAATTG TGCTGCTGACGGATGGGGAAGACAACACTATAAGTGGGTGCTTTAACGAGGTCAAACAAAGTGGTGCCATCATCCACACAGTCGCTTTGG GGCCCTCTGCAGCTCAAGAACTAGAGGAGCTGTCCAAAATGACAGGAGGTTTACAGACATATGCTTCAGATCAAGTTCAGAACAATGGCC TCATTGATGCTTTTGGGGCCCTTTCATCAGGAAATGGAGCTGTCTCTCAGCGCTCCATCCAGCTTGAGAGTAAGGGATTAACCCTCCAGA ACAGCCAGTGGATGAATGGCACAGTGATCGTGGACAGCACCGTGGGAAAGGACACTTTGTTTCTTATCACCTGGACAATGCAGCCTCCCC AAATCCTTCTCTGGGATCCCAGTGGACAGAAGCAAGGTGGCTTTGTAGTGGACAAAAACACCAAAATGGCCTACCTCCAAATCCCAGGCA TTGCTAAGGTTGGCACTTGGAAATACAGTCTGCAAGCAAGCTCACAAACCTTGACCCTGACTGTCACGTCCCGTGCGTCCAATGCTACCC TGCCTCCAATTACAGTGACTTCCAAAACGAACAAGGACACCAGCAAATTCCCCAGCCCTCTGGTAGTTTATGCAAATATTCGCCAAGGAG CCTCCCCAATTCTCAGGGCCAGTGTCACAGCCCTGATTGAATCAGTGAATGGAAAAACAGTTACCTTGGAACTACTGGATAATGGAGCAG GTGCTGATGCTACTAAGGATGACGGTGTCTACTCAAGGTATTTCACAACTTATGACACGAATGGTAGATACAGTGTAAAAGTGCGGGCTC TGGGAGGAGTTAACGCAGCCAGACGGAGAGTGATACCCCAGCAGAGTGGAGCACTGTACATACCTGGCTGGATTGAGAATGATGAAATAC AATGGAATCCACCAAGACCTGAAATTAATAAGGATGATGTTCAACACAAGCAAGTGTGTTTCAGCAGAACATCCTCGGGAGGCTCATTTG TGGCTTCTGATGTCCCAAATGCTCCCATACCTGATCTCTTCCCACCTGGCCAAATCACCGACCTGAAGGCGGAAATTCACGGGGGCAGTC TCATTAATCTGACTTGGACAGCTCCTGGGGATGATTATGACCATGGAACAGCTCACAAGTATATCATTCGAATAAGTACAAGTATTCTTG ATCTCAGAGACAAGTTCAATGAATCTCTTCAAGTGAATACTACTGCTCTCATCCCAAAGGAAGCCAACTCTGAGGAAGTCTTTTTGTTTA AACCAGAAAACATTACTTTTGAAAATGGCACAGATCTTTTCATTGCTATTCAGGCTGTTGATAAGGTCGATCTGAAATCAGAAATATCCA ACATTGCACGAGTATCTTTGTTTATTCCTCCACAGACTCCGCCAGAGACACCTAGTCCTGATGAAACGTCTGCTCCTTGTCCTAATATTC ATATCAACAGCACCATTCCTGGCATTCACATTTTAAAAATTATGTGGAAGTGGATAGGAGAACTGCAGCTGTCAATAGCCTAGGGCTGAA TTTTTGTCAGATAAATAAAATAAATCATTCATCCTTTTTTTTGATTATAAAATTTTCTAAAATGTATTTTAGACTTCCTGTAGGGGGCGA TATACTAAATGTATATAGTACATTTATACTAAATGTATTCCTGTAGGGGGCGATATACTAAATGTATTTTAGACTTCCTGTAGGGGGCGA >81479_81479_3_SH3GLB1-CLCA1_SH3GLB1_chr1_87181548_ENST00000370558_CLCA1_chr1_86934704_ENST00000234701_length(amino acids)=895AA_BP= MSNSLIQLNNNGYEGIVVAIDPNVPEDETLIQQIKDMVTQASLYLLEATGKRFYFKNVAILIPETWKTKADYVRPKLETYKNADVLVAES TPPGNDEPYTEQMGNCGEKGERIHLTPDFIAGKKLAEYGPQGRAFVHEWAHLRWGVFDEYNNDEKFYLSNGRIQAVRCSAGITGTNVVKK CQGGSCYTKRCTFNKVTGLYEKGCEFVLQSRQTEKASIMFAQHVDSIVEFCTEQNHNKEAPNKQNQKCNLRSTWEVIRDSEDFKKTTPMT TQPPNPTFSLLQIGQRIVCLVLDKSGSMATGNRLNRLNQAGQLFLLQTVELGSWVGMVTFDSAAHVQNELIQINSGSDRDTLAKRLPAAA SGGTSICSGLRSAFTVIRKKYPTDGSEIVLLTDGEDNTISGCFNEVKQSGAIIHTVALGPSAAQELEELSKMTGGLQTYASDQVQNNGLI DAFGALSSGNGAVSQRSIQLESKGLTLQNSQWMNGTVIVDSTVGKDTLFLITWTMQPPQILLWDPSGQKQGGFVVDKNTKMAYLQIPGIA KVGTWKYSLQASSQTLTLTVTSRASNATLPPITVTSKTNKDTSKFPSPLVVYANIRQGASPILRASVTALIESVNGKTVTLELLDNGAGA DATKDDGVYSRYFTTYDTNGRYSVKVRALGGVNAARRRVIPQQSGALYIPGWIENDEIQWNPPRPEINKDDVQHKQVCFSRTSSGGSFVA SDVPNAPIPDLFPPGQITDLKAEIHGGSLINLTWTAPGDDYDHGTAHKYIIRISTSILDLRDKFNESLQVNTTALIPKEANSEEVFLFKP -------------------------------------------------------------- >81479_81479_4_SH3GLB1-CLCA1_SH3GLB1_chr1_87181548_ENST00000482504_CLCA1_chr1_86934704_ENST00000234701_length(transcript)=3128nt_BP=219nt CTAGGATGAATATCATGGACTTCAACGTGAAGAAGCTGGCGGCCGACGCAGGCACCTTCCTCAGTCGCGCCGTGCAGTTCACAGAAGAAA AGCTTGGCCAGGCTGAGAAGACAGAATTGGATGCTCACTTAGAGAACCTCCTTAGCAAAGCTGAATGTACCAAAATATGGACAGAAAAAA TAATGAAACAAACTGAAGTGTTATTGCAGCCAAATCCAAAGGGGCCCTGAGTAATTCACTCATTCAGCTGAACAACAATGGCTATGAAGG CATTGTCGTTGCAATCGACCCCAATGTGCCAGAAGATGAAACACTCATTCAACAAATAAAGGACATGGTGACCCAGGCATCTCTGTATCT GCTTGAAGCTACAGGAAAGCGATTTTATTTCAAAAATGTTGCCATTTTGATTCCTGAAACATGGAAGACAAAGGCTGACTATGTGAGACC AAAACTTGAGACCTACAAAAATGCTGATGTTCTGGTTGCTGAGTCTACTCCTCCAGGTAATGATGAACCCTACACTGAGCAGATGGGCAA CTGTGGAGAGAAGGGTGAAAGGATCCACCTCACTCCTGATTTCATTGCAGGAAAAAAGTTAGCTGAATATGGACCACAAGGTAGGGCATT TGTCCATGAGTGGGCTCATCTACGATGGGGAGTATTTGACGAGTACAATAATGATGAGAAATTCTACTTATCCAATGGAAGAATACAAGC AGTAAGATGTTCAGCAGGTATTACTGGTACAAATGTAGTAAAGAAGTGTCAGGGAGGCAGCTGTTACACCAAAAGATGCACATTCAATAA AGTAACAGGACTCTATGAAAAAGGATGTGAGTTTGTTCTCCAATCCCGCCAGACGGAGAAGGCTTCTATAATGTTTGCACAACATGTTGA TTCTATAGTTGAATTCTGTACAGAACAAAACCACAACAAAGAAGCTCCAAACAAGCAAAATCAAAAATGCAATCTCCGAAGCACATGGGA AGTGATCCGTGATTCTGAGGACTTTAAGAAAACCACTCCTATGACAACACAGCCACCAAATCCCACCTTCTCATTGCTGCAGATTGGACA AAGAATTGTGTGTTTAGTCCTTGACAAATCTGGAAGCATGGCGACTGGTAACCGCCTCAATCGACTGAATCAAGCAGGCCAGCTTTTCCT GCTGCAGACAGTTGAGCTGGGGTCCTGGGTTGGGATGGTGACATTTGACAGTGCTGCCCATGTACAAAATGAACTCATACAGATAAACAG TGGCAGTGACAGGGACACACTCGCCAAAAGATTACCTGCAGCAGCTTCAGGAGGGACGTCCATCTGCAGCGGGCTTCGATCGGCATTTAC TGTGATTAGGAAGAAATATCCAACTGATGGATCTGAAATTGTGCTGCTGACGGATGGGGAAGACAACACTATAAGTGGGTGCTTTAACGA GGTCAAACAAAGTGGTGCCATCATCCACACAGTCGCTTTGGGGCCCTCTGCAGCTCAAGAACTAGAGGAGCTGTCCAAAATGACAGGAGG TTTACAGACATATGCTTCAGATCAAGTTCAGAACAATGGCCTCATTGATGCTTTTGGGGCCCTTTCATCAGGAAATGGAGCTGTCTCTCA GCGCTCCATCCAGCTTGAGAGTAAGGGATTAACCCTCCAGAACAGCCAGTGGATGAATGGCACAGTGATCGTGGACAGCACCGTGGGAAA GGACACTTTGTTTCTTATCACCTGGACAATGCAGCCTCCCCAAATCCTTCTCTGGGATCCCAGTGGACAGAAGCAAGGTGGCTTTGTAGT GGACAAAAACACCAAAATGGCCTACCTCCAAATCCCAGGCATTGCTAAGGTTGGCACTTGGAAATACAGTCTGCAAGCAAGCTCACAAAC CTTGACCCTGACTGTCACGTCCCGTGCGTCCAATGCTACCCTGCCTCCAATTACAGTGACTTCCAAAACGAACAAGGACACCAGCAAATT CCCCAGCCCTCTGGTAGTTTATGCAAATATTCGCCAAGGAGCCTCCCCAATTCTCAGGGCCAGTGTCACAGCCCTGATTGAATCAGTGAA TGGAAAAACAGTTACCTTGGAACTACTGGATAATGGAGCAGGTGCTGATGCTACTAAGGATGACGGTGTCTACTCAAGGTATTTCACAAC TTATGACACGAATGGTAGATACAGTGTAAAAGTGCGGGCTCTGGGAGGAGTTAACGCAGCCAGACGGAGAGTGATACCCCAGCAGAGTGG AGCACTGTACATACCTGGCTGGATTGAGAATGATGAAATACAATGGAATCCACCAAGACCTGAAATTAATAAGGATGATGTTCAACACAA GCAAGTGTGTTTCAGCAGAACATCCTCGGGAGGCTCATTTGTGGCTTCTGATGTCCCAAATGCTCCCATACCTGATCTCTTCCCACCTGG CCAAATCACCGACCTGAAGGCGGAAATTCACGGGGGCAGTCTCATTAATCTGACTTGGACAGCTCCTGGGGATGATTATGACCATGGAAC AGCTCACAAGTATATCATTCGAATAAGTACAAGTATTCTTGATCTCAGAGACAAGTTCAATGAATCTCTTCAAGTGAATACTACTGCTCT CATCCCAAAGGAAGCCAACTCTGAGGAAGTCTTTTTGTTTAAACCAGAAAACATTACTTTTGAAAATGGCACAGATCTTTTCATTGCTAT TCAGGCTGTTGATAAGGTCGATCTGAAATCAGAAATATCCAACATTGCACGAGTATCTTTGTTTATTCCTCCACAGACTCCGCCAGAGAC ACCTAGTCCTGATGAAACGTCTGCTCCTTGTCCTAATATTCATATCAACAGCACCATTCCTGGCATTCACATTTTAAAAATTATGTGGAA GTGGATAGGAGAACTGCAGCTGTCAATAGCCTAGGGCTGAATTTTTGTCAGATAAATAAAATAAATCATTCATCCTTTTTTTTGATTATA AAATTTTCTAAAATGTATTTTAGACTTCCTGTAGGGGGCGATATACTAAATGTATATAGTACATTTATACTAAATGTATTCCTGTAGGGG >81479_81479_4_SH3GLB1-CLCA1_SH3GLB1_chr1_87181548_ENST00000482504_CLCA1_chr1_86934704_ENST00000234701_length(amino acids)=895AA_BP= MSNSLIQLNNNGYEGIVVAIDPNVPEDETLIQQIKDMVTQASLYLLEATGKRFYFKNVAILIPETWKTKADYVRPKLETYKNADVLVAES TPPGNDEPYTEQMGNCGEKGERIHLTPDFIAGKKLAEYGPQGRAFVHEWAHLRWGVFDEYNNDEKFYLSNGRIQAVRCSAGITGTNVVKK CQGGSCYTKRCTFNKVTGLYEKGCEFVLQSRQTEKASIMFAQHVDSIVEFCTEQNHNKEAPNKQNQKCNLRSTWEVIRDSEDFKKTTPMT TQPPNPTFSLLQIGQRIVCLVLDKSGSMATGNRLNRLNQAGQLFLLQTVELGSWVGMVTFDSAAHVQNELIQINSGSDRDTLAKRLPAAA SGGTSICSGLRSAFTVIRKKYPTDGSEIVLLTDGEDNTISGCFNEVKQSGAIIHTVALGPSAAQELEELSKMTGGLQTYASDQVQNNGLI DAFGALSSGNGAVSQRSIQLESKGLTLQNSQWMNGTVIVDSTVGKDTLFLITWTMQPPQILLWDPSGQKQGGFVVDKNTKMAYLQIPGIA KVGTWKYSLQASSQTLTLTVTSRASNATLPPITVTSKTNKDTSKFPSPLVVYANIRQGASPILRASVTALIESVNGKTVTLELLDNGAGA DATKDDGVYSRYFTTYDTNGRYSVKVRALGGVNAARRRVIPQQSGALYIPGWIENDEIQWNPPRPEINKDDVQHKQVCFSRTSSGGSFVA SDVPNAPIPDLFPPGQITDLKAEIHGGSLINLTWTAPGDDYDHGTAHKYIIRISTSILDLRDKFNESLQVNTTALIPKEANSEEVFLFKP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SH3GLB1-CLCA1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SH3GLB1-CLCA1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SH3GLB1-CLCA1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
