
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SIL1-STK10 (FusionGDB2 ID:81895) |
Fusion Gene Summary for SIL1-STK10 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SIL1-STK10 | Fusion gene ID: 81895 | Hgene | Tgene | Gene symbol | SIL1 | STK10 | Gene ID | 64374 | 6793 |
| Gene name | SIL1 nucleotide exchange factor | serine/threonine kinase 10 | |
| Synonyms | BAP|MSS|ULG5 | LOK|PRO2729 | |
| Cytomap | 5q31.2 | 5q35.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | nucleotide exchange factor SIL1BiP-associated proteinSIL1 homolog, endoplasmic reticulum chaperoneSIL1-like protein endoplasmic reticulum chaperone | serine/threonine-protein kinase 10lymphocyte-oriented kinase | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000265195, ENST00000394817, ENST00000509534, ENST00000515008, | ENST00000517775, ENST00000176763, | |
| Fusion gene scores | * DoF score | 14 X 11 X 10=1540 | 16 X 15 X 7=1680 |
| # samples | 17 | 17 | |
| ** MAII score | log2(17/1540*10)=-3.17932369944456 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(17/1680*10)=-3.30485458152842 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SIL1 [Title/Abstract] AND STK10 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SIL1(138456724)-STK10(171583792), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | SIL1-STK10 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. SIL1-STK10 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. SIL1-STK10 seems lost the major protein functional domain in Tgene partner, which is a kinase due to the frame-shifted ORF. SIL1-STK10 seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | STK10 | GO:0046777 | protein autophosphorylation | 12639966|18239682 |
 Fusion gene breakpoints across SIL1 (5'-gene) Fusion gene breakpoints across SIL1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
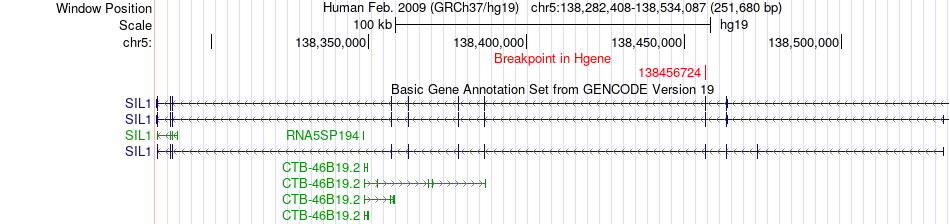 |
 Fusion gene breakpoints across STK10 (3'-gene) Fusion gene breakpoints across STK10 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
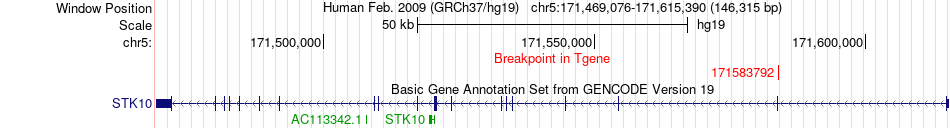 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-B6-A0IE-01A | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
Top |
Fusion Gene ORF analysis for SIL1-STK10 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000265195 | ENST00000517775 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| 5CDS-intron | ENST00000394817 | ENST00000517775 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| 5CDS-intron | ENST00000509534 | ENST00000517775 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| Frame-shift | ENST00000394817 | ENST00000176763 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| In-frame | ENST00000265195 | ENST00000176763 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| In-frame | ENST00000509534 | ENST00000176763 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| intron-3CDS | ENST00000515008 | ENST00000176763 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
| intron-intron | ENST00000515008 | ENST00000517775 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000265195 | SIL1 | chr5 | 138456724 | - | ENST00000176763 | STK10 | chr5 | 171583792 | - | 5950 | 390 | 333 | 3140 | 935 |
| ENST00000509534 | SIL1 | chr5 | 138456724 | - | ENST00000176763 | STK10 | chr5 | 171583792 | - | 6038 | 478 | 421 | 3228 | 935 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000265195 | ENST00000176763 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - | 0.004760886 | 0.99523914 |
| ENST00000509534 | ENST00000176763 | SIL1 | chr5 | 138456724 | - | STK10 | chr5 | 171583792 | - | 0.004820532 | 0.99517953 |
Top |
Fusion Genomic Features for SIL1-STK10 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
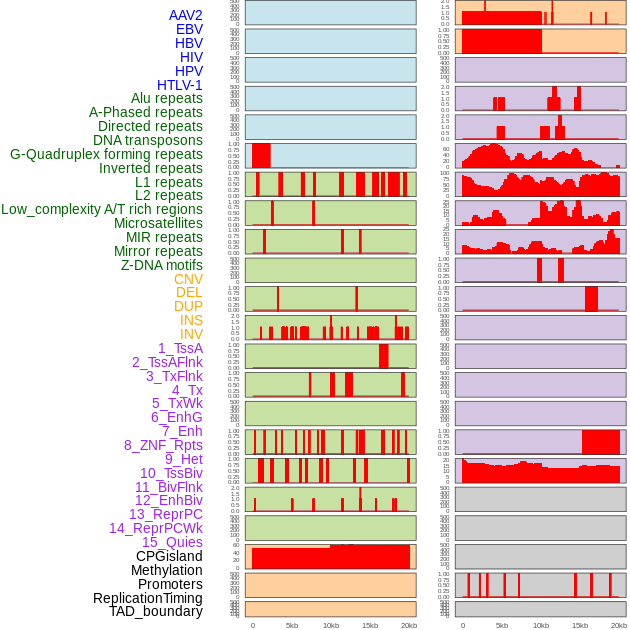 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for SIL1-STK10 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr5:138456724/chr5:171583792) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | STK10 | chr5:138456724 | chr5:171583792 | ENST00000176763 | 0 | 19 | 573_947 | 52 | 969.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | STK10 | chr5:138456724 | chr5:171583792 | ENST00000176763 | 0 | 19 | 750_884 | 52 | 969.0 | Compositional bias | Note=Gln-rich | |
| Tgene | STK10 | chr5:138456724 | chr5:171583792 | ENST00000176763 | 0 | 19 | 175_224 | 52 | 969.0 | Region | Note=Activation segment |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | STK10 | chr5:138456724 | chr5:171583792 | ENST00000176763 | 0 | 19 | 36_294 | 52 | 969.0 | Domain | Protein kinase | |
| Tgene | STK10 | chr5:138456724 | chr5:171583792 | ENST00000176763 | 0 | 19 | 42_50 | 52 | 969.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for SIL1-STK10 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >81895_81895_1_SIL1-STK10_SIL1_chr5_138456724_ENST00000265195_STK10_chr5_171583792_ENST00000176763_length(transcript)=5950nt_BP=390nt CTGGAATTGTGGGAGTTGTGTCTGCCACTCGGCTGCCGGAGGCCGAAGGCCAGCTATTACTTCTCTACTGAGGCCAACTGTGAGCCCTGG TGTGTTTCATTGGCAGGAGAAGAGGCTAATTAAGTGAAACTGAAAGGTCCCTGACTATGGCTCCCCAGAGCCTGCCTTCATCTAGGATGG CTCCTCTGGGCATGCTGCTTGGGCTGCTGATGGCCGCCTGCTTCACCTTCTGCCTCAGTCATCAGAACCTGAAGGAGTTTGCCCTGACCA ACCCAGAGAAGAGCAGCACCAAAGAAACAGAGAGAAAAGAAACCAAAGCCGAGGAGGAGCTGGATGCCGAAGTCCTGGAGGTGTTCCACC CGACGCATGAGTGGCAGGCCCTTCAGCCAGGCCAAGAATAAGGAGACGGGTGCTTTGGCTGCGGCCAAAGTCATTGAAACCAAGAGTGAG GAGGAGCTGGAGGACTACATCGTGGAGATTGAGATCCTGGCCACCTGCGACCACCCCTACATTGTGAAGCTCCTGGGAGCCTACTATCAC GACGGGAAGCTGTGGATCATGATTGAGTTCTGTCCAGGGGGAGCCGTGGACGCCATCATGCTGGAGCTGGACAGAGGCCTCACGGAGCCC CAGATACAGGTGGTTTGCCGCCAGATGCTAGAAGCCCTCAACTTCCTGCACAGCAAGAGGATCATCCACCGAGATCTGAAAGCTGGCAAC GTGCTGATGACCCTCGAGGGAGACATCAGGCTGGCTGACTTTGGTGTGTCTGCCAAGAATCTGAAGACTCTACAGAAACGAGATTCCTTC ATCGGCACGCCTTACTGGATGGCCCCCGAGGTGGTCATGTGTGAGACCATGAAAGACACGCCCTACGACTACAAAGCCGACATCTGGTCC CTGGGCATCACGCTGATTGAGATGGCCCAGATCGAGCCGCCACACCACGAGCTCAACCCCATGCGGGTCCTGCTAAAGATCGCCAAGTCG GACCCTCCCACGCTGCTCACGCCCTCTAAGTGGTCTGTAGAGTTCCGTGACTTCCTGAAGATAGCCCTGGATAAGAACCCAGAAACCCGA CCCAGTGCCGCGCAGCTGCTGGAGCATCCCTTCGTCAGCAGCATCACCAGTAACAAGGCTCTGCGGGAGCTGGTGGCTGAGGCCAAGGCC GAGGTGATGGAAGAGATCGAAGACGGCCGGGATGAGGGGGAAGAGGAGGACGCCGTGGATGCCGCCTCCACCCTGGAGAACCATACTCAG AACTCCTCTGAGGTGAGTCCGCCAAGCCTCAATGCTGACAAGCCTCTCGAGGAGTCACCTTCCACCCCGCTGGCACCCAGCCAGTCTCAG GACAGTGTGAATGAGCCCTGCAGCCAGCCCTCTGGGGACAGATCCCTCCAAACCACCAGTCCCCCAGTCGTGGCCCCTGGAAATGAGAAC GGCCTGGCAGTGCCTGTGCCCCTGCGGAAGTCCCGACCCGTGTCAATGGATGCCAGAATTCAGGTAGCCCAGGAGAAGCAAGTTGCTGAG CAGGGTGGGGACCTCAGCCCAGCAGCCAACAGATCTCAAAAGGCCAGCCAGAGCCGGCCCAACAGCAGCGCCCTGGAGACCTTGGGTGGG GAGAAGCTGGCCAATGGCAGCCTGGAGCCACCTGCCCAGGCAGCTCCAGGGCCTTCCAAGAGGGACTCGGACTGCAGCAGCCTCTGCACC TCTGAGAGCATGGACTATGGTACCAATCTCTCCACTGACCTGTCGCTGAACAAAGAGATGGGCTCTCTGTCCATCAAGGACCCGAAACTG TACAAAAAAACCCTCAAGCGGACACGCAAATTTGTGGTGGATGGTGTGGAGGTGAGCATCACCACCTCCAAGATCATCAGCGAAGATGAG AAGAAGGATGAGGAGATGAGATTTCTCAGGCGCCAGGAACTCCGAGAGCTTCGGCTGCTCCAGAAAGAAGAGCATCGGAACCAGACCCAG CTGAGTAACAAGCATGAGCTGCAGCTGGAGCAAATGCATAAACGTTTTGAACAGGAAATCAACGCCAAGAAGAAGTTCTTTGACACGGAA TTAGAGAACCTGGAGCGTCAGCAAAAGCAGCAAGTGGAGAAGATGGAGCAAGACCATGCCGTGCGCCGCCGGGAGGAGGCCAGGCGGATC CGCCTGGAGCAGGATCGGGACTACACCAGGTTCCAAGAGCAGCTCAAACTGATGAAGAAAGAGGTGAAGAACGAGGTGGAGAAGCTCCCC CGACAGCAGCGGAAGGAAAGCATGAAGCAGAAGATGGAGGAGCACACGCAGAAAAAGCAGCTTCTTGACCGGGACTTTGTAGCCAAGCAG AAGGAGGACCTGGAGCTGGCCATGAAGAGGCTCACCACCGACAACAGGCGGGAGATCTGTGACAAGGAGCGCGAGTGCCTCATGAAGAAG CAGGAGCTCCTTCGAGACCGGGAAGCAGCCCTGTGGGAGATGGAAGAGCACCAGCTGCAGGAGAGGCACCAGCTGGTGAAGCAGCAGCTC AAAGACCAGTACTTCCTCCAGCGGCACGAGCTGCTGCGCAAGCATGAGAAGGAGCGGGAGCAGATGCAGCGCTACAACCAGCGCATGATA GAGCAGCTGAAGGTGCGGCAGCAACAGGAAAAGGCGCGGCTGCCCAAGATCCAGAGGAGTGAGGGCAAGACGCGCATGGCCATGTACAAG AAGAGCCTCCACATCAACGGCGGGGGCAGCGCAGCTGAGCAGCGTGAGAAGATCAAGCAGTTCTCCCAGCAGGAGGAGAAGAGGCAGAAG TCGGAGCGGCTGCAGCAACAGCAGAAACACGAGAACCAGATGCGGGACATGCTGGCGCAGTGTGAGAGCAACATGAGCGAGCTGCAGCAG CTGCAGAATGAAAAGTGCCACCTCCTGGTAGAGCACGAAACCCAGAAACTGAAGGCCCTGGATGAGAGCCATAACCAGAACCTGAAGGAA TGGCGGGACAAGCTTCGGCCGCGCAAGAAGGCTCTGGAAGAGGATCTGAACCAGAAGAAGCGGGAGCAGGAGATGTTCTTCAAGCTGAGC GAGGAGGCGGAGTGCCCAAACCCCTCCACCCCAAGCAAGGCCGCCAAGTTCTTCCCCTACAGTTCTGCGGATGCTTCTTAACAACCGCCC GGGGCTGTGGCTGGCAGCTTGGTGGGCCCCAGGGCCTTCTCCTTCATTCTCTGTGAACATGTAACTCAGGACCCCTTTTCCCTCTTGCGT CTGTGCCAGCTCAAATCCAGCCCCTCGCCCTGTGCCACCCCAACTGTGCCTGATAGACCTGCCCCAGCGTTCCTGACTTCTTGCTGGCCT GTGGAGGGTGAGGTGTAATTATTTGTCACCTGAACCTAATGTATATTCTCCTTGAGCCCCAGATCCCTTCAAGCTGGAAGGGATGGGGCT GTTGGTGGGGTCAGGGTCCAAGAGGAATGGGTGTTCTGTGGCCTCGAGTCCTCTCCTGTTTGCGAAAATACCAGTTTTGCTCTCTGTGGG ACAAAGCACTGCTGATGAAGTCCCCGTGGGCTCATCCGGGCTGGAATTCTTGGTTTTTCAGCCATTCCCTGCAGAGTCACTCAATCATCA AGTCCCTCACAGCCATTTCTGTTCCCAGAGGAGCCAGGCCTGCAGCTGGCTGCTCAGGAGATGGCCTCATTCCTCCTGTTCTCCCAGTTT GCTTTCCACTTAAGACAAAGCCTTGTCTATGTGGGGGGCGGGGACCGGGGAAAGAGGGAGGCTGAAATGTTTATTCTGCTTCTCCCGTGT TTCATGCCATCTCGCGTCCCCCTTCCTGCACATGGGTGTGAATGCACACACATACGCGTACACACAGGACTTGGTCTGCTGGCCTGGCCT CTTCTGCCCAGGTGGGTTGGAACACGTTTGCTGCCTGAGCCTGTGCCACTGAGCATGTTAGGTGGAGCAGTTGGTGTGGCACGTGCGGGG TGTTGGCACCGGAGGCATGGAAAAGCACAGGCTGTACTGCCAGGCTGCGATGCGTGCTGGCCCCCGCACAGGCTCCTGTGTGCAGGGACT GATTCCTCAGCACACGAGGCTTCCACAACCCAGTCTGCTCCATAGCACTCTGGCCCACCCTGTCTGCAGGTGAAACAGGAGGGCTGTTTG CCTCTGCCCCATCCCCCCGACTGTGTTCAGGAGTCCCACCTTGCATTTCAGACCTGGGCTGGCAGTCTGTTGGACTTCTCTTCAGGAAGA AAAAGCATCAGGGGGAAATGGAATGCCCCTGCCCCAGGAACATGGCAGAAGCACAGGTTCTGTACCTCAGATGGACTCCTGCTGGGCCTT CGGGGTCTCAGTTGGCTTCCCCCAGATTCTGATTCTACAGCTGCAGAATGTATATAACGCAATAAAAGCAAATGTTTGAACCAGTATGCG GGTTTACTTACAAGGAAACCTTTTTTCACCCTGATTTCTAATTATTTTATTGAAACAACTATAAAATGTATGGTTTCTCCAGGAAGATAT CTGCTTTTTAAAACAACAAACAAGAACAACAACACAAAACTGGTAATGATTTGGAGTAATCATGCGGGCATATTGAGTCTGGGTAGTGTT TCGCTGGTGGTAGAGTGGTTGAGACTTCCTGGGAGGACTTTTTCCGCCTTCACTCCTACAGATTTAAGATTCTTTATTTTTAGGCCGGGC ATGGTGACTCACCTGAGGTGAGGAGTTCGAGACTAGCCTGGCCAACATGGCAAAACCCTGTCTCTACTAAAAGTACAAAAATTAGCCAGG CGTGGTGGTGCATGCCTATAATCCCAGCTACTCGGGAGGCTGAGGCAGGAGAATCGCTTGAACCCAGGAGGCAGAGGTTGCAGTAAGCCA AGGTCACACCACTGCACTCCAGCCTAGGTGACAGGGCGAGACTCTGTCTCTCAAAAAAAAAAAAAAAAAAAATTCTTTGAGACAGGGTCT CTAATATGTTGCCCAAGCTGATCTCAAACTCGTGGCTCAACTGATTCTCCCGCTTCAACCTTGAGTAGCTGGGACGACAGATACATGCTA CTGCACCCAGCTAGGTTAAGGTTCTGTATACTCTGTCTTTCTAAAGCAATTCCCTGGAAATTTTCTCAACATTGGTTATAACTGAATGCT TGGGAAGGGAGGGGCAGCCCTTCAACAGGAAAAGGGGACTGCCCAAATCTTAAATGCACAGTCAAGATTCTCTTGGAGCTAGCAATGGTT CTCAGGTTAGTGAGCATTTAGCTTGCCTGCTGCAGAATTTGACCCAGATGGGGGTGTTTTTTAGGTCTCTCACAAATGAGACAAGCGAAA CAATTGTCTCCTTTTATTCTCTTTGGTGCATTGGTGCTGGGGAAACATGAACTAGCGGCAGTGTAACTGCAGAACATAGACCCAGTTCTA CCAGGCCAGGCCAGCGTTGGGAACCACCAGACAGGGCTGCTTTGGGCTTTGCTTACAGTATTTCCATGTGTAGCCTGGCGTGTGAGAAAG TATTAGGTGAAATGCCAGTTTCATGGTTCAGGTGAAAGTCTGTGATCATTCCCCTCGTGGCTCGTCCTTCACATCACTTTTGCCCTTCAA GGAGTTGCCGCGTCCCCGCTCAGTGCCGCCTGAGCCCTCAGAGCTCCCCTGTGCTTTTCTGGATGGGGACTGGCGGGGTCACCTAGCCTC ACCGTGGAGCCACCGTGCAATGCCCATCTCTGAGAGGCCCACGCGGTATTCCTCGTGCCCTGTGTTAGTGCTTTCTGTATAAGGGACAGA CAGAACTGGGTTTTTTTCCTCTGTCTGGTTTTAGAGTAAATGTAACTAACTTTTATTTTCCCTTTATGAAAGATAGAAATTATTTTTATG TAGTTTCCAGACTTTATACAAAATTTTTGTAAAATGTTCTCTGGAAAGTTAACTACAAGAATGTAAATATGCTTCTAATAAAATAACAAG >81895_81895_1_SIL1-STK10_SIL1_chr5_138456724_ENST00000265195_STK10_chr5_171583792_ENST00000176763_length(amino acids)=935AA_BP=18 MPKSWRCSTRRMSGRPFSQAKNKETGALAAAKVIETKSEEELEDYIVEIEILATCDHPYIVKLLGAYYHDGKLWIMIEFCPGGAVDAIML ELDRGLTEPQIQVVCRQMLEALNFLHSKRIIHRDLKAGNVLMTLEGDIRLADFGVSAKNLKTLQKRDSFIGTPYWMAPEVVMCETMKDTP YDYKADIWSLGITLIEMAQIEPPHHELNPMRVLLKIAKSDPPTLLTPSKWSVEFRDFLKIALDKNPETRPSAAQLLEHPFVSSITSNKAL RELVAEAKAEVMEEIEDGRDEGEEEDAVDAASTLENHTQNSSEVSPPSLNADKPLEESPSTPLAPSQSQDSVNEPCSQPSGDRSLQTTSP PVVAPGNENGLAVPVPLRKSRPVSMDARIQVAQEKQVAEQGGDLSPAANRSQKASQSRPNSSALETLGGEKLANGSLEPPAQAAPGPSKR DSDCSSLCTSESMDYGTNLSTDLSLNKEMGSLSIKDPKLYKKTLKRTRKFVVDGVEVSITTSKIISEDEKKDEEMRFLRRQELRELRLLQ KEEHRNQTQLSNKHELQLEQMHKRFEQEINAKKKFFDTELENLERQQKQQVEKMEQDHAVRRREEARRIRLEQDRDYTRFQEQLKLMKKE VKNEVEKLPRQQRKESMKQKMEEHTQKKQLLDRDFVAKQKEDLELAMKRLTTDNRREICDKERECLMKKQELLRDREAALWEMEEHQLQE RHQLVKQQLKDQYFLQRHELLRKHEKEREQMQRYNQRMIEQLKVRQQQEKARLPKIQRSEGKTRMAMYKKSLHINGGGSAAEQREKIKQF SQQEEKRQKSERLQQQQKHENQMRDMLAQCESNMSELQQLQNEKCHLLVEHETQKLKALDESHNQNLKEWRDKLRPRKKALEEDLNQKKR -------------------------------------------------------------- >81895_81895_2_SIL1-STK10_SIL1_chr5_138456724_ENST00000509534_STK10_chr5_171583792_ENST00000176763_length(transcript)=6038nt_BP=478nt ATTACTTCTCTACTGAGGCCAACTGTGAGCCCTGGTGTGTTTCATTGGCAGGAGAAGAGGCTAATTAAGTGAAACTGAAAGAACAAATAA AATGATGAGTCCAGTTTCTGTGGCTGATTTCACAGACCCCTCGAGAGACAGCACGGCTATGGAAAGAGCATGGCTTTGGGGCCAAACTGG CTTTGAATTCTGACTCTACTGCTCTGTGGCTGCATGAACTCTGGCACGTCCCTGACTATGGCTCCCCAGAGCCTGCCTTCATCTAGGATG GCTCCTCTGGGCATGCTGCTTGGGCTGCTGATGGCCGCCTGCTTCACCTTCTGCCTCAGTCATCAGAACCTGGAGTTTGCCCTGACCAAC CCAGAGAAGAGCAGCACCAAAGAAACAGAGAGAAAAGAAACCAAAGCCGAGGAGGAGCTGGATGCCGAAGTCCTGGAGGTGTTCCACCCG ACGCATGAGTGGCAGGCCCTTCAGCCAGGCCAAGAATAAGGAGACGGGTGCTTTGGCTGCGGCCAAAGTCATTGAAACCAAGAGTGAGGA GGAGCTGGAGGACTACATCGTGGAGATTGAGATCCTGGCCACCTGCGACCACCCCTACATTGTGAAGCTCCTGGGAGCCTACTATCACGA CGGGAAGCTGTGGATCATGATTGAGTTCTGTCCAGGGGGAGCCGTGGACGCCATCATGCTGGAGCTGGACAGAGGCCTCACGGAGCCCCA GATACAGGTGGTTTGCCGCCAGATGCTAGAAGCCCTCAACTTCCTGCACAGCAAGAGGATCATCCACCGAGATCTGAAAGCTGGCAACGT GCTGATGACCCTCGAGGGAGACATCAGGCTGGCTGACTTTGGTGTGTCTGCCAAGAATCTGAAGACTCTACAGAAACGAGATTCCTTCAT CGGCACGCCTTACTGGATGGCCCCCGAGGTGGTCATGTGTGAGACCATGAAAGACACGCCCTACGACTACAAAGCCGACATCTGGTCCCT GGGCATCACGCTGATTGAGATGGCCCAGATCGAGCCGCCACACCACGAGCTCAACCCCATGCGGGTCCTGCTAAAGATCGCCAAGTCGGA CCCTCCCACGCTGCTCACGCCCTCTAAGTGGTCTGTAGAGTTCCGTGACTTCCTGAAGATAGCCCTGGATAAGAACCCAGAAACCCGACC CAGTGCCGCGCAGCTGCTGGAGCATCCCTTCGTCAGCAGCATCACCAGTAACAAGGCTCTGCGGGAGCTGGTGGCTGAGGCCAAGGCCGA GGTGATGGAAGAGATCGAAGACGGCCGGGATGAGGGGGAAGAGGAGGACGCCGTGGATGCCGCCTCCACCCTGGAGAACCATACTCAGAA CTCCTCTGAGGTGAGTCCGCCAAGCCTCAATGCTGACAAGCCTCTCGAGGAGTCACCTTCCACCCCGCTGGCACCCAGCCAGTCTCAGGA CAGTGTGAATGAGCCCTGCAGCCAGCCCTCTGGGGACAGATCCCTCCAAACCACCAGTCCCCCAGTCGTGGCCCCTGGAAATGAGAACGG CCTGGCAGTGCCTGTGCCCCTGCGGAAGTCCCGACCCGTGTCAATGGATGCCAGAATTCAGGTAGCCCAGGAGAAGCAAGTTGCTGAGCA GGGTGGGGACCTCAGCCCAGCAGCCAACAGATCTCAAAAGGCCAGCCAGAGCCGGCCCAACAGCAGCGCCCTGGAGACCTTGGGTGGGGA GAAGCTGGCCAATGGCAGCCTGGAGCCACCTGCCCAGGCAGCTCCAGGGCCTTCCAAGAGGGACTCGGACTGCAGCAGCCTCTGCACCTC TGAGAGCATGGACTATGGTACCAATCTCTCCACTGACCTGTCGCTGAACAAAGAGATGGGCTCTCTGTCCATCAAGGACCCGAAACTGTA CAAAAAAACCCTCAAGCGGACACGCAAATTTGTGGTGGATGGTGTGGAGGTGAGCATCACCACCTCCAAGATCATCAGCGAAGATGAGAA GAAGGATGAGGAGATGAGATTTCTCAGGCGCCAGGAACTCCGAGAGCTTCGGCTGCTCCAGAAAGAAGAGCATCGGAACCAGACCCAGCT GAGTAACAAGCATGAGCTGCAGCTGGAGCAAATGCATAAACGTTTTGAACAGGAAATCAACGCCAAGAAGAAGTTCTTTGACACGGAATT AGAGAACCTGGAGCGTCAGCAAAAGCAGCAAGTGGAGAAGATGGAGCAAGACCATGCCGTGCGCCGCCGGGAGGAGGCCAGGCGGATCCG CCTGGAGCAGGATCGGGACTACACCAGGTTCCAAGAGCAGCTCAAACTGATGAAGAAAGAGGTGAAGAACGAGGTGGAGAAGCTCCCCCG ACAGCAGCGGAAGGAAAGCATGAAGCAGAAGATGGAGGAGCACACGCAGAAAAAGCAGCTTCTTGACCGGGACTTTGTAGCCAAGCAGAA GGAGGACCTGGAGCTGGCCATGAAGAGGCTCACCACCGACAACAGGCGGGAGATCTGTGACAAGGAGCGCGAGTGCCTCATGAAGAAGCA GGAGCTCCTTCGAGACCGGGAAGCAGCCCTGTGGGAGATGGAAGAGCACCAGCTGCAGGAGAGGCACCAGCTGGTGAAGCAGCAGCTCAA AGACCAGTACTTCCTCCAGCGGCACGAGCTGCTGCGCAAGCATGAGAAGGAGCGGGAGCAGATGCAGCGCTACAACCAGCGCATGATAGA GCAGCTGAAGGTGCGGCAGCAACAGGAAAAGGCGCGGCTGCCCAAGATCCAGAGGAGTGAGGGCAAGACGCGCATGGCCATGTACAAGAA GAGCCTCCACATCAACGGCGGGGGCAGCGCAGCTGAGCAGCGTGAGAAGATCAAGCAGTTCTCCCAGCAGGAGGAGAAGAGGCAGAAGTC GGAGCGGCTGCAGCAACAGCAGAAACACGAGAACCAGATGCGGGACATGCTGGCGCAGTGTGAGAGCAACATGAGCGAGCTGCAGCAGCT GCAGAATGAAAAGTGCCACCTCCTGGTAGAGCACGAAACCCAGAAACTGAAGGCCCTGGATGAGAGCCATAACCAGAACCTGAAGGAATG GCGGGACAAGCTTCGGCCGCGCAAGAAGGCTCTGGAAGAGGATCTGAACCAGAAGAAGCGGGAGCAGGAGATGTTCTTCAAGCTGAGCGA GGAGGCGGAGTGCCCAAACCCCTCCACCCCAAGCAAGGCCGCCAAGTTCTTCCCCTACAGTTCTGCGGATGCTTCTTAACAACCGCCCGG GGCTGTGGCTGGCAGCTTGGTGGGCCCCAGGGCCTTCTCCTTCATTCTCTGTGAACATGTAACTCAGGACCCCTTTTCCCTCTTGCGTCT GTGCCAGCTCAAATCCAGCCCCTCGCCCTGTGCCACCCCAACTGTGCCTGATAGACCTGCCCCAGCGTTCCTGACTTCTTGCTGGCCTGT GGAGGGTGAGGTGTAATTATTTGTCACCTGAACCTAATGTATATTCTCCTTGAGCCCCAGATCCCTTCAAGCTGGAAGGGATGGGGCTGT TGGTGGGGTCAGGGTCCAAGAGGAATGGGTGTTCTGTGGCCTCGAGTCCTCTCCTGTTTGCGAAAATACCAGTTTTGCTCTCTGTGGGAC AAAGCACTGCTGATGAAGTCCCCGTGGGCTCATCCGGGCTGGAATTCTTGGTTTTTCAGCCATTCCCTGCAGAGTCACTCAATCATCAAG TCCCTCACAGCCATTTCTGTTCCCAGAGGAGCCAGGCCTGCAGCTGGCTGCTCAGGAGATGGCCTCATTCCTCCTGTTCTCCCAGTTTGC TTTCCACTTAAGACAAAGCCTTGTCTATGTGGGGGGCGGGGACCGGGGAAAGAGGGAGGCTGAAATGTTTATTCTGCTTCTCCCGTGTTT CATGCCATCTCGCGTCCCCCTTCCTGCACATGGGTGTGAATGCACACACATACGCGTACACACAGGACTTGGTCTGCTGGCCTGGCCTCT TCTGCCCAGGTGGGTTGGAACACGTTTGCTGCCTGAGCCTGTGCCACTGAGCATGTTAGGTGGAGCAGTTGGTGTGGCACGTGCGGGGTG TTGGCACCGGAGGCATGGAAAAGCACAGGCTGTACTGCCAGGCTGCGATGCGTGCTGGCCCCCGCACAGGCTCCTGTGTGCAGGGACTGA TTCCTCAGCACACGAGGCTTCCACAACCCAGTCTGCTCCATAGCACTCTGGCCCACCCTGTCTGCAGGTGAAACAGGAGGGCTGTTTGCC TCTGCCCCATCCCCCCGACTGTGTTCAGGAGTCCCACCTTGCATTTCAGACCTGGGCTGGCAGTCTGTTGGACTTCTCTTCAGGAAGAAA AAGCATCAGGGGGAAATGGAATGCCCCTGCCCCAGGAACATGGCAGAAGCACAGGTTCTGTACCTCAGATGGACTCCTGCTGGGCCTTCG GGGTCTCAGTTGGCTTCCCCCAGATTCTGATTCTACAGCTGCAGAATGTATATAACGCAATAAAAGCAAATGTTTGAACCAGTATGCGGG TTTACTTACAAGGAAACCTTTTTTCACCCTGATTTCTAATTATTTTATTGAAACAACTATAAAATGTATGGTTTCTCCAGGAAGATATCT GCTTTTTAAAACAACAAACAAGAACAACAACACAAAACTGGTAATGATTTGGAGTAATCATGCGGGCATATTGAGTCTGGGTAGTGTTTC GCTGGTGGTAGAGTGGTTGAGACTTCCTGGGAGGACTTTTTCCGCCTTCACTCCTACAGATTTAAGATTCTTTATTTTTAGGCCGGGCAT GGTGACTCACCTGAGGTGAGGAGTTCGAGACTAGCCTGGCCAACATGGCAAAACCCTGTCTCTACTAAAAGTACAAAAATTAGCCAGGCG TGGTGGTGCATGCCTATAATCCCAGCTACTCGGGAGGCTGAGGCAGGAGAATCGCTTGAACCCAGGAGGCAGAGGTTGCAGTAAGCCAAG GTCACACCACTGCACTCCAGCCTAGGTGACAGGGCGAGACTCTGTCTCTCAAAAAAAAAAAAAAAAAAAATTCTTTGAGACAGGGTCTCT AATATGTTGCCCAAGCTGATCTCAAACTCGTGGCTCAACTGATTCTCCCGCTTCAACCTTGAGTAGCTGGGACGACAGATACATGCTACT GCACCCAGCTAGGTTAAGGTTCTGTATACTCTGTCTTTCTAAAGCAATTCCCTGGAAATTTTCTCAACATTGGTTATAACTGAATGCTTG GGAAGGGAGGGGCAGCCCTTCAACAGGAAAAGGGGACTGCCCAAATCTTAAATGCACAGTCAAGATTCTCTTGGAGCTAGCAATGGTTCT CAGGTTAGTGAGCATTTAGCTTGCCTGCTGCAGAATTTGACCCAGATGGGGGTGTTTTTTAGGTCTCTCACAAATGAGACAAGCGAAACA ATTGTCTCCTTTTATTCTCTTTGGTGCATTGGTGCTGGGGAAACATGAACTAGCGGCAGTGTAACTGCAGAACATAGACCCAGTTCTACC AGGCCAGGCCAGCGTTGGGAACCACCAGACAGGGCTGCTTTGGGCTTTGCTTACAGTATTTCCATGTGTAGCCTGGCGTGTGAGAAAGTA TTAGGTGAAATGCCAGTTTCATGGTTCAGGTGAAAGTCTGTGATCATTCCCCTCGTGGCTCGTCCTTCACATCACTTTTGCCCTTCAAGG AGTTGCCGCGTCCCCGCTCAGTGCCGCCTGAGCCCTCAGAGCTCCCCTGTGCTTTTCTGGATGGGGACTGGCGGGGTCACCTAGCCTCAC CGTGGAGCCACCGTGCAATGCCCATCTCTGAGAGGCCCACGCGGTATTCCTCGTGCCCTGTGTTAGTGCTTTCTGTATAAGGGACAGACA GAACTGGGTTTTTTTCCTCTGTCTGGTTTTAGAGTAAATGTAACTAACTTTTATTTTCCCTTTATGAAAGATAGAAATTATTTTTATGTA GTTTCCAGACTTTATACAAAATTTTTGTAAAATGTTCTCTGGAAAGTTAACTACAAGAATGTAAATATGCTTCTAATAAAATAACAAGGT >81895_81895_2_SIL1-STK10_SIL1_chr5_138456724_ENST00000509534_STK10_chr5_171583792_ENST00000176763_length(amino acids)=935AA_BP=18 MPKSWRCSTRRMSGRPFSQAKNKETGALAAAKVIETKSEEELEDYIVEIEILATCDHPYIVKLLGAYYHDGKLWIMIEFCPGGAVDAIML ELDRGLTEPQIQVVCRQMLEALNFLHSKRIIHRDLKAGNVLMTLEGDIRLADFGVSAKNLKTLQKRDSFIGTPYWMAPEVVMCETMKDTP YDYKADIWSLGITLIEMAQIEPPHHELNPMRVLLKIAKSDPPTLLTPSKWSVEFRDFLKIALDKNPETRPSAAQLLEHPFVSSITSNKAL RELVAEAKAEVMEEIEDGRDEGEEEDAVDAASTLENHTQNSSEVSPPSLNADKPLEESPSTPLAPSQSQDSVNEPCSQPSGDRSLQTTSP PVVAPGNENGLAVPVPLRKSRPVSMDARIQVAQEKQVAEQGGDLSPAANRSQKASQSRPNSSALETLGGEKLANGSLEPPAQAAPGPSKR DSDCSSLCTSESMDYGTNLSTDLSLNKEMGSLSIKDPKLYKKTLKRTRKFVVDGVEVSITTSKIISEDEKKDEEMRFLRRQELRELRLLQ KEEHRNQTQLSNKHELQLEQMHKRFEQEINAKKKFFDTELENLERQQKQQVEKMEQDHAVRRREEARRIRLEQDRDYTRFQEQLKLMKKE VKNEVEKLPRQQRKESMKQKMEEHTQKKQLLDRDFVAKQKEDLELAMKRLTTDNRREICDKERECLMKKQELLRDREAALWEMEEHQLQE RHQLVKQQLKDQYFLQRHELLRKHEKEREQMQRYNQRMIEQLKVRQQQEKARLPKIQRSEGKTRMAMYKKSLHINGGGSAAEQREKIKQF SQQEEKRQKSERLQQQQKHENQMRDMLAQCESNMSELQQLQNEKCHLLVEHETQKLKALDESHNQNLKEWRDKLRPRKKALEEDLNQKKR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SIL1-STK10 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | SIL1 | chr5:138456724 | chr5:171583792 | ENST00000265195 | - | 4 | 11 | 1_256 | 81.33333333333333 | 462.0 | HSPA5 and localization to the endoplasmic reticulum |
| Hgene | SIL1 | chr5:138456724 | chr5:171583792 | ENST00000394817 | - | 3 | 10 | 1_256 | 81.33333333333333 | 462.0 | HSPA5 and localization to the endoplasmic reticulum |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SIL1-STK10 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SIL1-STK10 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
