
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SMAD3-IQCH (FusionGDB2 ID:83797) |
Fusion Gene Summary for SMAD3-IQCH |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SMAD3-IQCH | Fusion gene ID: 83797 | Hgene | Tgene | Gene symbol | SMAD3 | IQCH | Gene ID | 4088 | 128153 |
| Gene name | SMAD family member 3 | spermatogenesis associated 17 | |
| Synonyms | HSPC193|HsT17436|JV15-2|LDS1C|LDS3|MADH3 | CFAP305|FAP305|IQCH|MOT17|MSRG-11|MSRG11 | |
| Cytomap | 15q22.33 | 1q41 | |
| Type of gene | protein-coding | protein-coding | |
| Description | mothers against decapentaplegic homolog 3MAD homolog 3MAD, mothers against decapentaplegic homolog 3SMA- and MAD-related protein 3SMAD, mothers against DPP homolog 3hMAD-3hSMAD3mad homolog JV15-2mad protein homologmad3mothers against DPP homolog | spermatogenesis-associated protein 17IQ motif containing Hspermatogenesis-related protein 11 | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | . | Q86VS3 | |
| Ensembl transtripts involved in fusion gene | ENST00000327367, ENST00000439724, ENST00000537194, ENST00000540846, ENST00000559092, | ENST00000335894, ENST00000358767, ENST00000360277, ENST00000546225, ENST00000512104, ENST00000560790, | |
| Fusion gene scores | * DoF score | 17 X 13 X 12=2652 | 8 X 11 X 6=528 |
| # samples | 30 | 12 | |
| ** MAII score | log2(30/2652*10)=-3.14404636961671 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(12/528*10)=-2.13750352374993 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SMAD3 [Title/Abstract] AND IQCH [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SMAD3(67358698)-IQCH(67636401), # samples:1 SMAD3(67462942)-IQCH(67649683), # samples:1 SMAD3(67358698)-IQCH(67664449), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | SMAD3-IQCH seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. SMAD3-IQCH seems lost the major protein functional domain in Hgene partner, which is a transcription factor due to the frame-shifted ORF. SMAD3-IQCH seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SMAD3 | GO:0000122 | negative regulation of transcription by RNA polymerase II | 8774881 |
| Hgene | SMAD3 | GO:0006357 | regulation of transcription by RNA polymerase II | 21947082 |
| Hgene | SMAD3 | GO:0007179 | transforming growth factor beta receptor signaling pathway | 9732876|18548003|21947082 |
| Hgene | SMAD3 | GO:0007183 | SMAD protein complex assembly | 9111321|10823886 |
| Hgene | SMAD3 | GO:0010628 | positive regulation of gene expression | 21307346 |
| Hgene | SMAD3 | GO:0010718 | positive regulation of epithelial to mesenchymal transition | 21307346 |
| Hgene | SMAD3 | GO:0030308 | negative regulation of cell growth | 8774881 |
| Hgene | SMAD3 | GO:0045429 | positive regulation of nitric oxide biosynthetic process | 27038547 |
| Hgene | SMAD3 | GO:0045599 | negative regulation of fat cell differentiation | 19816956 |
| Hgene | SMAD3 | GO:0045893 | positive regulation of transcription, DNA-templated | 9111321|9311995|9732876 |
| Hgene | SMAD3 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 8774881|18832382 |
| Hgene | SMAD3 | GO:0051481 | negative regulation of cytosolic calcium ion concentration | 27038547 |
| Hgene | SMAD3 | GO:0071560 | cellular response to transforming growth factor beta stimulus | 12902338 |
| Hgene | SMAD3 | GO:1901203 | positive regulation of extracellular matrix assembly | 21307346 |
 Fusion gene breakpoints across SMAD3 (5'-gene) Fusion gene breakpoints across SMAD3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
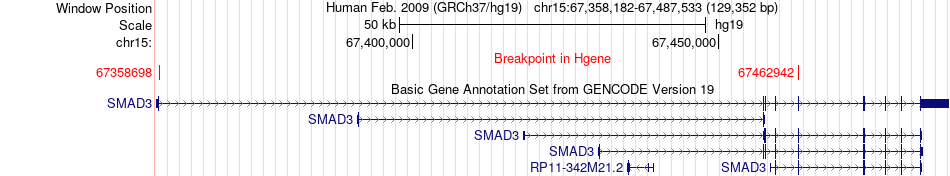 |
 Fusion gene breakpoints across IQCH (3'-gene) Fusion gene breakpoints across IQCH (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
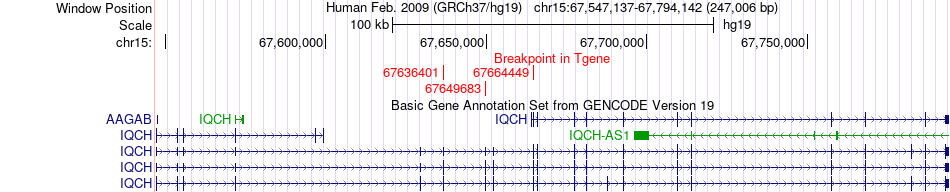 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-E9-A1RH-01A | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| ChimerDB4 | COAD | TCGA-AA-3489-01A | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| ChimerDB4 | STAD | TCGA-VQ-AA6D-01A | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
Top |
Fusion Gene ORF analysis for SMAD3-IQCH |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000327367 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000335894 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000546225 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000327367 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| 5CDS-5UTR | ENST00000439724 | ENST00000335894 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000439724 | ENST00000546225 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000537194 | ENST00000335894 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000537194 | ENST00000546225 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000540846 | ENST00000335894 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-5UTR | ENST00000540846 | ENST00000546225 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000327367 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| 5CDS-intron | ENST00000327367 | ENST00000360277 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000327367 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| 5CDS-intron | ENST00000327367 | ENST00000512104 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000327367 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| 5CDS-intron | ENST00000327367 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| 5CDS-intron | ENST00000327367 | ENST00000560790 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000327367 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| 5CDS-intron | ENST00000439724 | ENST00000360277 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000439724 | ENST00000512104 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000439724 | ENST00000560790 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000537194 | ENST00000360277 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000537194 | ENST00000512104 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000537194 | ENST00000560790 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000540846 | ENST00000360277 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000540846 | ENST00000512104 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| 5CDS-intron | ENST00000540846 | ENST00000560790 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| Frame-shift | ENST00000327367 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| In-frame | ENST00000327367 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| In-frame | ENST00000439724 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| In-frame | ENST00000537194 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| In-frame | ENST00000540846 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-3CDS | ENST00000439724 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-3CDS | ENST00000537194 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-3CDS | ENST00000540846 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-3CDS | ENST00000559092 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-3CDS | ENST00000559092 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000439724 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000439724 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000439724 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000439724 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000439724 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000439724 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000537194 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000537194 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000537194 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000537194 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000537194 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000537194 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000540846 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000540846 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000540846 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000540846 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000540846 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000540846 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000559092 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000559092 | ENST00000335894 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-5UTR | ENST00000559092 | ENST00000335894 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000559092 | ENST00000358767 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000559092 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-5UTR | ENST00000559092 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-5UTR | ENST00000559092 | ENST00000546225 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-5UTR | ENST00000559092 | ENST00000546225 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000439724 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000439724 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000439724 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000439724 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000439724 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000537194 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000537194 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000537194 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000537194 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000537194 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000540846 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000540846 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000540846 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000540846 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000540846 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000559092 | ENST00000360277 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000559092 | ENST00000360277 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-intron | ENST00000559092 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000559092 | ENST00000512104 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-intron | ENST00000559092 | ENST00000512104 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
| intron-intron | ENST00000559092 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636401 | + |
| intron-intron | ENST00000559092 | ENST00000560790 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + |
| intron-intron | ENST00000559092 | ENST00000560790 | SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664449 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000327367 | SMAD3 | chr15 | 67462942 | + | ENST00000358767 | IQCH | chr15 | 67649683 | + | 4137 | 968 | 310 | 2940 | 876 |
| ENST00000540846 | SMAD3 | chr15 | 67462942 | + | ENST00000358767 | IQCH | chr15 | 67649683 | + | 3890 | 721 | 273 | 2693 | 806 |
| ENST00000439724 | SMAD3 | chr15 | 67462942 | + | ENST00000358767 | IQCH | chr15 | 67649683 | + | 3721 | 552 | 26 | 2524 | 832 |
| ENST00000537194 | SMAD3 | chr15 | 67462942 | + | ENST00000358767 | IQCH | chr15 | 67649683 | + | 3470 | 301 | 195 | 2273 | 692 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000327367 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + | 0.000915179 | 0.99908483 |
| ENST00000540846 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + | 0.00048836 | 0.9995116 |
| ENST00000439724 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + | 0.000766676 | 0.99923337 |
| ENST00000537194 | ENST00000358767 | SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649683 | + | 0.00116281 | 0.9988372 |
Top |
Fusion Genomic Features for SMAD3-IQCH |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649682 | + | 0.9999831 | 1.69E-05 |
| SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636400 | + | 3.04E-06 | 0.9999969 |
| SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664448 | + | 1.21E-08 | 1 |
| SMAD3 | chr15 | 67462942 | + | IQCH | chr15 | 67649682 | + | 0.9999831 | 1.69E-05 |
| SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67636400 | + | 3.04E-06 | 0.9999969 |
| SMAD3 | chr15 | 67358698 | + | IQCH | chr15 | 67664448 | + | 1.21E-08 | 1 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
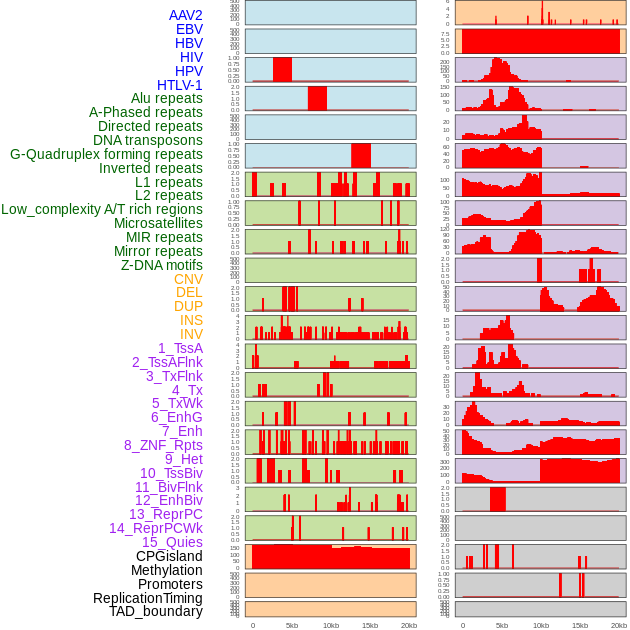 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
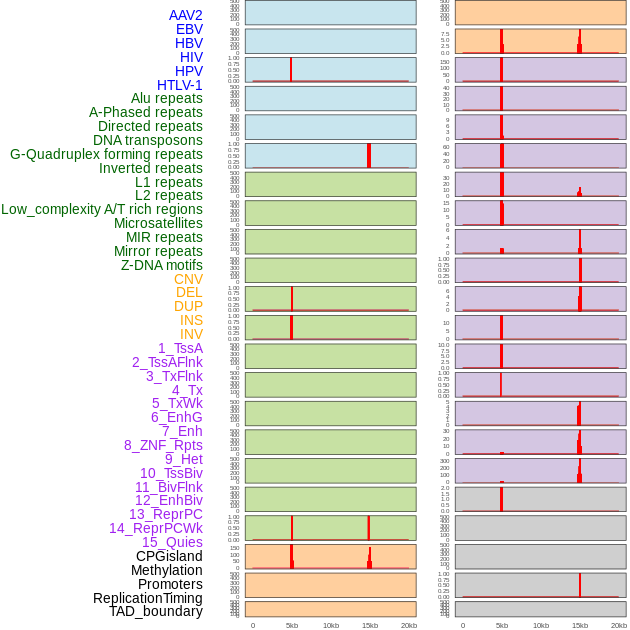 |
Top |
Fusion Protein Features for SMAD3-IQCH |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:67358698/chr15:67636401) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | IQCH |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: May play a regulatory role in spermatogenesis. {ECO:0000269|PubMed:15897968}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000327367 | + | 5 | 9 | 10_136 | 219 | 426.0 | Domain | MH1 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000439724 | + | 5 | 9 | 10_136 | 175 | 382.0 | Domain | MH1 |
| Tgene | IQCH | chr15:67462942 | chr15:67649683 | ENST00000335894 | 5 | 21 | 369_398 | 212 | 1028.0 | Domain | IQ | |
| Tgene | IQCH | chr15:67462942 | chr15:67649683 | ENST00000360277 | 0 | 12 | 369_398 | 0 | 1009.3333333333334 | Domain | IQ | |
| Tgene | IQCH | chr15:67462942 | chr15:67649683 | ENST00000512104 | 0 | 6 | 369_398 | 0 | 169.0 | Domain | IQ | |
| Tgene | IQCH | chr15:67462942 | chr15:67649683 | ENST00000546225 | 5 | 20 | 369_398 | 0 | 685.0 | Domain | IQ |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000327367 | + | 5 | 9 | 232_425 | 219 | 426.0 | Domain | MH2 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000439724 | + | 5 | 9 | 232_425 | 175 | 382.0 | Domain | MH2 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000537194 | + | 3 | 7 | 10_136 | 24 | 231.0 | Domain | MH1 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000537194 | + | 3 | 7 | 232_425 | 24 | 231.0 | Domain | MH2 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000540846 | + | 5 | 9 | 10_136 | 114 | 321.0 | Domain | MH1 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000540846 | + | 5 | 9 | 232_425 | 114 | 321.0 | Domain | MH2 |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000327367 | + | 5 | 9 | 137_231 | 219 | 426.0 | Region | Note=Linker |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000439724 | + | 5 | 9 | 137_231 | 175 | 382.0 | Region | Note=Linker |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000537194 | + | 3 | 7 | 137_231 | 24 | 231.0 | Region | Note=Linker |
| Hgene | SMAD3 | chr15:67462942 | chr15:67649683 | ENST00000540846 | + | 5 | 9 | 137_231 | 114 | 321.0 | Region | Note=Linker |
Top |
Fusion Gene Sequence for SMAD3-IQCH |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >83797_83797_1_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000327367_IQCH_chr15_67649683_ENST00000358767_length(transcript)=4137nt_BP=968nt ACTTGGAGTCTCGCGGCCGCCGCCTCCGCCCCGCGTTCGGGGCCTTCCCGACCCTGCACTGCTGCCGTCCGCCCGCCCGGCCGCTCTTCT CTTCGCCGTGGGAGCCGCTCCGGGCGCAGGGCCGCGCGCCGAGCCCCGCAGGCTGCAGCGCCGCGGCCCGGCCCGGCGCCCCGGCAACTT CGCCGAGAGTTGAGGCGAAGTTTGGGCGACCGCGGCAGGCCCCGGCCGAGCTCCCCTCTGCGCCCCCGGCGTCCCGTCGAGCCCAGCCCC GCCGGGGGCGCTCCTCGCCGCCCGCGCGCCCTCCCCAGCCATGTCGTCCATCCTGCCTTTCACTCCCCCGATCGTGAAGCGCCTGCTGGG CTGGAAGAAGGGCGAGCAGAACGGGCAGGAGGAGAAATGGTGCGAGAAGGCGGTCAAGAGCCTGGTCAAGAAACTCAAGAAGACGGGGCA GCTGGACGAGCTGGAGAAGGCCATCACCACGCAGAACGTCAACACCAAGTGCATCACCATCCCCAGGTCCCTGGATGGCCGGTTGCAGGT GTCCCATCGGAAGGGGCTCCCTCATGTCATCTACTGCCGCCTGTGGCGATGGCCAGACCTGCACAGCCACCACGAGCTACGGGCCATGGA GCTGTGTGAGTTCGCCTTCAATATGAAGAAGGACGAGGTCTGCGTGAATCCCTACCACTACCAGAGAGTAGAGACACCAGTTCTACCTCC TGTGTTGGTGCCACGCCACACAGAGATCCCGGCCGAGTTCCCCCCACTGGACGACTACAGCCATTCCATCCCCGAAAACACTAACTTCCC CGCAGGCATCGAGCCCCAGAGCAATATTCCAGAGACCCCACCCCCTGGCTACCTGAGTGAAGATGGAGAAACCAGTGACCACCAGATGAA CCACAGCATGGACGCAGGTTCTCCAAACCTATCCCCGAATCCGATGTCCCCAGCACATAATAACTTGGCCACTTTCACTATACCTCGGGA ACCACCTCCATCTCCAGCAGAAGTGAAGTTCTTTCCCAAGAAACAAAGATCAAAGGGGAAAAGCAGAAGGTCAAGAGGACATCATGATAG GAAGGCCATGAAAGTCAAAACACCTTTGAGAGCCCTGAAATCACTGTGGGATTATGACTTTTTAATTTATGATGGTGTCATAGACAATAC AGCCCCAGACTTCTTAGCATTCAAGGAACATTTTAGCTTAGCTTGGGGAGGTATTTTTTCTCTCTTGGAACACGTCGAGAAGTTTCTCAG GAACTATGCTATACCAGAAGTCAAAATAAAAGGGAATAATTTGGTGGCCCTCCTTCCAGAGTTTGAGCTGACGAATAAACTTACCAGATA TGACCTTCTCTCAGTGTTAGAGGACCCAGCTCATGTCCAAATGCTGATAAATCTTCCAGGGCAAAGGTACAAGGGCCAAGATGGAAATTC GGAGGCCGCCATGAAGATCCAAGCCACATGGAAATGCTACAAAGCAAGAAAATTCTTCCTCTTTTATCGCCAGCAGAAGTGGGCATCAGG TGTGATTGCCATTGCTTGGCTGTTATATTGCCATAAGACTCGACTAAAGAAGATACTAAAGGAATCACGTCAGAGACACCTGGAGAATTT TCGCATTCGAGCCAAGCATCTGGCAGCCAACTGGAATCGCATCAGGACCTCCAGGAGGACTATTATCCATATCCCATCATTAGGGTATTC CCAGCCTGTGAGAGAACATATTGCCGATTTCAACACACAGCAGAACATGCAGCTGGGGAGGCTGTGTGACATCTTAGATGCCAATGTGAA TGTCATCTACATCTGCTCCCATCATATGAATGACGAGTTAGTGCTGTATTACAAAAAAATCCTAAGTCTACATGCAGCCGTCAAATCTGG GAACCTTGAGGACAGAAGTGACCTGCAGGACAGGTTCAAAATTATCACACCTGAAGCTGTAAACATCTTCCCTATGATAGAGCAGCTGAG TCAGCTGATAACTGATCACCTGCAAATACAGCGTTGGCTCTTTAAAATGGACTCTGAGTTCCGAGGAAATGGGACTGCATTTTGTGATAT TCCTTCCTACCTAAAGTGCTACAAATGGGTGCTAAAGGAGAGTAGCAGATATGGCCTTGAAGACTGGAGAAAGAAATGGGCACAAGAGCC AGCTTTGGTGAAGATCTCTGAGGAGCTGGCGGGCATTTTAGCACAGCACGCACAGCCAGTCAATGAGAAACGGTTCCCGACGTGGAGGAA ATTCCTCCAAACATTTCTCAGTCAAGGGGGTGTGATCGAAGCATTCCCACCTGCAGACAATGTCACCAACCTCACAGTGGACATGCTGAT AGAGCCCAACGGGAAAATCAGCGTGCTGTCGACAGGGGACCAGCTTCATGCTGAAAGCCCCTTCATCTCCTCTGGTACCACCGTGCCTCA GACCTCAGTGGATCCCCAAGTTCTCACTTATTTGTGCCTCCAAATTGGAAAAGCCTGCAGAATGAGAGATGTGGTTGGTTACTTTTCGAT AGATCTGGTGACTTTTATAGATCCAAGCACCTTGGAACAACAGGTGTGGGCAACCGGCCTTAACCTCGCATATAGTGACCAGCTGGCCCT GACTCAACTCACCTTATACCTGACAAACGGCCATCTGGATTGCAGTTTGAGCACCCTGGAAGTGCCCCGCTTTGTTCCAAAGGAAAGGAA GAAAACCAAATGCATGAGTGCGCTGTCAATGCCGATGCTGGCAACCAGTCGCTATGCAGTGATGACCACCCAGCTAAGACACAGCAATCT CTCACTGGTTTTCCACTATGTTTTTCTCCAGATCTGTAGGGCCCATGGCATTGGCTATGATGTTGAGAACAATCGGCGAGGATCTCCAGG GGGTCCTCATGACCTTTGCTCGCCATCTCTTCATCATCCATCAAGAAATATCAGCACCTAATATGCAAGGCGAGACCAATTTTAAGACCA CCATTGCTGATATTGAAACTATTCTAAGAGTAACAAAGGAAAACAAAATGAGATTTGAAGAGGAGCAACAGTCCAAAGATGATAAAAACC TCTCTAAACCCAAGAAATGATCCTGGAATACAGTACATAACAATTTGGATCCCAGTCTGGAATAAAAAGGGCAATTTTTTTTTCTGTTAG AAATAAAAGCCAGGGGAAATTGGTTTGCTTTGTGTGCTAGGAGGTGAATCAGAACAGATTATAATGAAATGCTCTTTTTAAAACATTGTT TATTAAGTGATCTTATTTTATTTATTAAACCAAAACTTATTTGTGTTTTCATTTGAGAGTGTTGAACAATCCCTTCTTCTTCTCAAACTC AGAAAAAAGTAATCTGATAAAAGAAGAAAGTTAAAAGTCTTACTGATATCACCTCCGCATTTACTTCCTCATAGGCCTCAGGATTATGTA GCTTTTATTTTTTATGTTTTATAAAGTTTTCTCCTATTTCTAATAGTCCATCGATCTTCTGCATTTATAGGTTTGAATAAAGGCTTAAGA ATTGCTATTTGTCAAAACACAAATGCCTTTTTTTCAAAGCTTCAAATTATATTGAAATTATATTTATATGAATTTTTAAGTGACTATCAT TCATCGTGTTTTTGTTATACCAAGCCACTATTGTCTGCTATTGTTGCCATTTTTCAAAAAGTAGGATAAGCCAGCATCTAGTTAACCAAA GCATCAGAATGTATCTAGGTGGCAGTATCTGCTCTTAGTTTAAGATGGAGTGTAGCTCGAGCTCTGTAGCTTGAATAAACATTTGTATTC TCTAGTTCTCCATTTAATACAGCAGTAGGATTTTTTCAAGTTTAGTTCAACATTATACTACCTTTAAAAAGGCAGACTGTTAAGATTGTA GTAGTGCTGACTCTAACTTTCAGCATTCACTTCATAATTGTCCAAAAATATTTCCACATCAGTGAGCTTTACTGAAGGGTGAAATAAAAG CAAATTACAACACCAAAACCATTATTGTTTTAAAGAATGACCTTTATGTCTATAAAAATATTTTAGTGTATTGGGGAAAGGAATAGGATT >83797_83797_1_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000327367_IQCH_chr15_67649683_ENST00000358767_length(amino acids)=876AA_BP=0 MSSILPFTPPIVKRLLGWKKGEQNGQEEKWCEKAVKSLVKKLKKTGQLDELEKAITTQNVNTKCITIPRSLDGRLQVSHRKGLPHVIYCR LWRWPDLHSHHELRAMELCEFAFNMKKDEVCVNPYHYQRVETPVLPPVLVPRHTEIPAEFPPLDDYSHSIPENTNFPAGIEPQSNIPETP PPGYLSEDGETSDHQMNHSMDAGSPNLSPNPMSPAHNNLATFTIPREPPPSPAEVKFFPKKQRSKGKSRRSRGHHDRKAMKVKTPLRALK SLWDYDFLIYDGVIDNTAPDFLAFKEHFSLAWGGIFSLLEHVEKFLRNYAIPEVKIKGNNLVALLPEFELTNKLTRYDLLSVLEDPAHVQ MLINLPGQRYKGQDGNSEAAMKIQATWKCYKARKFFLFYRQQKWASGVIAIAWLLYCHKTRLKKILKESRQRHLENFRIRAKHLAANWNR IRTSRRTIIHIPSLGYSQPVREHIADFNTQQNMQLGRLCDILDANVNVIYICSHHMNDELVLYYKKILSLHAAVKSGNLEDRSDLQDRFK IITPEAVNIFPMIEQLSQLITDHLQIQRWLFKMDSEFRGNGTAFCDIPSYLKCYKWVLKESSRYGLEDWRKKWAQEPALVKISEELAGIL AQHAQPVNEKRFPTWRKFLQTFLSQGGVIEAFPPADNVTNLTVDMLIEPNGKISVLSTGDQLHAESPFISSGTTVPQTSVDPQVLTYLCL QIGKACRMRDVVGYFSIDLVTFIDPSTLEQQVWATGLNLAYSDQLALTQLTLYLTNGHLDCSLSTLEVPRFVPKERKKTKCMSALSMPML -------------------------------------------------------------- >83797_83797_2_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000439724_IQCH_chr15_67649683_ENST00000358767_length(transcript)=3721nt_BP=552nt CCCTGTGACCTCCCAACTTCACAAACATGTCTTGCCTGCACCCTAGGCAAACGTGGAAAGGCGCAGCTCTGGTACACCGGAAAGCATGGT GGATGGGGAGGTCCCTGGATGGCCGGTTGCAGGTGTCCCATCGGAAGGGGCTCCCTCATGTCATCTACTGCCGCCTGTGGCGATGGCCAG ACCTGCACAGCCACCACGAGCTACGGGCCATGGAGCTGTGTGAGTTCGCCTTCAATATGAAGAAGGACGAGGTCTGCGTGAATCCCTACC ACTACCAGAGAGTAGAGACACCAGTTCTACCTCCTGTGTTGGTGCCACGCCACACAGAGATCCCGGCCGAGTTCCCCCCACTGGACGACT ACAGCCATTCCATCCCCGAAAACACTAACTTCCCCGCAGGCATCGAGCCCCAGAGCAATATTCCAGAGACCCCACCCCCTGGCTACCTGA GTGAAGATGGAGAAACCAGTGACCACCAGATGAACCACAGCATGGACGCAGGTTCTCCAAACCTATCCCCGAATCCGATGTCCCCAGCAC ATAATAACTTGGCCACTTTCACTATACCTCGGGAACCACCTCCATCTCCAGCAGAAGTGAAGTTCTTTCCCAAGAAACAAAGATCAAAGG GGAAAAGCAGAAGGTCAAGAGGACATCATGATAGGAAGGCCATGAAAGTCAAAACACCTTTGAGAGCCCTGAAATCACTGTGGGATTATG ACTTTTTAATTTATGATGGTGTCATAGACAATACAGCCCCAGACTTCTTAGCATTCAAGGAACATTTTAGCTTAGCTTGGGGAGGTATTT TTTCTCTCTTGGAACACGTCGAGAAGTTTCTCAGGAACTATGCTATACCAGAAGTCAAAATAAAAGGGAATAATTTGGTGGCCCTCCTTC CAGAGTTTGAGCTGACGAATAAACTTACCAGATATGACCTTCTCTCAGTGTTAGAGGACCCAGCTCATGTCCAAATGCTGATAAATCTTC CAGGGCAAAGGTACAAGGGCCAAGATGGAAATTCGGAGGCCGCCATGAAGATCCAAGCCACATGGAAATGCTACAAAGCAAGAAAATTCT TCCTCTTTTATCGCCAGCAGAAGTGGGCATCAGGTGTGATTGCCATTGCTTGGCTGTTATATTGCCATAAGACTCGACTAAAGAAGATAC TAAAGGAATCACGTCAGAGACACCTGGAGAATTTTCGCATTCGAGCCAAGCATCTGGCAGCCAACTGGAATCGCATCAGGACCTCCAGGA GGACTATTATCCATATCCCATCATTAGGGTATTCCCAGCCTGTGAGAGAACATATTGCCGATTTCAACACACAGCAGAACATGCAGCTGG GGAGGCTGTGTGACATCTTAGATGCCAATGTGAATGTCATCTACATCTGCTCCCATCATATGAATGACGAGTTAGTGCTGTATTACAAAA AAATCCTAAGTCTACATGCAGCCGTCAAATCTGGGAACCTTGAGGACAGAAGTGACCTGCAGGACAGGTTCAAAATTATCACACCTGAAG CTGTAAACATCTTCCCTATGATAGAGCAGCTGAGTCAGCTGATAACTGATCACCTGCAAATACAGCGTTGGCTCTTTAAAATGGACTCTG AGTTCCGAGGAAATGGGACTGCATTTTGTGATATTCCTTCCTACCTAAAGTGCTACAAATGGGTGCTAAAGGAGAGTAGCAGATATGGCC TTGAAGACTGGAGAAAGAAATGGGCACAAGAGCCAGCTTTGGTGAAGATCTCTGAGGAGCTGGCGGGCATTTTAGCACAGCACGCACAGC CAGTCAATGAGAAACGGTTCCCGACGTGGAGGAAATTCCTCCAAACATTTCTCAGTCAAGGGGGTGTGATCGAAGCATTCCCACCTGCAG ACAATGTCACCAACCTCACAGTGGACATGCTGATAGAGCCCAACGGGAAAATCAGCGTGCTGTCGACAGGGGACCAGCTTCATGCTGAAA GCCCCTTCATCTCCTCTGGTACCACCGTGCCTCAGACCTCAGTGGATCCCCAAGTTCTCACTTATTTGTGCCTCCAAATTGGAAAAGCCT GCAGAATGAGAGATGTGGTTGGTTACTTTTCGATAGATCTGGTGACTTTTATAGATCCAAGCACCTTGGAACAACAGGTGTGGGCAACCG GCCTTAACCTCGCATATAGTGACCAGCTGGCCCTGACTCAACTCACCTTATACCTGACAAACGGCCATCTGGATTGCAGTTTGAGCACCC TGGAAGTGCCCCGCTTTGTTCCAAAGGAAAGGAAGAAAACCAAATGCATGAGTGCGCTGTCAATGCCGATGCTGGCAACCAGTCGCTATG CAGTGATGACCACCCAGCTAAGACACAGCAATCTCTCACTGGTTTTCCACTATGTTTTTCTCCAGATCTGTAGGGCCCATGGCATTGGCT ATGATGTTGAGAACAATCGGCGAGGATCTCCAGGGGGTCCTCATGACCTTTGCTCGCCATCTCTTCATCATCCATCAAGAAATATCAGCA CCTAATATGCAAGGCGAGACCAATTTTAAGACCACCATTGCTGATATTGAAACTATTCTAAGAGTAACAAAGGAAAACAAAATGAGATTT GAAGAGGAGCAACAGTCCAAAGATGATAAAAACCTCTCTAAACCCAAGAAATGATCCTGGAATACAGTACATAACAATTTGGATCCCAGT CTGGAATAAAAAGGGCAATTTTTTTTTCTGTTAGAAATAAAAGCCAGGGGAAATTGGTTTGCTTTGTGTGCTAGGAGGTGAATCAGAACA GATTATAATGAAATGCTCTTTTTAAAACATTGTTTATTAAGTGATCTTATTTTATTTATTAAACCAAAACTTATTTGTGTTTTCATTTGA GAGTGTTGAACAATCCCTTCTTCTTCTCAAACTCAGAAAAAAGTAATCTGATAAAAGAAGAAAGTTAAAAGTCTTACTGATATCACCTCC GCATTTACTTCCTCATAGGCCTCAGGATTATGTAGCTTTTATTTTTTATGTTTTATAAAGTTTTCTCCTATTTCTAATAGTCCATCGATC TTCTGCATTTATAGGTTTGAATAAAGGCTTAAGAATTGCTATTTGTCAAAACACAAATGCCTTTTTTTCAAAGCTTCAAATTATATTGAA ATTATATTTATATGAATTTTTAAGTGACTATCATTCATCGTGTTTTTGTTATACCAAGCCACTATTGTCTGCTATTGTTGCCATTTTTCA AAAAGTAGGATAAGCCAGCATCTAGTTAACCAAAGCATCAGAATGTATCTAGGTGGCAGTATCTGCTCTTAGTTTAAGATGGAGTGTAGC TCGAGCTCTGTAGCTTGAATAAACATTTGTATTCTCTAGTTCTCCATTTAATACAGCAGTAGGATTTTTTCAAGTTTAGTTCAACATTAT ACTACCTTTAAAAAGGCAGACTGTTAAGATTGTAGTAGTGCTGACTCTAACTTTCAGCATTCACTTCATAATTGTCCAAAAATATTTCCA CATCAGTGAGCTTTACTGAAGGGTGAAATAAAAGCAAATTACAACACCAAAACCATTATTGTTTTAAAGAATGACCTTTATGTCTATAAA AATATTTTAGTGTATTGGGGAAAGGAATAGGATTTTTACCTGTGGAATTTCAATAATTTGTTAACTGTGATTCAGCCACAGGCTAAGAGT >83797_83797_2_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000439724_IQCH_chr15_67649683_ENST00000358767_length(amino acids)=832AA_BP=0 MSCLHPRQTWKGAALVHRKAWWMGRSLDGRLQVSHRKGLPHVIYCRLWRWPDLHSHHELRAMELCEFAFNMKKDEVCVNPYHYQRVETPV LPPVLVPRHTEIPAEFPPLDDYSHSIPENTNFPAGIEPQSNIPETPPPGYLSEDGETSDHQMNHSMDAGSPNLSPNPMSPAHNNLATFTI PREPPPSPAEVKFFPKKQRSKGKSRRSRGHHDRKAMKVKTPLRALKSLWDYDFLIYDGVIDNTAPDFLAFKEHFSLAWGGIFSLLEHVEK FLRNYAIPEVKIKGNNLVALLPEFELTNKLTRYDLLSVLEDPAHVQMLINLPGQRYKGQDGNSEAAMKIQATWKCYKARKFFLFYRQQKW ASGVIAIAWLLYCHKTRLKKILKESRQRHLENFRIRAKHLAANWNRIRTSRRTIIHIPSLGYSQPVREHIADFNTQQNMQLGRLCDILDA NVNVIYICSHHMNDELVLYYKKILSLHAAVKSGNLEDRSDLQDRFKIITPEAVNIFPMIEQLSQLITDHLQIQRWLFKMDSEFRGNGTAF CDIPSYLKCYKWVLKESSRYGLEDWRKKWAQEPALVKISEELAGILAQHAQPVNEKRFPTWRKFLQTFLSQGGVIEAFPPADNVTNLTVD MLIEPNGKISVLSTGDQLHAESPFISSGTTVPQTSVDPQVLTYLCLQIGKACRMRDVVGYFSIDLVTFIDPSTLEQQVWATGLNLAYSDQ LALTQLTLYLTNGHLDCSLSTLEVPRFVPKERKKTKCMSALSMPMLATSRYAVMTTQLRHSNLSLVFHYVFLQICRAHGIGYDVENNRRG -------------------------------------------------------------- >83797_83797_3_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000537194_IQCH_chr15_67649683_ENST00000358767_length(transcript)=3470nt_BP=301nt AGTAATCCTGCTGCGTTCCTCTTAGAGCATTCTAATTTGGACGTGACGTTCATCCTGGGTTAGTTTACTGGGCTGAGATTGGCTACAGGC CTGTGTGCTTGGGTCAGAGTGTTAAGGAGGAGACCCACTGTCCAACCTTCTCAGATCCTTTGCGGGTAGCCCTGGCGTCCCGCGGAGACC CCACCCCCTGGCTACCTGAGTGAAGATGGAGAAACCAGTGACCACCAGATGAACCACAGCATGGACGCAGGTTCTCCAAACCTATCCCCG AATCCGATGTCCCCAGCACATAATAACTTGGCCACTTTCACTATACCTCGGGAACCACCTCCATCTCCAGCAGAAGTGAAGTTCTTTCCC AAGAAACAAAGATCAAAGGGGAAAAGCAGAAGGTCAAGAGGACATCATGATAGGAAGGCCATGAAAGTCAAAACACCTTTGAGAGCCCTG AAATCACTGTGGGATTATGACTTTTTAATTTATGATGGTGTCATAGACAATACAGCCCCAGACTTCTTAGCATTCAAGGAACATTTTAGC TTAGCTTGGGGAGGTATTTTTTCTCTCTTGGAACACGTCGAGAAGTTTCTCAGGAACTATGCTATACCAGAAGTCAAAATAAAAGGGAAT AATTTGGTGGCCCTCCTTCCAGAGTTTGAGCTGACGAATAAACTTACCAGATATGACCTTCTCTCAGTGTTAGAGGACCCAGCTCATGTC CAAATGCTGATAAATCTTCCAGGGCAAAGGTACAAGGGCCAAGATGGAAATTCGGAGGCCGCCATGAAGATCCAAGCCACATGGAAATGC TACAAAGCAAGAAAATTCTTCCTCTTTTATCGCCAGCAGAAGTGGGCATCAGGTGTGATTGCCATTGCTTGGCTGTTATATTGCCATAAG ACTCGACTAAAGAAGATACTAAAGGAATCACGTCAGAGACACCTGGAGAATTTTCGCATTCGAGCCAAGCATCTGGCAGCCAACTGGAAT CGCATCAGGACCTCCAGGAGGACTATTATCCATATCCCATCATTAGGGTATTCCCAGCCTGTGAGAGAACATATTGCCGATTTCAACACA CAGCAGAACATGCAGCTGGGGAGGCTGTGTGACATCTTAGATGCCAATGTGAATGTCATCTACATCTGCTCCCATCATATGAATGACGAG TTAGTGCTGTATTACAAAAAAATCCTAAGTCTACATGCAGCCGTCAAATCTGGGAACCTTGAGGACAGAAGTGACCTGCAGGACAGGTTC AAAATTATCACACCTGAAGCTGTAAACATCTTCCCTATGATAGAGCAGCTGAGTCAGCTGATAACTGATCACCTGCAAATACAGCGTTGG CTCTTTAAAATGGACTCTGAGTTCCGAGGAAATGGGACTGCATTTTGTGATATTCCTTCCTACCTAAAGTGCTACAAATGGGTGCTAAAG GAGAGTAGCAGATATGGCCTTGAAGACTGGAGAAAGAAATGGGCACAAGAGCCAGCTTTGGTGAAGATCTCTGAGGAGCTGGCGGGCATT TTAGCACAGCACGCACAGCCAGTCAATGAGAAACGGTTCCCGACGTGGAGGAAATTCCTCCAAACATTTCTCAGTCAAGGGGGTGTGATC GAAGCATTCCCACCTGCAGACAATGTCACCAACCTCACAGTGGACATGCTGATAGAGCCCAACGGGAAAATCAGCGTGCTGTCGACAGGG GACCAGCTTCATGCTGAAAGCCCCTTCATCTCCTCTGGTACCACCGTGCCTCAGACCTCAGTGGATCCCCAAGTTCTCACTTATTTGTGC CTCCAAATTGGAAAAGCCTGCAGAATGAGAGATGTGGTTGGTTACTTTTCGATAGATCTGGTGACTTTTATAGATCCAAGCACCTTGGAA CAACAGGTGTGGGCAACCGGCCTTAACCTCGCATATAGTGACCAGCTGGCCCTGACTCAACTCACCTTATACCTGACAAACGGCCATCTG GATTGCAGTTTGAGCACCCTGGAAGTGCCCCGCTTTGTTCCAAAGGAAAGGAAGAAAACCAAATGCATGAGTGCGCTGTCAATGCCGATG CTGGCAACCAGTCGCTATGCAGTGATGACCACCCAGCTAAGACACAGCAATCTCTCACTGGTTTTCCACTATGTTTTTCTCCAGATCTGT AGGGCCCATGGCATTGGCTATGATGTTGAGAACAATCGGCGAGGATCTCCAGGGGGTCCTCATGACCTTTGCTCGCCATCTCTTCATCAT CCATCAAGAAATATCAGCACCTAATATGCAAGGCGAGACCAATTTTAAGACCACCATTGCTGATATTGAAACTATTCTAAGAGTAACAAA GGAAAACAAAATGAGATTTGAAGAGGAGCAACAGTCCAAAGATGATAAAAACCTCTCTAAACCCAAGAAATGATCCTGGAATACAGTACA TAACAATTTGGATCCCAGTCTGGAATAAAAAGGGCAATTTTTTTTTCTGTTAGAAATAAAAGCCAGGGGAAATTGGTTTGCTTTGTGTGC TAGGAGGTGAATCAGAACAGATTATAATGAAATGCTCTTTTTAAAACATTGTTTATTAAGTGATCTTATTTTATTTATTAAACCAAAACT TATTTGTGTTTTCATTTGAGAGTGTTGAACAATCCCTTCTTCTTCTCAAACTCAGAAAAAAGTAATCTGATAAAAGAAGAAAGTTAAAAG TCTTACTGATATCACCTCCGCATTTACTTCCTCATAGGCCTCAGGATTATGTAGCTTTTATTTTTTATGTTTTATAAAGTTTTCTCCTAT TTCTAATAGTCCATCGATCTTCTGCATTTATAGGTTTGAATAAAGGCTTAAGAATTGCTATTTGTCAAAACACAAATGCCTTTTTTTCAA AGCTTCAAATTATATTGAAATTATATTTATATGAATTTTTAAGTGACTATCATTCATCGTGTTTTTGTTATACCAAGCCACTATTGTCTG CTATTGTTGCCATTTTTCAAAAAGTAGGATAAGCCAGCATCTAGTTAACCAAAGCATCAGAATGTATCTAGGTGGCAGTATCTGCTCTTA GTTTAAGATGGAGTGTAGCTCGAGCTCTGTAGCTTGAATAAACATTTGTATTCTCTAGTTCTCCATTTAATACAGCAGTAGGATTTTTTC AAGTTTAGTTCAACATTATACTACCTTTAAAAAGGCAGACTGTTAAGATTGTAGTAGTGCTGACTCTAACTTTCAGCATTCACTTCATAA TTGTCCAAAAATATTTCCACATCAGTGAGCTTTACTGAAGGGTGAAATAAAAGCAAATTACAACACCAAAACCATTATTGTTTTAAAGAA TGACCTTTATGTCTATAAAAATATTTTAGTGTATTGGGGAAAGGAATAGGATTTTTACCTGTGGAATTTCAATAATTTGTTAACTGTGAT >83797_83797_3_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000537194_IQCH_chr15_67649683_ENST00000358767_length(amino acids)=692AA_BP=1 MSEDGETSDHQMNHSMDAGSPNLSPNPMSPAHNNLATFTIPREPPPSPAEVKFFPKKQRSKGKSRRSRGHHDRKAMKVKTPLRALKSLWD YDFLIYDGVIDNTAPDFLAFKEHFSLAWGGIFSLLEHVEKFLRNYAIPEVKIKGNNLVALLPEFELTNKLTRYDLLSVLEDPAHVQMLIN LPGQRYKGQDGNSEAAMKIQATWKCYKARKFFLFYRQQKWASGVIAIAWLLYCHKTRLKKILKESRQRHLENFRIRAKHLAANWNRIRTS RRTIIHIPSLGYSQPVREHIADFNTQQNMQLGRLCDILDANVNVIYICSHHMNDELVLYYKKILSLHAAVKSGNLEDRSDLQDRFKIITP EAVNIFPMIEQLSQLITDHLQIQRWLFKMDSEFRGNGTAFCDIPSYLKCYKWVLKESSRYGLEDWRKKWAQEPALVKISEELAGILAQHA QPVNEKRFPTWRKFLQTFLSQGGVIEAFPPADNVTNLTVDMLIEPNGKISVLSTGDQLHAESPFISSGTTVPQTSVDPQVLTYLCLQIGK ACRMRDVVGYFSIDLVTFIDPSTLEQQVWATGLNLAYSDQLALTQLTLYLTNGHLDCSLSTLEVPRFVPKERKKTKCMSALSMPMLATSR -------------------------------------------------------------- >83797_83797_4_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000540846_IQCH_chr15_67649683_ENST00000358767_length(transcript)=3890nt_BP=721nt AAATATGAGCTTGTGCTTGCTGGAGGAGGATGACAGAGGAGCCTGCTGCTGAGTTCACTGGTGCTGGGGTTAGGTCACTGCTGGGCTGAA GCGCACTGACCATAAGAGCAACATGTGGGCAAGAGCCGCGGCACTGGGGTAATTTATTGCCGCCGCTCGCTTCACCAGGAACCCCACACG CTGGGTTCCCACAGGATGCGACATTCCCACAGGATGGGACAACTGCATGGAAACCCACACTCGGGCCTGTGTTGAGCAACCACGTTTGAG TCCCTGGATGGCCGGTTGCAGGTGTCCCATCGGAAGGGGCTCCCTCATGTCATCTACTGCCGCCTGTGGCGATGGCCAGACCTGCACAGC CACCACGAGCTACGGGCCATGGAGCTGTGTGAGTTCGCCTTCAATATGAAGAAGGACGAGGTCTGCGTGAATCCCTACCACTACCAGAGA GTAGAGACACCAGTTCTACCTCCTGTGTTGGTGCCACGCCACACAGAGATCCCGGCCGAGTTCCCCCCACTGGACGACTACAGCCATTCC ATCCCCGAAAACACTAACTTCCCCGCAGGCATCGAGCCCCAGAGCAATATTCCAGAGACCCCACCCCCTGGCTACCTGAGTGAAGATGGA GAAACCAGTGACCACCAGATGAACCACAGCATGGACGCAGGTTCTCCAAACCTATCCCCGAATCCGATGTCCCCAGCACATAATAACTTG GCCACTTTCACTATACCTCGGGAACCACCTCCATCTCCAGCAGAAGTGAAGTTCTTTCCCAAGAAACAAAGATCAAAGGGGAAAAGCAGA AGGTCAAGAGGACATCATGATAGGAAGGCCATGAAAGTCAAAACACCTTTGAGAGCCCTGAAATCACTGTGGGATTATGACTTTTTAATT TATGATGGTGTCATAGACAATACAGCCCCAGACTTCTTAGCATTCAAGGAACATTTTAGCTTAGCTTGGGGAGGTATTTTTTCTCTCTTG GAACACGTCGAGAAGTTTCTCAGGAACTATGCTATACCAGAAGTCAAAATAAAAGGGAATAATTTGGTGGCCCTCCTTCCAGAGTTTGAG CTGACGAATAAACTTACCAGATATGACCTTCTCTCAGTGTTAGAGGACCCAGCTCATGTCCAAATGCTGATAAATCTTCCAGGGCAAAGG TACAAGGGCCAAGATGGAAATTCGGAGGCCGCCATGAAGATCCAAGCCACATGGAAATGCTACAAAGCAAGAAAATTCTTCCTCTTTTAT CGCCAGCAGAAGTGGGCATCAGGTGTGATTGCCATTGCTTGGCTGTTATATTGCCATAAGACTCGACTAAAGAAGATACTAAAGGAATCA CGTCAGAGACACCTGGAGAATTTTCGCATTCGAGCCAAGCATCTGGCAGCCAACTGGAATCGCATCAGGACCTCCAGGAGGACTATTATC CATATCCCATCATTAGGGTATTCCCAGCCTGTGAGAGAACATATTGCCGATTTCAACACACAGCAGAACATGCAGCTGGGGAGGCTGTGT GACATCTTAGATGCCAATGTGAATGTCATCTACATCTGCTCCCATCATATGAATGACGAGTTAGTGCTGTATTACAAAAAAATCCTAAGT CTACATGCAGCCGTCAAATCTGGGAACCTTGAGGACAGAAGTGACCTGCAGGACAGGTTCAAAATTATCACACCTGAAGCTGTAAACATC TTCCCTATGATAGAGCAGCTGAGTCAGCTGATAACTGATCACCTGCAAATACAGCGTTGGCTCTTTAAAATGGACTCTGAGTTCCGAGGA AATGGGACTGCATTTTGTGATATTCCTTCCTACCTAAAGTGCTACAAATGGGTGCTAAAGGAGAGTAGCAGATATGGCCTTGAAGACTGG AGAAAGAAATGGGCACAAGAGCCAGCTTTGGTGAAGATCTCTGAGGAGCTGGCGGGCATTTTAGCACAGCACGCACAGCCAGTCAATGAG AAACGGTTCCCGACGTGGAGGAAATTCCTCCAAACATTTCTCAGTCAAGGGGGTGTGATCGAAGCATTCCCACCTGCAGACAATGTCACC AACCTCACAGTGGACATGCTGATAGAGCCCAACGGGAAAATCAGCGTGCTGTCGACAGGGGACCAGCTTCATGCTGAAAGCCCCTTCATC TCCTCTGGTACCACCGTGCCTCAGACCTCAGTGGATCCCCAAGTTCTCACTTATTTGTGCCTCCAAATTGGAAAAGCCTGCAGAATGAGA GATGTGGTTGGTTACTTTTCGATAGATCTGGTGACTTTTATAGATCCAAGCACCTTGGAACAACAGGTGTGGGCAACCGGCCTTAACCTC GCATATAGTGACCAGCTGGCCCTGACTCAACTCACCTTATACCTGACAAACGGCCATCTGGATTGCAGTTTGAGCACCCTGGAAGTGCCC CGCTTTGTTCCAAAGGAAAGGAAGAAAACCAAATGCATGAGTGCGCTGTCAATGCCGATGCTGGCAACCAGTCGCTATGCAGTGATGACC ACCCAGCTAAGACACAGCAATCTCTCACTGGTTTTCCACTATGTTTTTCTCCAGATCTGTAGGGCCCATGGCATTGGCTATGATGTTGAG AACAATCGGCGAGGATCTCCAGGGGGTCCTCATGACCTTTGCTCGCCATCTCTTCATCATCCATCAAGAAATATCAGCACCTAATATGCA AGGCGAGACCAATTTTAAGACCACCATTGCTGATATTGAAACTATTCTAAGAGTAACAAAGGAAAACAAAATGAGATTTGAAGAGGAGCA ACAGTCCAAAGATGATAAAAACCTCTCTAAACCCAAGAAATGATCCTGGAATACAGTACATAACAATTTGGATCCCAGTCTGGAATAAAA AGGGCAATTTTTTTTTCTGTTAGAAATAAAAGCCAGGGGAAATTGGTTTGCTTTGTGTGCTAGGAGGTGAATCAGAACAGATTATAATGA AATGCTCTTTTTAAAACATTGTTTATTAAGTGATCTTATTTTATTTATTAAACCAAAACTTATTTGTGTTTTCATTTGAGAGTGTTGAAC AATCCCTTCTTCTTCTCAAACTCAGAAAAAAGTAATCTGATAAAAGAAGAAAGTTAAAAGTCTTACTGATATCACCTCCGCATTTACTTC CTCATAGGCCTCAGGATTATGTAGCTTTTATTTTTTATGTTTTATAAAGTTTTCTCCTATTTCTAATAGTCCATCGATCTTCTGCATTTA TAGGTTTGAATAAAGGCTTAAGAATTGCTATTTGTCAAAACACAAATGCCTTTTTTTCAAAGCTTCAAATTATATTGAAATTATATTTAT ATGAATTTTTAAGTGACTATCATTCATCGTGTTTTTGTTATACCAAGCCACTATTGTCTGCTATTGTTGCCATTTTTCAAAAAGTAGGAT AAGCCAGCATCTAGTTAACCAAAGCATCAGAATGTATCTAGGTGGCAGTATCTGCTCTTAGTTTAAGATGGAGTGTAGCTCGAGCTCTGT AGCTTGAATAAACATTTGTATTCTCTAGTTCTCCATTTAATACAGCAGTAGGATTTTTTCAAGTTTAGTTCAACATTATACTACCTTTAA AAAGGCAGACTGTTAAGATTGTAGTAGTGCTGACTCTAACTTTCAGCATTCACTTCATAATTGTCCAAAAATATTTCCACATCAGTGAGC TTTACTGAAGGGTGAAATAAAAGCAAATTACAACACCAAAACCATTATTGTTTTAAAGAATGACCTTTATGTCTATAAAAATATTTTAGT GTATTGGGGAAAGGAATAGGATTTTTACCTGTGGAATTTCAATAATTTGTTAACTGTGATTCAGCCACAGGCTAAGAGTTTCTGTAATAA >83797_83797_4_SMAD3-IQCH_SMAD3_chr15_67462942_ENST00000540846_IQCH_chr15_67649683_ENST00000358767_length(amino acids)=806AA_BP=1 MDGRLQVSHRKGLPHVIYCRLWRWPDLHSHHELRAMELCEFAFNMKKDEVCVNPYHYQRVETPVLPPVLVPRHTEIPAEFPPLDDYSHSI PENTNFPAGIEPQSNIPETPPPGYLSEDGETSDHQMNHSMDAGSPNLSPNPMSPAHNNLATFTIPREPPPSPAEVKFFPKKQRSKGKSRR SRGHHDRKAMKVKTPLRALKSLWDYDFLIYDGVIDNTAPDFLAFKEHFSLAWGGIFSLLEHVEKFLRNYAIPEVKIKGNNLVALLPEFEL TNKLTRYDLLSVLEDPAHVQMLINLPGQRYKGQDGNSEAAMKIQATWKCYKARKFFLFYRQQKWASGVIAIAWLLYCHKTRLKKILKESR QRHLENFRIRAKHLAANWNRIRTSRRTIIHIPSLGYSQPVREHIADFNTQQNMQLGRLCDILDANVNVIYICSHHMNDELVLYYKKILSL HAAVKSGNLEDRSDLQDRFKIITPEAVNIFPMIEQLSQLITDHLQIQRWLFKMDSEFRGNGTAFCDIPSYLKCYKWVLKESSRYGLEDWR KKWAQEPALVKISEELAGILAQHAQPVNEKRFPTWRKFLQTFLSQGGVIEAFPPADNVTNLTVDMLIEPNGKISVLSTGDQLHAESPFIS SGTTVPQTSVDPQVLTYLCLQIGKACRMRDVVGYFSIDLVTFIDPSTLEQQVWATGLNLAYSDQLALTQLTLYLTNGHLDCSLSTLEVPR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SMAD3-IQCH |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SMAD3-IQCH |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SMAD3-IQCH |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
