
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SMARCC1-DOCK3 (FusionGDB2 ID:83941) |
Fusion Gene Summary for SMARCC1-DOCK3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SMARCC1-DOCK3 | Fusion gene ID: 83941 | Hgene | Tgene | Gene symbol | SMARCC1 | DOCK3 | Gene ID | 6599 | 1795 |
| Gene name | SWI/SNF related, matrix associated, actin dependent regulator of chromatin subfamily c member 1 | dedicator of cytokinesis 3 | |
| Synonyms | BAF155|CRACC1|Rsc8|SRG3|SWI3 | MOCA|NEDIDHA|PBP | |
| Cytomap | 3p21.31 | 3p21.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | SWI/SNF complex subunit SMARCC1BRG1-associated factor 155SWI/SNF complex 155 kDa subunitSWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1chromatin remodeling complex BAF155 subunitmammalian chromatin remode | dedicator of cytokinesis protein 3modifier of cell adhesionpresenilin-binding protein | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q8IZD9 | |
| Ensembl transtripts involved in fusion gene | ENST00000425518, ENST00000254480, | ENST00000266037, | |
| Fusion gene scores | * DoF score | 19 X 26 X 11=5434 | 12 X 10 X 7=840 |
| # samples | 37 | 12 | |
| ** MAII score | log2(37/5434*10)=-3.87641738970566 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(12/840*10)=-2.8073549220576 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SMARCC1 [Title/Abstract] AND DOCK3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | DOCK3(50712731)-SMARCC1(47663831), # samples:4 SMARCC1(47814307)-DOCK3(50927457), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | DOCK3-SMARCC1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. DOCK3-SMARCC1 seems lost the major protein functional domain in Tgene partner, which is a epigenetic factor due to the frame-shifted ORF. DOCK3-SMARCC1 seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SMARCC1 | GO:0006337 | nucleosome disassembly | 8895581 |
| Hgene | SMARCC1 | GO:0006338 | chromatin remodeling | 10078207|11018012|11726552 |
| Hgene | SMARCC1 | GO:0045893 | positive regulation of transcription, DNA-templated | 11018012 |
 Fusion gene breakpoints across SMARCC1 (5'-gene) Fusion gene breakpoints across SMARCC1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
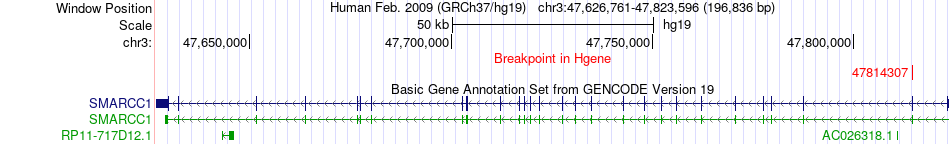 |
 Fusion gene breakpoints across DOCK3 (3'-gene) Fusion gene breakpoints across DOCK3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
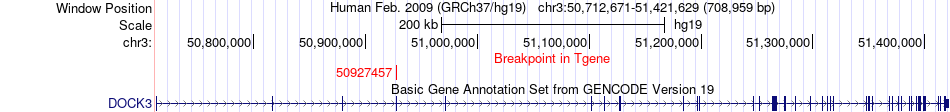 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BLCA | TCGA-BL-A3JM-01A | SMARCC1 | chr3 | 47814307 | - | DOCK3 | chr3 | 50927457 | + |
Top |
Fusion Gene ORF analysis for SMARCC1-DOCK3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5UTR-3CDS | ENST00000425518 | ENST00000266037 | SMARCC1 | chr3 | 47814307 | - | DOCK3 | chr3 | 50927457 | + |
| In-frame | ENST00000254480 | ENST00000266037 | SMARCC1 | chr3 | 47814307 | - | DOCK3 | chr3 | 50927457 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000254480 | SMARCC1 | chr3 | 47814307 | - | ENST00000266037 | DOCK3 | chr3 | 50927457 | + | 9005 | 435 | 96 | 6365 | 2089 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000254480 | ENST00000266037 | SMARCC1 | chr3 | 47814307 | - | DOCK3 | chr3 | 50927457 | + | 0.000654121 | 0.99934584 |
Top |
Fusion Genomic Features for SMARCC1-DOCK3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| SMARCC1 | chr3 | 47814306 | - | DOCK3 | chr3 | 50927456 | + | 1.93E-14 | 1 |
| SMARCC1 | chr3 | 47814306 | - | DOCK3 | chr3 | 50927456 | + | 1.93E-14 | 1 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for SMARCC1-DOCK3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr3:50712731/chr3:47663831) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | DOCK3 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Potential guanine nucleotide exchange factor (GEF). GEF proteins activate some small GTPases by exchanging bound GDP for free GTP. Its interaction with presenilin proteins as well as its ability to stimulate Tau/MAPT phosphorylation suggest that it may be involved in Alzheimer disease. Ectopic expression in nerve cells decreases the secretion of amyloid-beta APBA1 protein and lowers the rate of cell-substratum adhesion, suggesting that it may affect the function of some small GTPase involved in the regulation of actin cytoskeleton or cell adhesion receptors (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | DOCK3 | chr3:47814307 | chr3:50927457 | ENST00000266037 | 2 | 53 | 1731_1768 | 54 | 2031.0 | Compositional bias | Note=Ser-rich | |
| Tgene | DOCK3 | chr3:47814307 | chr3:50927457 | ENST00000266037 | 2 | 53 | 1228_1635 | 54 | 2031.0 | Domain | DOCKER | |
| Tgene | DOCK3 | chr3:47814307 | chr3:50927457 | ENST00000266037 | 2 | 53 | 421_599 | 54 | 2031.0 | Domain | C2 DOCK-type | |
| Tgene | DOCK3 | chr3:47814307 | chr3:50927457 | ENST00000266037 | 2 | 53 | 1970_1976 | 54 | 2031.0 | Motif | SH3-binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 914_946 | 105 | 1106.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 329_336 | 105 | 1106.0 | Compositional bias | Note=Poly-Pro |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 769_863 | 105 | 1106.0 | Compositional bias | Note=Glu-rich |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 867_878 | 105 | 1106.0 | Compositional bias | Note=Poly-Ala |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 977_1105 | 105 | 1106.0 | Compositional bias | Note=Pro-rich |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 449_546 | 105 | 1106.0 | Domain | SWIRM |
| Hgene | SMARCC1 | chr3:47814307 | chr3:50927457 | ENST00000254480 | - | 2 | 28 | 618_669 | 105 | 1106.0 | Domain | SANT |
| Tgene | DOCK3 | chr3:47814307 | chr3:50927457 | ENST00000266037 | 2 | 53 | 6_67 | 54 | 2031.0 | Domain | SH3 |
Top |
Fusion Gene Sequence for SMARCC1-DOCK3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >83941_83941_1_SMARCC1-DOCK3_SMARCC1_chr3_47814307_ENST00000254480_DOCK3_chr3_50927457_ENST00000266037_length(transcript)=9005nt_BP=435nt GGCTGGGCGGGGCCGGGAAGCGGCAGTGGCGGCTACGCGCGCGGGGGTGCGCGCGGGAACGACCGGGAAACACCGCGAGGGCCGGGGTGG GCCAGGCTGTGGGGACGACGGGCTGCGACGATGGCCGCAGCGGCGGGCGGCGGCGGGCCGGGGACAGCGGTAGGCGCCACGGGCTCGGGG ATTGCGGCGGCAGCCGCAGGCCTAGCTGTTTATCGACGGAAGGATGGGGGCCCGGCCACCAAGTTTTGGGAGAGCCCGGAGACGGTGTCC CAGCTGGATTCGGTGCGGGTCTGGCTGGGCAAGCACTACAAGAAGTATGTTCATGCGGATGCTCCTACCAATAAAACACTGGCTGGGCTG GTGGTGCAGCTTCTTCAGTTCCAGGAAGATGCCTTTGGGAAGCATGTCACCAACCCGGCCTTCACCAAACTCCCTGGGATCTTTCCTGCA AATTACATTCACTTGAAAAAGGCAATTGTCAGTAATAGGGGGCAGTATGAAACTGTGGTTCCACTTGAAGATTCTATTGTGACTGAGGTT ACAGCAACTCTACAAGAATGGGCAAGTTTGTGGAAACAGTTGTATGTGAAACACAAAGTAGATCTTTTCTACAAACTACGCCATGTGATG AATGAACTTATTGACCTGCGAAGGCAGCTACTGTCTGGTCACCTGACTCAGGATCAGGTGCGGGAGGTTAAGCGGCACATCACCGTGCGC CTGGACTGGGGTAATGAACATTTGGGCCTGGACCTGGTGCCTCGGAAGGACTTTGAAGTAGTGGACTCGGACCAGATTAGTGTCTCAGAT CTCTATAAGATGCATTTATCTAGCCGGCAGAGTGTACAGCAAAGCACATCCCAGGTAGATACAATGCGCCCACGTCATGGGGAAACATGT CGGATGCCAGTGCCACATCACTTCTTCCTCAGCCTGAAGAGTTTCACTTACAATACTATTGGGGAAGATACCGATGTCTTCTTTTCCTTA TATGACATGAGGGAAGGCAAGCAGATCAGTGAGCGGTTTCTGGTAAGACTGAACAAGAATGGTGGGCCGAGGAACCCAGAGAAGATAGAA CGAATGTGTGCCCTTTTTACAGATCTGAGCAGCAAAGATATGAAGAGAGATTTGTATATCGTTGCCCATGTGATCCGAATAGGCCGGATG CTCCTGAACGACTCAAAGAAAGGTCCTCCTCACCTGCACTACAGGCGACCATATGGCTGTGCGGTCCTAAGCATCTTGGATGTCCTACAG TCACTCACAGAAGTAAAGGAAGAAAAGGATTTTGTTCTTAAGGTTTACACGTGCAACAACGAGAGTGAGTGGTCCCAGATCCACGAGAAC ATCATCCGAAAGTCCAGTGCCAAGTACTCTGCCCCCAGCGCCAGCCATGGTCTTATCATTTCTCTGCAGCTTCTTCGTGGAGACATGGAA CAGATTCGGAGAGAAAATCCCATGATATTTAATAGGGGATTGGCAATTACAAGAAAATTGGGATTTCCTGATGTCATTATGCCAGGTGAT ATCCGCAATGACCTGTACCTAACCCTGGAGAAGGGGGATTTCGAGAGAGGAGGAAAGAGTGTACAAAAGAATATTGAAGTGACCATGTAT GTGCTTTATGCAGATGGAGAAATCTTGAAGGATTGCATCAGCTTGGGTTCAGGAGAGCCAAATAGGAGTTCCTACCACTCCTTTGTCCTC TACCACAGTAATAGTCCTCGCTGGGGAGAAATTATCAAATTGCCTATCCCCATTGACCGGTTCCGGGGCTCCCACCTGCGCTTTGAGTTC AGACATTGTTCCACAAAGGACAAAGGGGAAAAGAAACTCTTTGGCTTTGCATTCTCAACCCTGATGCGTGATGATGGCACCACCCTCTCA GATGATATTCACGAGCTTTATGTGTACAAGTGTGATGAGAATAGCACGTTTAATAACCATGCTCTGTACCTGGGCCTGCCCTGCTGCAAA GAGGACTACAATGGCTGCCCTAATATTCCTTCTAGCCTCATCTTCCAGCGCAGCACCAAAGAGTCTTTCTTCATCTCCACTCAGCTCTCC TCTACCAAACTCACCCAGAATGTGGACCTCCTAGCTCTGCTGAAGTGGAAAGCCTTCCCCGACCGGATCATGGATGTACTAGGGCGGCTG CGGCATGTCAGTGGGGAGGAAATTGTTAAGTTTCTGCAGGACATCTTAGATACACTCTTTGTGATTTTGGATGATAATACAGAGAAGTAC GGCCTGTTGGTTTTTCAGTCTCTGGTGTTCATCATCAACCTGCTCCGAGACATCAAGTATTTTCACTTTCGACCTGTGATGGACACGTAT ATCCAGAAGCACTTTGCTGGAGCTCTGGCATACAAGGAGCTCATCCGCTGTTTGAAGTGGTATATGGACTGCTCAGCAGAACTGATTCGA CAGGACCACATTCAAGAAGCTATGCGGGCCTTGGAGTACCTTTTCAAGTTCATTGTACAGTCACGGATCCTGTACTCACGAGCCACTTGT GGAATGGAAGAGGAACAATTCAGATCCAGTATCCAAGAACTTTTCCAGTCCATCCGGTTTGTGCTCAGTCTGGACAGCCGAAACTCAGAA ACACTCCTTTTTACTCAGGCTGCACTCCTCAATTCTTTCCCAACCATCTTTGATGAGCTTCTGCAAATGTTCACCGTGCAAGAGGTGGCA GAGTTTGTGAGAGGGACACTGGGGAGCATGCCCAGCACTGTGCACATTGGGCAGTCAATGGACGTGGTCAAGCTGCAGTCCATTGCCAGG ACAGTGGATAGCCGCCTGTTTTCTTTCTCAGAATCCCGCCGCATCCTGCTTCCTGTGGTTCTCCATCACATTCACCTTCACCTGAGGCAG CAGAAAGAGCTGCTAATTTGCTCAGGGATTCTTGGCAGCATCTTCTCCATCGTCAAGACCAGCTCTCTGGAGGCAGATGTCATGGAGGAA GTAGAGATGATGGTGGAGAGCCTCCTGGACGTGCTCTTGCAGACTCTGCTCACCATCATGAGCAAATCGCACGCTCAGGAGGCGGTAAGA GGGCAGCGGTGCCCGCAGTGCACAGCCGAGATCACTGGCGAGTATGTGTCCTGCCTTCTCTCACTGCTCCGCCAGATGTGTGACACCCAT TTCCAGCACCTCCTGGACAACTTCCAGAGCAAAGATGAACTCAAGGAATTTCTGCTGAAGATTTTTTGCGTGTTCCGGAACCTGATGAAG ATGAGTGTCTTCCCTCGGGACTGGATGGTAATGAGACTGCTCACAAGCAATATTATAGTCACTACTGTCCAGTACCTGTCCTCTGCACTG CACAAGAATTTCACAGAGACTGACTTTGACTTTAAGGTGTGGAATTCTTACTTTAGCCTGGCAGTTCTATTCATAAATCAGCCAAGCCTT CAGCTAGAAATTATCACCTCAGCCAAAAGGAAGAAGATTCTAGATAAGTATGGGGACATGCGTGTAATGATGGCCTATGAACTGTTCAGC ATGTGGCAGAATTTGGGTGAACATAAGATCCACTTTATTCCGGGAATGATTGGTCCTTTTCTGGGTGTGACACTGGTCCCACAGCCAGAA GTACGGAATATCATGATTCCCATCTTTCATGACATGATGGACTGGGAGCAGAGAAAAAATGGCAACTTCAAACAGGTGGAGGCCGAGTTG ATTGACAAGCTGGACAGCATGGTGTCAGAAGGGAAAGGTGACGAGAGCTACAGGGAGCTCTTCAGCCTACTAACCCAGCTGTTTGGGCCC TACCCCAGCCTGCTGGAGAAGGTTGAACAAGAAACATGGCGCGAGACCGGCATTTCCTTTGTGACCTCAGTCACCCGCCTCATGGAACGT CTTCTTGACTACAGGGACTGCATGAAAGGAGAGGAAACAGAGAATAAGAAGATAGGCTGCACTGTTAACCTGATGAATTTTTACAAATCT GAGATTAACAAGGAAGAAATGTATATCCGCTACATCCATAAGCTTTGTGACATGCACTTGCAGGCCGAAAACTACACAGAGGCCGCATTT ACCCTGCTCCTTTACTGTGAGCTGCTGCAGTGGGAGGACCGGCCACTACGGGAATTCCTCCACTACCCATCGCAGACAGAGTGGCAGCGG AAGGAGGGACTGTGCCGGAAGATCATTCACTACTTCAACAAAGGCAAGAGCTGGGAGTTTGGGATCCCACTGTGCAGGGAGCTGGCGTGT CAGTACGAGAGCCTCTATGATTACCAGAGCCTCAGCTGGATTCGGAAAATGGAGGCCAGCTACTATGACAACATTATGGAGCAGCAACGC CTGGAGCCTGAGTTCTTTCGGGTCGGCTTCTATGGCAGGAAGTTTCCTTTCTTTCTTCGGAACAAAGAATACGTGTGCCGTGGCCATGAC TACGAGAGGCTGGAGGCCTTCCAGCAGAGGATGCTCAGTGAGTTTCCGCAGGCTGTCGCCATGCAGCACCCCAACCATCCTGATGACGCC ATCCTACAGTGCGATGCCCAGTACTTGCAGATCTATGCAGTGACGCCCATTCCAGATTATGTGGATGTTCTGCAGATGGATAGGGTACCA GATCGAGTCAAGAGCTTCTATCGCGTCAACAATGTGAGGAAGTTCCGGTATGACAGGCCTTTTCACAAAGGCCCCAAGGACAAGGAGAAT GAATTCAAGAGCCTGTGGATTGAACGTACCACACTGACCCTGACCCACAGCTTGCCTGGCATCTCTCGGTGGTTTGAAGTGGAGAGGAGG GAACTGGTGGAGGTGAGCCCTCTGGAGAATGCCATCCAAGTGGTTGAGAATAAGAACCAGGAGCTACGCTCCCTGATCAGCCAGTATCAA CACAAGCAGGTGCATGGCAACATTAACCTGCTAAGCATGTGCCTGAATGGTGTCATTGATGCAGCTGTCAATGGAGGCATTGCACGCTAT CAGGAGGCCTTCTTTGATAAAGATTACATCAACAAGCACCCAGGAGATGCTGAGAAGATCACCCAGCTCAAGGAGCTTATGCAGGAGCAG GTTCATGTCCTTGGAGTTGGGCTAGCAGTTCATGAGAAGTTTGTGCACCCAGAAATGCGGCCTCTGCATAAGAAGCTAATTGATCAGTTC CAGATGATGCGGGCCAGTCTCTACCATGAGTTTCCAGGTTTGGATAAGCTAAGTCCTGCATGTTCAGGCACCAGCACCCCACGGGGAAAT GTTCTGGCATCCCATAGCCCCATGAGTCCGGAGAGCATCAAGATGACCCACCGGCACAGCCCCATGAACTTGATGGGCACAGGCCGCCAT TCATCATCCTCTCTCTCCTCACATGCGTCTAGTGAAGCAGGAAACATGGTGATGCTGGGTGACGGCTCCATGGGTGATGCTCCTGAGGAC CTGTACCACCACATGCAGCTCGCGTATCCCAACCCCAGGTACCAAGGCTCAGTCACCAACGTCTCTGTTCTGTCCTCGTCCCAGGCAAGC CCTTCTTCCTCCAGCCTGAGTTCCACTCACTCAGCACCATCCCAGATGATTACCTCTGCCCCTTCCAGTGCCCGAGGCTCTCCCTCTCTG CCAGATAAGTACCGCCATGCCCGTGAAATGATGTTGTTGCTGCCCACATACCGGGACCGCCCAAGCAGTGCCATGTATCCAGCAGCCATC CTGGAGAACGGACAGCCGCCGAATTTCCAGCGAGCCCTGTTCCAGCAAGTGGTCGGAGCCTGCAAACCCTGCAGTGATCCCAATCTGTCT GTGGCTGAAAAAGGTCATTACTCCCTACACTTTGACGCCTTCCACCACCCTCTGGGTGATACCCCCCCAGCCCTCCCTGCCCGGACCCTG CGCAAGTCTCCTCTCCACCCTATCCCAGCCTCCCCCACAAGCCCCCAGTCAGGTCTGGACGGCAGCAACTCTACGCTGTCCGGCAGTGCC AGCAGCGGCGTGTCCTCCTTGAGTGAGAGTAACTTTGGGCACTCCTCGGAGGCCCCACCTCGCACTGACACCATGGACTCCATGCCAAGT CAGGCCTGGAATGCTGACGAAGATCTTGAGCCACCCTACCTCCCTGTCCACTACAGCCTCTCTGAGTCTGCCGTCCTGGACTCCATCAAG GCCCAGCCATGCCGAAGCCACTCAGCCCCAGGGTGCGTCATCCCTCAGGACCCCATGGACCCGCCTGCGCTGCCGCCCAAGCCCTACCAC CCCCGCCTGCCGGCCCTGGAGCACGATGAGGGGGTGCTGCTGCGTGAAGAGACTGAGAGGCCTCGAGGCCTGCACCGCAAGGCTCCATTG CCTCCTGGGAGCGCTAAGGAGGAGCAGGCCCGCATGGCCTGGGAGCACGGCCGAGGGGAGCAGTGAGGGGCAACGAGGCGGCTGGGATGC CGCCCTCAGTAAGCAGCTTGCCAATCACTCCAGGTCTGAAAAGCAAGTCCCCCAGCCCCACCCCAGGGAGCCAGAGAGGCTTGCACTCAG GAGAGAACCACCCCCAAGTCTCCGTTCTACTGCCGTGAACTCATGTGTTGCCATGTACAGAGGCCACAGCAGCATGAAGGGTTGTGGCTT CCCTTTTTATTTTTTTACATTTCCATTTCTATGGGTTTTCCTTTCTTTCCTTTATGCAGTAGATGCTTTCTTCCTCCTGCAGTTCTGGAC CATGTGGAGCTACATGGAGAAATTGCACAGACAGAATCACTTTGCCACACTGCAGGGCCCAGGAGTGGGAGCCCAGGCCCTCCCTCCAGG GTAGAGATGCACCGAATGGAAATTGCACTAAGGACCTCTTCCTTTGCTGTATGGACAGAAGAGTTCATGGTTGCATTCAGGGATTGCAAG GACCATATAGACATGTCACAGGTGGAAAAAGGGCCACAAGCCGTTTTGCACAAATGCCTGCCCACCTGCCCCTCTTCCATCCAGGAGTGT GCTACTGAGGGGCTGGCTGGACCAACCTGCTGGCTCAGGCATGTGACTGGCCTAACTGCATGCCCAGGGCACTGCCAGGGTTCAATGGGC CAGCCGTCTCCTACTCCACCTTGGATTTTTCTCTCCATCACATCATTTCTGACCATTTGCTCCCTTCCATTCTTGAAACACCTCATGCCC CACAGGAGCCCCAGTGGTGACTGGATCTGCAAGGGTTATCTCCCGCCCTCTGGGGTTTGAGTGGAGGTATGGAGCCAGAGAGGCTATCCT AGCTACCTTCCTCCTAGGGGGAGCTGGTCCCCTTCCTATGTGGTAGAGAGACTGGTGTAAGAGTGTGGGGTGTGGCAGGCGCCGGAAATC CTCAAAGCTTTCAGAAGCATCCAGAATGCCTACCATCAGCTCCTGAATCAGGGATTTTCAGCACTACCTCTGAGCCATGGGGTTGGCTGC CCAGCCAGGCTTCCAGGCCCTGAGGACATGTGGTCAGGTTCAGCAGGCTCCTGGGTAAAGAGGGTGGGAGGGGGCTTCTCCTTTTGGAAG CAGATAGCCATGCAGGTAGAATTGTCATCTCCCAAGTGGCCACATACAGCAACCCACTCCAGGCCATGGGGACCACCACTCAGGGTTCGG GGCTGGCAGGAGGGTAGTTTCTCCAGTCTCCCCAGTCTGTTGGTGTCTTTATAGGAAACTATAAAGCAGAGCAGTGGTGTCTTTAGAGGA AACTCATTCAAGTCAGGGCACTGATTTTCCTCTCAGTTTATTATTTGGGGGGATAGGGTCAGGTGGGTATAGTAGTACTTTACCAGGGTG CTTTTAAGTTACTTTAAAAAAAGCCTACAAAATATTTTTTTTCTTTTATTTGCTCGAGTTCACATTAATGATGGTCACAAGGCTGCCTTG TTGGGCAGGCGTATTGCCCCCAGTTCCTCTTCTGCTGGCCTGTATGACCTCCACAGCCAGGCCCTGGGCCCAAGCCCCTTGTCCCTTCTC CACTGCCCCTCTTTCCAGACAGTAAAGGCCATGGTCAGTGTGTTTTTCTCTTGTAAACAAACCCCAGCTTGTTTAACAGAAATGCTAATA ACCTACTGGGAAAGATGGAGGTCTAAATTACCTCCAGGGTTTTTCTGGGGGTTTATCACCAGTGTGGGTCCCTTCTGATACCACCAGGTT CACTCCAGGCAGAGTGGGGCGGAAGGCTGCTGAGGATATGGGTCAGTTACAGCAGCCCTCACCTCAAAGGGCTGGCCTGCTTCTCAGCCT ACATTCATTTGCAAGCTTCAATCTCTGGACCATCTGGTGTTCACAGGTGTTAGAGGGTTAGGGGTTAGGGGCTAGTTTTGGATTTGATTC ATAGGTAGGAGGGCTTAGATTTTAAGGCACTTCTGAAAGTCAATCCCTGGACAAGGCAGTCATCACATAAGAACAGCTACCTTCTCCACT TGGTGGCACAAGAGGTAGGGAGGGGAGTATGGGTTCATTTGGCTTCGCATTATGCAAGGTGAAACCGTTTGTTTTCCCTCTCCATTTTCC CTAACTAAATGAAAAGGACACATTCTGAAATCCCTTTTGTTGGAGAATAAGTCAGTCTGAGGGGAAATGGGAGGCCAGAGATGAGAACCC TTTGAAAAGATTGTAAAATACTGATTTTCATTCTTTCAAGCTTATTTGTAAATACCTATTTGAATGCTGTGTATTTGTACAGGAATTTGA GCAAAAAATGTATAGAGTGTGATGTCCAATTGGTATTCAGCACTATAAATGTGTTTTTAACCTCCCGCATTCTGTGCTTATTTAAAACAA GGAAACTTCTAACCATTTCTTTTGTGTATTCATGTTTAAAGAAAAAAAGTGATTTAAAAATGATCTTACCTGTACCAGAAAAGCAAAGTT AAAGGAAACAAAATTTGTACCATTGTCCCAAGAGGTATTTTACTGTATATATTGTGGTAGCATGTTCAAAATCCAACAAGTAATGTGAAT TTTAGATGTAAATATCTGCCACTTGATTTTTTTTCCCCCTTTCCCCACTTCCTTGACTGCTGTGATGTGAATTAAAGATAAATACGTGAT >83941_83941_1_SMARCC1-DOCK3_SMARCC1_chr3_47814307_ENST00000254480_DOCK3_chr3_50927457_ENST00000266037_length(amino acids)=2089AA_BP=9 MWGRRAATMAAAAGGGGPGTAVGATGSGIAAAAAGLAVYRRKDGGPATKFWESPETVSQLDSVRVWLGKHYKKYVHADAPTNKTLAGLVV QLLQFQEDAFGKHVTNPAFTKLPGIFPANYIHLKKAIVSNRGQYETVVPLEDSIVTEVTATLQEWASLWKQLYVKHKVDLFYKLRHVMNE LIDLRRQLLSGHLTQDQVREVKRHITVRLDWGNEHLGLDLVPRKDFEVVDSDQISVSDLYKMHLSSRQSVQQSTSQVDTMRPRHGETCRM PVPHHFFLSLKSFTYNTIGEDTDVFFSLYDMREGKQISERFLVRLNKNGGPRNPEKIERMCALFTDLSSKDMKRDLYIVAHVIRIGRMLL NDSKKGPPHLHYRRPYGCAVLSILDVLQSLTEVKEEKDFVLKVYTCNNESEWSQIHENIIRKSSAKYSAPSASHGLIISLQLLRGDMEQI RRENPMIFNRGLAITRKLGFPDVIMPGDIRNDLYLTLEKGDFERGGKSVQKNIEVTMYVLYADGEILKDCISLGSGEPNRSSYHSFVLYH SNSPRWGEIIKLPIPIDRFRGSHLRFEFRHCSTKDKGEKKLFGFAFSTLMRDDGTTLSDDIHELYVYKCDENSTFNNHALYLGLPCCKED YNGCPNIPSSLIFQRSTKESFFISTQLSSTKLTQNVDLLALLKWKAFPDRIMDVLGRLRHVSGEEIVKFLQDILDTLFVILDDNTEKYGL LVFQSLVFIINLLRDIKYFHFRPVMDTYIQKHFAGALAYKELIRCLKWYMDCSAELIRQDHIQEAMRALEYLFKFIVQSRILYSRATCGM EEEQFRSSIQELFQSIRFVLSLDSRNSETLLFTQAALLNSFPTIFDELLQMFTVQEVAEFVRGTLGSMPSTVHIGQSMDVVKLQSIARTV DSRLFSFSESRRILLPVVLHHIHLHLRQQKELLICSGILGSIFSIVKTSSLEADVMEEVEMMVESLLDVLLQTLLTIMSKSHAQEAVRGQ RCPQCTAEITGEYVSCLLSLLRQMCDTHFQHLLDNFQSKDELKEFLLKIFCVFRNLMKMSVFPRDWMVMRLLTSNIIVTTVQYLSSALHK NFTETDFDFKVWNSYFSLAVLFINQPSLQLEIITSAKRKKILDKYGDMRVMMAYELFSMWQNLGEHKIHFIPGMIGPFLGVTLVPQPEVR NIMIPIFHDMMDWEQRKNGNFKQVEAELIDKLDSMVSEGKGDESYRELFSLLTQLFGPYPSLLEKVEQETWRETGISFVTSVTRLMERLL DYRDCMKGEETENKKIGCTVNLMNFYKSEINKEEMYIRYIHKLCDMHLQAENYTEAAFTLLLYCELLQWEDRPLREFLHYPSQTEWQRKE GLCRKIIHYFNKGKSWEFGIPLCRELACQYESLYDYQSLSWIRKMEASYYDNIMEQQRLEPEFFRVGFYGRKFPFFLRNKEYVCRGHDYE RLEAFQQRMLSEFPQAVAMQHPNHPDDAILQCDAQYLQIYAVTPIPDYVDVLQMDRVPDRVKSFYRVNNVRKFRYDRPFHKGPKDKENEF KSLWIERTTLTLTHSLPGISRWFEVERRELVEVSPLENAIQVVENKNQELRSLISQYQHKQVHGNINLLSMCLNGVIDAAVNGGIARYQE AFFDKDYINKHPGDAEKITQLKELMQEQVHVLGVGLAVHEKFVHPEMRPLHKKLIDQFQMMRASLYHEFPGLDKLSPACSGTSTPRGNVL ASHSPMSPESIKMTHRHSPMNLMGTGRHSSSSLSSHASSEAGNMVMLGDGSMGDAPEDLYHHMQLAYPNPRYQGSVTNVSVLSSSQASPS SSSLSSTHSAPSQMITSAPSSARGSPSLPDKYRHAREMMLLLPTYRDRPSSAMYPAAILENGQPPNFQRALFQQVVGACKPCSDPNLSVA EKGHYSLHFDAFHHPLGDTPPALPARTLRKSPLHPIPASPTSPQSGLDGSNSTLSGSASSGVSSLSESNFGHSSEAPPRTDTMDSMPSQA WNADEDLEPPYLPVHYSLSESAVLDSIKAQPCRSHSAPGCVIPQDPMDPPALPPKPYHPRLPALEHDEGVLLREETERPRGLHRKAPLPP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SMARCC1-DOCK3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SMARCC1-DOCK3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SMARCC1-DOCK3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
