
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SMCHD1-NDC80 (FusionGDB2 ID:84030) |
Fusion Gene Summary for SMCHD1-NDC80 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SMCHD1-NDC80 | Fusion gene ID: 84030 | Hgene | Tgene | Gene symbol | SMCHD1 | NDC80 | Gene ID | 23347 | 10403 |
| Gene name | structural maintenance of chromosomes flexible hinge domain containing 1 | NDC80 kinetochore complex component | |
| Synonyms | BAMS|FSHD2 | HEC|HEC1|HsHec1|KNTC2|TID3|hsNDC80 | |
| Cytomap | 18p11.32 | 18p11.32 | |
| Type of gene | protein-coding | protein-coding | |
| Description | structural maintenance of chromosomes flexible hinge domain-containing protein 1SMC hinge domain-containing protein 1 | kinetochore protein NDC80 homologNDC80 homolog, kinetochore complex componentNDC80 kinetochore complex component homologhighly expressed in cancer proteinhighly expressed in cancer, rich in leucine heptad repeatskinetochore associated 2kinetochore p | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | . | O14777 | |
| Ensembl transtripts involved in fusion gene | ENST00000261598, ENST00000320876, ENST00000609587, | ENST00000261597, | |
| Fusion gene scores | * DoF score | 6 X 9 X 6=324 | 3 X 8 X 5=120 |
| # samples | 11 | 8 | |
| ** MAII score | log2(11/324*10)=-1.55849028935997 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(8/120*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SMCHD1 [Title/Abstract] AND NDC80 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SMCHD1(2656260)-NDC80(2601395), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SMCHD1 | GO:0009048 | dosage compensation by inactivation of X chromosome | 23542155 |
 Fusion gene breakpoints across SMCHD1 (5'-gene) Fusion gene breakpoints across SMCHD1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
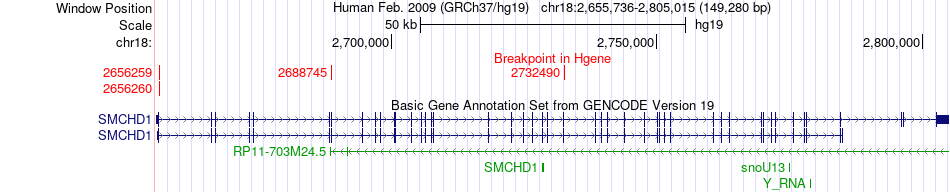 |
 Fusion gene breakpoints across NDC80 (3'-gene) Fusion gene breakpoints across NDC80 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
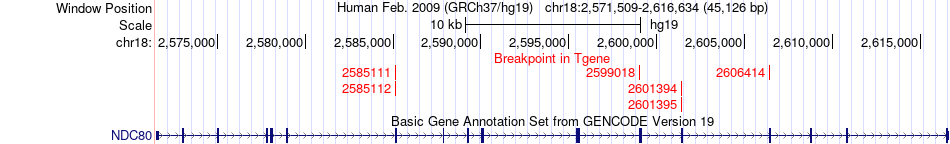 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BLCA | TCGA-ZF-A9RD-01A | SMCHD1 | chr18 | 2656260 | - | NDC80 | chr18 | 2599018 | + |
| ChimerDB4 | BLCA | TCGA-ZF-A9RD | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2599018 | + |
| ChimerDB4 | LUAD | TCGA-62-A46U-01A | SMCHD1 | chr18 | 2688745 | - | NDC80 | chr18 | 2585112 | + |
| ChimerDB4 | LUAD | TCGA-62-A46U-01A | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585112 | + |
| ChimerDB4 | LUAD | TCGA-62-A46U | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + |
| ChimerDB4 | SARC | TCGA-IF-A4AJ-01A | SMCHD1 | chr18 | 2732490 | - | NDC80 | chr18 | 2599018 | + |
| ChimerDB4 | UCEC | TCGA-2E-A9G8-01A | SMCHD1 | chr18 | 2656259 | + | NDC80 | chr18 | 2601394 | + |
| ChimerDB4 | UCEC | TCGA-2E-A9G8-01A | SMCHD1 | chr18 | 2656260 | - | NDC80 | chr18 | 2601395 | + |
| ChimerDB4 | UCEC | TCGA-2E-A9G8-01A | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601395 | + |
| ChimerDB4 | UCEC | TCGA-2E-A9G8-01A | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606414 | + |
| ChimerDB4 | UCEC | TCGA-2E-A9G8 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601395 | + |
| ChimerDB4 | UCEC | TCGA-2E-A9G8 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606414 | + |
Top |
Fusion Gene ORF analysis for SMCHD1-NDC80 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2599018 | + |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585112 | + |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2732490 | - | NDC80 | chr18 | 2599018 | + |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2656259 | + | NDC80 | chr18 | 2601394 | + |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601395 | + |
| In-frame | ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606414 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2599018 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585112 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2732490 | - | NDC80 | chr18 | 2599018 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2656259 | + | NDC80 | chr18 | 2601394 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601395 | + |
| In-frame | ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606414 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2599018 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585112 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2732490 | - | NDC80 | chr18 | 2599018 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2656259 | + | NDC80 | chr18 | 2601394 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601395 | + |
| intron-3CDS | ENST00000609587 | ENST00000261597 | SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606414 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000320876 | SMCHD1 | chr18 | 2688745 | + | ENST00000261597 | NDC80 | chr18 | 2585111 | + | 2622 | 1211 | 314 | 2560 | 748 |
| ENST00000261598 | SMCHD1 | chr18 | 2688745 | + | ENST00000261597 | NDC80 | chr18 | 2585111 | + | 2473 | 1062 | 165 | 2411 | 748 |
| ENST00000320876 | SMCHD1 | chr18 | 2688745 | + | ENST00000261597 | NDC80 | chr18 | 2585112 | + | 2622 | 1211 | 314 | 2560 | 748 |
| ENST00000261598 | SMCHD1 | chr18 | 2688745 | + | ENST00000261597 | NDC80 | chr18 | 2585112 | + | 2473 | 1062 | 165 | 2411 | 748 |
| ENST00000320876 | SMCHD1 | chr18 | 2732490 | - | ENST00000261597 | NDC80 | chr18 | 2599018 | + | 4383 | 3614 | 314 | 4321 | 1335 |
| ENST00000261598 | SMCHD1 | chr18 | 2732490 | - | ENST00000261597 | NDC80 | chr18 | 2599018 | + | 4234 | 3465 | 165 | 4172 | 1335 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + | 0.000515443 | 0.99948454 |
| ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + | 0.000576081 | 0.999424 |
| ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585112 | + | 0.000515443 | 0.99948454 |
| ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585112 | + | 0.000576081 | 0.999424 |
| ENST00000320876 | ENST00000261597 | SMCHD1 | chr18 | 2732490 | - | NDC80 | chr18 | 2599018 | + | 0.000123148 | 0.99987686 |
| ENST00000261598 | ENST00000261597 | SMCHD1 | chr18 | 2732490 | - | NDC80 | chr18 | 2599018 | + | 0.000113859 | 0.99988616 |
Top |
Fusion Genomic Features for SMCHD1-NDC80 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601394 | + | 0.001469484 | 0.9985305 |
| SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2599017 | + | 0.000106196 | 0.9998938 |
| SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606413 | + | 2.84E-06 | 0.99999714 |
| SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + | 1.22E-06 | 0.9999988 |
| SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + | 1.22E-06 | 0.9999988 |
| SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2601394 | + | 0.001469484 | 0.9985305 |
| SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2599017 | + | 0.000106196 | 0.9998938 |
| SMCHD1 | chr18 | 2656260 | + | NDC80 | chr18 | 2606413 | + | 2.84E-06 | 0.99999714 |
| SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + | 1.22E-06 | 0.9999988 |
| SMCHD1 | chr18 | 2688745 | + | NDC80 | chr18 | 2585111 | + | 1.22E-06 | 0.9999988 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
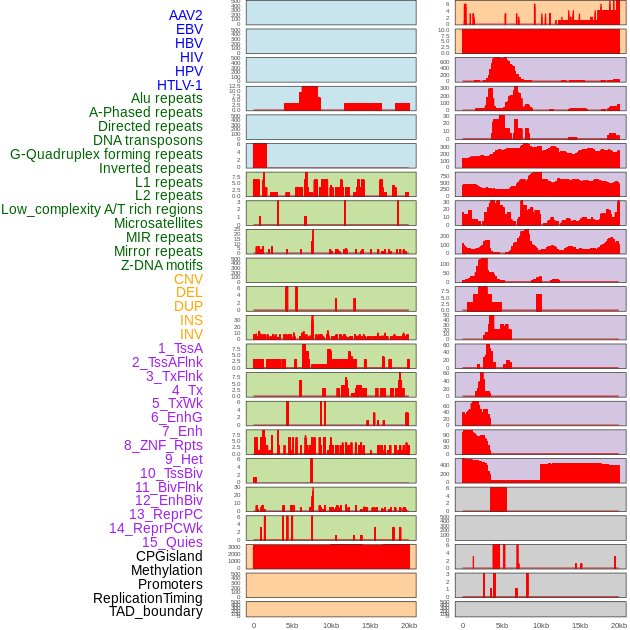 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
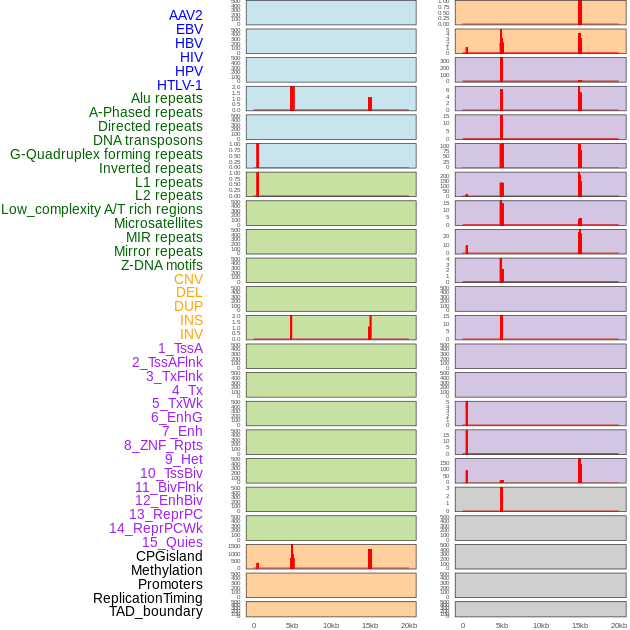 |
Top |
Fusion Protein Features for SMCHD1-NDC80 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr18:2656260/chr18:2601395) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | NDC80 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Acts as a component of the essential kinetochore-associated NDC80 complex, which is required for chromosome segregation and spindle checkpoint activity (PubMed:9315664, PubMed:12351790, PubMed:14654001, PubMed:14699129, PubMed:15062103, PubMed:15235793, PubMed:15239953, PubMed:15548592, PubMed:16732327). Required for kinetochore integrity and the organization of stable microtubule binding sites in the outer plate of the kinetochore (PubMed:15548592). The NDC80 complex synergistically enhances the affinity of the SKA1 complex for microtubules and may allow the NDC80 complex to track depolymerizing microtubules (PubMed:23085020). Plays a role in chromosome congression and is essential for the end-on attachment of the kinetochores to spindle microtubules (PubMed:25743205, PubMed:23891108). {ECO:0000269|PubMed:12351790, ECO:0000269|PubMed:14654001, ECO:0000269|PubMed:14699129, ECO:0000269|PubMed:15062103, ECO:0000269|PubMed:15235793, ECO:0000269|PubMed:15239953, ECO:0000269|PubMed:15548592, ECO:0000269|PubMed:16732327, ECO:0000269|PubMed:23085020, ECO:0000269|PubMed:23891108, ECO:0000269|PubMed:25743205, ECO:0000269|PubMed:9315664}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMCHD1 | chr18:2732490 | chr18:2599018 | ENST00000261598 | - | 25 | 46 | 111_702 | 1092 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2732490 | chr18:2599018 | ENST00000320876 | - | 25 | 48 | 111_702 | 1092 | 2006.0 | Region | ATPase activity domain |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 458_642 | 458 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 458_642 | 407 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 458_642 | 458 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 261_403 | 193 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 458_642 | 193 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 261_403 | 193 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 458_642 | 193 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 458_642 | 407 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SMCHD1 | chr18:2656259 | chr18:2601394 | ENST00000261598 | + | 1 | 46 | 1720_1847 | 62 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656259 | chr18:2601394 | ENST00000320876 | + | 1 | 48 | 1720_1847 | 62 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2599018 | ENST00000261598 | + | 1 | 46 | 1720_1847 | 62 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2599018 | ENST00000320876 | + | 1 | 48 | 1720_1847 | 62 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2601395 | ENST00000261598 | + | 1 | 46 | 1720_1847 | 62 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2601395 | ENST00000320876 | + | 1 | 48 | 1720_1847 | 62 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2606414 | ENST00000261598 | + | 1 | 46 | 1720_1847 | 62 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2606414 | ENST00000320876 | + | 1 | 48 | 1720_1847 | 62 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585111 | ENST00000261598 | + | 7 | 46 | 1720_1847 | 291 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585111 | ENST00000320876 | + | 7 | 48 | 1720_1847 | 291 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585112 | ENST00000261598 | + | 7 | 46 | 1720_1847 | 291 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585112 | ENST00000320876 | + | 7 | 48 | 1720_1847 | 291 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2732490 | chr18:2599018 | ENST00000261598 | - | 25 | 46 | 1720_1847 | 1092 | 1918.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2732490 | chr18:2599018 | ENST00000320876 | - | 25 | 48 | 1720_1847 | 1092 | 2006.0 | Domain | SMC hinge |
| Hgene | SMCHD1 | chr18:2656259 | chr18:2601394 | ENST00000261598 | + | 1 | 46 | 111_702 | 62 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656259 | chr18:2601394 | ENST00000320876 | + | 1 | 48 | 111_702 | 62 | 2006.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2599018 | ENST00000261598 | + | 1 | 46 | 111_702 | 62 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2599018 | ENST00000320876 | + | 1 | 48 | 111_702 | 62 | 2006.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2601395 | ENST00000261598 | + | 1 | 46 | 111_702 | 62 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2601395 | ENST00000320876 | + | 1 | 48 | 111_702 | 62 | 2006.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2606414 | ENST00000261598 | + | 1 | 46 | 111_702 | 62 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2656260 | chr18:2606414 | ENST00000320876 | + | 1 | 48 | 111_702 | 62 | 2006.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585111 | ENST00000261598 | + | 7 | 46 | 111_702 | 291 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585111 | ENST00000320876 | + | 7 | 48 | 111_702 | 291 | 2006.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585112 | ENST00000261598 | + | 7 | 46 | 111_702 | 291 | 1918.0 | Region | ATPase activity domain |
| Hgene | SMCHD1 | chr18:2688745 | chr18:2585112 | ENST00000320876 | + | 7 | 48 | 111_702 | 291 | 2006.0 | Region | ATPase activity domain |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 261_403 | 458 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 261_403 | 407 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 261_403 | 458 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 261_403 | 488 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 458_642 | 488 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 261_403 | 407 | 643.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 1_250 | 458 | 643.0 | Region | Note=Nuclear localization | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 1_250 | 407 | 643.0 | Region | Note=Nuclear localization | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 1_250 | 458 | 643.0 | Region | Note=Nuclear localization | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 1_250 | 488 | 643.0 | Region | Note=Nuclear localization | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 1_250 | 193 | 643.0 | Region | Note=Nuclear localization | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 1_250 | 193 | 643.0 | Region | Note=Nuclear localization | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 1_250 | 407 | 643.0 | Region | Note=Nuclear localization |
Top |
Fusion Gene Sequence for SMCHD1-NDC80 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >84030_84030_1_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000261598_NDC80_chr18_2585111_ENST00000261597_length(transcript)=2473nt_BP=1062nt GAATCGGTTCCCGGGTGATCCTCGCGCCTGCCGCTGCTCGGCCGCCGCCGCTGACGAGGAGCTGCAGCGCGCCGGGCCGAGGCCTCGAGC CGCCCCGGGAGCTGGAGCTGAAGGCGCCGCGCGGAGCGCGCACCTCAGCCCTGAGCCCGGCGGCGGCAGGCGTCGCTGTCTTTTCTCCTT TTCCCCAATATGGCAGCGGCGGACGGCGGCGGGCCTGGTGGGGCCTCTGTGGGGACTGAGGAGGATGGCGGAGGCGTCGGCCACAGGACG GTGTACTTGTTTGATCGGCGCGAAAAGGAGTCCGAGCTCGGGGACCGGCCTCTGCAGGTCGGGGAGCGCTCGGACTACGCGGGATTTCGC GCGTGTGTGTGTCAGACACTTGGCATTTCACCTGAAGAAAAATTTGTTATTACAACAACAAGTAGGAAAGAAATTACCTGTGATAATTTT GATGAAACTGTTAAAGATGGAGTCACCTTATACCTGCTACAGTCGGTCAATCAGTTACTACTGACAGCTACGAAAGAACGAATTGACTTC TTACCTCACTATGACACACTGGTTAAAAGTGGCATGTATGAATATTATGCCAGTGAAGGACAAAATCCTTTGCCATTTGCTCTTGCGGAA TTAATTGACAATTCATTGTCTGCTACTTCTCGTAACATTGGGGTTAGAAGAATACAGATCAAATTGCTTTTTGATGAAACACAAGGAAAA CCTGCTGTTGCAGTGATAGATAATGGAAGAGGAATGACCTCTAAACAGCTTAACAACTGGGCCGTGTATAGGTTGTCAAAATTCACAAGG CAAGGTGACTTTGAAAGTGATCATTCAGGATATGTTCGTCCAGTACCAGTGCCACGCAGTTTAAATAGTGATATTTCCTATTTTGGTGTT GGGGGCAAGCAAGCTGTCTTCTTTGTTGGACAATCAGCCAGAATGATAAGCAAACCTGCAGATTCCCAAGATGTTCACGAGCTTGTGCTT TCTAAAGAAGATTTTGAGAAGAAGGAGAAAAATAAAGAGGCAATATATAGTGGATATATTAGAAACAGAAAGATACATACTGCCATGAAA GAAAGCTCACCTTTATTTGATGATGGGCAGCCTTGGGGAGAAGAAACTGAAGATGGAATTATGCATAATAAGTTGTTTTTGGACTACACC ATAAAATGCTATGAGAGTTTTATGAGTGGTGCCGACAGCTTTGATGAGATGAATGCAGAGCTGCAGTCAAAACTGAAGGATTTATTTAAT GTGGATGCTTTTAAGCTGGAATCATTAGAAGCAAAAAACAGAGCATTGAATGAACAGATTGCAAGATTGGAACAAGAAAGAGAAAAAGAA CCGAATCGTCTAGAGTCGTTGAGAAAACTGAAGGCTTCCTTACAAGGAGATGTTCAAAAGTATCAGGCATACATGAGCAATTTGGAGTCT CATTCAGCCATTCTTGACCAGAAATTAAATGGTCTCAATGAGGAAATTGCTAGAGTAGAACTAGAATGTGAAACAATAAAACAGGAGAAC ACTCGACTACAGAATATCATTGACAACCAGAAGTACTCAGTTGCAGACATTGAGCGAATAAATCATGAAAGAAATGAATTGCAGCAGACT ATTAATAAATTAACCAAGGACCTGGAAGCTGAACAACAGAAGTTGTGGAATGAGGAGTTAAAATATGCCAGAGGCAAAGAAGCGATTGAA ACACAATTAGCAGAGTATCACAAATTGGCTAGAAAATTAAAACTTATTCCTAAAGGTGCTGAGAATTCCAAAGGTTATGACTTTGAAATT AAGTTTAATCCCGAGGCTGGTGCCAACTGCCTTGTCAAATACAGGGCTCAAGTTTATGTACCTCTTAAGGAACTCCTGAATGAAACTGAA GAAGAAATTAATAAAGCCCTAAATAAAAAAATGGGTTTGGAGGATACTTTAGAACAATTGAATGCAATGATAACAGAAAGCAAGAGAAGT GTGAGAACTCTGAAAGAAGAAGTTCAAAAGCTGGATGATCTTTACCAACAAAAAATTAAGGAAGCAGAGGAAGAGGATGAAAAATGTGCC AGTGAGCTTGAGTCCTTGGAGAAACACAAGCACCTGCTAGAAAGTACTGTTAACCAGGGGCTCAGTGAAGCTATGAATGAATTAGATGCT GTTCAGCGGGAATACCAACTAGTTGTGCAAACCACGACTGAAGAAAGACGAAAAGTGGGAAATAACTTGCAACGTCTGTTAGAGATGGTT GCTACACATGTTGGGTCTGTAGAGAAACATCTTGAGGAGCAGATTGCTAAAGTTGATAGAGAATATGAAGAATGCATGTCAGAAGATCTC TCGGAAAATATTAAAGAGATTAGAGATAAGTATGAGAAGAAAGCTACTCTAATTAAGTCTTCTGAAGAATGAAGATAAAATGTTGATCAT >84030_84030_1_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000261598_NDC80_chr18_2585111_ENST00000261597_length(amino acids)=748AA_BP=299 MSFLLFPNMAAADGGGPGGASVGTEEDGGGVGHRTVYLFDRREKESELGDRPLQVGERSDYAGFRACVCQTLGISPEEKFVITTTSRKEI TCDNFDETVKDGVTLYLLQSVNQLLLTATKERIDFLPHYDTLVKSGMYEYYASEGQNPLPFALAELIDNSLSATSRNIGVRRIQIKLLFD ETQGKPAVAVIDNGRGMTSKQLNNWAVYRLSKFTRQGDFESDHSGYVRPVPVPRSLNSDISYFGVGGKQAVFFVGQSARMISKPADSQDV HELVLSKEDFEKKEKNKEAIYSGYIRNRKIHTAMKESSPLFDDGQPWGEETEDGIMHNKLFLDYTIKCYESFMSGADSFDEMNAELQSKL KDLFNVDAFKLESLEAKNRALNEQIARLEQEREKEPNRLESLRKLKASLQGDVQKYQAYMSNLESHSAILDQKLNGLNEEIARVELECET IKQENTRLQNIIDNQKYSVADIERINHERNELQQTINKLTKDLEAEQQKLWNEELKYARGKEAIETQLAEYHKLARKLKLIPKGAENSKG YDFEIKFNPEAGANCLVKYRAQVYVPLKELLNETEEEINKALNKKMGLEDTLEQLNAMITESKRSVRTLKEEVQKLDDLYQQKIKEAEEE DEKCASELESLEKHKHLLESTVNQGLSEAMNELDAVQREYQLVVQTTTEERRKVGNNLQRLLEMVATHVGSVEKHLEEQIAKVDREYEEC -------------------------------------------------------------- >84030_84030_2_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000261598_NDC80_chr18_2585112_ENST00000261597_length(transcript)=2473nt_BP=1062nt GAATCGGTTCCCGGGTGATCCTCGCGCCTGCCGCTGCTCGGCCGCCGCCGCTGACGAGGAGCTGCAGCGCGCCGGGCCGAGGCCTCGAGC CGCCCCGGGAGCTGGAGCTGAAGGCGCCGCGCGGAGCGCGCACCTCAGCCCTGAGCCCGGCGGCGGCAGGCGTCGCTGTCTTTTCTCCTT TTCCCCAATATGGCAGCGGCGGACGGCGGCGGGCCTGGTGGGGCCTCTGTGGGGACTGAGGAGGATGGCGGAGGCGTCGGCCACAGGACG GTGTACTTGTTTGATCGGCGCGAAAAGGAGTCCGAGCTCGGGGACCGGCCTCTGCAGGTCGGGGAGCGCTCGGACTACGCGGGATTTCGC GCGTGTGTGTGTCAGACACTTGGCATTTCACCTGAAGAAAAATTTGTTATTACAACAACAAGTAGGAAAGAAATTACCTGTGATAATTTT GATGAAACTGTTAAAGATGGAGTCACCTTATACCTGCTACAGTCGGTCAATCAGTTACTACTGACAGCTACGAAAGAACGAATTGACTTC TTACCTCACTATGACACACTGGTTAAAAGTGGCATGTATGAATATTATGCCAGTGAAGGACAAAATCCTTTGCCATTTGCTCTTGCGGAA TTAATTGACAATTCATTGTCTGCTACTTCTCGTAACATTGGGGTTAGAAGAATACAGATCAAATTGCTTTTTGATGAAACACAAGGAAAA CCTGCTGTTGCAGTGATAGATAATGGAAGAGGAATGACCTCTAAACAGCTTAACAACTGGGCCGTGTATAGGTTGTCAAAATTCACAAGG CAAGGTGACTTTGAAAGTGATCATTCAGGATATGTTCGTCCAGTACCAGTGCCACGCAGTTTAAATAGTGATATTTCCTATTTTGGTGTT GGGGGCAAGCAAGCTGTCTTCTTTGTTGGACAATCAGCCAGAATGATAAGCAAACCTGCAGATTCCCAAGATGTTCACGAGCTTGTGCTT TCTAAAGAAGATTTTGAGAAGAAGGAGAAAAATAAAGAGGCAATATATAGTGGATATATTAGAAACAGAAAGATACATACTGCCATGAAA GAAAGCTCACCTTTATTTGATGATGGGCAGCCTTGGGGAGAAGAAACTGAAGATGGAATTATGCATAATAAGTTGTTTTTGGACTACACC ATAAAATGCTATGAGAGTTTTATGAGTGGTGCCGACAGCTTTGATGAGATGAATGCAGAGCTGCAGTCAAAACTGAAGGATTTATTTAAT GTGGATGCTTTTAAGCTGGAATCATTAGAAGCAAAAAACAGAGCATTGAATGAACAGATTGCAAGATTGGAACAAGAAAGAGAAAAAGAA CCGAATCGTCTAGAGTCGTTGAGAAAACTGAAGGCTTCCTTACAAGGAGATGTTCAAAAGTATCAGGCATACATGAGCAATTTGGAGTCT CATTCAGCCATTCTTGACCAGAAATTAAATGGTCTCAATGAGGAAATTGCTAGAGTAGAACTAGAATGTGAAACAATAAAACAGGAGAAC ACTCGACTACAGAATATCATTGACAACCAGAAGTACTCAGTTGCAGACATTGAGCGAATAAATCATGAAAGAAATGAATTGCAGCAGACT ATTAATAAATTAACCAAGGACCTGGAAGCTGAACAACAGAAGTTGTGGAATGAGGAGTTAAAATATGCCAGAGGCAAAGAAGCGATTGAA ACACAATTAGCAGAGTATCACAAATTGGCTAGAAAATTAAAACTTATTCCTAAAGGTGCTGAGAATTCCAAAGGTTATGACTTTGAAATT AAGTTTAATCCCGAGGCTGGTGCCAACTGCCTTGTCAAATACAGGGCTCAAGTTTATGTACCTCTTAAGGAACTCCTGAATGAAACTGAA GAAGAAATTAATAAAGCCCTAAATAAAAAAATGGGTTTGGAGGATACTTTAGAACAATTGAATGCAATGATAACAGAAAGCAAGAGAAGT GTGAGAACTCTGAAAGAAGAAGTTCAAAAGCTGGATGATCTTTACCAACAAAAAATTAAGGAAGCAGAGGAAGAGGATGAAAAATGTGCC AGTGAGCTTGAGTCCTTGGAGAAACACAAGCACCTGCTAGAAAGTACTGTTAACCAGGGGCTCAGTGAAGCTATGAATGAATTAGATGCT GTTCAGCGGGAATACCAACTAGTTGTGCAAACCACGACTGAAGAAAGACGAAAAGTGGGAAATAACTTGCAACGTCTGTTAGAGATGGTT GCTACACATGTTGGGTCTGTAGAGAAACATCTTGAGGAGCAGATTGCTAAAGTTGATAGAGAATATGAAGAATGCATGTCAGAAGATCTC TCGGAAAATATTAAAGAGATTAGAGATAAGTATGAGAAGAAAGCTACTCTAATTAAGTCTTCTGAAGAATGAAGATAAAATGTTGATCAT >84030_84030_2_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000261598_NDC80_chr18_2585112_ENST00000261597_length(amino acids)=748AA_BP=299 MSFLLFPNMAAADGGGPGGASVGTEEDGGGVGHRTVYLFDRREKESELGDRPLQVGERSDYAGFRACVCQTLGISPEEKFVITTTSRKEI TCDNFDETVKDGVTLYLLQSVNQLLLTATKERIDFLPHYDTLVKSGMYEYYASEGQNPLPFALAELIDNSLSATSRNIGVRRIQIKLLFD ETQGKPAVAVIDNGRGMTSKQLNNWAVYRLSKFTRQGDFESDHSGYVRPVPVPRSLNSDISYFGVGGKQAVFFVGQSARMISKPADSQDV HELVLSKEDFEKKEKNKEAIYSGYIRNRKIHTAMKESSPLFDDGQPWGEETEDGIMHNKLFLDYTIKCYESFMSGADSFDEMNAELQSKL KDLFNVDAFKLESLEAKNRALNEQIARLEQEREKEPNRLESLRKLKASLQGDVQKYQAYMSNLESHSAILDQKLNGLNEEIARVELECET IKQENTRLQNIIDNQKYSVADIERINHERNELQQTINKLTKDLEAEQQKLWNEELKYARGKEAIETQLAEYHKLARKLKLIPKGAENSKG YDFEIKFNPEAGANCLVKYRAQVYVPLKELLNETEEEINKALNKKMGLEDTLEQLNAMITESKRSVRTLKEEVQKLDDLYQQKIKEAEEE DEKCASELESLEKHKHLLESTVNQGLSEAMNELDAVQREYQLVVQTTTEERRKVGNNLQRLLEMVATHVGSVEKHLEEQIAKVDREYEEC -------------------------------------------------------------- >84030_84030_3_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000320876_NDC80_chr18_2585111_ENST00000261597_length(transcript)=2622nt_BP=1211nt GCGGCGCTGGTCGCGGGTCGCACCGTGAGGCTCGCGTGGGCGGCGATAGGCGCTGGGCCCGGGCCCGGTGAGGAGCGCGCCGCGCGTCCC CTTCTCCTCAGGAGTGGCGGGCCGCGGAAGTGACGTGGTGCACGGGCAGGAGCGCGTTTGAATCGGTTCCCGGGTGATCCTCGCGCCTGC CGCTGCTCGGCCGCCGCCGCTGACGAGGAGCTGCAGCGCGCCGGGCCGAGGCCTCGAGCCGCCCCGGGAGCTGGAGCTGAAGGCGCCGCG CGGAGCGCGCACCTCAGCCCTGAGCCCGGCGGCGGCAGGCGTCGCTGTCTTTTCTCCTTTTCCCCAATATGGCAGCGGCGGACGGCGGCG GGCCTGGTGGGGCCTCTGTGGGGACTGAGGAGGATGGCGGAGGCGTCGGCCACAGGACGGTGTACTTGTTTGATCGGCGCGAAAAGGAGT CCGAGCTCGGGGACCGGCCTCTGCAGGTCGGGGAGCGCTCGGACTACGCGGGATTTCGCGCGTGTGTGTGTCAGACACTTGGCATTTCAC CTGAAGAAAAATTTGTTATTACAACAACAAGTAGGAAAGAAATTACCTGTGATAATTTTGATGAAACTGTTAAAGATGGAGTCACCTTAT ACCTGCTACAGTCGGTCAATCAGTTACTACTGACAGCTACGAAAGAACGAATTGACTTCTTACCTCACTATGACACACTGGTTAAAAGTG GCATGTATGAATATTATGCCAGTGAAGGACAAAATCCTTTGCCATTTGCTCTTGCGGAATTAATTGACAATTCATTGTCTGCTACTTCTC GTAACATTGGGGTTAGAAGAATACAGATCAAATTGCTTTTTGATGAAACACAAGGAAAACCTGCTGTTGCAGTGATAGATAATGGAAGAG GAATGACCTCTAAACAGCTTAACAACTGGGCCGTGTATAGGTTGTCAAAATTCACAAGGCAAGGTGACTTTGAAAGTGATCATTCAGGAT ATGTTCGTCCAGTACCAGTGCCACGCAGTTTAAATAGTGATATTTCCTATTTTGGTGTTGGGGGCAAGCAAGCTGTCTTCTTTGTTGGAC AATCAGCCAGAATGATAAGCAAACCTGCAGATTCCCAAGATGTTCACGAGCTTGTGCTTTCTAAAGAAGATTTTGAGAAGAAGGAGAAAA ATAAAGAGGCAATATATAGTGGATATATTAGAAACAGAAAGATACATACTGCCATGAAAGAAAGCTCACCTTTATTTGATGATGGGCAGC CTTGGGGAGAAGAAACTGAAGATGGAATTATGCATAATAAGTTGTTTTTGGACTACACCATAAAATGCTATGAGAGTTTTATGAGTGGTG CCGACAGCTTTGATGAGATGAATGCAGAGCTGCAGTCAAAACTGAAGGATTTATTTAATGTGGATGCTTTTAAGCTGGAATCATTAGAAG CAAAAAACAGAGCATTGAATGAACAGATTGCAAGATTGGAACAAGAAAGAGAAAAAGAACCGAATCGTCTAGAGTCGTTGAGAAAACTGA AGGCTTCCTTACAAGGAGATGTTCAAAAGTATCAGGCATACATGAGCAATTTGGAGTCTCATTCAGCCATTCTTGACCAGAAATTAAATG GTCTCAATGAGGAAATTGCTAGAGTAGAACTAGAATGTGAAACAATAAAACAGGAGAACACTCGACTACAGAATATCATTGACAACCAGA AGTACTCAGTTGCAGACATTGAGCGAATAAATCATGAAAGAAATGAATTGCAGCAGACTATTAATAAATTAACCAAGGACCTGGAAGCTG AACAACAGAAGTTGTGGAATGAGGAGTTAAAATATGCCAGAGGCAAAGAAGCGATTGAAACACAATTAGCAGAGTATCACAAATTGGCTA GAAAATTAAAACTTATTCCTAAAGGTGCTGAGAATTCCAAAGGTTATGACTTTGAAATTAAGTTTAATCCCGAGGCTGGTGCCAACTGCC TTGTCAAATACAGGGCTCAAGTTTATGTACCTCTTAAGGAACTCCTGAATGAAACTGAAGAAGAAATTAATAAAGCCCTAAATAAAAAAA TGGGTTTGGAGGATACTTTAGAACAATTGAATGCAATGATAACAGAAAGCAAGAGAAGTGTGAGAACTCTGAAAGAAGAAGTTCAAAAGC TGGATGATCTTTACCAACAAAAAATTAAGGAAGCAGAGGAAGAGGATGAAAAATGTGCCAGTGAGCTTGAGTCCTTGGAGAAACACAAGC ACCTGCTAGAAAGTACTGTTAACCAGGGGCTCAGTGAAGCTATGAATGAATTAGATGCTGTTCAGCGGGAATACCAACTAGTTGTGCAAA CCACGACTGAAGAAAGACGAAAAGTGGGAAATAACTTGCAACGTCTGTTAGAGATGGTTGCTACACATGTTGGGTCTGTAGAGAAACATC TTGAGGAGCAGATTGCTAAAGTTGATAGAGAATATGAAGAATGCATGTCAGAAGATCTCTCGGAAAATATTAAAGAGATTAGAGATAAGT ATGAGAAGAAAGCTACTCTAATTAAGTCTTCTGAAGAATGAAGATAAAATGTTGATCATGTATATATATCCATAGTGAATAAAATTGTCT >84030_84030_3_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000320876_NDC80_chr18_2585111_ENST00000261597_length(amino acids)=748AA_BP=299 MSFLLFPNMAAADGGGPGGASVGTEEDGGGVGHRTVYLFDRREKESELGDRPLQVGERSDYAGFRACVCQTLGISPEEKFVITTTSRKEI TCDNFDETVKDGVTLYLLQSVNQLLLTATKERIDFLPHYDTLVKSGMYEYYASEGQNPLPFALAELIDNSLSATSRNIGVRRIQIKLLFD ETQGKPAVAVIDNGRGMTSKQLNNWAVYRLSKFTRQGDFESDHSGYVRPVPVPRSLNSDISYFGVGGKQAVFFVGQSARMISKPADSQDV HELVLSKEDFEKKEKNKEAIYSGYIRNRKIHTAMKESSPLFDDGQPWGEETEDGIMHNKLFLDYTIKCYESFMSGADSFDEMNAELQSKL KDLFNVDAFKLESLEAKNRALNEQIARLEQEREKEPNRLESLRKLKASLQGDVQKYQAYMSNLESHSAILDQKLNGLNEEIARVELECET IKQENTRLQNIIDNQKYSVADIERINHERNELQQTINKLTKDLEAEQQKLWNEELKYARGKEAIETQLAEYHKLARKLKLIPKGAENSKG YDFEIKFNPEAGANCLVKYRAQVYVPLKELLNETEEEINKALNKKMGLEDTLEQLNAMITESKRSVRTLKEEVQKLDDLYQQKIKEAEEE DEKCASELESLEKHKHLLESTVNQGLSEAMNELDAVQREYQLVVQTTTEERRKVGNNLQRLLEMVATHVGSVEKHLEEQIAKVDREYEEC -------------------------------------------------------------- >84030_84030_4_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000320876_NDC80_chr18_2585112_ENST00000261597_length(transcript)=2622nt_BP=1211nt GCGGCGCTGGTCGCGGGTCGCACCGTGAGGCTCGCGTGGGCGGCGATAGGCGCTGGGCCCGGGCCCGGTGAGGAGCGCGCCGCGCGTCCC CTTCTCCTCAGGAGTGGCGGGCCGCGGAAGTGACGTGGTGCACGGGCAGGAGCGCGTTTGAATCGGTTCCCGGGTGATCCTCGCGCCTGC CGCTGCTCGGCCGCCGCCGCTGACGAGGAGCTGCAGCGCGCCGGGCCGAGGCCTCGAGCCGCCCCGGGAGCTGGAGCTGAAGGCGCCGCG CGGAGCGCGCACCTCAGCCCTGAGCCCGGCGGCGGCAGGCGTCGCTGTCTTTTCTCCTTTTCCCCAATATGGCAGCGGCGGACGGCGGCG GGCCTGGTGGGGCCTCTGTGGGGACTGAGGAGGATGGCGGAGGCGTCGGCCACAGGACGGTGTACTTGTTTGATCGGCGCGAAAAGGAGT CCGAGCTCGGGGACCGGCCTCTGCAGGTCGGGGAGCGCTCGGACTACGCGGGATTTCGCGCGTGTGTGTGTCAGACACTTGGCATTTCAC CTGAAGAAAAATTTGTTATTACAACAACAAGTAGGAAAGAAATTACCTGTGATAATTTTGATGAAACTGTTAAAGATGGAGTCACCTTAT ACCTGCTACAGTCGGTCAATCAGTTACTACTGACAGCTACGAAAGAACGAATTGACTTCTTACCTCACTATGACACACTGGTTAAAAGTG GCATGTATGAATATTATGCCAGTGAAGGACAAAATCCTTTGCCATTTGCTCTTGCGGAATTAATTGACAATTCATTGTCTGCTACTTCTC GTAACATTGGGGTTAGAAGAATACAGATCAAATTGCTTTTTGATGAAACACAAGGAAAACCTGCTGTTGCAGTGATAGATAATGGAAGAG GAATGACCTCTAAACAGCTTAACAACTGGGCCGTGTATAGGTTGTCAAAATTCACAAGGCAAGGTGACTTTGAAAGTGATCATTCAGGAT ATGTTCGTCCAGTACCAGTGCCACGCAGTTTAAATAGTGATATTTCCTATTTTGGTGTTGGGGGCAAGCAAGCTGTCTTCTTTGTTGGAC AATCAGCCAGAATGATAAGCAAACCTGCAGATTCCCAAGATGTTCACGAGCTTGTGCTTTCTAAAGAAGATTTTGAGAAGAAGGAGAAAA ATAAAGAGGCAATATATAGTGGATATATTAGAAACAGAAAGATACATACTGCCATGAAAGAAAGCTCACCTTTATTTGATGATGGGCAGC CTTGGGGAGAAGAAACTGAAGATGGAATTATGCATAATAAGTTGTTTTTGGACTACACCATAAAATGCTATGAGAGTTTTATGAGTGGTG CCGACAGCTTTGATGAGATGAATGCAGAGCTGCAGTCAAAACTGAAGGATTTATTTAATGTGGATGCTTTTAAGCTGGAATCATTAGAAG CAAAAAACAGAGCATTGAATGAACAGATTGCAAGATTGGAACAAGAAAGAGAAAAAGAACCGAATCGTCTAGAGTCGTTGAGAAAACTGA AGGCTTCCTTACAAGGAGATGTTCAAAAGTATCAGGCATACATGAGCAATTTGGAGTCTCATTCAGCCATTCTTGACCAGAAATTAAATG GTCTCAATGAGGAAATTGCTAGAGTAGAACTAGAATGTGAAACAATAAAACAGGAGAACACTCGACTACAGAATATCATTGACAACCAGA AGTACTCAGTTGCAGACATTGAGCGAATAAATCATGAAAGAAATGAATTGCAGCAGACTATTAATAAATTAACCAAGGACCTGGAAGCTG AACAACAGAAGTTGTGGAATGAGGAGTTAAAATATGCCAGAGGCAAAGAAGCGATTGAAACACAATTAGCAGAGTATCACAAATTGGCTA GAAAATTAAAACTTATTCCTAAAGGTGCTGAGAATTCCAAAGGTTATGACTTTGAAATTAAGTTTAATCCCGAGGCTGGTGCCAACTGCC TTGTCAAATACAGGGCTCAAGTTTATGTACCTCTTAAGGAACTCCTGAATGAAACTGAAGAAGAAATTAATAAAGCCCTAAATAAAAAAA TGGGTTTGGAGGATACTTTAGAACAATTGAATGCAATGATAACAGAAAGCAAGAGAAGTGTGAGAACTCTGAAAGAAGAAGTTCAAAAGC TGGATGATCTTTACCAACAAAAAATTAAGGAAGCAGAGGAAGAGGATGAAAAATGTGCCAGTGAGCTTGAGTCCTTGGAGAAACACAAGC ACCTGCTAGAAAGTACTGTTAACCAGGGGCTCAGTGAAGCTATGAATGAATTAGATGCTGTTCAGCGGGAATACCAACTAGTTGTGCAAA CCACGACTGAAGAAAGACGAAAAGTGGGAAATAACTTGCAACGTCTGTTAGAGATGGTTGCTACACATGTTGGGTCTGTAGAGAAACATC TTGAGGAGCAGATTGCTAAAGTTGATAGAGAATATGAAGAATGCATGTCAGAAGATCTCTCGGAAAATATTAAAGAGATTAGAGATAAGT ATGAGAAGAAAGCTACTCTAATTAAGTCTTCTGAAGAATGAAGATAAAATGTTGATCATGTATATATATCCATAGTGAATAAAATTGTCT >84030_84030_4_SMCHD1-NDC80_SMCHD1_chr18_2688745_ENST00000320876_NDC80_chr18_2585112_ENST00000261597_length(amino acids)=748AA_BP=299 MSFLLFPNMAAADGGGPGGASVGTEEDGGGVGHRTVYLFDRREKESELGDRPLQVGERSDYAGFRACVCQTLGISPEEKFVITTTSRKEI TCDNFDETVKDGVTLYLLQSVNQLLLTATKERIDFLPHYDTLVKSGMYEYYASEGQNPLPFALAELIDNSLSATSRNIGVRRIQIKLLFD ETQGKPAVAVIDNGRGMTSKQLNNWAVYRLSKFTRQGDFESDHSGYVRPVPVPRSLNSDISYFGVGGKQAVFFVGQSARMISKPADSQDV HELVLSKEDFEKKEKNKEAIYSGYIRNRKIHTAMKESSPLFDDGQPWGEETEDGIMHNKLFLDYTIKCYESFMSGADSFDEMNAELQSKL KDLFNVDAFKLESLEAKNRALNEQIARLEQEREKEPNRLESLRKLKASLQGDVQKYQAYMSNLESHSAILDQKLNGLNEEIARVELECET IKQENTRLQNIIDNQKYSVADIERINHERNELQQTINKLTKDLEAEQQKLWNEELKYARGKEAIETQLAEYHKLARKLKLIPKGAENSKG YDFEIKFNPEAGANCLVKYRAQVYVPLKELLNETEEEINKALNKKMGLEDTLEQLNAMITESKRSVRTLKEEVQKLDDLYQQKIKEAEEE DEKCASELESLEKHKHLLESTVNQGLSEAMNELDAVQREYQLVVQTTTEERRKVGNNLQRLLEMVATHVGSVEKHLEEQIAKVDREYEEC -------------------------------------------------------------- >84030_84030_5_SMCHD1-NDC80_SMCHD1_chr18_2732490_ENST00000261598_NDC80_chr18_2599018_ENST00000261597_length(transcript)=4234nt_BP=3465nt GAATCGGTTCCCGGGTGATCCTCGCGCCTGCCGCTGCTCGGCCGCCGCCGCTGACGAGGAGCTGCAGCGCGCCGGGCCGAGGCCTCGAGC CGCCCCGGGAGCTGGAGCTGAAGGCGCCGCGCGGAGCGCGCACCTCAGCCCTGAGCCCGGCGGCGGCAGGCGTCGCTGTCTTTTCTCCTT TTCCCCAATATGGCAGCGGCGGACGGCGGCGGGCCTGGTGGGGCCTCTGTGGGGACTGAGGAGGATGGCGGAGGCGTCGGCCACAGGACG GTGTACTTGTTTGATCGGCGCGAAAAGGAGTCCGAGCTCGGGGACCGGCCTCTGCAGGTCGGGGAGCGCTCGGACTACGCGGGATTTCGC GCGTGTGTGTGTCAGACACTTGGCATTTCACCTGAAGAAAAATTTGTTATTACAACAACAAGTAGGAAAGAAATTACCTGTGATAATTTT GATGAAACTGTTAAAGATGGAGTCACCTTATACCTGCTACAGTCGGTCAATCAGTTACTACTGACAGCTACGAAAGAACGAATTGACTTC TTACCTCACTATGACACACTGGTTAAAAGTGGCATGTATGAATATTATGCCAGTGAAGGACAAAATCCTTTGCCATTTGCTCTTGCGGAA TTAATTGACAATTCATTGTCTGCTACTTCTCGTAACATTGGGGTTAGAAGAATACAGATCAAATTGCTTTTTGATGAAACACAAGGAAAA CCTGCTGTTGCAGTGATAGATAATGGAAGAGGAATGACCTCTAAACAGCTTAACAACTGGGCCGTGTATAGGTTGTCAAAATTCACAAGG CAAGGTGACTTTGAAAGTGATCATTCAGGATATGTTCGTCCAGTACCAGTGCCACGCAGTTTAAATAGTGATATTTCCTATTTTGGTGTT GGGGGCAAGCAAGCTGTCTTCTTTGTTGGACAATCAGCCAGAATGATAAGCAAACCTGCAGATTCCCAAGATGTTCACGAGCTTGTGCTT TCTAAAGAAGATTTTGAGAAGAAGGAGAAAAATAAAGAGGCAATATATAGTGGATATATTAGAAACAGAAAGCCCTCTGATTCTGTTCAC ATTACAAATGATGATGAAAGATTTCTACATCATCTTATCATAGAGGAGAAGGAAAAAGATAGCTTTACTGCTGTGGTTATCACAGGGGTA CAACCAGAACACATACAGTACTTGAAAAATTATTTCCACCTTTGGACACGACAGTTAGCGCATATTTATCACTACTATATTCATGGCCCA AAAGGAAATGAAATACGAACATCAAAAGAAGTTGAACCTTTCAACAATATTGATATTGAAATTTCTATGTTTGAAAAAGGGAAGGTACCT AAGATTGTCAACCTAAGGGAAATACAAGACGACATGCAGACGTTGTATGTAAACACAGCAGCTGATAGTTTTGAATTCAAAGCTCATGTT GAAGGAGATGGTGTAGTGGAAGGGATTATCCGTTATCATCCATTCTTATATGATAGAGAAACTTACCCTGATGATCCATGCTTTCCATCA AAATTAAAAGATGAAGATGATGAAGATGATTGTTTCATACTTGAGAAAGCAGCTAGAGGGAAAAGGCCTATTTTTGAATGTTTTTGGAAT GGACGATTAATACCATATACATCAGTTGAAGACTTTGACTGGTGTACTCCTCCTAAGAAGAGAGGGCTTGCACCAATTGAATGCTACAAT AGGATATCTGGTGCATTATTCACTAATGACAAATTCCAGGTCAGCACAAATAAATTGACGTTTATGGATCTTGAGCTAAAATTGAAAGAT AAGAACACCCTTTTTACAAGGATTTTAAATGGACAGGAACAGCGAATGAAAATTGACAGAGAATTTGCTTTGTGGCTGAAGGACTGTCAT GAGAAGTATGATAAACAAATAAAATTTACACTTTTTAAGGGAGTAATTACACGTCCTGATCTTCCTTCTAAAAAGCAAGGTCCCTGGGCA ACATATGCAGCAATAGAATGGGATGGAAAGATATACAAAGCAGGACAGCTGGTCAAGACAATCAAGACACTTCCCCTCTTTTATGGAAGC ATAGTAAGATTTTTTCTTTATGGCGATCATGATGGAGAAGTATATGCTACAGGAGGAGAGGTTCAAATTGCAATGGAACCTCAGGCACTA TATGATGAAGTAAGAACTGTGCCAATTGCAAAGCTGGATAGGACAGTTGCTGAGAAAGCTGTTAAAAAATATGTAGAAGATGAAATGGCA AGGCTCCCTGATAGATTGTCAGTAACTTGGCCTGAAGGAGATGAATTATTGCCTAATGAGGTTAGGCCTGCTGGAACCCCTATTGGTGCG TTAAGAATTGAAATACTGAATAAAAAAGGGGAAGCAATGCAAAAGCTTCCAGGAACAAGCCATGGAGGGTCAAAGAAACTCCTGGTTGAG CTCAAAGTTATTTTACATTCTTCAAGTGGAAATAAAGAGATTATTTCGCATATTAGTCAACATGGAGGAAAATGGCCTTACTGGTTTAAA AAAATGGAAAATATTCAGAAGTTGGGGAATTATACCTTGAAATTACAAGTTGTGTTGAATGAAAGTAATGCAGACACTTATGCAGGAAGA CCACTACCATCTAAAGCAATTAAGTTTTCTGTTAAAGAGGGTAAGCCAGAGAAATTTTCATTTGGTCTTCTGGATCTTCCTTTTCGTGTT GGAGTTCCATTTAATATCCCTCTGGAGTTTCAGGATGAATTTGGTCATACCAGTCAACTAGTAACTGATATTCAGCCAGTTCTTGAAGCA AGTGGTTTATCTTTACATTATGAAGAAATAACCAAAGGACCAAATTGTGTAATTCGAGGTGTTACAGCCAAGGGCCCTGTAAACTCTTGT CAAGGCAAGAATTATAATCTGAAGGTTACTCTGCCTGGCTTAAAAGAAGACTCACAGATTTTGAAAATTAGATTACTACCTGGTCACCCT CGTCGACTGAAAGTGAAACCTGATTCTGAAATTTTAGTTATAGAAAATGGAACAGCTTTCCCATTTCAGGTGGAAGTTTTAGATGAATCA GACAACATAACAGCACAACCAAAATTGATTGTTCATTGTAAGTTTTCAGGTGCTCCAAACCTTCCAGTCTATGTTGTAGATTGCAGTAGT TCTGGAACCAGTATTTTAACAGGATCTGCAATTCAAGTTCAGAATATTAAAAAAGACCAGACGCTTAAAGCAAGAATTGAAATACCTAGT TGTAAAGATGTGGCACCTGTGGAGAAGACTATTAAGTTGCTTCCCAGTAGCCATGTTGCAAGACTACAAATATTCAGTGTAGAAGGACAA AAGGCAATTCAGATCAAACATCAGGATGAGGTTAATTGGATAGCGGGTGATATTATGCATAATCTTATTTTTCAAATGTATGATGAAGGA GAAAGAGAAATCAATATAACATCAGCTTTAGCAGAAAAAATTAAAATTGAAACACAATTAGCAGAGTATCACAAATTGGCTAGAAAATTA AAACTTATTCCTAAAGGTGCTGAGAATTCCAAAGGTTATGACTTTGAAATTAAGTTTAATCCCGAGGCTGGTGCCAACTGCCTTGTCAAA TACAGGGCTCAAGTTTATGTACCTCTTAAGGAACTCCTGAATGAAACTGAAGAAGAAATTAATAAAGCCCTAAATAAAAAAATGGGTTTG GAGGATACTTTAGAACAATTGAATGCAATGATAACAGAAAGCAAGAGAAGTGTGAGAACTCTGAAAGAAGAAGTTCAAAAGCTGGATGAT CTTTACCAACAAAAAATTAAGGAAGCAGAGGAAGAGGATGAAAAATGTGCCAGTGAGCTTGAGTCCTTGGAGAAACACAAGCACCTGCTA GAAAGTACTGTTAACCAGGGGCTCAGTGAAGCTATGAATGAATTAGATGCTGTTCAGCGGGAATACCAACTAGTTGTGCAAACCACGACT GAAGAAAGACGAAAAGTGGGAAATAACTTGCAACGTCTGTTAGAGATGGTTGCTACACATGTTGGGTCTGTAGAGAAACATCTTGAGGAG CAGATTGCTAAAGTTGATAGAGAATATGAAGAATGCATGTCAGAAGATCTCTCGGAAAATATTAAAGAGATTAGAGATAAGTATGAGAAG AAAGCTACTCTAATTAAGTCTTCTGAAGAATGAAGATAAAATGTTGATCATGTATATATATCCATAGTGAATAAAATTGTCTCAGTAAAG >84030_84030_5_SMCHD1-NDC80_SMCHD1_chr18_2732490_ENST00000261598_NDC80_chr18_2599018_ENST00000261597_length(amino acids)=1335AA_BP=1100 MSFLLFPNMAAADGGGPGGASVGTEEDGGGVGHRTVYLFDRREKESELGDRPLQVGERSDYAGFRACVCQTLGISPEEKFVITTTSRKEI TCDNFDETVKDGVTLYLLQSVNQLLLTATKERIDFLPHYDTLVKSGMYEYYASEGQNPLPFALAELIDNSLSATSRNIGVRRIQIKLLFD ETQGKPAVAVIDNGRGMTSKQLNNWAVYRLSKFTRQGDFESDHSGYVRPVPVPRSLNSDISYFGVGGKQAVFFVGQSARMISKPADSQDV HELVLSKEDFEKKEKNKEAIYSGYIRNRKPSDSVHITNDDERFLHHLIIEEKEKDSFTAVVITGVQPEHIQYLKNYFHLWTRQLAHIYHY YIHGPKGNEIRTSKEVEPFNNIDIEISMFEKGKVPKIVNLREIQDDMQTLYVNTAADSFEFKAHVEGDGVVEGIIRYHPFLYDRETYPDD PCFPSKLKDEDDEDDCFILEKAARGKRPIFECFWNGRLIPYTSVEDFDWCTPPKKRGLAPIECYNRISGALFTNDKFQVSTNKLTFMDLE LKLKDKNTLFTRILNGQEQRMKIDREFALWLKDCHEKYDKQIKFTLFKGVITRPDLPSKKQGPWATYAAIEWDGKIYKAGQLVKTIKTLP LFYGSIVRFFLYGDHDGEVYATGGEVQIAMEPQALYDEVRTVPIAKLDRTVAEKAVKKYVEDEMARLPDRLSVTWPEGDELLPNEVRPAG TPIGALRIEILNKKGEAMQKLPGTSHGGSKKLLVELKVILHSSSGNKEIISHISQHGGKWPYWFKKMENIQKLGNYTLKLQVVLNESNAD TYAGRPLPSKAIKFSVKEGKPEKFSFGLLDLPFRVGVPFNIPLEFQDEFGHTSQLVTDIQPVLEASGLSLHYEEITKGPNCVIRGVTAKG PVNSCQGKNYNLKVTLPGLKEDSQILKIRLLPGHPRRLKVKPDSEILVIENGTAFPFQVEVLDESDNITAQPKLIVHCKFSGAPNLPVYV VDCSSSGTSILTGSAIQVQNIKKDQTLKARIEIPSCKDVAPVEKTIKLLPSSHVARLQIFSVEGQKAIQIKHQDEVNWIAGDIMHNLIFQ MYDEGEREINITSALAEKIKIETQLAEYHKLARKLKLIPKGAENSKGYDFEIKFNPEAGANCLVKYRAQVYVPLKELLNETEEEINKALN KKMGLEDTLEQLNAMITESKRSVRTLKEEVQKLDDLYQQKIKEAEEEDEKCASELESLEKHKHLLESTVNQGLSEAMNELDAVQREYQLV -------------------------------------------------------------- >84030_84030_6_SMCHD1-NDC80_SMCHD1_chr18_2732490_ENST00000320876_NDC80_chr18_2599018_ENST00000261597_length(transcript)=4383nt_BP=3614nt GCGGCGCTGGTCGCGGGTCGCACCGTGAGGCTCGCGTGGGCGGCGATAGGCGCTGGGCCCGGGCCCGGTGAGGAGCGCGCCGCGCGTCCC CTTCTCCTCAGGAGTGGCGGGCCGCGGAAGTGACGTGGTGCACGGGCAGGAGCGCGTTTGAATCGGTTCCCGGGTGATCCTCGCGCCTGC CGCTGCTCGGCCGCCGCCGCTGACGAGGAGCTGCAGCGCGCCGGGCCGAGGCCTCGAGCCGCCCCGGGAGCTGGAGCTGAAGGCGCCGCG CGGAGCGCGCACCTCAGCCCTGAGCCCGGCGGCGGCAGGCGTCGCTGTCTTTTCTCCTTTTCCCCAATATGGCAGCGGCGGACGGCGGCG GGCCTGGTGGGGCCTCTGTGGGGACTGAGGAGGATGGCGGAGGCGTCGGCCACAGGACGGTGTACTTGTTTGATCGGCGCGAAAAGGAGT CCGAGCTCGGGGACCGGCCTCTGCAGGTCGGGGAGCGCTCGGACTACGCGGGATTTCGCGCGTGTGTGTGTCAGACACTTGGCATTTCAC CTGAAGAAAAATTTGTTATTACAACAACAAGTAGGAAAGAAATTACCTGTGATAATTTTGATGAAACTGTTAAAGATGGAGTCACCTTAT ACCTGCTACAGTCGGTCAATCAGTTACTACTGACAGCTACGAAAGAACGAATTGACTTCTTACCTCACTATGACACACTGGTTAAAAGTG GCATGTATGAATATTATGCCAGTGAAGGACAAAATCCTTTGCCATTTGCTCTTGCGGAATTAATTGACAATTCATTGTCTGCTACTTCTC GTAACATTGGGGTTAGAAGAATACAGATCAAATTGCTTTTTGATGAAACACAAGGAAAACCTGCTGTTGCAGTGATAGATAATGGAAGAG GAATGACCTCTAAACAGCTTAACAACTGGGCCGTGTATAGGTTGTCAAAATTCACAAGGCAAGGTGACTTTGAAAGTGATCATTCAGGAT ATGTTCGTCCAGTACCAGTGCCACGCAGTTTAAATAGTGATATTTCCTATTTTGGTGTTGGGGGCAAGCAAGCTGTCTTCTTTGTTGGAC AATCAGCCAGAATGATAAGCAAACCTGCAGATTCCCAAGATGTTCACGAGCTTGTGCTTTCTAAAGAAGATTTTGAGAAGAAGGAGAAAA ATAAAGAGGCAATATATAGTGGATATATTAGAAACAGAAAGCCCTCTGATTCTGTTCACATTACAAATGATGATGAAAGATTTCTACATC ATCTTATCATAGAGGAGAAGGAAAAAGATAGCTTTACTGCTGTGGTTATCACAGGGGTACAACCAGAACACATACAGTACTTGAAAAATT ATTTCCACCTTTGGACACGACAGTTAGCGCATATTTATCACTACTATATTCATGGCCCAAAAGGAAATGAAATACGAACATCAAAAGAAG TTGAACCTTTCAACAATATTGATATTGAAATTTCTATGTTTGAAAAAGGGAAGGTACCTAAGATTGTCAACCTAAGGGAAATACAAGACG ACATGCAGACGTTGTATGTAAACACAGCAGCTGATAGTTTTGAATTCAAAGCTCATGTTGAAGGAGATGGTGTAGTGGAAGGGATTATCC GTTATCATCCATTCTTATATGATAGAGAAACTTACCCTGATGATCCATGCTTTCCATCAAAATTAAAAGATGAAGATGATGAAGATGATT GTTTCATACTTGAGAAAGCAGCTAGAGGGAAAAGGCCTATTTTTGAATGTTTTTGGAATGGACGATTAATACCATATACATCAGTTGAAG ACTTTGACTGGTGTACTCCTCCTAAGAAGAGAGGGCTTGCACCAATTGAATGCTACAATAGGATATCTGGTGCATTATTCACTAATGACA AATTCCAGGTCAGCACAAATAAATTGACGTTTATGGATCTTGAGCTAAAATTGAAAGATAAGAACACCCTTTTTACAAGGATTTTAAATG GACAGGAACAGCGAATGAAAATTGACAGAGAATTTGCTTTGTGGCTGAAGGACTGTCATGAGAAGTATGATAAACAAATAAAATTTACAC TTTTTAAGGGAGTAATTACACGTCCTGATCTTCCTTCTAAAAAGCAAGGTCCCTGGGCAACATATGCAGCAATAGAATGGGATGGAAAGA TATACAAAGCAGGACAGCTGGTCAAGACAATCAAGACACTTCCCCTCTTTTATGGAAGCATAGTAAGATTTTTTCTTTATGGCGATCATG ATGGAGAAGTATATGCTACAGGAGGAGAGGTTCAAATTGCAATGGAACCTCAGGCACTATATGATGAAGTAAGAACTGTGCCAATTGCAA AGCTGGATAGGACAGTTGCTGAGAAAGCTGTTAAAAAATATGTAGAAGATGAAATGGCAAGGCTCCCTGATAGATTGTCAGTAACTTGGC CTGAAGGAGATGAATTATTGCCTAATGAGGTTAGGCCTGCTGGAACCCCTATTGGTGCGTTAAGAATTGAAATACTGAATAAAAAAGGGG AAGCAATGCAAAAGCTTCCAGGAACAAGCCATGGAGGGTCAAAGAAACTCCTGGTTGAGCTCAAAGTTATTTTACATTCTTCAAGTGGAA ATAAAGAGATTATTTCGCATATTAGTCAACATGGAGGAAAATGGCCTTACTGGTTTAAAAAAATGGAAAATATTCAGAAGTTGGGGAATT ATACCTTGAAATTACAAGTTGTGTTGAATGAAAGTAATGCAGACACTTATGCAGGAAGACCACTACCATCTAAAGCAATTAAGTTTTCTG TTAAAGAGGGTAAGCCAGAGAAATTTTCATTTGGTCTTCTGGATCTTCCTTTTCGTGTTGGAGTTCCATTTAATATCCCTCTGGAGTTTC AGGATGAATTTGGTCATACCAGTCAACTAGTAACTGATATTCAGCCAGTTCTTGAAGCAAGTGGTTTATCTTTACATTATGAAGAAATAA CCAAAGGACCAAATTGTGTAATTCGAGGTGTTACAGCCAAGGGCCCTGTAAACTCTTGTCAAGGCAAGAATTATAATCTGAAGGTTACTC TGCCTGGCTTAAAAGAAGACTCACAGATTTTGAAAATTAGATTACTACCTGGTCACCCTCGTCGACTGAAAGTGAAACCTGATTCTGAAA TTTTAGTTATAGAAAATGGAACAGCTTTCCCATTTCAGGTGGAAGTTTTAGATGAATCAGACAACATAACAGCACAACCAAAATTGATTG TTCATTGTAAGTTTTCAGGTGCTCCAAACCTTCCAGTCTATGTTGTAGATTGCAGTAGTTCTGGAACCAGTATTTTAACAGGATCTGCAA TTCAAGTTCAGAATATTAAAAAAGACCAGACGCTTAAAGCAAGAATTGAAATACCTAGTTGTAAAGATGTGGCACCTGTGGAGAAGACTA TTAAGTTGCTTCCCAGTAGCCATGTTGCAAGACTACAAATATTCAGTGTAGAAGGACAAAAGGCAATTCAGATCAAACATCAGGATGAGG TTAATTGGATAGCGGGTGATATTATGCATAATCTTATTTTTCAAATGTATGATGAAGGAGAAAGAGAAATCAATATAACATCAGCTTTAG CAGAAAAAATTAAAATTGAAACACAATTAGCAGAGTATCACAAATTGGCTAGAAAATTAAAACTTATTCCTAAAGGTGCTGAGAATTCCA AAGGTTATGACTTTGAAATTAAGTTTAATCCCGAGGCTGGTGCCAACTGCCTTGTCAAATACAGGGCTCAAGTTTATGTACCTCTTAAGG AACTCCTGAATGAAACTGAAGAAGAAATTAATAAAGCCCTAAATAAAAAAATGGGTTTGGAGGATACTTTAGAACAATTGAATGCAATGA TAACAGAAAGCAAGAGAAGTGTGAGAACTCTGAAAGAAGAAGTTCAAAAGCTGGATGATCTTTACCAACAAAAAATTAAGGAAGCAGAGG AAGAGGATGAAAAATGTGCCAGTGAGCTTGAGTCCTTGGAGAAACACAAGCACCTGCTAGAAAGTACTGTTAACCAGGGGCTCAGTGAAG CTATGAATGAATTAGATGCTGTTCAGCGGGAATACCAACTAGTTGTGCAAACCACGACTGAAGAAAGACGAAAAGTGGGAAATAACTTGC AACGTCTGTTAGAGATGGTTGCTACACATGTTGGGTCTGTAGAGAAACATCTTGAGGAGCAGATTGCTAAAGTTGATAGAGAATATGAAG AATGCATGTCAGAAGATCTCTCGGAAAATATTAAAGAGATTAGAGATAAGTATGAGAAGAAAGCTACTCTAATTAAGTCTTCTGAAGAAT >84030_84030_6_SMCHD1-NDC80_SMCHD1_chr18_2732490_ENST00000320876_NDC80_chr18_2599018_ENST00000261597_length(amino acids)=1335AA_BP=1100 MSFLLFPNMAAADGGGPGGASVGTEEDGGGVGHRTVYLFDRREKESELGDRPLQVGERSDYAGFRACVCQTLGISPEEKFVITTTSRKEI TCDNFDETVKDGVTLYLLQSVNQLLLTATKERIDFLPHYDTLVKSGMYEYYASEGQNPLPFALAELIDNSLSATSRNIGVRRIQIKLLFD ETQGKPAVAVIDNGRGMTSKQLNNWAVYRLSKFTRQGDFESDHSGYVRPVPVPRSLNSDISYFGVGGKQAVFFVGQSARMISKPADSQDV HELVLSKEDFEKKEKNKEAIYSGYIRNRKPSDSVHITNDDERFLHHLIIEEKEKDSFTAVVITGVQPEHIQYLKNYFHLWTRQLAHIYHY YIHGPKGNEIRTSKEVEPFNNIDIEISMFEKGKVPKIVNLREIQDDMQTLYVNTAADSFEFKAHVEGDGVVEGIIRYHPFLYDRETYPDD PCFPSKLKDEDDEDDCFILEKAARGKRPIFECFWNGRLIPYTSVEDFDWCTPPKKRGLAPIECYNRISGALFTNDKFQVSTNKLTFMDLE LKLKDKNTLFTRILNGQEQRMKIDREFALWLKDCHEKYDKQIKFTLFKGVITRPDLPSKKQGPWATYAAIEWDGKIYKAGQLVKTIKTLP LFYGSIVRFFLYGDHDGEVYATGGEVQIAMEPQALYDEVRTVPIAKLDRTVAEKAVKKYVEDEMARLPDRLSVTWPEGDELLPNEVRPAG TPIGALRIEILNKKGEAMQKLPGTSHGGSKKLLVELKVILHSSSGNKEIISHISQHGGKWPYWFKKMENIQKLGNYTLKLQVVLNESNAD TYAGRPLPSKAIKFSVKEGKPEKFSFGLLDLPFRVGVPFNIPLEFQDEFGHTSQLVTDIQPVLEASGLSLHYEEITKGPNCVIRGVTAKG PVNSCQGKNYNLKVTLPGLKEDSQILKIRLLPGHPRRLKVKPDSEILVIENGTAFPFQVEVLDESDNITAQPKLIVHCKFSGAPNLPVYV VDCSSSGTSILTGSAIQVQNIKKDQTLKARIEIPSCKDVAPVEKTIKLLPSSHVARLQIFSVEGQKAIQIKHQDEVNWIAGDIMHNLIFQ MYDEGEREINITSALAEKIKIETQLAEYHKLARKLKLIPKGAENSKGYDFEIKFNPEAGANCLVKYRAQVYVPLKELLNETEEEINKALN KKMGLEDTLEQLNAMITESKRSVRTLKEEVQKLDDLYQQKIKEAEEEDEKCASELESLEKHKHLLESTVNQGLSEAMNELDAVQREYQLV -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SMCHD1-NDC80 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 251_618 | 193.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 251_618 | 193.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 361_547 | 193.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 361_547 | 193.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 251_431 | 193.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 251_431 | 193.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 446_642 | 407.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 446_642 | 193.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 446_642 | 193.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 446_642 | 407.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 251_618 | 458.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 251_618 | 407.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 251_618 | 458.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 251_618 | 488.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 251_618 | 407.0 | 643.0 | NEK2 and ZWINT | |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 361_547 | 458.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 361_547 | 407.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 361_547 | 458.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 361_547 | 488.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 361_547 | 407.0 | 643.0 | PSMC2 and SMC1A | |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 128_251 | 458.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 128_251 | 407.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 128_251 | 458.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 128_251 | 488.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 128_251 | 193.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 128_251 | 193.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 128_251 | 407.0 | 643.0 | RB1 | |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 251_431 | 458.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 251_431 | 407.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 251_431 | 458.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 251_431 | 488.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 251_431 | 407.0 | 643.0 | SMC1A | |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 446_642 | 458.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 446_642 | 458.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 446_642 | 488.0 | 643.0 | the C-terminus of CDCA1 and the SPBC24-SPBC25 subcomplex | |
| Tgene | NDC80 | chr18:2656259 | chr18:2601394 | ENST00000261597 | 11 | 17 | 1_445 | 458.0 | 643.0 | the N-terminus of CDCA1 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2599018 | ENST00000261597 | 10 | 17 | 1_445 | 407.0 | 643.0 | the N-terminus of CDCA1 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2601395 | ENST00000261597 | 11 | 17 | 1_445 | 458.0 | 643.0 | the N-terminus of CDCA1 | |
| Tgene | NDC80 | chr18:2656260 | chr18:2606414 | ENST00000261597 | 12 | 17 | 1_445 | 488.0 | 643.0 | the N-terminus of CDCA1 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585111 | ENST00000261597 | 5 | 17 | 1_445 | 193.0 | 643.0 | the N-terminus of CDCA1 | |
| Tgene | NDC80 | chr18:2688745 | chr18:2585112 | ENST00000261597 | 5 | 17 | 1_445 | 193.0 | 643.0 | the N-terminus of CDCA1 | |
| Tgene | NDC80 | chr18:2732490 | chr18:2599018 | ENST00000261597 | 10 | 17 | 1_445 | 407.0 | 643.0 | the N-terminus of CDCA1 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SMCHD1-NDC80 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SMCHD1-NDC80 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
