
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:SSH1-SART3 (FusionGDB2 ID:86759) |
Fusion Gene Summary for SSH1-SART3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: SSH1-SART3 | Fusion gene ID: 86759 | Hgene | Tgene | Gene symbol | SSH1 | SART3 | Gene ID | 54434 | 9733 |
| Gene name | slingshot protein phosphatase 1 | spliceosome associated factor 3, U4/U6 recycling protein | |
| Synonyms | SSH1L | DSAP1|P100|RP11-13G14|TIP110|p110|p110(nrb) | |
| Cytomap | 12q24.11 | 12q23.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein phosphatase Slingshot homolog 1SSH-like protein 1hSSH-1Lslingshot homolog 1 | squamous cell carcinoma antigen recognized by T-cells 3HIV-1 Tat-interacting protein of 110kDaPRP24 homologSART-3hSART-3p110 nuclear RNA-binding proteinsquamous cell carcinoma antigen recognized by T cells 3tat-interacting protein of 110 kDa | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000326470, ENST00000326495, ENST00000360239, ENST00000551165, ENST00000546812, | ENST00000546611, ENST00000552221, ENST00000228284, ENST00000431469, | |
| Fusion gene scores | * DoF score | 9 X 9 X 8=648 | 6 X 7 X 2=84 |
| # samples | 13 | 7 | |
| ** MAII score | log2(13/648*10)=-2.31748218985617 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/84*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: SSH1 [Title/Abstract] AND SART3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | SSH1(109198832)-SART3(108932865), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | SSH1-SART3 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. SSH1-SART3 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | SSH1 | GO:0032268 | regulation of cellular protein metabolic process | 19000834 |
| Hgene | SSH1 | GO:0071318 | cellular response to ATP | 19000834 |
| Tgene | SART3 | GO:0000244 | spliceosomal tri-snRNP complex assembly | 12032085|20595234 |
| Tgene | SART3 | GO:0000387 | spliceosomal snRNP assembly | 14749385|15314151 |
| Tgene | SART3 | GO:0000398 | mRNA splicing, via spliceosome | 15314151 |
| Tgene | SART3 | GO:0006334 | nucleosome assembly | 24526689 |
| Tgene | SART3 | GO:0010468 | regulation of gene expression | 21447833 |
| Tgene | SART3 | GO:1903586 | positive regulation of histone deubiquitination | 24526689 |
 Fusion gene breakpoints across SSH1 (5'-gene) Fusion gene breakpoints across SSH1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
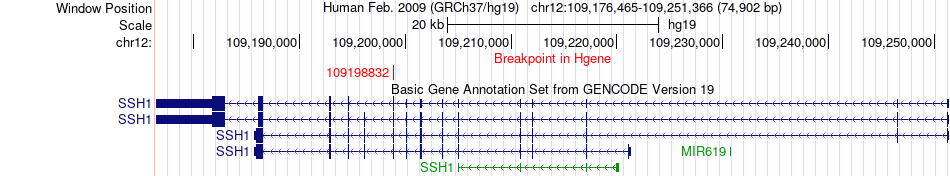 |
 Fusion gene breakpoints across SART3 (3'-gene) Fusion gene breakpoints across SART3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
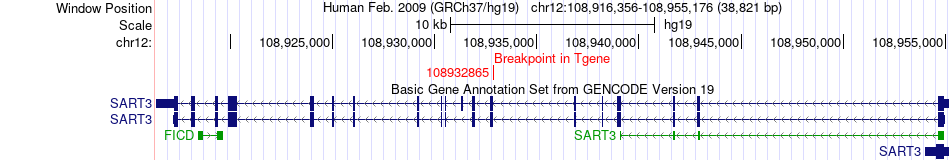 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SKCM | TCGA-EB-A1NK-01A | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
Top |
Fusion Gene ORF analysis for SSH1-SART3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000326470 | ENST00000546611 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000326470 | ENST00000552221 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000326495 | ENST00000546611 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000326495 | ENST00000552221 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000360239 | ENST00000546611 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000360239 | ENST00000552221 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000551165 | ENST00000546611 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| 5CDS-intron | ENST00000551165 | ENST00000552221 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| Frame-shift | ENST00000360239 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| Frame-shift | ENST00000360239 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| In-frame | ENST00000326470 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| In-frame | ENST00000326470 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| In-frame | ENST00000326495 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| In-frame | ENST00000326495 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| In-frame | ENST00000551165 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| In-frame | ENST00000551165 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| intron-3CDS | ENST00000546812 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| intron-3CDS | ENST00000546812 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| intron-intron | ENST00000546812 | ENST00000546611 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
| intron-intron | ENST00000546812 | ENST00000552221 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000326495 | SSH1 | chr12 | 109198832 | - | ENST00000228284 | SART3 | chr12 | 108932865 | - | 3911 | 1048 | 94 | 3033 | 979 |
| ENST00000326495 | SSH1 | chr12 | 109198832 | - | ENST00000431469 | SART3 | chr12 | 108932865 | - | 2971 | 1048 | 94 | 2925 | 943 |
| ENST00000551165 | SSH1 | chr12 | 109198832 | - | ENST00000228284 | SART3 | chr12 | 108932865 | - | 3911 | 1048 | 94 | 3033 | 979 |
| ENST00000551165 | SSH1 | chr12 | 109198832 | - | ENST00000431469 | SART3 | chr12 | 108932865 | - | 2971 | 1048 | 94 | 2925 | 943 |
| ENST00000326470 | SSH1 | chr12 | 109198832 | - | ENST00000228284 | SART3 | chr12 | 108932865 | - | 3996 | 1133 | 116 | 3118 | 1000 |
| ENST00000326470 | SSH1 | chr12 | 109198832 | - | ENST00000431469 | SART3 | chr12 | 108932865 | - | 3056 | 1133 | 116 | 3010 | 964 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000326495 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - | 0.003484471 | 0.9965155 |
| ENST00000326495 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - | 0.013241838 | 0.9867582 |
| ENST00000551165 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - | 0.003484471 | 0.9965155 |
| ENST00000551165 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - | 0.013241838 | 0.9867582 |
| ENST00000326470 | ENST00000228284 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - | 0.002944537 | 0.9970554 |
| ENST00000326470 | ENST00000431469 | SSH1 | chr12 | 109198832 | - | SART3 | chr12 | 108932865 | - | 0.009898527 | 0.99010146 |
Top |
Fusion Genomic Features for SSH1-SART3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
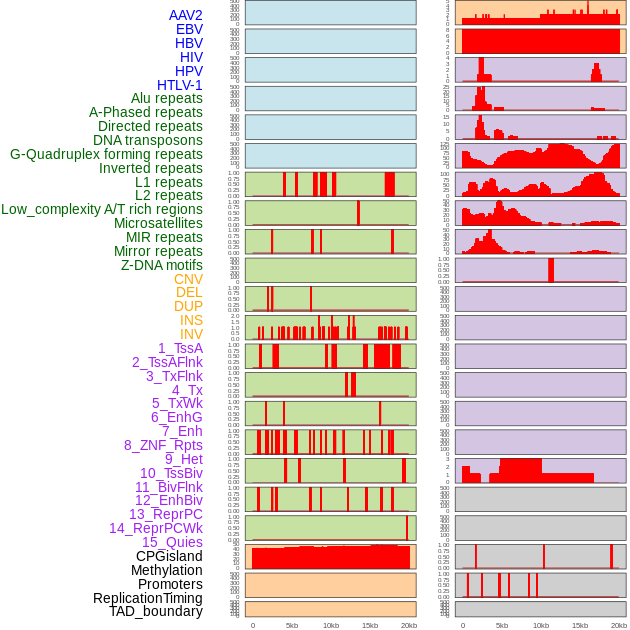 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for SSH1-SART3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:109198832/chr12:108932865) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 559_619 | 302 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 21_46 | 0 | 130.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 559_619 | 0 | 130.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 82_110 | 0 | 130.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 612_616 | 302 | 964.0 | Compositional bias | Note=Poly-Lys | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 612_616 | 0 | 130.0 | Compositional bias | Note=Poly-Lys | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 89_92 | 0 | 130.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 704_782 | 302 | 964.0 | Domain | RRM 1 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 801_878 | 302 | 964.0 | Domain | RRM 2 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 704_782 | 0 | 130.0 | Domain | RRM 1 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 801_878 | 0 | 130.0 | Domain | RRM 2 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 601_617 | 302 | 964.0 | Motif | Nuclear localization signal | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 601_617 | 0 | 130.0 | Motif | Nuclear localization signal | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 537_953 | 302 | 964.0 | Region | Necessary and sufficient for U6 snRNA binding | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 600_670 | 302 | 964.0 | Region | Required for nuclear localization | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 537_953 | 0 | 130.0 | Region | Necessary and sufficient for U6 snRNA binding | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 600_670 | 0 | 130.0 | Region | Required for nuclear localization | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 324_356 | 302 | 964.0 | Repeat | Note=HAT 5 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 359_391 | 302 | 964.0 | Repeat | Note=HAT 6 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 394_430 | 302 | 964.0 | Repeat | Note=HAT 7 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 126_158 | 0 | 130.0 | Repeat | Note=HAT 1 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 164_195 | 0 | 130.0 | Repeat | Note=HAT 2 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 201_237 | 0 | 130.0 | Repeat | Note=HAT 3 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 242_275 | 0 | 130.0 | Repeat | Note=HAT 4 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 324_356 | 0 | 130.0 | Repeat | Note=HAT 5 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 359_391 | 0 | 130.0 | Repeat | Note=HAT 6 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000546611 | 0 | 1 | 394_430 | 0 | 130.0 | Repeat | Note=HAT 7 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000326470 | - | 9 | 13 | 308_449 | 329 | 704.0 | Domain | Tyrosine-protein phosphatase |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000326495 | - | 10 | 15 | 308_449 | 318 | 1050.0 | Domain | Tyrosine-protein phosphatase |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000360239 | - | 10 | 14 | 308_449 | 21 | 738.0 | Domain | Tyrosine-protein phosphatase |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000551165 | - | 10 | 14 | 308_449 | 318 | 693.0 | Domain | Tyrosine-protein phosphatase |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 21_46 | 302 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 82_110 | 302 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 89_92 | 302 | 964.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 126_158 | 302 | 964.0 | Repeat | Note=HAT 1 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 164_195 | 302 | 964.0 | Repeat | Note=HAT 2 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 201_237 | 302 | 964.0 | Repeat | Note=HAT 3 | |
| Tgene | SART3 | chr12:109198832 | chr12:108932865 | ENST00000228284 | 5 | 19 | 242_275 | 302 | 964.0 | Repeat | Note=HAT 4 |
Top |
Fusion Gene Sequence for SSH1-SART3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >86759_86759_1_SSH1-SART3_SSH1_chr12_109198832_ENST00000326470_SART3_chr12_108932865_ENST00000228284_length(transcript)=3996nt_BP=1133nt TTTTTCAGCCAAAATATCAGAGGAAGACAAGAGTTCCACAAGAGCCATTTGCTTTTCGGGTAGGACTCTGAGGAGGAGTTGGCTCCCCAG CCCTCGTGGAAGCACTCGGCAGCCTCCTGGCTTTTCCTGATCTCCATAGGGAGCAGATGGCCAGAGCTCGGAGGGCAGTTGTAGGGAGCG TTAGAGATGTGAGCACTGCAGCCACGAACCTCTTTTATTTTACCGACTTCTGCATCTTCCTTCAGCCAACCCACTGCTTCTGCTGTCCAG AGGTTTCCAGCTCAAATTACTTAAGTGAGAGCTTTTTCATGGTGAAAGGCGCAGCCCTCTTCTTACAACAGGGAAGCAGCCCTCAAGGCC AGCGGAGTCTTCAGCACCCCCACAAGCATGCAGGTGATCTGCCTCAACATCTTCAGGTGATGATCAACCTTCTGCGTTGCGAAGACAGAA TCAAGCTGGCAGTGCGCCTGGAGAGCGCCTGGGCGGACCGGGTCCGGTACATGGTGGTGGTGTACAGCAGCGGGCGCCAGGACACCGAGG AGAATATCTTGCTGGGAGTGGACTTTTCCAGTAAGGAAAGTAAAAGCTGCACCATTGGGATGGTTCTCCGACTGTGGAGCGACACGAAAA TCCACCTTGATGGAGATGGTGGGTTCAGCGTGAGCACAGCAGGAAGGATGCACATATTTAAGCCTGTGTCTGTCCAGGCCATGTGGTCTG CCCTGCAGGTGCTTCACAAGGCCTGCGAAGTGGCCCGGAGGCACAACTACTTCCCCGGGGGTGTAGCTCTCATCTGGGCTACCTACTATG AGAGCTGCATCAGCTCCGAGCAGAGCTGCATCAACGAGTGGAACGCCATGCAGGACCTGGAGTCTACGCGGCCCGACTCCCCCGCGCTAT TTGTGGACAAGCCCACTGAAGGGGAAAGGACCGAGCGCCTCATCAAAGCCAAGCTCCGAAGCATCATGATGAGCCAGGATCTAGAAAATG TGACTTCCAAAGAGATTCGTAATGAATTAGAGAAACAGATGAATTGTAACTTGAAGGAACTCAAGGAATTTATAGACAATGAGATGCTAC TTATCTTGGGACAGATGGACAAGCCCTCCCTTATCTTCGATCATCTTTATCTCTTGCAGGCAGAGGCACCAAGGCTGGCAGAATATCAAG CATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGTCCCAG ACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAGAAACT GCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGTAACCTTCGAGA AAGCTTTGAATGCCGGCTTCATCCAGGCCACTGATTATGTGGAGATTTGGCAGGCATACCTTGATTACCTGAGGAGAAGGGTTGATTTCA AACAAGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGAGCGTTTCA ATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGCTCGGGAAC TCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGAGCTCATGGTGACACCC AGCACTGCCGGAAGGCTCTGCACCGGGCCGTCCAGTGCACCAGTGACTACCCAGAGCACGTCTGCGAAGTGTTACTCACCATGGAGAGGA CAGAAGGTTCTTTAGAAGATTGGGATATAGCTGTTCAGAAAACTGAAACCCGATTAGCTCGTGTCAATGAGCAGAGAATGAAGGCTGCAG AGAAGGAAGCAGCCCTTGTGCAGCAAGAAGAAGAAAAGGCTGAACAACGGAAAAGAGCTCGGGCTGAGAAGAAAGCGTTAAAAAAGAAGA AAAAGATCAGAGGCCCAGAGAAGCGCGGAGCAGATGAGGATGATGAGAAAGAGTGGGGCGATGATGAAGAAGAGCAGCCTTCCAAACGCA GAAGGGTCGAGAACAGCATCCCTGCAGCTGGAGAAACACAAAATGTAGAAGTAGCAGCAGGGCCCGCTGGGAAATGTGCTGCCGTAGATG TGGAGCCCCCTTCGAAGCAGAAGGAGAAGGCAGCCTCCCTGAAGAGGGACATGCCCAAGGTGCTGCACGACAGCAGCAAGGACAGCATCA CCGTCTTTGTCAGCAACCTGCCCTACAGCATGCAGGAGCCGGACACGAAGCTCAGGCCACTCTTCGAGGCCTGTGGGGAGGTGGTCCAGA TCCGACCCATCTTCAGCAACCGTGGGGATTTCCGAGGTTACTGCTACGTGGAGTTTAAAGAAGAGAAATCAGCCCTTCAGGCACTGGAGA TGGACCGGAAAAGTGTAGAAGGGAGGCCAATGTTTGTTTCCCCCTGTGTGGATAAGAGCAAAAACCCCGATTTTAAGGTGTTCAGGTACA GCACTTCCCTAGAGAAACACAAGCTGTTCATCTCAGGCCTGCCTTTCTCCTGTACTAAAGAGGAACTAGAAGAAATCTGTAAGGCTCATG GCACCGTGAAGGACCTCAGGCTGGTCACCAACCGGGCTGGCAAACCAAAGGGCCTGGCCTACGTGGAGTATGAAAATGAATCCCAGGCGT CGCAGGCTGTGATGAAGATGGACGGCATGACTATCAAAGAGAACATCATCAAAGTGGCAATCAGCAACCCTCCTCAGAGGAAAGTTCCAG AGAAGCCGGAGACCAGGAAGGCACCAGGTGGCCCCATGCTTTTGCCGCAGACATACGGAGCGAGGGGGAAGGGAAGGACGCAGCTGTCTC TACTGCCTCGTGCCCTGCAGCGCCCAAGTGCTGCAGCTCCTCAGGCTGAGAACGGCCCTGCCGCGGCTCCTGCAGTTGCCGCCCCAGCAG CCACCGAGGCACCCAAGATGTCCAATGCCGATTTTGCCAAGCTGTTTCTGAGAAAGTGAACGGGACGCTGGGAGACAGGAAATGCCTTAC TTCACTCTGGCCCGGCGGACCTCCCACCACCCAGCAGTGCACTGGGGATGGACAGGCCTGGTGTGCTGCGTGCTCGCAACCACAGATGGC TCCTCGGCTTTAGACAGAAAGGGGAAGGGGTTCTAAGTCAAGAGCCTTTCAGTGCTCCCTCATATTGAGGGCAGTGGCAGAAAAGTGACC ACTCTGCAGGCTGGGCCCAGGATGTGGTGTCCTGAGATAGTTTTGTATCTTAAAGACTGAGGCACAGAAGCGAAACGAGAACACACTGTT TTTGAGACACAGTTGTCCAAATGTTTCTGGCCAGCTCCGGCCCCTTTTTGTATGACACTTCTCTTCCACCCTGCACAGCACATGTGCCCG TCATTCTTTTAATTTTAAAAGATGAAATGGCAGATGCTAGTAATTCACAGAATGGCCTCTTGTGGGGGTGGGTCTGAGGGAAGTCAGCTA TAAAACATTTGCTGGAGTTTTGTTCAATGGGGCTGTGCATTTTTATATTATGTGTTTGTAAATGACATGTCAGCCATTGTTTCATGTTTC CTAAAAGCAGAATATTTGCAACATTTGTTTTGTATAGGAATTATTTGTGCCACCTGCTGTGGACTGTTTTCTTTGCCTAGTGACTAGTGA CCTGTGTTGTCTAAACATGAGTTTCAGCCCTTTGGTTTTGTTTAATACCATGTCAAATGCAAACTTCAATTCTCCCCATTTAGCTTTATT AAACTGACGTTCTCTTCAAAACTTCTTGCTGAATGGTACTCAGATGTGCATTCACATACAGATGTGTTTTGAAGTGGGTGTACCTTGCTT >86759_86759_1_SSH1-SART3_SSH1_chr12_109198832_ENST00000326470_SART3_chr12_108932865_ENST00000228284_length(amino acids)=1000AA_BP=1 MAFPDLHREQMARARRAVVGSVRDVSTAATNLFYFTDFCIFLQPTHCFCCPEVSSSNYLSESFFMVKGAALFLQQGSSPQGQRSLQHPHK HAGDLPQHLQVMINLLRCEDRIKLAVRLESAWADRVRYMVVVYSSGRQDTEENILLGVDFSSKESKSCTIGMVLRLWSDTKIHLDGDGGF SVSTAGRMHIFKPVSVQAMWSALQVLHKACEVARRHNYFPGGVALIWATYYESCISSEQSCINEWNAMQDLESTRPDSPALFVDKPTEGE RTERLIKAKLRSIMMSQDLENVTSKEIRNELEKQMNCNLKELKEFIDNEMLLILGQMDKPSLIFDHLYLLQAEAPRLAEYQAYIDFEMKI GDPARIQLIFERALVENCLVPDLWIRYSQYLDRQLKVKDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISVTFEKALNAGFIQ ATDYVEIWQAYLDYLRRRVDFKQDSSKELEELRAAFTRALEYLKQEVEERFNESGDPSCVIMQNWARIEARLCNNMQKARELWDSIMTRG NAKYANMWLEYYNLERAHGDTQHCRKALHRAVQCTSDYPEHVCEVLLTMERTEGSLEDWDIAVQKTETRLARVNEQRMKAAEKEAALVQQ EEEKAEQRKRARAEKKALKKKKKIRGPEKRGADEDDEKEWGDDEEEQPSKRRRVENSIPAAGETQNVEVAAGPAGKCAAVDVEPPSKQKE KAASLKRDMPKVLHDSSKDSITVFVSNLPYSMQEPDTKLRPLFEACGEVVQIRPIFSNRGDFRGYCYVEFKEEKSALQALEMDRKSVEGR PMFVSPCVDKSKNPDFKVFRYSTSLEKHKLFISGLPFSCTKEELEEICKAHGTVKDLRLVTNRAGKPKGLAYVEYENESQASQAVMKMDG MTIKENIIKVAISNPPQRKVPEKPETRKAPGGPMLLPQTYGARGKGRTQLSLLPRALQRPSAAAPQAENGPAAAPAVAAPAATEAPKMSN -------------------------------------------------------------- >86759_86759_2_SSH1-SART3_SSH1_chr12_109198832_ENST00000326470_SART3_chr12_108932865_ENST00000431469_length(transcript)=3056nt_BP=1133nt TTTTTCAGCCAAAATATCAGAGGAAGACAAGAGTTCCACAAGAGCCATTTGCTTTTCGGGTAGGACTCTGAGGAGGAGTTGGCTCCCCAG CCCTCGTGGAAGCACTCGGCAGCCTCCTGGCTTTTCCTGATCTCCATAGGGAGCAGATGGCCAGAGCTCGGAGGGCAGTTGTAGGGAGCG TTAGAGATGTGAGCACTGCAGCCACGAACCTCTTTTATTTTACCGACTTCTGCATCTTCCTTCAGCCAACCCACTGCTTCTGCTGTCCAG AGGTTTCCAGCTCAAATTACTTAAGTGAGAGCTTTTTCATGGTGAAAGGCGCAGCCCTCTTCTTACAACAGGGAAGCAGCCCTCAAGGCC AGCGGAGTCTTCAGCACCCCCACAAGCATGCAGGTGATCTGCCTCAACATCTTCAGGTGATGATCAACCTTCTGCGTTGCGAAGACAGAA TCAAGCTGGCAGTGCGCCTGGAGAGCGCCTGGGCGGACCGGGTCCGGTACATGGTGGTGGTGTACAGCAGCGGGCGCCAGGACACCGAGG AGAATATCTTGCTGGGAGTGGACTTTTCCAGTAAGGAAAGTAAAAGCTGCACCATTGGGATGGTTCTCCGACTGTGGAGCGACACGAAAA TCCACCTTGATGGAGATGGTGGGTTCAGCGTGAGCACAGCAGGAAGGATGCACATATTTAAGCCTGTGTCTGTCCAGGCCATGTGGTCTG CCCTGCAGGTGCTTCACAAGGCCTGCGAAGTGGCCCGGAGGCACAACTACTTCCCCGGGGGTGTAGCTCTCATCTGGGCTACCTACTATG AGAGCTGCATCAGCTCCGAGCAGAGCTGCATCAACGAGTGGAACGCCATGCAGGACCTGGAGTCTACGCGGCCCGACTCCCCCGCGCTAT TTGTGGACAAGCCCACTGAAGGGGAAAGGACCGAGCGCCTCATCAAAGCCAAGCTCCGAAGCATCATGATGAGCCAGGATCTAGAAAATG TGACTTCCAAAGAGATTCGTAATGAATTAGAGAAACAGATGAATTGTAACTTGAAGGAACTCAAGGAATTTATAGACAATGAGATGCTAC TTATCTTGGGACAGATGGACAAGCCCTCCCTTATCTTCGATCATCTTTATCTCTTGCAGGCAGAGGCACCAAGGCTGGCAGAATATCAAG CATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGTCCCAG ACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAGAAACT GCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGACTCCAGTAAAG AGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGAGCGTTTCAATGAGAGTGGTGATCCAA GCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGCTCGGGAACTCTGGGATAGCATCATGA CCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGAGCTCATGGTGACACCCAGCACTGCCGGAAGGCTC TGCACCGGGCCGTCCAGTGCACCAGTGACTACCCAGAGCACGTCTGCGAAGTGTTACTCACCATGGAGAGGACAGAAGGTTCTTTAGAAG ATTGGGATATAGCTGTTCAGAAAACTGAAACCCGATTAGCTCGTGTCAATGAGCAGAGAATGAAGGCTGCAGAGAAGGAAGCAGCCCTTG TGCAGCAAGAAGAAGAAAAGGCTGAACAACGGAAAAGAGCTCGGGCTGAGAAGAAAGCGTTAAAAAAGAAGAAAAAGATCAGAGGCCCAG AGAAGCGCGGAGCAGATGAGGATGATGAGAAAGAGTGGGGCGATGATGAAGAAGAGCAGCCTTCCAAACGCAGAAGGGTCGAGAACAGCA TCCCTGCAGCTGGAGAAACACAAAATGTAGAAGTAGCAGCAGGGCCCGCTGGGAAATGTGCTGCCGTAGATGTGGAGCCCCCTTCGAAGC AGAAGGAGAAGGCAGCCTCCCTGAAGAGGGACATGCCCAAGGTGCTGCACGACAGCAGCAAGGACAGCATCACCGTCTTTGTCAGCAACC TGCCCTACAGCATGCAGGAGCCGGACACGAAGCTCAGGCCACTCTTCGAGGCCTGTGGGGAGGTGGTCCAGATCCGACCCATCTTCAGCA ACCGTGGGGATTTCCGAGGTTACTGCTACGTGGAGTTTAAAGAAGAGAAATCAGCCCTTCAGGCACTGGAGATGGACCGGAAAAGTGTAG AAGGGAGGCCAATGTTTGTTTCCCCCTGTGTGGATAAGAGCAAAAACCCCGATTTTAAGGTGTTCAGGTACAGCACTTCCCTAGAGAAAC ACAAGCTGTTCATCTCAGGCCTGCCTTTCTCCTGTACTAAAGAGGAACTAGAAGAAATCTGTAAGGCTCATGGCACCGTGAAGGACCTCA GGCTGGTCACCAACCGGGCTGGCAAACCAAAGGGCCTGGCCTACGTGGAGTATGAAAATGAATCCCAGGCGTCGCAGGCTGTGATGAAGA TGGACGGCATGACTATCAAAGAGAACATCATCAAAGTGGCAATCAGCAACCCTCCTCAGAGGAAAGTTCCAGAGAAGCCGGAGACCAGGA AGGCACCAGGTGGCCCCATGCTTTTGCCGCAGACATACGGAGCGAGGGGGAAGGGAAGGACGCAGCTGTCTCTACTGCCTCGTGCCCTGC AGCGCCCAAGTGCTGCAGCTCCTCAGGCTGAGAACGGCCCTGCCGCGGCTCCTGCAGTTGCCGCCCCAGCAGCCACCGAGGCACCCAAGA >86759_86759_2_SSH1-SART3_SSH1_chr12_109198832_ENST00000326470_SART3_chr12_108932865_ENST00000431469_length(amino acids)=964AA_BP=1 MAFPDLHREQMARARRAVVGSVRDVSTAATNLFYFTDFCIFLQPTHCFCCPEVSSSNYLSESFFMVKGAALFLQQGSSPQGQRSLQHPHK HAGDLPQHLQVMINLLRCEDRIKLAVRLESAWADRVRYMVVVYSSGRQDTEENILLGVDFSSKESKSCTIGMVLRLWSDTKIHLDGDGGF SVSTAGRMHIFKPVSVQAMWSALQVLHKACEVARRHNYFPGGVALIWATYYESCISSEQSCINEWNAMQDLESTRPDSPALFVDKPTEGE RTERLIKAKLRSIMMSQDLENVTSKEIRNELEKQMNCNLKELKEFIDNEMLLILGQMDKPSLIFDHLYLLQAEAPRLAEYQAYIDFEMKI GDPARIQLIFERALVENCLVPDLWIRYSQYLDRQLKVKDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISDSSKELEELRAAF TRALEYLKQEVEERFNESGDPSCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERAHGDTQHCRKALHRAVQCTS DYPEHVCEVLLTMERTEGSLEDWDIAVQKTETRLARVNEQRMKAAEKEAALVQQEEEKAEQRKRARAEKKALKKKKKIRGPEKRGADEDD EKEWGDDEEEQPSKRRRVENSIPAAGETQNVEVAAGPAGKCAAVDVEPPSKQKEKAASLKRDMPKVLHDSSKDSITVFVSNLPYSMQEPD TKLRPLFEACGEVVQIRPIFSNRGDFRGYCYVEFKEEKSALQALEMDRKSVEGRPMFVSPCVDKSKNPDFKVFRYSTSLEKHKLFISGLP FSCTKEELEEICKAHGTVKDLRLVTNRAGKPKGLAYVEYENESQASQAVMKMDGMTIKENIIKVAISNPPQRKVPEKPETRKAPGGPMLL -------------------------------------------------------------- >86759_86759_3_SSH1-SART3_SSH1_chr12_109198832_ENST00000326495_SART3_chr12_108932865_ENST00000228284_length(transcript)=3911nt_BP=1048nt CGGCTCCAGGAGGCCCCGGGCCCGGCCCGGGCGGCGGTGGCGGTGGCGGCTCTAGCTCGAGACGTCTGTGGCGCCCTCGCACCGCGCCGC AGCCATGGCCCTGGTGACCCTGCAGCGCTCGCCCACGCCCAGCGCCGCCTCCTCCTCGGCCAGCAACAGCGAGTTGGAGGCTGGCAGCGA AGAAGATCGAAAATTAAACCTCAGCTTAAGTGAGAGCTTTTTCATGGTGAAAGGCGCAGCCCTCTTCTTACAACAGGGAAGCAGCCCTCA AGGCCAGCGGAGTCTTCAGCACCCCCACAAGCATGCAGGTGATCTGCCTCAACATCTTCAGGTGATGATCAACCTTCTGCGTTGCGAAGA CAGAATCAAGCTGGCAGTGCGCCTGGAGAGCGCCTGGGCGGACCGGGTCCGGTACATGGTGGTGGTGTACAGCAGCGGGCGCCAGGACAC CGAGGAGAATATCTTGCTGGGAGTGGACTTTTCCAGTAAGGAAAGTAAAAGCTGCACCATTGGGATGGTTCTCCGACTGTGGAGCGACAC GAAAATCCACCTTGATGGAGATGGTGGGTTCAGCGTGAGCACAGCAGGAAGGATGCACATATTTAAGCCTGTGTCTGTCCAGGCCATGTG GTCTGCCCTGCAGGTGCTTCACAAGGCCTGCGAAGTGGCCCGGAGGCACAACTACTTCCCCGGGGGTGTAGCTCTCATCTGGGCTACCTA CTATGAGAGCTGCATCAGCTCCGAGCAGAGCTGCATCAACGAGTGGAACGCCATGCAGGACCTGGAGTCTACGCGGCCCGACTCCCCCGC GCTATTTGTGGACAAGCCCACTGAAGGGGAAAGGACCGAGCGCCTCATCAAAGCCAAGCTCCGAAGCATCATGATGAGCCAGGATCTAGA AAATGTGACTTCCAAAGAGATTCGTAATGAATTAGAGAAACAGATGAATTGTAACTTGAAGGAACTCAAGGAATTTATAGACAATGAGAT GCTACTTATCTTGGGACAGATGGACAAGCCCTCCCTTATCTTCGATCATCTTTATCTCTTGCAGGCAGAGGCACCAAGGCTGGCAGAATA TCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGT CCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAG AAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGTAACCTT CGAGAAAGCTTTGAATGCCGGCTTCATCCAGGCCACTGATTATGTGGAGATTTGGCAGGCATACCTTGATTACCTGAGGAGAAGGGTTGA TTTCAAACAAGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGAGCG TTTCAATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGCTCG GGAACTCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGAGCTCATGGTGA CACCCAGCACTGCCGGAAGGCTCTGCACCGGGCCGTCCAGTGCACCAGTGACTACCCAGAGCACGTCTGCGAAGTGTTACTCACCATGGA GAGGACAGAAGGTTCTTTAGAAGATTGGGATATAGCTGTTCAGAAAACTGAAACCCGATTAGCTCGTGTCAATGAGCAGAGAATGAAGGC TGCAGAGAAGGAAGCAGCCCTTGTGCAGCAAGAAGAAGAAAAGGCTGAACAACGGAAAAGAGCTCGGGCTGAGAAGAAAGCGTTAAAAAA GAAGAAAAAGATCAGAGGCCCAGAGAAGCGCGGAGCAGATGAGGATGATGAGAAAGAGTGGGGCGATGATGAAGAAGAGCAGCCTTCCAA ACGCAGAAGGGTCGAGAACAGCATCCCTGCAGCTGGAGAAACACAAAATGTAGAAGTAGCAGCAGGGCCCGCTGGGAAATGTGCTGCCGT AGATGTGGAGCCCCCTTCGAAGCAGAAGGAGAAGGCAGCCTCCCTGAAGAGGGACATGCCCAAGGTGCTGCACGACAGCAGCAAGGACAG CATCACCGTCTTTGTCAGCAACCTGCCCTACAGCATGCAGGAGCCGGACACGAAGCTCAGGCCACTCTTCGAGGCCTGTGGGGAGGTGGT CCAGATCCGACCCATCTTCAGCAACCGTGGGGATTTCCGAGGTTACTGCTACGTGGAGTTTAAAGAAGAGAAATCAGCCCTTCAGGCACT GGAGATGGACCGGAAAAGTGTAGAAGGGAGGCCAATGTTTGTTTCCCCCTGTGTGGATAAGAGCAAAAACCCCGATTTTAAGGTGTTCAG GTACAGCACTTCCCTAGAGAAACACAAGCTGTTCATCTCAGGCCTGCCTTTCTCCTGTACTAAAGAGGAACTAGAAGAAATCTGTAAGGC TCATGGCACCGTGAAGGACCTCAGGCTGGTCACCAACCGGGCTGGCAAACCAAAGGGCCTGGCCTACGTGGAGTATGAAAATGAATCCCA GGCGTCGCAGGCTGTGATGAAGATGGACGGCATGACTATCAAAGAGAACATCATCAAAGTGGCAATCAGCAACCCTCCTCAGAGGAAAGT TCCAGAGAAGCCGGAGACCAGGAAGGCACCAGGTGGCCCCATGCTTTTGCCGCAGACATACGGAGCGAGGGGGAAGGGAAGGACGCAGCT GTCTCTACTGCCTCGTGCCCTGCAGCGCCCAAGTGCTGCAGCTCCTCAGGCTGAGAACGGCCCTGCCGCGGCTCCTGCAGTTGCCGCCCC AGCAGCCACCGAGGCACCCAAGATGTCCAATGCCGATTTTGCCAAGCTGTTTCTGAGAAAGTGAACGGGACGCTGGGAGACAGGAAATGC CTTACTTCACTCTGGCCCGGCGGACCTCCCACCACCCAGCAGTGCACTGGGGATGGACAGGCCTGGTGTGCTGCGTGCTCGCAACCACAG ATGGCTCCTCGGCTTTAGACAGAAAGGGGAAGGGGTTCTAAGTCAAGAGCCTTTCAGTGCTCCCTCATATTGAGGGCAGTGGCAGAAAAG TGACCACTCTGCAGGCTGGGCCCAGGATGTGGTGTCCTGAGATAGTTTTGTATCTTAAAGACTGAGGCACAGAAGCGAAACGAGAACACA CTGTTTTTGAGACACAGTTGTCCAAATGTTTCTGGCCAGCTCCGGCCCCTTTTTGTATGACACTTCTCTTCCACCCTGCACAGCACATGT GCCCGTCATTCTTTTAATTTTAAAAGATGAAATGGCAGATGCTAGTAATTCACAGAATGGCCTCTTGTGGGGGTGGGTCTGAGGGAAGTC AGCTATAAAACATTTGCTGGAGTTTTGTTCAATGGGGCTGTGCATTTTTATATTATGTGTTTGTAAATGACATGTCAGCCATTGTTTCAT GTTTCCTAAAAGCAGAATATTTGCAACATTTGTTTTGTATAGGAATTATTTGTGCCACCTGCTGTGGACTGTTTTCTTTGCCTAGTGACT AGTGACCTGTGTTGTCTAAACATGAGTTTCAGCCCTTTGGTTTTGTTTAATACCATGTCAAATGCAAACTTCAATTCTCCCCATTTAGCT TTATTAAACTGACGTTCTCTTCAAAACTTCTTGCTGAATGGTACTCAGATGTGCATTCACATACAGATGTGTTTTGAAGTGGGTGTACCT >86759_86759_3_SSH1-SART3_SSH1_chr12_109198832_ENST00000326495_SART3_chr12_108932865_ENST00000228284_length(amino acids)=979AA_BP=0 MALVTLQRSPTPSAASSSASNSELEAGSEEDRKLNLSLSESFFMVKGAALFLQQGSSPQGQRSLQHPHKHAGDLPQHLQVMINLLRCEDR IKLAVRLESAWADRVRYMVVVYSSGRQDTEENILLGVDFSSKESKSCTIGMVLRLWSDTKIHLDGDGGFSVSTAGRMHIFKPVSVQAMWS ALQVLHKACEVARRHNYFPGGVALIWATYYESCISSEQSCINEWNAMQDLESTRPDSPALFVDKPTEGERTERLIKAKLRSIMMSQDLEN VTSKEIRNELEKQMNCNLKELKEFIDNEMLLILGQMDKPSLIFDHLYLLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVP DLWIRYSQYLDRQLKVKDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISVTFEKALNAGFIQATDYVEIWQAYLDYLRRRVDF KQDSSKELEELRAAFTRALEYLKQEVEERFNESGDPSCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERAHGDT QHCRKALHRAVQCTSDYPEHVCEVLLTMERTEGSLEDWDIAVQKTETRLARVNEQRMKAAEKEAALVQQEEEKAEQRKRARAEKKALKKK KKIRGPEKRGADEDDEKEWGDDEEEQPSKRRRVENSIPAAGETQNVEVAAGPAGKCAAVDVEPPSKQKEKAASLKRDMPKVLHDSSKDSI TVFVSNLPYSMQEPDTKLRPLFEACGEVVQIRPIFSNRGDFRGYCYVEFKEEKSALQALEMDRKSVEGRPMFVSPCVDKSKNPDFKVFRY STSLEKHKLFISGLPFSCTKEELEEICKAHGTVKDLRLVTNRAGKPKGLAYVEYENESQASQAVMKMDGMTIKENIIKVAISNPPQRKVP -------------------------------------------------------------- >86759_86759_4_SSH1-SART3_SSH1_chr12_109198832_ENST00000326495_SART3_chr12_108932865_ENST00000431469_length(transcript)=2971nt_BP=1048nt CGGCTCCAGGAGGCCCCGGGCCCGGCCCGGGCGGCGGTGGCGGTGGCGGCTCTAGCTCGAGACGTCTGTGGCGCCCTCGCACCGCGCCGC AGCCATGGCCCTGGTGACCCTGCAGCGCTCGCCCACGCCCAGCGCCGCCTCCTCCTCGGCCAGCAACAGCGAGTTGGAGGCTGGCAGCGA AGAAGATCGAAAATTAAACCTCAGCTTAAGTGAGAGCTTTTTCATGGTGAAAGGCGCAGCCCTCTTCTTACAACAGGGAAGCAGCCCTCA AGGCCAGCGGAGTCTTCAGCACCCCCACAAGCATGCAGGTGATCTGCCTCAACATCTTCAGGTGATGATCAACCTTCTGCGTTGCGAAGA CAGAATCAAGCTGGCAGTGCGCCTGGAGAGCGCCTGGGCGGACCGGGTCCGGTACATGGTGGTGGTGTACAGCAGCGGGCGCCAGGACAC CGAGGAGAATATCTTGCTGGGAGTGGACTTTTCCAGTAAGGAAAGTAAAAGCTGCACCATTGGGATGGTTCTCCGACTGTGGAGCGACAC GAAAATCCACCTTGATGGAGATGGTGGGTTCAGCGTGAGCACAGCAGGAAGGATGCACATATTTAAGCCTGTGTCTGTCCAGGCCATGTG GTCTGCCCTGCAGGTGCTTCACAAGGCCTGCGAAGTGGCCCGGAGGCACAACTACTTCCCCGGGGGTGTAGCTCTCATCTGGGCTACCTA CTATGAGAGCTGCATCAGCTCCGAGCAGAGCTGCATCAACGAGTGGAACGCCATGCAGGACCTGGAGTCTACGCGGCCCGACTCCCCCGC GCTATTTGTGGACAAGCCCACTGAAGGGGAAAGGACCGAGCGCCTCATCAAAGCCAAGCTCCGAAGCATCATGATGAGCCAGGATCTAGA AAATGTGACTTCCAAAGAGATTCGTAATGAATTAGAGAAACAGATGAATTGTAACTTGAAGGAACTCAAGGAATTTATAGACAATGAGAT GCTACTTATCTTGGGACAGATGGACAAGCCCTCCCTTATCTTCGATCATCTTTATCTCTTGCAGGCAGAGGCACCAAGGCTGGCAGAATA TCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGT CCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAG AAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGACTCCAG TAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGAGCGTTTCAATGAGAGTGGTGA TCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGCTCGGGAACTCTGGGATAGCAT CATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGAGCTCATGGTGACACCCAGCACTGCCGGAA GGCTCTGCACCGGGCCGTCCAGTGCACCAGTGACTACCCAGAGCACGTCTGCGAAGTGTTACTCACCATGGAGAGGACAGAAGGTTCTTT AGAAGATTGGGATATAGCTGTTCAGAAAACTGAAACCCGATTAGCTCGTGTCAATGAGCAGAGAATGAAGGCTGCAGAGAAGGAAGCAGC CCTTGTGCAGCAAGAAGAAGAAAAGGCTGAACAACGGAAAAGAGCTCGGGCTGAGAAGAAAGCGTTAAAAAAGAAGAAAAAGATCAGAGG CCCAGAGAAGCGCGGAGCAGATGAGGATGATGAGAAAGAGTGGGGCGATGATGAAGAAGAGCAGCCTTCCAAACGCAGAAGGGTCGAGAA CAGCATCCCTGCAGCTGGAGAAACACAAAATGTAGAAGTAGCAGCAGGGCCCGCTGGGAAATGTGCTGCCGTAGATGTGGAGCCCCCTTC GAAGCAGAAGGAGAAGGCAGCCTCCCTGAAGAGGGACATGCCCAAGGTGCTGCACGACAGCAGCAAGGACAGCATCACCGTCTTTGTCAG CAACCTGCCCTACAGCATGCAGGAGCCGGACACGAAGCTCAGGCCACTCTTCGAGGCCTGTGGGGAGGTGGTCCAGATCCGACCCATCTT CAGCAACCGTGGGGATTTCCGAGGTTACTGCTACGTGGAGTTTAAAGAAGAGAAATCAGCCCTTCAGGCACTGGAGATGGACCGGAAAAG TGTAGAAGGGAGGCCAATGTTTGTTTCCCCCTGTGTGGATAAGAGCAAAAACCCCGATTTTAAGGTGTTCAGGTACAGCACTTCCCTAGA GAAACACAAGCTGTTCATCTCAGGCCTGCCTTTCTCCTGTACTAAAGAGGAACTAGAAGAAATCTGTAAGGCTCATGGCACCGTGAAGGA CCTCAGGCTGGTCACCAACCGGGCTGGCAAACCAAAGGGCCTGGCCTACGTGGAGTATGAAAATGAATCCCAGGCGTCGCAGGCTGTGAT GAAGATGGACGGCATGACTATCAAAGAGAACATCATCAAAGTGGCAATCAGCAACCCTCCTCAGAGGAAAGTTCCAGAGAAGCCGGAGAC CAGGAAGGCACCAGGTGGCCCCATGCTTTTGCCGCAGACATACGGAGCGAGGGGGAAGGGAAGGACGCAGCTGTCTCTACTGCCTCGTGC CCTGCAGCGCCCAAGTGCTGCAGCTCCTCAGGCTGAGAACGGCCCTGCCGCGGCTCCTGCAGTTGCCGCCCCAGCAGCCACCGAGGCACC CAAGATGTCCAATGCCGATTTTGCCAAGCTGTTTCTGAGAAAGTGAACGGGACGCTGGGAGACAGGAAATGCCTTACTTCACTCTGGCCC >86759_86759_4_SSH1-SART3_SSH1_chr12_109198832_ENST00000326495_SART3_chr12_108932865_ENST00000431469_length(amino acids)=943AA_BP=0 MALVTLQRSPTPSAASSSASNSELEAGSEEDRKLNLSLSESFFMVKGAALFLQQGSSPQGQRSLQHPHKHAGDLPQHLQVMINLLRCEDR IKLAVRLESAWADRVRYMVVVYSSGRQDTEENILLGVDFSSKESKSCTIGMVLRLWSDTKIHLDGDGGFSVSTAGRMHIFKPVSVQAMWS ALQVLHKACEVARRHNYFPGGVALIWATYYESCISSEQSCINEWNAMQDLESTRPDSPALFVDKPTEGERTERLIKAKLRSIMMSQDLEN VTSKEIRNELEKQMNCNLKELKEFIDNEMLLILGQMDKPSLIFDHLYLLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVP DLWIRYSQYLDRQLKVKDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISDSSKELEELRAAFTRALEYLKQEVEERFNESGDP SCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERAHGDTQHCRKALHRAVQCTSDYPEHVCEVLLTMERTEGSLE DWDIAVQKTETRLARVNEQRMKAAEKEAALVQQEEEKAEQRKRARAEKKALKKKKKIRGPEKRGADEDDEKEWGDDEEEQPSKRRRVENS IPAAGETQNVEVAAGPAGKCAAVDVEPPSKQKEKAASLKRDMPKVLHDSSKDSITVFVSNLPYSMQEPDTKLRPLFEACGEVVQIRPIFS NRGDFRGYCYVEFKEEKSALQALEMDRKSVEGRPMFVSPCVDKSKNPDFKVFRYSTSLEKHKLFISGLPFSCTKEELEEICKAHGTVKDL RLVTNRAGKPKGLAYVEYENESQASQAVMKMDGMTIKENIIKVAISNPPQRKVPEKPETRKAPGGPMLLPQTYGARGKGRTQLSLLPRAL -------------------------------------------------------------- >86759_86759_5_SSH1-SART3_SSH1_chr12_109198832_ENST00000551165_SART3_chr12_108932865_ENST00000228284_length(transcript)=3911nt_BP=1048nt CGGCTCCAGGAGGCCCCGGGCCCGGCCCGGGCGGCGGTGGCGGTGGCGGCTCTAGCTCGAGACGTCTGTGGCGCCCTCGCACCGCGCCGC AGCCATGGCCCTGGTGACCCTGCAGCGCTCGCCCACGCCCAGCGCCGCCTCCTCCTCGGCCAGCAACAGCGAGTTGGAGGCTGGCAGCGA AGAAGATCGAAAATTAAACCTCAGCTTAAGTGAGAGCTTTTTCATGGTGAAAGGCGCAGCCCTCTTCTTACAACAGGGAAGCAGCCCTCA AGGCCAGCGGAGTCTTCAGCACCCCCACAAGCATGCAGGTGATCTGCCTCAACATCTTCAGGTGATGATCAACCTTCTGCGTTGCGAAGA CAGAATCAAGCTGGCAGTGCGCCTGGAGAGCGCCTGGGCGGACCGGGTCCGGTACATGGTGGTGGTGTACAGCAGCGGGCGCCAGGACAC CGAGGAGAATATCTTGCTGGGAGTGGACTTTTCCAGTAAGGAAAGTAAAAGCTGCACCATTGGGATGGTTCTCCGACTGTGGAGCGACAC GAAAATCCACCTTGATGGAGATGGTGGGTTCAGCGTGAGCACAGCAGGAAGGATGCACATATTTAAGCCTGTGTCTGTCCAGGCCATGTG GTCTGCCCTGCAGGTGCTTCACAAGGCCTGCGAAGTGGCCCGGAGGCACAACTACTTCCCCGGGGGTGTAGCTCTCATCTGGGCTACCTA CTATGAGAGCTGCATCAGCTCCGAGCAGAGCTGCATCAACGAGTGGAACGCCATGCAGGACCTGGAGTCTACGCGGCCCGACTCCCCCGC GCTATTTGTGGACAAGCCCACTGAAGGGGAAAGGACCGAGCGCCTCATCAAAGCCAAGCTCCGAAGCATCATGATGAGCCAGGATCTAGA AAATGTGACTTCCAAAGAGATTCGTAATGAATTAGAGAAACAGATGAATTGTAACTTGAAGGAACTCAAGGAATTTATAGACAATGAGAT GCTACTTATCTTGGGACAGATGGACAAGCCCTCCCTTATCTTCGATCATCTTTATCTCTTGCAGGCAGAGGCACCAAGGCTGGCAGAATA TCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGT CCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAG AAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGTAACCTT CGAGAAAGCTTTGAATGCCGGCTTCATCCAGGCCACTGATTATGTGGAGATTTGGCAGGCATACCTTGATTACCTGAGGAGAAGGGTTGA TTTCAAACAAGACTCCAGTAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGAGCG TTTCAATGAGAGTGGTGATCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGCTCG GGAACTCTGGGATAGCATCATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGAGCTCATGGTGA CACCCAGCACTGCCGGAAGGCTCTGCACCGGGCCGTCCAGTGCACCAGTGACTACCCAGAGCACGTCTGCGAAGTGTTACTCACCATGGA GAGGACAGAAGGTTCTTTAGAAGATTGGGATATAGCTGTTCAGAAAACTGAAACCCGATTAGCTCGTGTCAATGAGCAGAGAATGAAGGC TGCAGAGAAGGAAGCAGCCCTTGTGCAGCAAGAAGAAGAAAAGGCTGAACAACGGAAAAGAGCTCGGGCTGAGAAGAAAGCGTTAAAAAA GAAGAAAAAGATCAGAGGCCCAGAGAAGCGCGGAGCAGATGAGGATGATGAGAAAGAGTGGGGCGATGATGAAGAAGAGCAGCCTTCCAA ACGCAGAAGGGTCGAGAACAGCATCCCTGCAGCTGGAGAAACACAAAATGTAGAAGTAGCAGCAGGGCCCGCTGGGAAATGTGCTGCCGT AGATGTGGAGCCCCCTTCGAAGCAGAAGGAGAAGGCAGCCTCCCTGAAGAGGGACATGCCCAAGGTGCTGCACGACAGCAGCAAGGACAG CATCACCGTCTTTGTCAGCAACCTGCCCTACAGCATGCAGGAGCCGGACACGAAGCTCAGGCCACTCTTCGAGGCCTGTGGGGAGGTGGT CCAGATCCGACCCATCTTCAGCAACCGTGGGGATTTCCGAGGTTACTGCTACGTGGAGTTTAAAGAAGAGAAATCAGCCCTTCAGGCACT GGAGATGGACCGGAAAAGTGTAGAAGGGAGGCCAATGTTTGTTTCCCCCTGTGTGGATAAGAGCAAAAACCCCGATTTTAAGGTGTTCAG GTACAGCACTTCCCTAGAGAAACACAAGCTGTTCATCTCAGGCCTGCCTTTCTCCTGTACTAAAGAGGAACTAGAAGAAATCTGTAAGGC TCATGGCACCGTGAAGGACCTCAGGCTGGTCACCAACCGGGCTGGCAAACCAAAGGGCCTGGCCTACGTGGAGTATGAAAATGAATCCCA GGCGTCGCAGGCTGTGATGAAGATGGACGGCATGACTATCAAAGAGAACATCATCAAAGTGGCAATCAGCAACCCTCCTCAGAGGAAAGT TCCAGAGAAGCCGGAGACCAGGAAGGCACCAGGTGGCCCCATGCTTTTGCCGCAGACATACGGAGCGAGGGGGAAGGGAAGGACGCAGCT GTCTCTACTGCCTCGTGCCCTGCAGCGCCCAAGTGCTGCAGCTCCTCAGGCTGAGAACGGCCCTGCCGCGGCTCCTGCAGTTGCCGCCCC AGCAGCCACCGAGGCACCCAAGATGTCCAATGCCGATTTTGCCAAGCTGTTTCTGAGAAAGTGAACGGGACGCTGGGAGACAGGAAATGC CTTACTTCACTCTGGCCCGGCGGACCTCCCACCACCCAGCAGTGCACTGGGGATGGACAGGCCTGGTGTGCTGCGTGCTCGCAACCACAG ATGGCTCCTCGGCTTTAGACAGAAAGGGGAAGGGGTTCTAAGTCAAGAGCCTTTCAGTGCTCCCTCATATTGAGGGCAGTGGCAGAAAAG TGACCACTCTGCAGGCTGGGCCCAGGATGTGGTGTCCTGAGATAGTTTTGTATCTTAAAGACTGAGGCACAGAAGCGAAACGAGAACACA CTGTTTTTGAGACACAGTTGTCCAAATGTTTCTGGCCAGCTCCGGCCCCTTTTTGTATGACACTTCTCTTCCACCCTGCACAGCACATGT GCCCGTCATTCTTTTAATTTTAAAAGATGAAATGGCAGATGCTAGTAATTCACAGAATGGCCTCTTGTGGGGGTGGGTCTGAGGGAAGTC AGCTATAAAACATTTGCTGGAGTTTTGTTCAATGGGGCTGTGCATTTTTATATTATGTGTTTGTAAATGACATGTCAGCCATTGTTTCAT GTTTCCTAAAAGCAGAATATTTGCAACATTTGTTTTGTATAGGAATTATTTGTGCCACCTGCTGTGGACTGTTTTCTTTGCCTAGTGACT AGTGACCTGTGTTGTCTAAACATGAGTTTCAGCCCTTTGGTTTTGTTTAATACCATGTCAAATGCAAACTTCAATTCTCCCCATTTAGCT TTATTAAACTGACGTTCTCTTCAAAACTTCTTGCTGAATGGTACTCAGATGTGCATTCACATACAGATGTGTTTTGAAGTGGGTGTACCT >86759_86759_5_SSH1-SART3_SSH1_chr12_109198832_ENST00000551165_SART3_chr12_108932865_ENST00000228284_length(amino acids)=979AA_BP=0 MALVTLQRSPTPSAASSSASNSELEAGSEEDRKLNLSLSESFFMVKGAALFLQQGSSPQGQRSLQHPHKHAGDLPQHLQVMINLLRCEDR IKLAVRLESAWADRVRYMVVVYSSGRQDTEENILLGVDFSSKESKSCTIGMVLRLWSDTKIHLDGDGGFSVSTAGRMHIFKPVSVQAMWS ALQVLHKACEVARRHNYFPGGVALIWATYYESCISSEQSCINEWNAMQDLESTRPDSPALFVDKPTEGERTERLIKAKLRSIMMSQDLEN VTSKEIRNELEKQMNCNLKELKEFIDNEMLLILGQMDKPSLIFDHLYLLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVP DLWIRYSQYLDRQLKVKDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISVTFEKALNAGFIQATDYVEIWQAYLDYLRRRVDF KQDSSKELEELRAAFTRALEYLKQEVEERFNESGDPSCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERAHGDT QHCRKALHRAVQCTSDYPEHVCEVLLTMERTEGSLEDWDIAVQKTETRLARVNEQRMKAAEKEAALVQQEEEKAEQRKRARAEKKALKKK KKIRGPEKRGADEDDEKEWGDDEEEQPSKRRRVENSIPAAGETQNVEVAAGPAGKCAAVDVEPPSKQKEKAASLKRDMPKVLHDSSKDSI TVFVSNLPYSMQEPDTKLRPLFEACGEVVQIRPIFSNRGDFRGYCYVEFKEEKSALQALEMDRKSVEGRPMFVSPCVDKSKNPDFKVFRY STSLEKHKLFISGLPFSCTKEELEEICKAHGTVKDLRLVTNRAGKPKGLAYVEYENESQASQAVMKMDGMTIKENIIKVAISNPPQRKVP -------------------------------------------------------------- >86759_86759_6_SSH1-SART3_SSH1_chr12_109198832_ENST00000551165_SART3_chr12_108932865_ENST00000431469_length(transcript)=2971nt_BP=1048nt CGGCTCCAGGAGGCCCCGGGCCCGGCCCGGGCGGCGGTGGCGGTGGCGGCTCTAGCTCGAGACGTCTGTGGCGCCCTCGCACCGCGCCGC AGCCATGGCCCTGGTGACCCTGCAGCGCTCGCCCACGCCCAGCGCCGCCTCCTCCTCGGCCAGCAACAGCGAGTTGGAGGCTGGCAGCGA AGAAGATCGAAAATTAAACCTCAGCTTAAGTGAGAGCTTTTTCATGGTGAAAGGCGCAGCCCTCTTCTTACAACAGGGAAGCAGCCCTCA AGGCCAGCGGAGTCTTCAGCACCCCCACAAGCATGCAGGTGATCTGCCTCAACATCTTCAGGTGATGATCAACCTTCTGCGTTGCGAAGA CAGAATCAAGCTGGCAGTGCGCCTGGAGAGCGCCTGGGCGGACCGGGTCCGGTACATGGTGGTGGTGTACAGCAGCGGGCGCCAGGACAC CGAGGAGAATATCTTGCTGGGAGTGGACTTTTCCAGTAAGGAAAGTAAAAGCTGCACCATTGGGATGGTTCTCCGACTGTGGAGCGACAC GAAAATCCACCTTGATGGAGATGGTGGGTTCAGCGTGAGCACAGCAGGAAGGATGCACATATTTAAGCCTGTGTCTGTCCAGGCCATGTG GTCTGCCCTGCAGGTGCTTCACAAGGCCTGCGAAGTGGCCCGGAGGCACAACTACTTCCCCGGGGGTGTAGCTCTCATCTGGGCTACCTA CTATGAGAGCTGCATCAGCTCCGAGCAGAGCTGCATCAACGAGTGGAACGCCATGCAGGACCTGGAGTCTACGCGGCCCGACTCCCCCGC GCTATTTGTGGACAAGCCCACTGAAGGGGAAAGGACCGAGCGCCTCATCAAAGCCAAGCTCCGAAGCATCATGATGAGCCAGGATCTAGA AAATGTGACTTCCAAAGAGATTCGTAATGAATTAGAGAAACAGATGAATTGTAACTTGAAGGAACTCAAGGAATTTATAGACAATGAGAT GCTACTTATCTTGGGACAGATGGACAAGCCCTCCCTTATCTTCGATCATCTTTATCTCTTGCAGGCAGAGGCACCAAGGCTGGCAGAATA TCAAGCATATATCGATTTTGAGATGAAAATTGGCGATCCTGCTCGCATTCAGTTGATCTTTGAGCGCGCCCTGGTCGAGAACTGCCTTGT CCCAGACTTATGGATCCGTTACAGTCAGTACCTAGATCGACAACTGAAAGTAAAGGATTTGGTTTTATCTGTACATAACCGCGCTATTAG AAACTGCCCCTGGACAGTTGCCTTATGGAGTCGGTACCTCTTGGCCATGGAGAGACATGGAGTTGATCATCAAGTAATTTCTGACTCCAG TAAAGAGCTGGAGGAGTTGAGGGCCGCCTTTACTCGTGCCTTGGAGTATCTGAAGCAGGAGGTGGAAGAGCGTTTCAATGAGAGTGGTGA TCCAAGCTGCGTGATTATGCAGAACTGGGCTAGGATTGAGGCTCGACTGTGCAATAACATGCAGAAAGCTCGGGAACTCTGGGATAGCAT CATGACCAGAGGAAATGCCAAGTACGCCAACATGTGGCTAGAGTATTACAACCTGGAAAGAGCTCATGGTGACACCCAGCACTGCCGGAA GGCTCTGCACCGGGCCGTCCAGTGCACCAGTGACTACCCAGAGCACGTCTGCGAAGTGTTACTCACCATGGAGAGGACAGAAGGTTCTTT AGAAGATTGGGATATAGCTGTTCAGAAAACTGAAACCCGATTAGCTCGTGTCAATGAGCAGAGAATGAAGGCTGCAGAGAAGGAAGCAGC CCTTGTGCAGCAAGAAGAAGAAAAGGCTGAACAACGGAAAAGAGCTCGGGCTGAGAAGAAAGCGTTAAAAAAGAAGAAAAAGATCAGAGG CCCAGAGAAGCGCGGAGCAGATGAGGATGATGAGAAAGAGTGGGGCGATGATGAAGAAGAGCAGCCTTCCAAACGCAGAAGGGTCGAGAA CAGCATCCCTGCAGCTGGAGAAACACAAAATGTAGAAGTAGCAGCAGGGCCCGCTGGGAAATGTGCTGCCGTAGATGTGGAGCCCCCTTC GAAGCAGAAGGAGAAGGCAGCCTCCCTGAAGAGGGACATGCCCAAGGTGCTGCACGACAGCAGCAAGGACAGCATCACCGTCTTTGTCAG CAACCTGCCCTACAGCATGCAGGAGCCGGACACGAAGCTCAGGCCACTCTTCGAGGCCTGTGGGGAGGTGGTCCAGATCCGACCCATCTT CAGCAACCGTGGGGATTTCCGAGGTTACTGCTACGTGGAGTTTAAAGAAGAGAAATCAGCCCTTCAGGCACTGGAGATGGACCGGAAAAG TGTAGAAGGGAGGCCAATGTTTGTTTCCCCCTGTGTGGATAAGAGCAAAAACCCCGATTTTAAGGTGTTCAGGTACAGCACTTCCCTAGA GAAACACAAGCTGTTCATCTCAGGCCTGCCTTTCTCCTGTACTAAAGAGGAACTAGAAGAAATCTGTAAGGCTCATGGCACCGTGAAGGA CCTCAGGCTGGTCACCAACCGGGCTGGCAAACCAAAGGGCCTGGCCTACGTGGAGTATGAAAATGAATCCCAGGCGTCGCAGGCTGTGAT GAAGATGGACGGCATGACTATCAAAGAGAACATCATCAAAGTGGCAATCAGCAACCCTCCTCAGAGGAAAGTTCCAGAGAAGCCGGAGAC CAGGAAGGCACCAGGTGGCCCCATGCTTTTGCCGCAGACATACGGAGCGAGGGGGAAGGGAAGGACGCAGCTGTCTCTACTGCCTCGTGC CCTGCAGCGCCCAAGTGCTGCAGCTCCTCAGGCTGAGAACGGCCCTGCCGCGGCTCCTGCAGTTGCCGCCCCAGCAGCCACCGAGGCACC CAAGATGTCCAATGCCGATTTTGCCAAGCTGTTTCTGAGAAAGTGAACGGGACGCTGGGAGACAGGAAATGCCTTACTTCACTCTGGCCC >86759_86759_6_SSH1-SART3_SSH1_chr12_109198832_ENST00000551165_SART3_chr12_108932865_ENST00000431469_length(amino acids)=943AA_BP=0 MALVTLQRSPTPSAASSSASNSELEAGSEEDRKLNLSLSESFFMVKGAALFLQQGSSPQGQRSLQHPHKHAGDLPQHLQVMINLLRCEDR IKLAVRLESAWADRVRYMVVVYSSGRQDTEENILLGVDFSSKESKSCTIGMVLRLWSDTKIHLDGDGGFSVSTAGRMHIFKPVSVQAMWS ALQVLHKACEVARRHNYFPGGVALIWATYYESCISSEQSCINEWNAMQDLESTRPDSPALFVDKPTEGERTERLIKAKLRSIMMSQDLEN VTSKEIRNELEKQMNCNLKELKEFIDNEMLLILGQMDKPSLIFDHLYLLQAEAPRLAEYQAYIDFEMKIGDPARIQLIFERALVENCLVP DLWIRYSQYLDRQLKVKDLVLSVHNRAIRNCPWTVALWSRYLLAMERHGVDHQVISDSSKELEELRAAFTRALEYLKQEVEERFNESGDP SCVIMQNWARIEARLCNNMQKARELWDSIMTRGNAKYANMWLEYYNLERAHGDTQHCRKALHRAVQCTSDYPEHVCEVLLTMERTEGSLE DWDIAVQKTETRLARVNEQRMKAAEKEAALVQQEEEKAEQRKRARAEKKALKKKKKIRGPEKRGADEDDEKEWGDDEEEQPSKRRRVENS IPAAGETQNVEVAAGPAGKCAAVDVEPPSKQKEKAASLKRDMPKVLHDSSKDSITVFVSNLPYSMQEPDTKLRPLFEACGEVVQIRPIFS NRGDFRGYCYVEFKEEKSALQALEMDRKSVEGRPMFVSPCVDKSKNPDFKVFRYSTSLEKHKLFISGLPFSCTKEELEEICKAHGTVKDL RLVTNRAGKPKGLAYVEYENESQASQAVMKMDGMTIKENIIKVAISNPPQRKVPEKPETRKAPGGPMLLPQTYGARGKGRTQLSLLPRAL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for SSH1-SART3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000326470 | - | 9 | 13 | 897_1049 | 329.0 | 704.0 | YWHAG |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000326495 | - | 10 | 15 | 897_1049 | 318.0 | 1050.0 | YWHAG |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000360239 | - | 10 | 14 | 897_1049 | 21.666666666666668 | 738.0 | YWHAG |
| Hgene | SSH1 | chr12:109198832 | chr12:108932865 | ENST00000551165 | - | 10 | 14 | 897_1049 | 318.0 | 693.0 | YWHAG |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for SSH1-SART3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for SSH1-SART3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
