
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:STXBP2-PIDD (FusionGDB2 ID:87880) |
Fusion Gene Summary for STXBP2-PIDD |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: STXBP2-PIDD | Fusion gene ID: 87880 | Hgene | Tgene | Gene symbol | STXBP2 | PIDD | Gene ID | 6813 | 55367 |
| Gene name | syntaxin binding protein 2 | p53-induced death domain protein 1 | |
| Synonyms | FHL5|Hunc18b|MUNC18-2|UNC18-2|UNC18B|pp10122 | LRDD|PIDD | |
| Cytomap | 19p13.2 | 11p15.5 | |
| Type of gene | protein-coding | protein-coding | |
| Description | syntaxin-binding protein 2protein unc-18 homolog Bunc-18B | p53-induced death domain-containing protein 1leucine-rich repeat and death domain-containing proteinleucine-rich repeats and death domain containingp53-induced protein with a death domain | |
| Modification date | 20200328 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000221283, ENST00000414284, ENST00000441779, ENST00000602355, | ENST00000347755, ENST00000534649, ENST00000411829, | |
| Fusion gene scores | * DoF score | 5 X 5 X 4=100 | 2 X 1 X 1=2 |
| # samples | 5 | 3 | |
| ** MAII score | log2(5/100*10)=-1 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/2*10)=3.90689059560852 | |
| Context | PubMed: STXBP2 [Title/Abstract] AND PIDD [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | STXBP2(7708131)-PIDD(804463), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | PIDD | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process | 17159900 |
| Tgene | PIDD | GO:0006974 | cellular response to DNA damage stimulus | 21726810 |
| Tgene | PIDD | GO:0016540 | protein autoprocessing | 17159900 |
 Fusion gene breakpoints across STXBP2 (5'-gene) Fusion gene breakpoints across STXBP2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
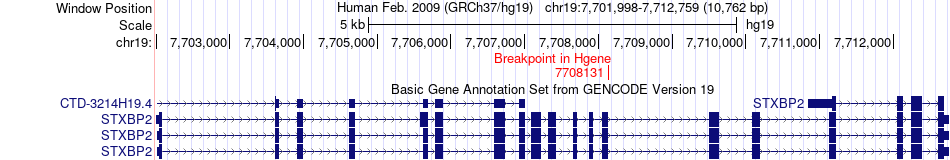 |
 Fusion gene breakpoints across PIDD (3'-gene) Fusion gene breakpoints across PIDD (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
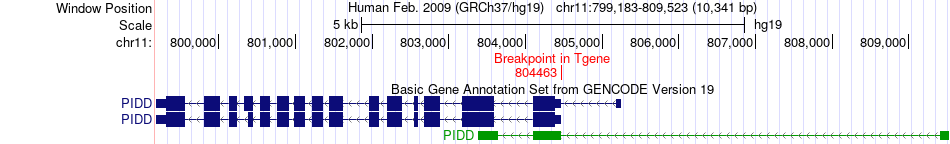 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-D7-8576 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
Top |
Fusion Gene ORF analysis for STXBP2-PIDD |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000221283 | ENST00000347755 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| 5CDS-5UTR | ENST00000221283 | ENST00000534649 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| 5CDS-5UTR | ENST00000414284 | ENST00000347755 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| 5CDS-5UTR | ENST00000414284 | ENST00000534649 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| 5CDS-5UTR | ENST00000441779 | ENST00000347755 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| 5CDS-5UTR | ENST00000441779 | ENST00000534649 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| In-frame | ENST00000221283 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| In-frame | ENST00000414284 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| In-frame | ENST00000441779 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| intron-3CDS | ENST00000602355 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| intron-5UTR | ENST00000602355 | ENST00000347755 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
| intron-5UTR | ENST00000602355 | ENST00000534649 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000441779 | STXBP2 | chr19 | 7708131 | + | ENST00000411829 | PIDD | chr11 | 804463 | - | 4063 | 1177 | 37 | 3939 | 1300 |
| ENST00000221283 | STXBP2 | chr19 | 7708131 | + | ENST00000411829 | PIDD | chr11 | 804463 | - | 4024 | 1138 | 31 | 3900 | 1289 |
| ENST00000414284 | STXBP2 | chr19 | 7708131 | + | ENST00000411829 | PIDD | chr11 | 804463 | - | 4007 | 1121 | 23 | 3883 | 1286 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000441779 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - | 0.004077309 | 0.9959227 |
| ENST00000221283 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - | 0.004105437 | 0.99589455 |
| ENST00000414284 | ENST00000411829 | STXBP2 | chr19 | 7708131 | + | PIDD | chr11 | 804463 | - | 0.004093534 | 0.9959065 |
Top |
Fusion Genomic Features for STXBP2-PIDD |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
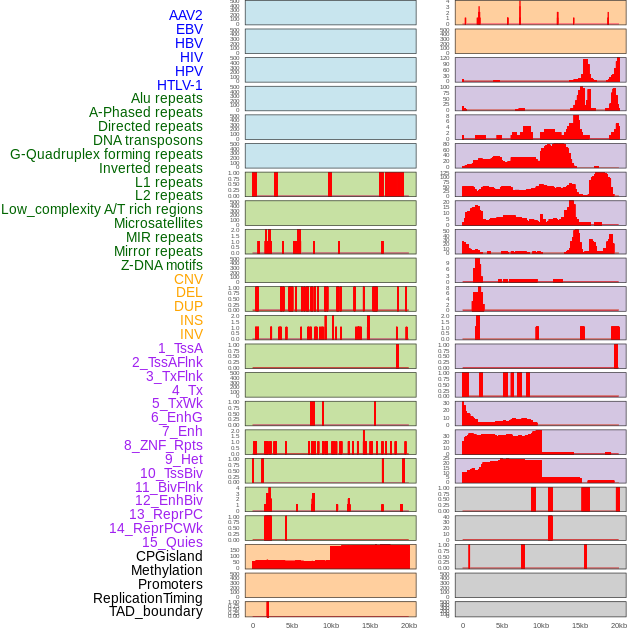 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for STXBP2-PIDD |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:7708131/chr11:804463) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 322_454 | 0 | 911.0 | Domain | ZU5 1 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 455_596 | 0 | 911.0 | Domain | ZU5 2 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 788_873 | 0 | 911.0 | Domain | Death | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 322_454 | 98 | 894.0 | Domain | ZU5 1 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 455_596 | 98 | 894.0 | Domain | ZU5 2 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 788_873 | 98 | 894.0 | Domain | Death | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 423_452 | 0 | 911.0 | Region | Peptidase S68 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 566_594 | 0 | 911.0 | Region | Peptidase S68 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 580_716 | 0 | 911.0 | Region | UPA domain | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 423_452 | 98 | 894.0 | Region | Peptidase S68 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 566_594 | 98 | 894.0 | Region | Peptidase S68 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 580_716 | 98 | 894.0 | Region | UPA domain | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 126_147 | 0 | 911.0 | Repeat | Note=LRR 1 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 149_171 | 0 | 911.0 | Repeat | Note=LRR 2 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 172_194 | 0 | 911.0 | Repeat | Note=LRR 3 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 195_216 | 0 | 911.0 | Repeat | Note=LRR 4 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 218_240 | 0 | 911.0 | Repeat | Note=LRR 5 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 241_263 | 0 | 911.0 | Repeat | Note=LRR 6 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000347755 | 0 | 16 | 264_285 | 0 | 911.0 | Repeat | Note=LRR 7 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 126_147 | 98 | 894.0 | Repeat | Note=LRR 1 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 149_171 | 98 | 894.0 | Repeat | Note=LRR 2 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 172_194 | 98 | 894.0 | Repeat | Note=LRR 3 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 195_216 | 98 | 894.0 | Repeat | Note=LRR 4 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 218_240 | 98 | 894.0 | Repeat | Note=LRR 5 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 241_263 | 98 | 894.0 | Repeat | Note=LRR 6 | |
| Tgene | PIDD | chr19:7708131 | chr11:804463 | ENST00000411829 | -1 | 15 | 264_285 | 98 | 894.0 | Repeat | Note=LRR 7 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for STXBP2-PIDD |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >87880_87880_1_STXBP2-PIDD_STXBP2_chr19_7708131_ENST00000221283_PIDD_chr11_804463_ENST00000411829_length(transcript)=4024nt_BP=1138nt CCGGAAGCGGCGGCGGCGCCCCTCGGGGAAGATGGCGCCCTCGGGGCTGAAGGCGGTGGTGGGGGAAAAAATTCTGAGCGGAGTTATTCG GAGTGTCAAGAAGGATGGGGAGTGGAAGGTGCTTATCATGGATCACCCAAGCATGCGCATCTTGTCTTCCTGCTGCAAAATGTCAGATAT CCTGGCTGAGGGCATCACCATTGTTGAAGACATCAACAAACGGCGGGAACCCATTCCCAGTCTGGAGGCCATTTATTTGCTGAGCCCCAC GGAGAAGTCGGTTCAGGCCCTGATCAAAGACTTCCAGGGGACCCCGACTTTCACCTACAAAGCGGCCCATATCTTCTTCACCGACACCTG CCCCGAGCCCCTGTTCAGTGAGCTAGGCCGCTCTCGTCTGGCAAAGGTGGTGAAGACGTTGAAGGAGATTCACCTTGCCTTCCTCCCCTA CGAGGCCCAGGTGTTCTCCCTCGATGCTCCCCACAGCACCTACAACCTCTACTGCCCCTTCCGGGCAGAGGAGCGCACGCGGCAGCTCGA GGTGCTGGCCCAGCAGATTGCCACGCTGTGCGCCACCCTGCAGGAGTACCCGGCCATCCGCTACCGCAAGGGCCCAGAGGACACAGCCCA GTTGGCCCACGCCGTCCTGGCCAAGCTGAACGCCTTCAAGGCAGACACTCCCAGTCTGGGCGAGGGCCCAGAGAAAACCCGCTCCCAGCT GCTGATAATGGACCGGGCAGCTGACCCCGTGTCCCCACTACTGCATGAGCTCACGTTCCAGGCCATGGCGTATGATCTGCTGGACATAGA GCAGGACACATACAGGTATGAGACCACCGGGCTGAGCGAGGCGCGGGAGAAGGCCGTCTTGCTGGACGAGGACGATGACTTGTGGGTGGA GCTTCGCCACATGCATATCGCAGATGTGTCCAAGAAGGTCACGGAGCTCCTGAGGACCTTCTGTGAGAGCAAGAGGCTGACCACGGACAA GGCGAACATCAAAGACCTATCCCAGATCCTGAAAAAGATGCCGCAGTACCAGAAGGAGCTGAATAAGTATTCTACGCACCTGCATCTAGC AGATGATTGTATGAAGCACTTCAAGGGCTCGGTGGAGAAGCTGTGTAGTGTGGAGCAGCTGCAGCTTCCCCGGGCGCTGCCTGGACAGGC CTGCCTGCGTGCTGGGACATGTCTGGCCTCCAAGGACCGTCGGTGGGCGATGGCTGCAACGGTGGAGGGGCCAGAGCTGGAGGCAGCTGC TGCCGCAGGAGATGCTTCAGAGGATTCGGACGCAGGGTCCAGGGCGCTGCCTTTCCTGGGCGGCAACCGGCTGAGCTTGGACCTGTACCC CGGGGGCTGCCAGCAGCTGCTGCACCTGTGTGTCCAGCAGCCTCTGCAGCTGCTGCAGGTGGAATTCTTGCGTCTGAGCACTCACGAGGA CCCTCAGCTGCTGGAGGCCACCCTGGCCCAGCTGCCTCAGAGCCTGTCCTGCCTCCGCTCCCTGGTCCTCAAAGGAGGGCAACGCCGGGA CACACTGGGTGCCTGTCTCCGGGGTGCCCTGACCAACCTGCCCGCTGGTCTGAGTGGCCTGGCCCATCTGGCCCACCTGGACCTGAGCTT CAACAGCCTGGAGACACTGCCGGCCTGTGTCCTGCAGATGCGAGGTCTGGGTGCGCTCTTGCTGTCTCACAACTGCCTCTCTGAGCTGCC TGAGGCTCTGGGGGCCCTCCCCGCCCTCACCTTCCTCACAGTGACACACAACCGCCTGCAGACGCTGCCCCCAGCACTGGGGGCCCTATC CACCCTGCAGCGCCTCGATCTCTCTCAGAATCTGCTGGACACGCTACCTCCTGAGATTGGAGGCCTGGGCAGCCTCCTGGAGCTCAACCT GGCCTCCAACCGGCTGCAGAGCCTCCCAGCCTCTCTGGCGGGACTTCGGTCCTTGCGGCTCCTTGTCCTGCACAGCAACCTCCTGGCCTC TGTGCCAGCTGACTTGGCCCGCCTTCCACTCCTCACCCGGCTCGACCTGAGGGACAACCAGCTCCGGGACCTGCCCCCTGAGCTGCTAGA CGCCCCCTTTGTGCGCCTGCAGGGGAACCCCCTGGGTGAGGCCTCGCCAGACGCCCCGAGTTCACCAGTGGCAGCCCTCATTCCAGAAAT GCCCAGACTGTTCCTGACCTCAGATTTGGACAGCTTTCCTGTGACCCCTCAAGGCTGCTCAGTGACCCTGGCCTGTGGCGTCCGCCTGCA GTTCCCAGCGGGAGCCACCGCCACCCCCATCACCATCCGCTATCGGCTGCTGCTGCCGGAGCCAGGCCTCGTCCCCCTGGGTCCTCATGA CGCCCTGCTCAGCCATGTGCTGGAGCTGCAGCCCCATGGGGTGGCCTTCCAGCAGGATGTGGGGCTGTGGCTGCTCTTCACCCCACCGCA GGCCCGGCGCTGCCGTGAAGTGGTGGTCAGGACCCGGAATGACAACAGCTGGGGTGACCTGGAGACCTACCTGGAGGAAGAGGCACCCCA GCGGCTCTGGGCTCACTGCCAGGTGCCCCACTTCTCCTGGTTCCTTGTGGTTTCCCGCCCTGTGTCCAATGCCTGCCTGGTGCCACCGGA GGGGACACTGCTGTGCTCCTCGGGTCATCCTGGGGTCAAAGTCATCTTCCCCCCTGGGGCCACTGAGGAGCCTCGTCGAGTCTCCATGCA GGTGGTGCGCATGGCTGGCCGAGAGCTGCAGGCCCTCCTGGGAGAACCAGAGGCTGCAGTGAGCCCCCTGCTGTGCCTGTCACAGAGCGG TCCCCCCAGCTTCCTCCAACCGGTCACCGTGCAGCTGCCTCTGCCCTCTGGCATCACAGGCCTCAGTCTGGACCGCTCCCGCCTGCACCT GTTGTACTGGGCCCCTCCTGCAGCCACCTGGGATGACATCACAGCTCAGGTGGTCCTGGAGCTCACCCACCTGTACGCACGCTTCCAGGT CACACACTTCTCCTGGTACTGGCTCTGGTACACCACCAAGAACTGTGTGGGAGGCCTGGCTCGGAAGGCCTGGGAGCGGCTGCGGCTGCA CCGTGTGAACCTCATCGCTCTGCAGCGGCGCCGGGACCCTGAGCAGGTCCTGCTGCAGTGCCTGCCCCGAAACAAGGTGGACGCCACCCT TCGGCGGCTGCTGGAGCGGTACCGGGGCCCCGAGCCCTCTGACACGGTGGAGATGTTCGAGGGCGAAGAGTTCTTTGCGGCCTTCGAGCG CGGCATCGACGTGGATGCTGACCGCCCTGACTGTGTGGAGGGCAGAATCTGCTTTGTCTTCTACTCGCACCTGAAGAATGTGAAGGAGGT GTCCTTCTACCGTGGCGCGGTGCCTGTGCGGGTGCCCGAGGAGGCTGAGGCTGCCCGGCAGAGGAAGGGCGCAGACGCCCTGTGGATGGC CACTCTGCCCATCAAGCTGCCGAGACTTCGAGGGTCCGAGGGGCCACGGCGGGGGGCTGGCCTCTCCTTGGCACCCTTGAATCTGGGAGA TGCCGAGACCGGCTTTCTGACGCAGAGCAACCTGCTGAGTGTGGCTGGGCGTCTGGGTCTGGACTGGCCAGCCGTGGCCCTGCACCTGGG GGTGTCCTACCGGGAGGTGCAGCGCATCCGGCACGAGTTCCGGGATGATCTGGATGAGCAGATCCGTCACATGCTCTTCTCCTGGGCTGA GCGCCAGGCTGGGCAGCCAGGGGCTGTGGGGCTCCTGGTGCAGGCCCTGGAGCAGAGTGACCGGCAGGACGTGGCTGAAGAGGTGCGCGC AGTCTTGGAGCTCGGCCGCCGCAAGTACCAGGACAGCATCCGACGCATGGGCTTGGCCCCCAAGGACCCCGCTCTGCCTGGCTCCTCGGC TCCACAGCCCCCAGAGCCTGCCCAGGCCTAGGCCCCACAGACTTTTAGGCTGGCCCAGATATTCCCCAGTGGATGGGCAGAGCCCCCACC >87880_87880_1_STXBP2-PIDD_STXBP2_chr19_7708131_ENST00000221283_PIDD_chr11_804463_ENST00000411829_length(amino acids)=1289AA_BP=369 MAPSGLKAVVGEKILSGVIRSVKKDGEWKVLIMDHPSMRILSSCCKMSDILAEGITIVEDINKRREPIPSLEAIYLLSPTEKSVQALIKD FQGTPTFTYKAAHIFFTDTCPEPLFSELGRSRLAKVVKTLKEIHLAFLPYEAQVFSLDAPHSTYNLYCPFRAEERTRQLEVLAQQIATLC ATLQEYPAIRYRKGPEDTAQLAHAVLAKLNAFKADTPSLGEGPEKTRSQLLIMDRAADPVSPLLHELTFQAMAYDLLDIEQDTYRYETTG LSEAREKAVLLDEDDDLWVELRHMHIADVSKKVTELLRTFCESKRLTTDKANIKDLSQILKKMPQYQKELNKYSTHLHLADDCMKHFKGS VEKLCSVEQLQLPRALPGQACLRAGTCLASKDRRWAMAATVEGPELEAAAAAGDASEDSDAGSRALPFLGGNRLSLDLYPGGCQQLLHLC VQQPLQLLQVEFLRLSTHEDPQLLEATLAQLPQSLSCLRSLVLKGGQRRDTLGACLRGALTNLPAGLSGLAHLAHLDLSFNSLETLPACV LQMRGLGALLLSHNCLSELPEALGALPALTFLTVTHNRLQTLPPALGALSTLQRLDLSQNLLDTLPPEIGGLGSLLELNLASNRLQSLPA SLAGLRSLRLLVLHSNLLASVPADLARLPLLTRLDLRDNQLRDLPPELLDAPFVRLQGNPLGEASPDAPSSPVAALIPEMPRLFLTSDLD SFPVTPQGCSVTLACGVRLQFPAGATATPITIRYRLLLPEPGLVPLGPHDALLSHVLELQPHGVAFQQDVGLWLLFTPPQARRCREVVVR TRNDNSWGDLETYLEEEAPQRLWAHCQVPHFSWFLVVSRPVSNACLVPPEGTLLCSSGHPGVKVIFPPGATEEPRRVSMQVVRMAGRELQ ALLGEPEAAVSPLLCLSQSGPPSFLQPVTVQLPLPSGITGLSLDRSRLHLLYWAPPAATWDDITAQVVLELTHLYARFQVTHFSWYWLWY TTKNCVGGLARKAWERLRLHRVNLIALQRRRDPEQVLLQCLPRNKVDATLRRLLERYRGPEPSDTVEMFEGEEFFAAFERGIDVDADRPD CVEGRICFVFYSHLKNVKEVSFYRGAVPVRVPEEAEAARQRKGADALWMATLPIKLPRLRGSEGPRRGAGLSLAPLNLGDAETGFLTQSN LLSVAGRLGLDWPAVALHLGVSYREVQRIRHEFRDDLDEQIRHMLFSWAERQAGQPGAVGLLVQALEQSDRQDVAEEVRAVLELGRRKYQ -------------------------------------------------------------- >87880_87880_2_STXBP2-PIDD_STXBP2_chr19_7708131_ENST00000414284_PIDD_chr11_804463_ENST00000411829_length(transcript)=4007nt_BP=1121nt GGCGGCGGCGCCCCTCGGGGAAGATGGCGCCCTCGGGGCTGAAGGCGGTGGTGGGGGAAAAAATTCTGAGCGGAGTTATTCGGAGTGTCA AGAAGGATGGGGAGTGGAAGGTGCTTATCATGGATCACCCAAGCATGCGCATCTTGTCTTCCTGCTGCAAAATGTCAGATATCCTGGCTG AGGGCATCACCATTGTTGAAGACATCAACAAACGGCGGGAACCCATTCCCAGTCTGGAGGCCATTTATTTGCTGAGCCCCACGGAGAAGG CCCTGATCAAAGACTTCCAGGGGACCCCGACTTTCACCTACAAAGCGGCCCATATCTTCTTCACCGACACCTGCCCCGAGCCCCTGTTCA GTGAGCTAGGCCGCTCTCGTCTGGCAAAGGTGGTGAAGACGTTGAAGGAGATTCACCTTGCCTTCCTCCCCTACGAGGCCCAGGTGTTCT CCCTCGATGCTCCCCACAGCACCTACAACCTCTACTGCCCCTTCCGGGCAGAGGAGCGCACGCGGCAGCTCGAGGTGCTGGCCCAGCAGA TTGCCACGCTGTGCGCCACCCTGCAGGAGTACCCGGCCATCCGCTACCGCAAGGGCCCAGAGGACACAGCCCAGTTGGCCCACGCCGTCC TGGCCAAGCTGAACGCCTTCAAGGCAGACACTCCCAGTCTGGGCGAGGGCCCAGAGAAAACCCGCTCCCAGCTGCTGATAATGGACCGGG CAGCTGACCCCGTGTCCCCACTACTGCATGAGCTCACGTTCCAGGCCATGGCGTATGATCTGCTGGACATAGAGCAGGACACATACAGGT ATGAGACCACCGGGCTGAGCGAGGCGCGGGAGAAGGCCGTCTTGCTGGACGAGGACGATGACTTGTGGGTGGAGCTTCGCCACATGCATA TCGCAGATGTGTCCAAGAAGGTCACGGAGCTCCTGAGGACCTTCTGTGAGAGCAAGAGGCTGACCACGGACAAGGCGAACATCAAAGACC TATCCCAGATCCTGAAAAAGATGCCGCAGTACCAGAAGGAGCTGAATAAGTATTCTACGCACCTGCATCTAGCAGATGATTGTATGAAGC ACTTCAAGGGCTCGGTGGAGAAGCTGTGTAGTGTGGAGCAGCTGCAGCTTCCCCGGGCGCTGCCTGGACAGGCCTGCCTGCGTGCTGGGA CATGTCTGGCCTCCAAGGACCGTCGGTGGGCGATGGCTGCAACGGTGGAGGGGCCAGAGCTGGAGGCAGCTGCTGCCGCAGGAGATGCTT CAGAGGATTCGGACGCAGGGTCCAGGGCGCTGCCTTTCCTGGGCGGCAACCGGCTGAGCTTGGACCTGTACCCCGGGGGCTGCCAGCAGC TGCTGCACCTGTGTGTCCAGCAGCCTCTGCAGCTGCTGCAGGTGGAATTCTTGCGTCTGAGCACTCACGAGGACCCTCAGCTGCTGGAGG CCACCCTGGCCCAGCTGCCTCAGAGCCTGTCCTGCCTCCGCTCCCTGGTCCTCAAAGGAGGGCAACGCCGGGACACACTGGGTGCCTGTC TCCGGGGTGCCCTGACCAACCTGCCCGCTGGTCTGAGTGGCCTGGCCCATCTGGCCCACCTGGACCTGAGCTTCAACAGCCTGGAGACAC TGCCGGCCTGTGTCCTGCAGATGCGAGGTCTGGGTGCGCTCTTGCTGTCTCACAACTGCCTCTCTGAGCTGCCTGAGGCTCTGGGGGCCC TCCCCGCCCTCACCTTCCTCACAGTGACACACAACCGCCTGCAGACGCTGCCCCCAGCACTGGGGGCCCTATCCACCCTGCAGCGCCTCG ATCTCTCTCAGAATCTGCTGGACACGCTACCTCCTGAGATTGGAGGCCTGGGCAGCCTCCTGGAGCTCAACCTGGCCTCCAACCGGCTGC AGAGCCTCCCAGCCTCTCTGGCGGGACTTCGGTCCTTGCGGCTCCTTGTCCTGCACAGCAACCTCCTGGCCTCTGTGCCAGCTGACTTGG CCCGCCTTCCACTCCTCACCCGGCTCGACCTGAGGGACAACCAGCTCCGGGACCTGCCCCCTGAGCTGCTAGACGCCCCCTTTGTGCGCC TGCAGGGGAACCCCCTGGGTGAGGCCTCGCCAGACGCCCCGAGTTCACCAGTGGCAGCCCTCATTCCAGAAATGCCCAGACTGTTCCTGA CCTCAGATTTGGACAGCTTTCCTGTGACCCCTCAAGGCTGCTCAGTGACCCTGGCCTGTGGCGTCCGCCTGCAGTTCCCAGCGGGAGCCA CCGCCACCCCCATCACCATCCGCTATCGGCTGCTGCTGCCGGAGCCAGGCCTCGTCCCCCTGGGTCCTCATGACGCCCTGCTCAGCCATG TGCTGGAGCTGCAGCCCCATGGGGTGGCCTTCCAGCAGGATGTGGGGCTGTGGCTGCTCTTCACCCCACCGCAGGCCCGGCGCTGCCGTG AAGTGGTGGTCAGGACCCGGAATGACAACAGCTGGGGTGACCTGGAGACCTACCTGGAGGAAGAGGCACCCCAGCGGCTCTGGGCTCACT GCCAGGTGCCCCACTTCTCCTGGTTCCTTGTGGTTTCCCGCCCTGTGTCCAATGCCTGCCTGGTGCCACCGGAGGGGACACTGCTGTGCT CCTCGGGTCATCCTGGGGTCAAAGTCATCTTCCCCCCTGGGGCCACTGAGGAGCCTCGTCGAGTCTCCATGCAGGTGGTGCGCATGGCTG GCCGAGAGCTGCAGGCCCTCCTGGGAGAACCAGAGGCTGCAGTGAGCCCCCTGCTGTGCCTGTCACAGAGCGGTCCCCCCAGCTTCCTCC AACCGGTCACCGTGCAGCTGCCTCTGCCCTCTGGCATCACAGGCCTCAGTCTGGACCGCTCCCGCCTGCACCTGTTGTACTGGGCCCCTC CTGCAGCCACCTGGGATGACATCACAGCTCAGGTGGTCCTGGAGCTCACCCACCTGTACGCACGCTTCCAGGTCACACACTTCTCCTGGT ACTGGCTCTGGTACACCACCAAGAACTGTGTGGGAGGCCTGGCTCGGAAGGCCTGGGAGCGGCTGCGGCTGCACCGTGTGAACCTCATCG CTCTGCAGCGGCGCCGGGACCCTGAGCAGGTCCTGCTGCAGTGCCTGCCCCGAAACAAGGTGGACGCCACCCTTCGGCGGCTGCTGGAGC GGTACCGGGGCCCCGAGCCCTCTGACACGGTGGAGATGTTCGAGGGCGAAGAGTTCTTTGCGGCCTTCGAGCGCGGCATCGACGTGGATG CTGACCGCCCTGACTGTGTGGAGGGCAGAATCTGCTTTGTCTTCTACTCGCACCTGAAGAATGTGAAGGAGGTGTCCTTCTACCGTGGCG CGGTGCCTGTGCGGGTGCCCGAGGAGGCTGAGGCTGCCCGGCAGAGGAAGGGCGCAGACGCCCTGTGGATGGCCACTCTGCCCATCAAGC TGCCGAGACTTCGAGGGTCCGAGGGGCCACGGCGGGGGGCTGGCCTCTCCTTGGCACCCTTGAATCTGGGAGATGCCGAGACCGGCTTTC TGACGCAGAGCAACCTGCTGAGTGTGGCTGGGCGTCTGGGTCTGGACTGGCCAGCCGTGGCCCTGCACCTGGGGGTGTCCTACCGGGAGG TGCAGCGCATCCGGCACGAGTTCCGGGATGATCTGGATGAGCAGATCCGTCACATGCTCTTCTCCTGGGCTGAGCGCCAGGCTGGGCAGC CAGGGGCTGTGGGGCTCCTGGTGCAGGCCCTGGAGCAGAGTGACCGGCAGGACGTGGCTGAAGAGGTGCGCGCAGTCTTGGAGCTCGGCC GCCGCAAGTACCAGGACAGCATCCGACGCATGGGCTTGGCCCCCAAGGACCCCGCTCTGCCTGGCTCCTCGGCTCCACAGCCCCCAGAGC CTGCCCAGGCCTAGGCCCCACAGACTTTTAGGCTGGCCCAGATATTCCCCAGTGGATGGGCAGAGCCCCCACCTTCAAGTCTCTCCAGTG >87880_87880_2_STXBP2-PIDD_STXBP2_chr19_7708131_ENST00000414284_PIDD_chr11_804463_ENST00000411829_length(amino acids)=1286AA_BP=366 MAPSGLKAVVGEKILSGVIRSVKKDGEWKVLIMDHPSMRILSSCCKMSDILAEGITIVEDINKRREPIPSLEAIYLLSPTEKALIKDFQG TPTFTYKAAHIFFTDTCPEPLFSELGRSRLAKVVKTLKEIHLAFLPYEAQVFSLDAPHSTYNLYCPFRAEERTRQLEVLAQQIATLCATL QEYPAIRYRKGPEDTAQLAHAVLAKLNAFKADTPSLGEGPEKTRSQLLIMDRAADPVSPLLHELTFQAMAYDLLDIEQDTYRYETTGLSE AREKAVLLDEDDDLWVELRHMHIADVSKKVTELLRTFCESKRLTTDKANIKDLSQILKKMPQYQKELNKYSTHLHLADDCMKHFKGSVEK LCSVEQLQLPRALPGQACLRAGTCLASKDRRWAMAATVEGPELEAAAAAGDASEDSDAGSRALPFLGGNRLSLDLYPGGCQQLLHLCVQQ PLQLLQVEFLRLSTHEDPQLLEATLAQLPQSLSCLRSLVLKGGQRRDTLGACLRGALTNLPAGLSGLAHLAHLDLSFNSLETLPACVLQM RGLGALLLSHNCLSELPEALGALPALTFLTVTHNRLQTLPPALGALSTLQRLDLSQNLLDTLPPEIGGLGSLLELNLASNRLQSLPASLA GLRSLRLLVLHSNLLASVPADLARLPLLTRLDLRDNQLRDLPPELLDAPFVRLQGNPLGEASPDAPSSPVAALIPEMPRLFLTSDLDSFP VTPQGCSVTLACGVRLQFPAGATATPITIRYRLLLPEPGLVPLGPHDALLSHVLELQPHGVAFQQDVGLWLLFTPPQARRCREVVVRTRN DNSWGDLETYLEEEAPQRLWAHCQVPHFSWFLVVSRPVSNACLVPPEGTLLCSSGHPGVKVIFPPGATEEPRRVSMQVVRMAGRELQALL GEPEAAVSPLLCLSQSGPPSFLQPVTVQLPLPSGITGLSLDRSRLHLLYWAPPAATWDDITAQVVLELTHLYARFQVTHFSWYWLWYTTK NCVGGLARKAWERLRLHRVNLIALQRRRDPEQVLLQCLPRNKVDATLRRLLERYRGPEPSDTVEMFEGEEFFAAFERGIDVDADRPDCVE GRICFVFYSHLKNVKEVSFYRGAVPVRVPEEAEAARQRKGADALWMATLPIKLPRLRGSEGPRRGAGLSLAPLNLGDAETGFLTQSNLLS VAGRLGLDWPAVALHLGVSYREVQRIRHEFRDDLDEQIRHMLFSWAERQAGQPGAVGLLVQALEQSDRQDVAEEVRAVLELGRRKYQDSI -------------------------------------------------------------- >87880_87880_3_STXBP2-PIDD_STXBP2_chr19_7708131_ENST00000441779_PIDD_chr11_804463_ENST00000411829_length(transcript)=4063nt_BP=1177nt ACACACCCGGAAGCGGCGGCGGCGCCCCTCGGGGAAGATGGCGCCCTCGGGGCTGAAGGCGGTGGTGGGGGAAAAAATTCTGAGCGGAGT TATTCGGAGTGTCAAGAAGGATGGGGAGTGGAAGGTGCTTATCATGGATCACCCAAGCATGCGCATCTTGTCTTCCTGCTGCAAAATGTC AGATATCCTGGCTGAGGGCATCACCATTGTTGAAGACATCAACAAACGGCGGGAACCCATTCCCAGTCTGGAGGCCATTTATTTGCTGAG CCCCACGGAGAAGGCTCAGGCCCAGAGAGTGATCCACCTTCCCCAGTCGGTTCAGGCCCTGATCAAAGACTTCCAGGGGACCCCGACTTT CACCTACAAAGCGGCCCATATCTTCTTCACCGACACCTGCCCCGAGCCCCTGTTCAGTGAGCTAGGCCGCTCTCGTCTGGCAAAGGTGGT GAAGACGTTGAAGGAGATTCACCTTGCCTTCCTCCCCTACGAGGCCCAGGTGTTCTCCCTCGATGCTCCCCACAGCACCTACAACCTCTA CTGCCCCTTCCGGGCAGAGGAGCGCACGCGGCAGCTCGAGGTGCTGGCCCAGCAGATTGCCACGCTGTGCGCCACCCTGCAGGAGTACCC GGCCATCCGCTACCGCAAGGGCCCAGAGGACACAGCCCAGTTGGCCCACGCCGTCCTGGCCAAGCTGAACGCCTTCAAGGCAGACACTCC CAGTCTGGGCGAGGGCCCAGAGAAAACCCGCTCCCAGCTGCTGATAATGGACCGGGCAGCTGACCCCGTGTCCCCACTACTGCATGAGCT CACGTTCCAGGCCATGGCGTATGATCTGCTGGACATAGAGCAGGACACATACAGGTATGAGACCACCGGGCTGAGCGAGGCGCGGGAGAA GGCCGTCTTGCTGGACGAGGACGATGACTTGTGGGTGGAGCTTCGCCACATGCATATCGCAGATGTGTCCAAGAAGGTCACGGAGCTCCT GAGGACCTTCTGTGAGAGCAAGAGGCTGACCACGGACAAGGCGAACATCAAAGACCTATCCCAGATCCTGAAAAAGATGCCGCAGTACCA GAAGGAGCTGAATAAGTATTCTACGCACCTGCATCTAGCAGATGATTGTATGAAGCACTTCAAGGGCTCGGTGGAGAAGCTGTGTAGTGT GGAGCAGCTGCAGCTTCCCCGGGCGCTGCCTGGACAGGCCTGCCTGCGTGCTGGGACATGTCTGGCCTCCAAGGACCGTCGGTGGGCGAT GGCTGCAACGGTGGAGGGGCCAGAGCTGGAGGCAGCTGCTGCCGCAGGAGATGCTTCAGAGGATTCGGACGCAGGGTCCAGGGCGCTGCC TTTCCTGGGCGGCAACCGGCTGAGCTTGGACCTGTACCCCGGGGGCTGCCAGCAGCTGCTGCACCTGTGTGTCCAGCAGCCTCTGCAGCT GCTGCAGGTGGAATTCTTGCGTCTGAGCACTCACGAGGACCCTCAGCTGCTGGAGGCCACCCTGGCCCAGCTGCCTCAGAGCCTGTCCTG CCTCCGCTCCCTGGTCCTCAAAGGAGGGCAACGCCGGGACACACTGGGTGCCTGTCTCCGGGGTGCCCTGACCAACCTGCCCGCTGGTCT GAGTGGCCTGGCCCATCTGGCCCACCTGGACCTGAGCTTCAACAGCCTGGAGACACTGCCGGCCTGTGTCCTGCAGATGCGAGGTCTGGG TGCGCTCTTGCTGTCTCACAACTGCCTCTCTGAGCTGCCTGAGGCTCTGGGGGCCCTCCCCGCCCTCACCTTCCTCACAGTGACACACAA CCGCCTGCAGACGCTGCCCCCAGCACTGGGGGCCCTATCCACCCTGCAGCGCCTCGATCTCTCTCAGAATCTGCTGGACACGCTACCTCC TGAGATTGGAGGCCTGGGCAGCCTCCTGGAGCTCAACCTGGCCTCCAACCGGCTGCAGAGCCTCCCAGCCTCTCTGGCGGGACTTCGGTC CTTGCGGCTCCTTGTCCTGCACAGCAACCTCCTGGCCTCTGTGCCAGCTGACTTGGCCCGCCTTCCACTCCTCACCCGGCTCGACCTGAG GGACAACCAGCTCCGGGACCTGCCCCCTGAGCTGCTAGACGCCCCCTTTGTGCGCCTGCAGGGGAACCCCCTGGGTGAGGCCTCGCCAGA CGCCCCGAGTTCACCAGTGGCAGCCCTCATTCCAGAAATGCCCAGACTGTTCCTGACCTCAGATTTGGACAGCTTTCCTGTGACCCCTCA AGGCTGCTCAGTGACCCTGGCCTGTGGCGTCCGCCTGCAGTTCCCAGCGGGAGCCACCGCCACCCCCATCACCATCCGCTATCGGCTGCT GCTGCCGGAGCCAGGCCTCGTCCCCCTGGGTCCTCATGACGCCCTGCTCAGCCATGTGCTGGAGCTGCAGCCCCATGGGGTGGCCTTCCA GCAGGATGTGGGGCTGTGGCTGCTCTTCACCCCACCGCAGGCCCGGCGCTGCCGTGAAGTGGTGGTCAGGACCCGGAATGACAACAGCTG GGGTGACCTGGAGACCTACCTGGAGGAAGAGGCACCCCAGCGGCTCTGGGCTCACTGCCAGGTGCCCCACTTCTCCTGGTTCCTTGTGGT TTCCCGCCCTGTGTCCAATGCCTGCCTGGTGCCACCGGAGGGGACACTGCTGTGCTCCTCGGGTCATCCTGGGGTCAAAGTCATCTTCCC CCCTGGGGCCACTGAGGAGCCTCGTCGAGTCTCCATGCAGGTGGTGCGCATGGCTGGCCGAGAGCTGCAGGCCCTCCTGGGAGAACCAGA GGCTGCAGTGAGCCCCCTGCTGTGCCTGTCACAGAGCGGTCCCCCCAGCTTCCTCCAACCGGTCACCGTGCAGCTGCCTCTGCCCTCTGG CATCACAGGCCTCAGTCTGGACCGCTCCCGCCTGCACCTGTTGTACTGGGCCCCTCCTGCAGCCACCTGGGATGACATCACAGCTCAGGT GGTCCTGGAGCTCACCCACCTGTACGCACGCTTCCAGGTCACACACTTCTCCTGGTACTGGCTCTGGTACACCACCAAGAACTGTGTGGG AGGCCTGGCTCGGAAGGCCTGGGAGCGGCTGCGGCTGCACCGTGTGAACCTCATCGCTCTGCAGCGGCGCCGGGACCCTGAGCAGGTCCT GCTGCAGTGCCTGCCCCGAAACAAGGTGGACGCCACCCTTCGGCGGCTGCTGGAGCGGTACCGGGGCCCCGAGCCCTCTGACACGGTGGA GATGTTCGAGGGCGAAGAGTTCTTTGCGGCCTTCGAGCGCGGCATCGACGTGGATGCTGACCGCCCTGACTGTGTGGAGGGCAGAATCTG CTTTGTCTTCTACTCGCACCTGAAGAATGTGAAGGAGGTGTCCTTCTACCGTGGCGCGGTGCCTGTGCGGGTGCCCGAGGAGGCTGAGGC TGCCCGGCAGAGGAAGGGCGCAGACGCCCTGTGGATGGCCACTCTGCCCATCAAGCTGCCGAGACTTCGAGGGTCCGAGGGGCCACGGCG GGGGGCTGGCCTCTCCTTGGCACCCTTGAATCTGGGAGATGCCGAGACCGGCTTTCTGACGCAGAGCAACCTGCTGAGTGTGGCTGGGCG TCTGGGTCTGGACTGGCCAGCCGTGGCCCTGCACCTGGGGGTGTCCTACCGGGAGGTGCAGCGCATCCGGCACGAGTTCCGGGATGATCT GGATGAGCAGATCCGTCACATGCTCTTCTCCTGGGCTGAGCGCCAGGCTGGGCAGCCAGGGGCTGTGGGGCTCCTGGTGCAGGCCCTGGA GCAGAGTGACCGGCAGGACGTGGCTGAAGAGGTGCGCGCAGTCTTGGAGCTCGGCCGCCGCAAGTACCAGGACAGCATCCGACGCATGGG CTTGGCCCCCAAGGACCCCGCTCTGCCTGGCTCCTCGGCTCCACAGCCCCCAGAGCCTGCCCAGGCCTAGGCCCCACAGACTTTTAGGCT GGCCCAGATATTCCCCAGTGGATGGGCAGAGCCCCCACCTTCAAGTCTCTCCAGTGTGTGGGGACGGGTCCCTGTGAGCAACAAAACTGC >87880_87880_3_STXBP2-PIDD_STXBP2_chr19_7708131_ENST00000441779_PIDD_chr11_804463_ENST00000411829_length(amino acids)=1300AA_BP=380 MAPSGLKAVVGEKILSGVIRSVKKDGEWKVLIMDHPSMRILSSCCKMSDILAEGITIVEDINKRREPIPSLEAIYLLSPTEKAQAQRVIH LPQSVQALIKDFQGTPTFTYKAAHIFFTDTCPEPLFSELGRSRLAKVVKTLKEIHLAFLPYEAQVFSLDAPHSTYNLYCPFRAEERTRQL EVLAQQIATLCATLQEYPAIRYRKGPEDTAQLAHAVLAKLNAFKADTPSLGEGPEKTRSQLLIMDRAADPVSPLLHELTFQAMAYDLLDI EQDTYRYETTGLSEAREKAVLLDEDDDLWVELRHMHIADVSKKVTELLRTFCESKRLTTDKANIKDLSQILKKMPQYQKELNKYSTHLHL ADDCMKHFKGSVEKLCSVEQLQLPRALPGQACLRAGTCLASKDRRWAMAATVEGPELEAAAAAGDASEDSDAGSRALPFLGGNRLSLDLY PGGCQQLLHLCVQQPLQLLQVEFLRLSTHEDPQLLEATLAQLPQSLSCLRSLVLKGGQRRDTLGACLRGALTNLPAGLSGLAHLAHLDLS FNSLETLPACVLQMRGLGALLLSHNCLSELPEALGALPALTFLTVTHNRLQTLPPALGALSTLQRLDLSQNLLDTLPPEIGGLGSLLELN LASNRLQSLPASLAGLRSLRLLVLHSNLLASVPADLARLPLLTRLDLRDNQLRDLPPELLDAPFVRLQGNPLGEASPDAPSSPVAALIPE MPRLFLTSDLDSFPVTPQGCSVTLACGVRLQFPAGATATPITIRYRLLLPEPGLVPLGPHDALLSHVLELQPHGVAFQQDVGLWLLFTPP QARRCREVVVRTRNDNSWGDLETYLEEEAPQRLWAHCQVPHFSWFLVVSRPVSNACLVPPEGTLLCSSGHPGVKVIFPPGATEEPRRVSM QVVRMAGRELQALLGEPEAAVSPLLCLSQSGPPSFLQPVTVQLPLPSGITGLSLDRSRLHLLYWAPPAATWDDITAQVVLELTHLYARFQ VTHFSWYWLWYTTKNCVGGLARKAWERLRLHRVNLIALQRRRDPEQVLLQCLPRNKVDATLRRLLERYRGPEPSDTVEMFEGEEFFAAFE RGIDVDADRPDCVEGRICFVFYSHLKNVKEVSFYRGAVPVRVPEEAEAARQRKGADALWMATLPIKLPRLRGSEGPRRGAGLSLAPLNLG DAETGFLTQSNLLSVAGRLGLDWPAVALHLGVSYREVQRIRHEFRDDLDEQIRHMLFSWAERQAGQPGAVGLLVQALEQSDRQDVAEEVR -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for STXBP2-PIDD |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for STXBP2-PIDD |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for STXBP2-PIDD |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
