
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:BAZ1A-LRRC9 (FusionGDB2 ID:9013) |
Fusion Gene Summary for BAZ1A-LRRC9 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: BAZ1A-LRRC9 | Fusion gene ID: 9013 | Hgene | Tgene | Gene symbol | BAZ1A | LRRC9 | Gene ID | 11177 | 341883 |
| Gene name | bromodomain adjacent to zinc finger domain 1A | leucine rich repeat containing 9 | |
| Synonyms | ACF1|WALp1|WCRF180|hACF1 | - | |
| Cytomap | 14q13.1-q13.2 | 14q23.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | bromodomain adjacent to zinc finger domain protein 1AATP-dependent chromatin remodeling proteinATP-utilizing chromatin assembly and remodeling factor 1CHRAC subunit ACF1hWALp1williams syndrome transcription factor-related chromatin-remodeling factor | leucine-rich repeat-containing protein 9 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q9NRL2 | Q6ZRR7 | |
| Ensembl transtripts involved in fusion gene | ENST00000358716, ENST00000360310, ENST00000382422, ENST00000553853, | ENST00000454474, ENST00000445360, | |
| Fusion gene scores | * DoF score | 12 X 9 X 5=540 | 5 X 5 X 3=75 |
| # samples | 12 | 4 | |
| ** MAII score | log2(12/540*10)=-2.16992500144231 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/75*10)=-0.906890595608519 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: BAZ1A [Title/Abstract] AND LRRC9 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | BAZ1A(35331250)-LRRC9(60444735), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | BAZ1A-LRRC9 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. BAZ1A-LRRC9 seems lost the major protein functional domain in Hgene partner, which is a epigenetic factor due to the frame-shifted ORF. BAZ1A-LRRC9 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. BAZ1A-LRRC9 seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. BAZ1A-LRRC9 seems lost the major protein functional domain in Hgene partner, which is a kinase due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | BAZ1A | GO:0006261 | DNA-dependent DNA replication | 12434153 |
 Fusion gene breakpoints across BAZ1A (5'-gene) Fusion gene breakpoints across BAZ1A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
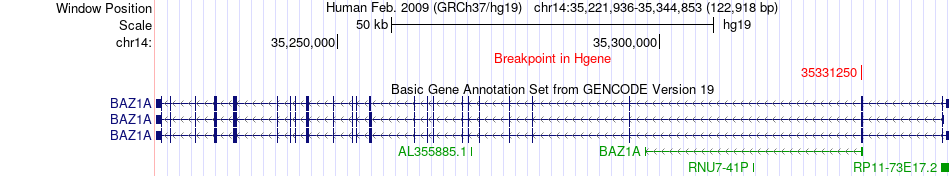 |
 Fusion gene breakpoints across LRRC9 (3'-gene) Fusion gene breakpoints across LRRC9 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
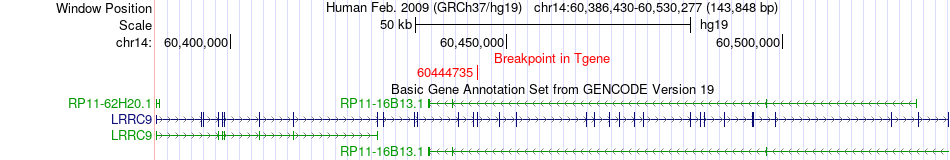 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUAD | TCGA-78-8660-01A | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
Top |
Fusion Gene ORF analysis for BAZ1A-LRRC9 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000358716 | ENST00000454474 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| 5CDS-intron | ENST00000360310 | ENST00000454474 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| 5CDS-intron | ENST00000382422 | ENST00000454474 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| 5UTR-3CDS | ENST00000553853 | ENST00000445360 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| 5UTR-intron | ENST00000553853 | ENST00000454474 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| Frame-shift | ENST00000382422 | ENST00000445360 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| In-frame | ENST00000358716 | ENST00000445360 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
| In-frame | ENST00000360310 | ENST00000445360 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000358716 | BAZ1A | chr14 | 35331250 | - | ENST00000445360 | LRRC9 | chr14 | 60444735 | + | 3711 | 960 | 872 | 3559 | 895 |
| ENST00000360310 | BAZ1A | chr14 | 35331250 | - | ENST00000445360 | LRRC9 | chr14 | 60444735 | + | 3711 | 960 | 872 | 3559 | 895 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000358716 | ENST00000445360 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + | 0.00031719 | 0.9996828 |
| ENST00000360310 | ENST00000445360 | BAZ1A | chr14 | 35331250 | - | LRRC9 | chr14 | 60444735 | + | 0.00031719 | 0.9996828 |
Top |
Fusion Genomic Features for BAZ1A-LRRC9 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| BAZ1A | chr14 | 35331249 | - | LRRC9 | chr14 | 60444734 | + | 0.000267364 | 0.9997327 |
| BAZ1A | chr14 | 35331249 | - | LRRC9 | chr14 | 60444734 | + | 0.000267364 | 0.9997327 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
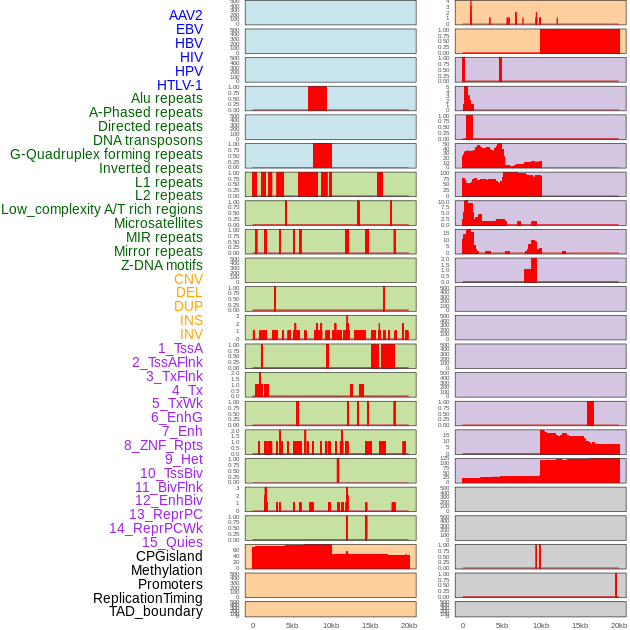 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
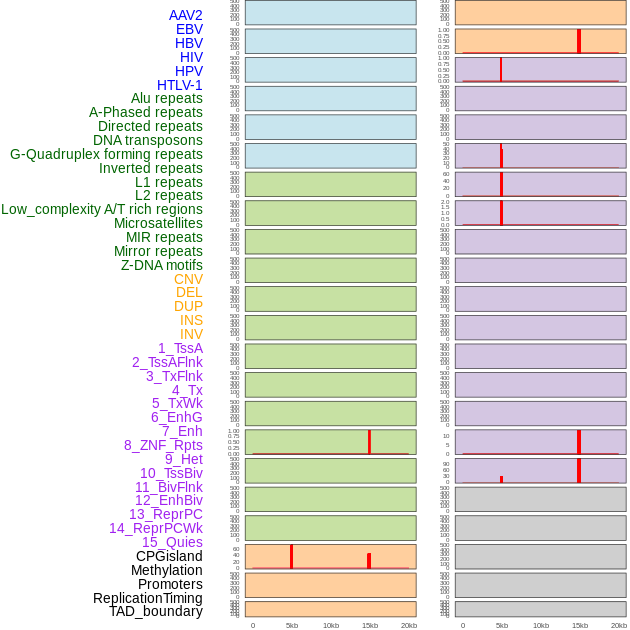 |
Top |
Fusion Protein Features for BAZ1A-LRRC9 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr14:35331250/chr14:60444735) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| BAZ1A | LRRC9 |
| FUNCTION: Component of the ACF complex, an ATP-dependent chromatin remodeling complex, that regulates spacing of nucleosomes using ATP to generate evenly spaced nucleosomes along the chromatin. The ATPase activity of the complex is regulated by the length of flanking DNA. Also involved in facilitating the DNA replication process. BAZ1A is the accessory, non-catalytic subunit of the complex which can enhance and direct the process provided by the ATPase subunit, SMARCA5, probably through targeting pericentromeric heterochromatin in late S phase. Moves end-positioned nucleosomes to a predominantly central position. May have a role in nuclear receptor-mediated transcription repression.; FUNCTION: Component of the histone-fold protein complex CHRAC complex which facilitates nucleosome sliding by the ACF complex and enhances ACF-mediated chromatin assembly. The C-terminal regions of both CHRAC1 and POLE1 are required for these functions. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 22_128 | 130 | 1525.0 | Domain | WAC |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 22_128 | 130 | 1557.0 | Domain | WAC |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 22_128 | 130 | 1557.0 | Domain | WAC |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 1_128 | 130 | 1525.0 | Region | Note=Required for association with the CHRAC1/POLE3 complex |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 1_128 | 130 | 1557.0 | Region | Note=Required for association with the CHRAC1/POLE3 complex |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 1_128 | 130 | 1557.0 | Region | Note=Required for association with the CHRAC1/POLE3 complex |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1023_1048 | 587 | 1454.0 | Repeat | Note=LRR 20 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1092_1115 | 587 | 1454.0 | Repeat | Note=LRR 21 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1116_1138 | 587 | 1454.0 | Repeat | Note=LRR 22 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1139_1161 | 587 | 1454.0 | Repeat | Note=LRR 23 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1201_1224 | 587 | 1454.0 | Repeat | Note=LRR 24 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1225_1247 | 587 | 1454.0 | Repeat | Note=LRR 25 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1248_1270 | 587 | 1454.0 | Repeat | Note=LRR 26 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1272_1292 | 587 | 1454.0 | Repeat | Note=LRR 27 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1293_1317 | 587 | 1454.0 | Repeat | Note=LRR 28 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1319_1345 | 587 | 1454.0 | Repeat | Note=LRR 29 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 1365_1388 | 587 | 1454.0 | Repeat | Note=LRR 30 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 671_693 | 587 | 1454.0 | Repeat | Note=LRR 9 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 694_715 | 587 | 1454.0 | Repeat | Note=LRR 10 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 716_737 | 587 | 1454.0 | Repeat | Note=LRR 11 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 739_758 | 587 | 1454.0 | Repeat | Note=LRR 12 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 759_784 | 587 | 1454.0 | Repeat | Note=LRR 13 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 786_812 | 587 | 1454.0 | Repeat | Note=LRR 14 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 886_908 | 587 | 1454.0 | Repeat | Note=LRR 15 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 909_930 | 587 | 1454.0 | Repeat | Note=LRR 16 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 931_952 | 587 | 1454.0 | Repeat | Note=LRR 17 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 953_975 | 587 | 1454.0 | Repeat | Note=LRR 18 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 976_1001 | 587 | 1454.0 | Repeat | Note=LRR 19 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 306_397 | 130 | 1525.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 634_709 | 130 | 1525.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 306_397 | 130 | 1557.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 634_709 | 130 | 1557.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 306_397 | 130 | 1557.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 634_709 | 130 | 1557.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 1239_1257 | 130 | 1525.0 | Compositional bias | Note=Glu-rich |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 487_491 | 130 | 1525.0 | Compositional bias | Note=Poly-Glu |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 1239_1257 | 130 | 1557.0 | Compositional bias | Note=Glu-rich |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 487_491 | 130 | 1557.0 | Compositional bias | Note=Poly-Glu |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 1239_1257 | 130 | 1557.0 | Compositional bias | Note=Glu-rich |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 487_491 | 130 | 1557.0 | Compositional bias | Note=Poly-Glu |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 1446_1516 | 130 | 1525.0 | Domain | Bromo |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 422_487 | 130 | 1525.0 | Domain | DDT |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 1446_1516 | 130 | 1557.0 | Domain | Bromo |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 422_487 | 130 | 1557.0 | Domain | DDT |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 1446_1516 | 130 | 1557.0 | Domain | Bromo |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 422_487 | 130 | 1557.0 | Domain | DDT |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 1148_1198 | 130 | 1525.0 | Zinc finger | PHD-type |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 1148_1198 | 130 | 1557.0 | Zinc finger | PHD-type |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 1148_1198 | 130 | 1557.0 | Zinc finger | PHD-type |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 120_141 | 587 | 1454.0 | Repeat | Note=LRR 3 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 142_164 | 587 | 1454.0 | Repeat | Note=LRR 4 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 166_188 | 587 | 1454.0 | Repeat | Note=LRR 5 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 224_248 | 587 | 1454.0 | Repeat | Note=LRR 6 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 264_287 | 587 | 1454.0 | Repeat | Note=LRR 7 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 344_367 | 587 | 1454.0 | Repeat | Note=LRR 8 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 53_78 | 587 | 1454.0 | Repeat | Note=LRR 1 | |
| Tgene | LRRC9 | chr14:35331250 | chr14:60444735 | ENST00000445360 | 13 | 32 | 97_119 | 587 | 1454.0 | Repeat | Note=LRR 2 |
Top |
Fusion Gene Sequence for BAZ1A-LRRC9 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >9013_9013_1_BAZ1A-LRRC9_BAZ1A_chr14_35331250_ENST00000358716_LRRC9_chr14_60444735_ENST00000445360_length(transcript)=3711nt_BP=960nt CTTTTCCCATCGTGTAGTCAAGAGTCTGTGCCAGACTTGAAGGCTTTACTTTGTTAGCCATGTGTTTATGAACCCCCAGCGCTTTCCCTA GATCTTTTGGCTGATAATCTCAAACATGGAGGATGCTTCTGAATCTTCACGAGGGGTTGCTCCATTAATTAATAATGTAGTTCTCCCAGG CTCTCCGCTGTCTCTTCCTGTATCAGTGACAGGCTGTAAAAGTCATCGAGTAGCCAATAAAAAGGTAGAAGCGAGGAGTGAAAAGCTCCT CCCAACAGCTCTTCCTCCTTCAGAGCCGAAAGTAGATCAGAAACTTCCCAGGAGCTCCGAGAGGCGGGGAAGTGGCGGTGGGACGCAATT CCCCGCGCGGAGTCGGGCAGTGGCAGCGGGAGAAGCGGCAGCCAGGGGCGCGGCGGGGCCGGAGAGAGGCGGTCCCCTGGGAGGACGGGG TCTCCCCTCGTTGCCTTTGTAGTGGAGAAGGTGGACAAGTGGCAGTCGGCGTGATCGCAGGGAAGCGGGGCCGGCGCGGGCGGCCGAGGG TCCAGGCGAGCCCGCGGGCGGACGGGAGATGCCGCTGCTACACCGAAAGCCGTTTGTGAGACAGAAGCCGCCCGCGGACCTGCGGCCCGA CGAGGAAGTTTTCTACTGTAAAGTCACCAACGAGATCTTCCGCCACTACGATGACTTTTTTGAACGAACCATTCTGTGCAACAGCCTTGT GTGGAGTTGTGCTGTGACGGGTAGACCTGGACTGACGTATCAGGAAGCACTTGAGTCAGAAAAAAAAGCAAGACAGAATCTTCAGAGTTT TCCAGAACCACTAATTATTCCAGTTTTATACTTGACCAGCCTTACCCATCGTTCGCGCTTACATGAAATTTGTGATGATATCTTTGCATA TGTCAAGGATCGATATTTTGTCGAAGAAACTGTGGAAGTCATTAGGAACAATGGTGCAAGATTCTGTCATGGGACAAAGAAACTGTGATT GCAGTGTTCGGCAGTGCAAGTGGTTTGTCTTTGATCATGACCTTGTTTTGCCGGAATATGTTGTTGAATTTGAGTATATTACAATGGTCA AGGCACCCTCTTTATTTTCTGTATTCAACAATGTTATTCTAGAAGAAAGCAAAAAAAACCCAGAAGTATCAGTATTTTCCAAGGATTTGA AATTTGATGATGAAGTTATAAAAATGGAGCCCAGAATCAAGGCCCGACCAAAACTGATTAGTTTGGATGATAAAACAATACTTTCCCTTG CAAAGACTAGCGTTTACAGTCATATTGTGAGTCTGAATCTACATGGAAACAGCTTGAGTAAATTGAGAGATCTCTCCAAGTTAACAGGAC TTCGAAAACTAAACATTAGCTTTAATGAATTTACCTGTTTAGATGATGTATACCACTTGTATAACCTTGAATATTTGGATGCAAGCCATA ACCATGTGATAACCCTTGAGGGATTTAGAGGTCTGATGAAACTGAAGCACTTAGACTTGAGCTGGAATCAACTGAAAAAATCTGGCAATG AAATAAATATGTTATGCAAACACACCACAAGCCTTCTCACTCTTGATATTCAACATAATCCATGGCAAAAGCCAGCCACATTGAGGCTGA GTGTTATTGGCAGATTAAAGACTCTTACCCATTTAAATGGAGTCTTCATTTCTGAAGAAGAAGCAACAGCAGCTATGAAATTTATTGCAG GAACAAGGATTACTCAGTTATCTCTCTTACGGCATTCTAGTACTAAAGAGGAGAGACCACGGATACTTAGTATATGGCCTTCTGCCAAAA TTCTGACGCAGGTTTCAAAGTTAGGACCTCATTTACATCTGAGTGGAAACTGCTACTTGAAGATAACTGCCTTAAATTTAGATGGACAAC ATCTTTTTGAAATCACAAATTTAGAAAAATTGGAAAATTTGAAATGGGCATCATTCAGCAATAATAATCTCACTAAAATGGAGGGTCTGG AATCCTGTATTAACTTGGAAGAGCTCACATTAGATGGAAACTGCATCTCAAAGATAGAAGGTATTTCCAAGATGACTAAATTAACTCGCC TCAGCATAAATAATAATCTTCTCACTGGTTGGGAAGAGCATACCTTTGATAACATGCTTCATCTTCACTCGCTTTCTCTTGAGAACAACA GAATCACTTCATTGAGTGGTTTACAAAAATCTTTTACTCTTGTTGAGTTATACATAAGCAATAACTATATTGCTGTCAATCAGGAGATGC ACAATTTAAAGGGTTTATGCAACTTGGTCATTCTAGACATGTGTGGGAACATCATTATATGGAATCAAGAAAACTACCGGTTGTTTGTAA TATTTCATCTTCCAGAACTGAAAGCTTTGGATGGGATACCAATTGAACCATCAGAGACTGACAGTGCAAAAGATTTGTTTGGTGGTAGAC TTACTTCTGACATGATTGCAGAACGACAAGGACATTCCAACTTCAAACAAATGCAAGAACTAAATTGGACTTCATCATCTATTAGAACAG TGGATTTGATTCCAGTCGACCAATTTAGAAATGTGTGTAATGTGAATCTACAGAACAATCATCTTACCTCATTCAGTGGATTAATCTATC TACCTAATGTGAAGGTCTTATGCCTTAACTATAATCATATTGAATCAATCATGCCCAGACTAAAGCCTCAGACTCACTTAACCAGTAGAC AACTACTGTACCAGAAAGTGCCTTCAAGTGGCTATGGACAGCAAGGAATTTCAAAAACAAACAGAGATATAATGAGCAGTGAAAATTTGC CTCCAATAATGCACAGTTTAGAAGTTCTTCATTTGGGCTACAATGGAATTTGTAATTTGATCCAGCTACAACTTAACAGACTAAGAAATT TAAAATTCCTCTTTCTACAGGGTAATGAAATCAGCCAAGTTGAAGGGCTTGACAACTTAGTAGTCCTTCAAGAATTGGTAGTGGACCATA ACCGCATCCGATCATTTAATGACAGTGCTTTTGCCAAACCAAGTTCTTTATTGGCACTTCACTTGGAGGAAAACAGACTACGAGAACTGG GCAAATTACAATCTTTGGTAAAACTAGAGAAACTCTTCCTAGGATATAATAAAATCCAGGATATCACAGAACTGGAAAAACTTGACGTTA TCTCTACTCTCAGGGAGCTTACAGTGTATGGCAATCCAATTTGTAGAAAAATGCTACATCGCCACATGCTTATATTTCGACTGCCTAACT TACAGATGTTAGATGGAAGTCCTGTGAATTCAGATGATAGGGCAAAAGCTGAATTTCACCTCGCTGAACTACAAGCAAAGAAAAACTCGC TTATTCCTGTTACCCACTCTCCTATGGATGGCAGATCATTTGGCCAAGTGAAAACTCCTCCCATAGAGATAACAAATGTACTTCTGCCTA GTGGATTCAGCCATTACTTGGGATCTGACGTCACTTTAACTCCTGAAGTTGAAGAATTTTTGGGAGCAACTTTCCAAGATCAAATCGAAT GTAACTGCCTAAAGAGAAATGAGCATACACCAAGAAACTCACCTGTTTAGTAGTTGTACAAAAACTAGCGGACAGTGGTCAGTTAAAAAA AAAAAAAAAAGATAATCATGTTACTTAAGTTACTGAAGCACTTCAGTTCCATATGAAGATAATCTATTACCCTTCATGAACTTGATTTAA >9013_9013_1_BAZ1A-LRRC9_BAZ1A_chr14_35331250_ENST00000358716_LRRC9_chr14_60444735_ENST00000445360_length(amino acids)=895AA_BP=29 MKFVMISLHMSRIDILSKKLWKSLGTMVQDSVMGQRNCDCSVRQCKWFVFDHDLVLPEYVVEFEYITMVKAPSLFSVFNNVILEESKKNP EVSVFSKDLKFDDEVIKMEPRIKARPKLISLDDKTILSLAKTSVYSHIVSLNLHGNSLSKLRDLSKLTGLRKLNISFNEFTCLDDVYHLY NLEYLDASHNHVITLEGFRGLMKLKHLDLSWNQLKKSGNEINMLCKHTTSLLTLDIQHNPWQKPATLRLSVIGRLKTLTHLNGVFISEEE ATAAMKFIAGTRITQLSLLRHSSTKEERPRILSIWPSAKILTQVSKLGPHLHLSGNCYLKITALNLDGQHLFEITNLEKLENLKWASFSN NNLTKMEGLESCINLEELTLDGNCISKIEGISKMTKLTRLSINNNLLTGWEEHTFDNMLHLHSLSLENNRITSLSGLQKSFTLVELYISN NYIAVNQEMHNLKGLCNLVILDMCGNIIIWNQENYRLFVIFHLPELKALDGIPIEPSETDSAKDLFGGRLTSDMIAERQGHSNFKQMQEL NWTSSSIRTVDLIPVDQFRNVCNVNLQNNHLTSFSGLIYLPNVKVLCLNYNHIESIMPRLKPQTHLTSRQLLYQKVPSSGYGQQGISKTN RDIMSSENLPPIMHSLEVLHLGYNGICNLIQLQLNRLRNLKFLFLQGNEISQVEGLDNLVVLQELVVDHNRIRSFNDSAFAKPSSLLALH LEENRLRELGKLQSLVKLEKLFLGYNKIQDITELEKLDVISTLRELTVYGNPICRKMLHRHMLIFRLPNLQMLDGSPVNSDDRAKAEFHL -------------------------------------------------------------- >9013_9013_2_BAZ1A-LRRC9_BAZ1A_chr14_35331250_ENST00000360310_LRRC9_chr14_60444735_ENST00000445360_length(transcript)=3711nt_BP=960nt CTTTTCCCATCGTGTAGTCAAGAGTCTGTGCCAGACTTGAAGGCTTTACTTTGTTAGCCATGTGTTTATGAACCCCCAGCGCTTTCCCTA GATCTTTTGGCTGATAATCTCAAACATGGAGGATGCTTCTGAATCTTCACGAGGGGTTGCTCCATTAATTAATAATGTAGTTCTCCCAGG CTCTCCGCTGTCTCTTCCTGTATCAGTGACAGGCTGTAAAAGTCATCGAGTAGCCAATAAAAAGGTAGAAGCGAGGAGTGAAAAGCTCCT CCCAACAGCTCTTCCTCCTTCAGAGCCGAAAGTAGATCAGAAACTTCCCAGGAGCTCCGAGAGGCGGGGAAGTGGCGGTGGGACGCAATT CCCCGCGCGGAGTCGGGCAGTGGCAGCGGGAGAAGCGGCAGCCAGGGGCGCGGCGGGGCCGGAGAGAGGCGGTCCCCTGGGAGGACGGGG TCTCCCCTCGTTGCCTTTGTAGTGGAGAAGGTGGACAAGTGGCAGTCGGCGTGATCGCAGGGAAGCGGGGCCGGCGCGGGCGGCCGAGGG TCCAGGCGAGCCCGCGGGCGGACGGGAGATGCCGCTGCTACACCGAAAGCCGTTTGTGAGACAGAAGCCGCCCGCGGACCTGCGGCCCGA CGAGGAAGTTTTCTACTGTAAAGTCACCAACGAGATCTTCCGCCACTACGATGACTTTTTTGAACGAACCATTCTGTGCAACAGCCTTGT GTGGAGTTGTGCTGTGACGGGTAGACCTGGACTGACGTATCAGGAAGCACTTGAGTCAGAAAAAAAAGCAAGACAGAATCTTCAGAGTTT TCCAGAACCACTAATTATTCCAGTTTTATACTTGACCAGCCTTACCCATCGTTCGCGCTTACATGAAATTTGTGATGATATCTTTGCATA TGTCAAGGATCGATATTTTGTCGAAGAAACTGTGGAAGTCATTAGGAACAATGGTGCAAGATTCTGTCATGGGACAAAGAAACTGTGATT GCAGTGTTCGGCAGTGCAAGTGGTTTGTCTTTGATCATGACCTTGTTTTGCCGGAATATGTTGTTGAATTTGAGTATATTACAATGGTCA AGGCACCCTCTTTATTTTCTGTATTCAACAATGTTATTCTAGAAGAAAGCAAAAAAAACCCAGAAGTATCAGTATTTTCCAAGGATTTGA AATTTGATGATGAAGTTATAAAAATGGAGCCCAGAATCAAGGCCCGACCAAAACTGATTAGTTTGGATGATAAAACAATACTTTCCCTTG CAAAGACTAGCGTTTACAGTCATATTGTGAGTCTGAATCTACATGGAAACAGCTTGAGTAAATTGAGAGATCTCTCCAAGTTAACAGGAC TTCGAAAACTAAACATTAGCTTTAATGAATTTACCTGTTTAGATGATGTATACCACTTGTATAACCTTGAATATTTGGATGCAAGCCATA ACCATGTGATAACCCTTGAGGGATTTAGAGGTCTGATGAAACTGAAGCACTTAGACTTGAGCTGGAATCAACTGAAAAAATCTGGCAATG AAATAAATATGTTATGCAAACACACCACAAGCCTTCTCACTCTTGATATTCAACATAATCCATGGCAAAAGCCAGCCACATTGAGGCTGA GTGTTATTGGCAGATTAAAGACTCTTACCCATTTAAATGGAGTCTTCATTTCTGAAGAAGAAGCAACAGCAGCTATGAAATTTATTGCAG GAACAAGGATTACTCAGTTATCTCTCTTACGGCATTCTAGTACTAAAGAGGAGAGACCACGGATACTTAGTATATGGCCTTCTGCCAAAA TTCTGACGCAGGTTTCAAAGTTAGGACCTCATTTACATCTGAGTGGAAACTGCTACTTGAAGATAACTGCCTTAAATTTAGATGGACAAC ATCTTTTTGAAATCACAAATTTAGAAAAATTGGAAAATTTGAAATGGGCATCATTCAGCAATAATAATCTCACTAAAATGGAGGGTCTGG AATCCTGTATTAACTTGGAAGAGCTCACATTAGATGGAAACTGCATCTCAAAGATAGAAGGTATTTCCAAGATGACTAAATTAACTCGCC TCAGCATAAATAATAATCTTCTCACTGGTTGGGAAGAGCATACCTTTGATAACATGCTTCATCTTCACTCGCTTTCTCTTGAGAACAACA GAATCACTTCATTGAGTGGTTTACAAAAATCTTTTACTCTTGTTGAGTTATACATAAGCAATAACTATATTGCTGTCAATCAGGAGATGC ACAATTTAAAGGGTTTATGCAACTTGGTCATTCTAGACATGTGTGGGAACATCATTATATGGAATCAAGAAAACTACCGGTTGTTTGTAA TATTTCATCTTCCAGAACTGAAAGCTTTGGATGGGATACCAATTGAACCATCAGAGACTGACAGTGCAAAAGATTTGTTTGGTGGTAGAC TTACTTCTGACATGATTGCAGAACGACAAGGACATTCCAACTTCAAACAAATGCAAGAACTAAATTGGACTTCATCATCTATTAGAACAG TGGATTTGATTCCAGTCGACCAATTTAGAAATGTGTGTAATGTGAATCTACAGAACAATCATCTTACCTCATTCAGTGGATTAATCTATC TACCTAATGTGAAGGTCTTATGCCTTAACTATAATCATATTGAATCAATCATGCCCAGACTAAAGCCTCAGACTCACTTAACCAGTAGAC AACTACTGTACCAGAAAGTGCCTTCAAGTGGCTATGGACAGCAAGGAATTTCAAAAACAAACAGAGATATAATGAGCAGTGAAAATTTGC CTCCAATAATGCACAGTTTAGAAGTTCTTCATTTGGGCTACAATGGAATTTGTAATTTGATCCAGCTACAACTTAACAGACTAAGAAATT TAAAATTCCTCTTTCTACAGGGTAATGAAATCAGCCAAGTTGAAGGGCTTGACAACTTAGTAGTCCTTCAAGAATTGGTAGTGGACCATA ACCGCATCCGATCATTTAATGACAGTGCTTTTGCCAAACCAAGTTCTTTATTGGCACTTCACTTGGAGGAAAACAGACTACGAGAACTGG GCAAATTACAATCTTTGGTAAAACTAGAGAAACTCTTCCTAGGATATAATAAAATCCAGGATATCACAGAACTGGAAAAACTTGACGTTA TCTCTACTCTCAGGGAGCTTACAGTGTATGGCAATCCAATTTGTAGAAAAATGCTACATCGCCACATGCTTATATTTCGACTGCCTAACT TACAGATGTTAGATGGAAGTCCTGTGAATTCAGATGATAGGGCAAAAGCTGAATTTCACCTCGCTGAACTACAAGCAAAGAAAAACTCGC TTATTCCTGTTACCCACTCTCCTATGGATGGCAGATCATTTGGCCAAGTGAAAACTCCTCCCATAGAGATAACAAATGTACTTCTGCCTA GTGGATTCAGCCATTACTTGGGATCTGACGTCACTTTAACTCCTGAAGTTGAAGAATTTTTGGGAGCAACTTTCCAAGATCAAATCGAAT GTAACTGCCTAAAGAGAAATGAGCATACACCAAGAAACTCACCTGTTTAGTAGTTGTACAAAAACTAGCGGACAGTGGTCAGTTAAAAAA AAAAAAAAAAGATAATCATGTTACTTAAGTTACTGAAGCACTTCAGTTCCATATGAAGATAATCTATTACCCTTCATGAACTTGATTTAA >9013_9013_2_BAZ1A-LRRC9_BAZ1A_chr14_35331250_ENST00000360310_LRRC9_chr14_60444735_ENST00000445360_length(amino acids)=895AA_BP=29 MKFVMISLHMSRIDILSKKLWKSLGTMVQDSVMGQRNCDCSVRQCKWFVFDHDLVLPEYVVEFEYITMVKAPSLFSVFNNVILEESKKNP EVSVFSKDLKFDDEVIKMEPRIKARPKLISLDDKTILSLAKTSVYSHIVSLNLHGNSLSKLRDLSKLTGLRKLNISFNEFTCLDDVYHLY NLEYLDASHNHVITLEGFRGLMKLKHLDLSWNQLKKSGNEINMLCKHTTSLLTLDIQHNPWQKPATLRLSVIGRLKTLTHLNGVFISEEE ATAAMKFIAGTRITQLSLLRHSSTKEERPRILSIWPSAKILTQVSKLGPHLHLSGNCYLKITALNLDGQHLFEITNLEKLENLKWASFSN NNLTKMEGLESCINLEELTLDGNCISKIEGISKMTKLTRLSINNNLLTGWEEHTFDNMLHLHSLSLENNRITSLSGLQKSFTLVELYISN NYIAVNQEMHNLKGLCNLVILDMCGNIIIWNQENYRLFVIFHLPELKALDGIPIEPSETDSAKDLFGGRLTSDMIAERQGHSNFKQMQEL NWTSSSIRTVDLIPVDQFRNVCNVNLQNNHLTSFSGLIYLPNVKVLCLNYNHIESIMPRLKPQTHLTSRQLLYQKVPSSGYGQQGISKTN RDIMSSENLPPIMHSLEVLHLGYNGICNLIQLQLNRLRNLKFLFLQGNEISQVEGLDNLVVLQELVVDHNRIRSFNDSAFAKPSSLLALH LEENRLRELGKLQSLVKLEKLFLGYNKIQDITELEKLDVISTLRELTVYGNPICRKMLHRHMLIFRLPNLQMLDGSPVNSDDRAKAEFHL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for BAZ1A-LRRC9 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000358716 | - | 3 | 26 | 667_933 | 130.66666666666666 | 1525.0 | SMARCA5 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000360310 | - | 3 | 27 | 667_933 | 130.66666666666666 | 1557.0 | SMARCA5 |
| Hgene | BAZ1A | chr14:35331250 | chr14:60444735 | ENST00000382422 | - | 2 | 26 | 667_933 | 130.66666666666666 | 1557.0 | SMARCA5 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for BAZ1A-LRRC9 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for BAZ1A-LRRC9 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
