
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:TGFB1-TBC1D16 (FusionGDB2 ID:90431) |
Fusion Gene Summary for TGFB1-TBC1D16 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: TGFB1-TBC1D16 | Fusion gene ID: 90431 | Hgene | Tgene | Gene symbol | TGFB1 | TBC1D16 | Gene ID | 7040 | 125058 |
| Gene name | transforming growth factor beta 1 | TBC1 domain family member 16 | |
| Synonyms | CED|DPD1|IBDIMDE|LAP|TGF-beta1|TGFB|TGFbeta | - | |
| Cytomap | 19q13.2 | 17q25.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | transforming growth factor beta-1 proproteinTGF-beta-1latency-associated peptideprepro-transforming growth factor beta-1transforming growth factor beta1 | TBC1 domain family member 16CTD-2529O21.1 | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000221930, | ENST00000340848, ENST00000570373, ENST00000572862, ENST00000576768, ENST00000310924, | |
| Fusion gene scores | * DoF score | 7 X 7 X 6=294 | 7 X 6 X 4=168 |
| # samples | 9 | 7 | |
| ** MAII score | log2(9/294*10)=-1.70781924850669 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/168*10)=-1.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: TGFB1 [Title/Abstract] AND TBC1D16 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | TGFB1(41847787)-TBC1D16(77926617), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | TGFB1 | GO:0001837 | epithelial to mesenchymal transition | 25893292|29529050 |
| Hgene | TGFB1 | GO:0001933 | negative regulation of protein phosphorylation | 8053900 |
| Hgene | TGFB1 | GO:0001934 | positive regulation of protein phosphorylation | 18625725 |
| Hgene | TGFB1 | GO:0002062 | chondrocyte differentiation | 15040835 |
| Hgene | TGFB1 | GO:0002244 | hematopoietic progenitor cell differentiation | 15451575 |
| Hgene | TGFB1 | GO:0006611 | protein export from nucleus | 9770491|17438144|18588859 |
| Hgene | TGFB1 | GO:0006754 | ATP biosynthetic process | 10513816 |
| Hgene | TGFB1 | GO:0006796 | phosphate-containing compound metabolic process | 10513816 |
| Hgene | TGFB1 | GO:0006954 | inflammatory response | 21147091 |
| Hgene | TGFB1 | GO:0007050 | cell cycle arrest | 14555988 |
| Hgene | TGFB1 | GO:0007093 | mitotic cell cycle checkpoint | 15334054 |
| Hgene | TGFB1 | GO:0007173 | epidermal growth factor receptor signaling pathway | 18625725 |
| Hgene | TGFB1 | GO:0007179 | transforming growth factor beta receptor signaling pathway | 9389648|11157754 |
| Hgene | TGFB1 | GO:0007182 | common-partner SMAD protein phosphorylation | 20573232 |
| Hgene | TGFB1 | GO:0007183 | SMAD protein complex assembly | 17438144 |
| Hgene | TGFB1 | GO:0008284 | positive regulation of cell proliferation | 10513816|14633705 |
| Hgene | TGFB1 | GO:0008285 | negative regulation of cell proliferation | 15334054 |
| Hgene | TGFB1 | GO:0010628 | positive regulation of gene expression | 18625725|18832382|18941241|19913496|25322725|26634652|26687115|27162619|29167509 |
| Hgene | TGFB1 | GO:0010629 | negative regulation of gene expression | 19913496|20067797|22269326|25163461|26634652|29167509|29529050 |
| Hgene | TGFB1 | GO:0010718 | positive regulation of epithelial to mesenchymal transition | 17999987|18505915 |
| Hgene | TGFB1 | GO:0010763 | positive regulation of fibroblast migration | 18555217 |
| Hgene | TGFB1 | GO:0010800 | positive regulation of peptidyl-threonine phosphorylation | 18625725|19736306 |
| Hgene | TGFB1 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation | 9389648|19736306|26634652 |
| Hgene | TGFB1 | GO:0010936 | negative regulation of macrophage cytokine production | 20875417 |
| Hgene | TGFB1 | GO:0016477 | cell migration | 25893292 |
| Hgene | TGFB1 | GO:0017015 | regulation of transforming growth factor beta receptor signaling pathway | 15334054 |
| Hgene | TGFB1 | GO:0022408 | negative regulation of cell-cell adhesion | 18593713 |
| Hgene | TGFB1 | GO:0030214 | hyaluronan catabolic process | 17324121 |
| Hgene | TGFB1 | GO:0030308 | negative regulation of cell growth | 15334054 |
| Hgene | TGFB1 | GO:0030335 | positive regulation of cell migration | 19736306 |
| Hgene | TGFB1 | GO:0031293 | membrane protein intracellular domain proteolysis | 25310401 |
| Hgene | TGFB1 | GO:0031334 | positive regulation of protein complex assembly | 19366691 |
| Hgene | TGFB1 | GO:0031663 | lipopolysaccharide-mediated signaling pathway | 21147091 |
| Hgene | TGFB1 | GO:0032270 | positive regulation of cellular protein metabolic process | 15219857 |
| Hgene | TGFB1 | GO:0032355 | response to estradiol | 18039789 |
| Hgene | TGFB1 | GO:0032570 | response to progesterone | 18039789 |
| Hgene | TGFB1 | GO:0032740 | positive regulation of interleukin-17 production | 18453574 |
| Hgene | TGFB1 | GO:0032801 | receptor catabolic process | 17878231 |
| Hgene | TGFB1 | GO:0032930 | positive regulation of superoxide anion generation | 22073128 |
| Hgene | TGFB1 | GO:0032967 | positive regulation of collagen biosynthetic process | 19734317|22269326|25310401 |
| Hgene | TGFB1 | GO:0033138 | positive regulation of peptidyl-serine phosphorylation | 18625725|19736306 |
| Hgene | TGFB1 | GO:0035307 | positive regulation of protein dephosphorylation | 14555988 |
| Hgene | TGFB1 | GO:0042307 | positive regulation of protein import into nucleus | 19366691 |
| Hgene | TGFB1 | GO:0043117 | positive regulation of vascular permeability | 21168935 |
| Hgene | TGFB1 | GO:0043406 | positive regulation of MAP kinase activity | 18625725 |
| Hgene | TGFB1 | GO:0043536 | positive regulation of blood vessel endothelial cell migration | 18555217 |
| Hgene | TGFB1 | GO:0043537 | negative regulation of blood vessel endothelial cell migration | 18555217 |
| Hgene | TGFB1 | GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity | 18625725 |
| Hgene | TGFB1 | GO:0045216 | cell-cell junction organization | 18505915 |
| Hgene | TGFB1 | GO:0045599 | negative regulation of fat cell differentiation | 15040835 |
| Hgene | TGFB1 | GO:0045662 | negative regulation of myoblast differentiation | 9770491 |
| Hgene | TGFB1 | GO:0045786 | negative regulation of cell cycle | 11502704 |
| Hgene | TGFB1 | GO:0045892 | negative regulation of transcription, DNA-templated | 15702480|18832382 |
| Hgene | TGFB1 | GO:0045893 | positive regulation of transcription, DNA-templated | 9389648|14517293|15334054|16816361 |
| Hgene | TGFB1 | GO:0045918 | negative regulation of cytolysis | 24586048 |
| Hgene | TGFB1 | GO:0045930 | negative regulation of mitotic cell cycle | 14555988 |
| Hgene | TGFB1 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 18832382 |
| Hgene | TGFB1 | GO:0048298 | positive regulation of isotype switching to IgA isotypes | 14988498 |
| Hgene | TGFB1 | GO:0048642 | negative regulation of skeletal muscle tissue development | 9770491 |
| Hgene | TGFB1 | GO:0050680 | negative regulation of epithelial cell proliferation | 9950587 |
| Hgene | TGFB1 | GO:0050714 | positive regulation of protein secretion | 18505915 |
| Hgene | TGFB1 | GO:0050731 | positive regulation of peptidyl-tyrosine phosphorylation | 21168935 |
| Hgene | TGFB1 | GO:0050921 | positive regulation of chemotaxis | 18555217 |
| Hgene | TGFB1 | GO:0051897 | positive regulation of protein kinase B signaling | 18625725 |
| Hgene | TGFB1 | GO:0060389 | pathway-restricted SMAD protein phosphorylation | 11157754|17999987|18453574|25893292 |
| Hgene | TGFB1 | GO:0060390 | regulation of SMAD protein signal transduction | 25893292 |
| Hgene | TGFB1 | GO:0060391 | positive regulation of SMAD protein signal transduction | 9389648|19366691|29167509 |
| Hgene | TGFB1 | GO:0070168 | negative regulation of biomineral tissue development | 26634652 |
| Hgene | TGFB1 | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | 25310401 |
| Hgene | TGFB1 | GO:0070723 | response to cholesterol | 17878231 |
| Hgene | TGFB1 | GO:0071407 | cellular response to organic cyclic compound | 21147091 |
| Hgene | TGFB1 | GO:0071560 | cellular response to transforming growth factor beta stimulus | 19736306|22269326 |
| Hgene | TGFB1 | GO:0085029 | extracellular matrix assembly | 19734317 |
| Hgene | TGFB1 | GO:0090263 | positive regulation of canonical Wnt signaling pathway | 12893825|15040835 |
| Hgene | TGFB1 | GO:0097191 | extrinsic apoptotic signaling pathway | 15334054 |
| Hgene | TGFB1 | GO:1900126 | negative regulation of hyaluronan biosynthetic process | 17324121 |
| Hgene | TGFB1 | GO:1900182 | positive regulation of protein localization to nucleus | 26634652 |
| Hgene | TGFB1 | GO:1901666 | positive regulation of NAD+ ADP-ribosyltransferase activity | 22073128 |
| Hgene | TGFB1 | GO:1902895 | positive regulation of pri-miRNA transcription by RNA polymerase II | 26311719|26493107 |
| Hgene | TGFB1 | GO:1903077 | negative regulation of protein localization to plasma membrane | 21168935|24586048 |
| Hgene | TGFB1 | GO:1903800 | positive regulation of production of miRNAs involved in gene silencing by miRNA | 18548003 |
| Hgene | TGFB1 | GO:2000679 | positive regulation of transcription regulatory region DNA binding | 22073128 |
| Hgene | TGFB1 | GO:2000727 | positive regulation of cardiac muscle cell differentiation | 25163461 |
| Tgene | TBC1D16 | GO:0001919 | regulation of receptor recycling | 23019362 |
 Fusion gene breakpoints across TGFB1 (5'-gene) Fusion gene breakpoints across TGFB1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
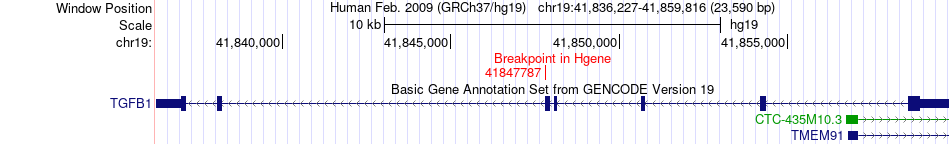 |
 Fusion gene breakpoints across TBC1D16 (3'-gene) Fusion gene breakpoints across TBC1D16 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
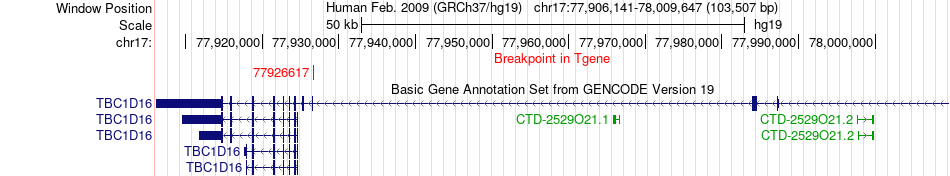 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-BR-8485 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - |
Top |
Fusion Gene ORF analysis for TGFB1-TBC1D16 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000221930 | ENST00000340848 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - |
| 5CDS-intron | ENST00000221930 | ENST00000570373 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - |
| 5CDS-intron | ENST00000221930 | ENST00000572862 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - |
| 5CDS-intron | ENST00000221930 | ENST00000576768 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - |
| In-frame | ENST00000221930 | ENST00000310924 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000221930 | TGFB1 | chr19 | 41847787 | - | ENST00000310924 | TBC1D16 | chr17 | 77926617 | - | 11768 | 1727 | 171 | 3251 | 1026 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000221930 | ENST00000310924 | TGFB1 | chr19 | 41847787 | - | TBC1D16 | chr17 | 77926617 | - | 0.003232018 | 0.996768 |
Top |
Fusion Genomic Features for TGFB1-TBC1D16 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
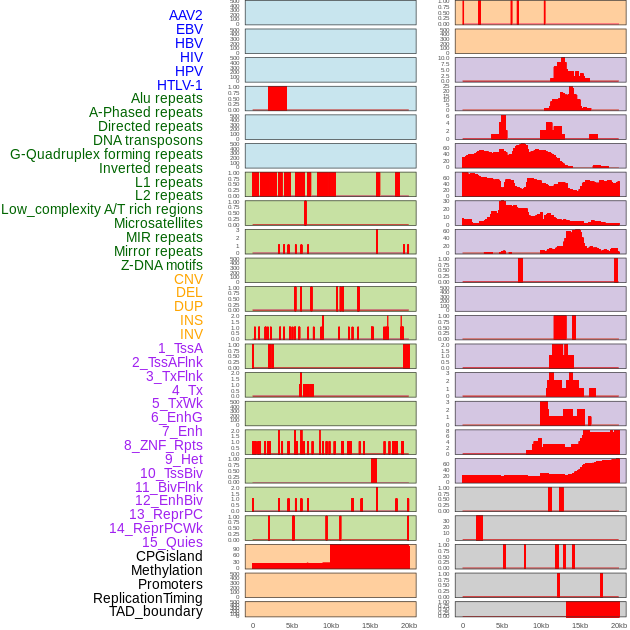 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for TGFB1-TBC1D16 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:41847787/chr17:77926617) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TGFB1 | chr19:41847787 | chr17:77926617 | ENST00000221930 | - | 5 | 7 | 244_246 | 286 | 391.0 | Motif | Cell attachment site |
| Hgene | TGFB1 | chr19:41847787 | chr17:77926617 | ENST00000221930 | - | 5 | 7 | 226_252 | 286 | 391.0 | Region | Bowtie tail |
| Hgene | TGFB1 | chr19:41847787 | chr17:77926617 | ENST00000221930 | - | 5 | 7 | 30_74 | 286 | 391.0 | Region | Straightjacket domain |
| Hgene | TGFB1 | chr19:41847787 | chr17:77926617 | ENST00000221930 | - | 5 | 7 | 75_271 | 286 | 391.0 | Region | Arm domain |
| Tgene | TBC1D16 | chr19:41847787 | chr17:77926617 | ENST00000310924 | 2 | 12 | 425_635 | 259 | 768.0 | Domain | Rab-GAP TBC |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | TBC1D16 | chr19:41847787 | chr17:77926617 | ENST00000310924 | 2 | 12 | 212_265 | 259 | 768.0 | Compositional bias | Note=Ser-rich |
Top |
Fusion Gene Sequence for TGFB1-TBC1D16 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >90431_90431_1_TGFB1-TBC1D16_TGFB1_chr19_41847787_ENST00000221930_TBC1D16_chr17_77926617_ENST00000310924_length(transcript)=11768nt_BP=1727nt CCTTCGCGCCCTGGGCCATCTCCCTCCCACCTCCCTCCGCGGAGCAGCCAGACAGCGAGGGCCCCGGCCGGGGGCAGGGGGGACGCCCCG TCCGGGGCACCCCCCCGGCTCTGAGCCGCCCGCGGGGCCGGCCTCGGCCCGGAGCGGAGGAAGGAGTCGCCGAGGAGCAGCCTGAGGCCC CAGAGTCTGAGACGAGCCGCCGCCGCCCCCGCCACTGCGGGGAGGAGGGGGAGGAGGAGCGGGAGGAGGGACGAGCTGGTCGGGAGAAGA GGAAAAAAACTTTTGAGACTTTTCCGTTGCCGCTGGGAGCCGGAGGCGCGGGGACCTCTTGGCGCGACGCTGCCCCGCGAGGAGGCAGGA CTTGGGGACCCCAGACCGCCTCCCTTTGCCGCCGGGGACGCTTGCTCCCTCCCTGCCCCCTACACGGCGTCCCTCAGGCGCCCCCATTCC GGACCAGCCCTCGGGAGTCGCCGACCCGGCCTCCCGCAAAGACTTTTCCCCAGACCTCGGGCGCACCCCCTGCACGCCGCCTTCATCCCC GGCCTGTCTCCTGAGCCCCCGCGCATCCTAGACCCTTTCTCCTCCAGGAGACGGATCTCTCTCCGACCTGCCACAGATCCCCTATTCAAG ACCACCCACCTTCTGGTACCAGATCGCGCCCATCTAGGTTATTTCCGTGGGATACTGAGACACCCCCGGTCCAAGCCTCCCCTCCACCAC TGCGCCCTTCTCCCTGAGGACCTCAGCTTTCCCTCGAGGCCCTCCTACCTTTTGCCGGGAGACCCCCAGCCCCTGCAGGGGCGGGGCCTC CCCACCACACCAGCCCTGTTCGCGCTCTCGGCAGTGCCGGGGGGCGCCGCCTCCCCCATGCCGCCCTCCGGGCTGCGGCTGCTGCCGCTG CTGCTACCGCTGCTGTGGCTACTGGTGCTGACGCCTGGCCGGCCGGCCGCGGGACTATCCACCTGCAAGACTATCGACATGGAGCTGGTG AAGCGGAAGCGCATCGAGGCCATCCGCGGCCAGATCCTGTCCAAGCTGCGGCTCGCCAGCCCCCCGAGCCAGGGGGAGGTGCCGCCCGGC CCGCTGCCCGAGGCCGTGCTCGCCCTGTACAACAGCACCCGCGACCGGGTGGCCGGGGAGAGTGCAGAACCGGAGCCCGAGCCTGAGGCC GACTACTACGCCAAGGAGGTCACCCGCGTGCTAATGGTGGAAACCCACAACGAAATCTATGACAAGTTCAAGCAGAGTACACACAGCATA TATATGTTCTTCAACACATCAGAGCTCCGAGAAGCGGTACCTGAACCCGTGTTGCTCTCCCGGGCAGAGCTGCGTCTGCTGAGGCTCAAG TTAAAAGTGGAGCAGCACGTGGAGCTGTACCAGAAATACAGCAACAATTCCTGGCGATACCTCAGCAACCGGCTGCTGGCACCCAGCGAC TCGCCAGAGTGGTTATCTTTTGATGTCACCGGAGTTGTGCGGCAGTGGTTGAGCCGTGGAGGGGAAATTGAGGGCTTTCGCCTTAGCGCC CACTGCTCCTGTGACAGCAGGGATAACACACTGCAAGTGGACATCAACGGGTTCACTACCGGCCGCCGAGGTGACCTGGCCACCATTCAT GGCATGAACCGGCCTTTCCTGCTTCTCATGGCCACCCCGCTGGAGAGGGCCCAGCATCTGCAAAGCTCCCGGCACCGCCGAGCCCTGGAC ACCAACTATTGCTTCAGCCCCCCGTCCAGCTCCGACGCCGGCCTGCGGTTCCCGGACAGCAACGGCCTCCTGCAGACCCCACGCTGGGAC GAGCCGCAGCGGGTGTGCGCCCTGGAGCAGATTTGCGGCGTGTTCCGCGTGGACCTGGGCCACATGCGCTCCCTCCGCCTTTTCTTCAGC GACGAGGCCTGCACCAGCGGCCAGCTGGTCGTTGCCAGCCGAGAGAGCCAGTACAAGGTTTTCCACTTCCACCACGGCGGCCTGGACAAG CTGTCTGACGTGTTCCAGCAGTGGAAATACTGCACCGAGATGCAGCTCAAAGACCAGCAGGTCGCCCCCGATAAGACATGCATGCAGTTC TCCATCCGCCGCCCCAAGCTGCCGTCCTCCGAGACGCACCCCGAGGAGAGCATGTACAAGAGGCTCGGCGTCTCCGCCTGGCTCAACCAC CTGAATGAGCTGGGCCAGGTGGAGGAGGAGTACAAGCTGCGGAAGGCCATTTTCTTTGGCGGTATTGATGTGTCAATCCGCGGGGAGGTC TGGCCCTTCCTGCTGCGCTATTACAGCCACGAGTCCACGTCGGAGGAGCGGGAGGCGCTGCGGCTGCAGAAGCGAAAGGAGTACTCTGAG ATCCAGCAGAAAAGGCTCTCCATGACTCCCGAGGAGCACAGAGCGTTCTGGCGTAATGTGCAGTTCACTGTGGACAAAGACGTGGTCCGG ACAGATCGGAACAACCAGTTCTTCCGGGGGGAAGACAATCCCAATGTGGAGAGCATGAGGAGGATCCTGCTGAACTACGCCGTGTACAAC CCTGCCGTCGGCTATTCCCAAGGGATGTCGGACCTGGTGGCGCCCATCTTGGCCGAGGTCCTGGATGAGTCAGACACCTTCTGGTGCTTT GTGGGTTTGATGCAGAACACGATCTTCGTCAGCTCACCCCGGGACGAGGACATGGAGAAACAACTGCTGTACCTGCGCGAGCTGCTGCGG CTGACGCACGTGCGCTTCTACCAGCACCTGGTCTCGCTGGGCGAGGACGGCCTGCAGATGCTCTTCTGCCACCGCTGGCTCCTTCTGTGC TTCAAGCGGGAGTTCCCCGAGGCCGAAGCGCTGCGGATCTGGGAGGCCTGCTGGGCCCACTACCAGACGGACTACTTCCACCTTTTCATC TGCGTGGCCATCGTGGCCATCTACGGGGATGACGTCATCGAGCAGCAGCTGGCCACGGACCAGATGCTCCTGCACTTCGGAAACCTGGCC ATGCACATGAACGGGGAGCTCGTTCTCCGGAAGGCGAGGAGTTTGCTGTACCAGTTCCGCCTCCTGCCCCGGATCCCCTGCAGCCTGCAC GATCTGTGTAAGCTGTGCGGGTCAGGCATGTGGGACAGCGGCTCCATGCCCGCGGTGGAGTGCACCGGCCACCATCCCGGCTCGGAGAGC TGTCCCTACGGGGGCACGGTGGAGATGCCTTCCCCCAAGTCCCTGAGGGAAGGCAAGAAGGGCCCAAAGACGCCGCAGGACGGCTTCGGC TTCCGCAGATAGGTCGGGCCCCCGACACCGGACAGGGGTTGAGGGGACCTCCTCAGAGGCCCTGGGCACGGGAGGGGGTGGGGCTGGGCG TGAAGGGGACAGGGGACGGTAGAAACCTAAGGAAAATGCTTTTGGGCAACATGAGAGGAACCTTTTCATATTAATGACAAAATTAGAGTC TGGAAGTGACAGAAGTCAGATCTACAGCCACCCAGAGGAAAGTCAGCTCCTGAAACGCTGCAGTGGAACGCGCAGCCACCGCACCTGAGA CGCAGGCTGGCTGGGCTCTCCTGCTGGCTGCCCTGGAGGATTTCAACATGTCCCAGGATTTGCTCCACCCTCGAGGGCAGCCAGACAGCG TCGCCAGGCAATGAGGAAAGCAGAGACAGGAGAGGAAGGCCTCACTCACCCACTGCGTCGAGGGCTGCAGAACACAGCGGGGTCCTGTCC AGGCCCAGGGACATCTTTGCAAGCCAGACACACTTCCTCTTGAGACCTCGTTCTCTCGGAGTGAGCCAAACACACTTCCCAAAACGTCCC CAGCCACAGCTGGGATGCCGATGGAAAGGCATCTGCCATAAAAGAAAAGCAAAAGATAAAAAGCCCAACCGATGTGGGGATAGAGAGGCG GAAGAGCAGTCAGGCTTGAGGAGCTGGCGCTTGTAATGTTTATCCGTTTAAACATTTCGTCCTCCTGGTACACGAAGGGAACTGTCTGCC CAGGAGCCTGAGCCTCAGGCTGTTGGAGAAGCATCTGATGCCTTTTTCTTTGCTGGGGGTCTTCTACGTGAGGTTCCTTGGCGTTGTTTA AGGTCAACTCCACCAAATACAGCAACCAGCTGGGGCTTGAATGGGTCGATGTGTGCGTGTGTCCCTGCGTGCATGCATGCATGCACACGT GCGTGCGTGTTTGTGTTTGCAAGATCCTCTGACAATGATCTGCTTTGTCCCAGGGACGTGGTAGTGTATTAGCTGACCAACCAAGACAAG GACACCTGGAAATAGTTAGACTCTTCCACTTCCCTTCAGGACCTGTGAACTCAGATATGTAAGTCAGCTTCTCAGGTGGTCGGACGTGAG AGCGACCACAGTGAAGGAGCCAGCACTCAGGGCTCTCTGCCTTCTATGTGGGAATGAGGTCTTCCCAACAGACTCTCCCTCTTCCAAAAG ATTGTGTATCAGTTGCTCCTTCTGGGATTTAGAGATGGACAGAAAAGGGTGTGCGATCTATGCTGTGAGAACTCCCAGCCAGCTTAGAAG TGTCGAGCCATTTGCACAGAAAGCCACTCCTTGAGCGAGGAGACCAAATCCCTCCTGAAATCCTCCATCGCGTCCTTCTGGGGAGAAAAA CCCTTGATGTGCTGAGAACCATCATGGGGACCAGGATAGAAGGCTTCTTCCCACTCAAAGCTTTTCTCCCTGGAGGGTGGGCACTGCTGG GCCATGCCACTTCAAAGCAGTGTTCCTCAGCAGGAAAGCGGAGGTCACCACTTACCGGCCTCCTCCACCTTCTCGGCTTCTCTTTTCTCC ATGAACCCAGGTCGTCCAGCAGGTACTTCCAAGTTCCCAGGTCTGTCTGCCTAAGAGCCTTTTGAGGAGACCGTCCTGGAGCCCCATCAG TGCCCAGATCCTGGGGTACCGACCATTGCTGTCTAGCAGTGGGGGATCCTGTGGTGGGAATGGGGTGGGCTTCTCATCCATGTTGCTTCT GGGAAGAGAGGGTTGCCTTTCTGGGCTAGGGAGGTGGCTGGAGCTTCTGCCCTGACCCTCCGCTAGAAACCAGTTATATCCATTGCCACA GCAATACTGTGTAACAAATCCGCCAACACTCGGTGGCCTGCAACAGTCAGCACTGATCTAGGGCAGGAGTCAGCAGTCTGGGCAGGGTGA TTCTTCTGGTCTAGGCTGGGCTTGTTTGTTTAGGGCCAGTGGGTTGTTAAGTCCCAGGGGATGCTCATGGTGCAGAGGTCGGACCTGGCC CTGCTCTGTGGTCTCTTGCATCCCTCCTGCAGGCCAGCCATGGCACAGCCAAAGGCACGGAGGGAGAGACAGCAAGTGCCTTTCCAAGCC TTGGTGGACACCACACCTGCAGCTCTCCCCTTGGCCACAGCAAGTCACGTGGCTGGGCTCAGCATTAAGGGGTGGGGGACAGGCCTGCCA CTTTATTGTCCGCTCTGCACAGTCGGGTGGCAGTAGGCTGAGAATTTGGAGGGCTAAACAACTGGGACTCTGGAGAGTCTGTGTCCTAAT AATGCCTGCTTTGGAGCACTCCTTCCCCACCCGCCTGCTCCAGGGGAATAGCGTCCCTGGTCCTAGCATTTCACTAGATACCTCGTGCCT TTGAACCGCTGTGTTTGGGAGGGAGGAAGAGGATAGACAGGTCCAGGGCTCCCCTCACTTGGGAAGGTCCTAGTAGAAGGCATTCCTTCT GAGTCTCCTGGCCCTACAGCTTCTCACCCCTGCTGCTCCCCAAGACCTGACCCGGACCAATCAGCTGCATCTCTGCACCAAGTGCCACCC CCACCGTGCACTGCTCGCACCTCACGCTCCCCATGGGCTGGCCCGGACCCCAGTGAGGTCTGCACCTCTCGGACCAGCCAGGACCTCGGA AGCCATGAGCTGGGTGTTCAGGGCTTGGCGGAGGAGACGCTGCTCCGCGGGGGATGGAGAAGCCCCGCCCCTCCACAACTCCCTTAATCT GGATGCAATTAGAGGAAACCAGCCAAAAACCCCGCGGCAGCTGTTGCCCCGGGATTATTCCCAGCCAGCGAGAGGCATTAGGGCGATAAT TCAGTCATTAGACTTCTAGCCTTCCCACCGCCAGAACGTGCAAATAGGGGATAATCAGCCCTGCACGGCCATCCTCAGCAATGCTGCTTA TTTAACCCATTTTTTAAAATGACACTTAATGTTATTGGCTGGCGTACGTGTGTGCGCATACTTAAAAGTTGATGCGTGGTGCAGCTCTGC TGCTAATGTCTGCTGACAGCGTCGCCCGATGTTGCTCAGGGCCTCTGGGAAGAGGGGACACCTTTGATGTGGGCGTGCTGGCACCCAAGA GGCAACGTGGTCAGGCTCGATGCTTGCCAGCCCTGGAAGGTCCCGCCAGGGTTGGGGGGTCGGGTGAGCACTCAGAGATCTGGGGAGGAA GCGAGGAGCCTGGTGTCCTCACCAGGTGCAGAGGGGAAATGCGTACTGCTCCAGAAGGTTCTGACTCAAGTCTCGGCCCTGAGGACTCAG AACTCACAAGTCATGGATCAAGTACAGAGGGAGGGAGGGCCCCAGCCCACCCCTGCGGGGGAGGAGGAGGGTCTGAAGGTGGGAGGGCAG CTCCATGAGCAGCCACTGTTCCTGACCAGCCGAAGTGCCAGGCAGGGGGAGGTTTCCATGTCCAGGAGGGAGAGGACCTCCACCCTCTGA CCTTTCAGTTTTCCATAGCAGCCCAGTGTGAAGCTCGGCGGGGGTGGGCAGGGCATCAGGACAAGATGAAGATGCTGGCTCTGGAATGAG GACTCCCCTTGGGCTAAGGGGTAGTGGCTGGCAGAGGTTGCCCAGCTCTTTTGGAGGCCACAGGGAATAGAGGTGGGGTGGTGCTTCCGT TCCTGTGAGGCCCGCCCCCCTGAGCCCCCTCGTCTGCTTCTTGTTTGGCAGAAATAGATGTGGGGCACAGCCCCTAGATGATGGGAATCT GCCCAAGCTGTGCCCTCTGGGGAATGAAATCTTGCTCCTCGTGCAGCTTCCCCTTTCCCTTGCGGGGTGGGGGCTGGGGGTGCCAAGGCT CCACCCTTGGAGGGGGTTGCTGGGGGTGCCAAGGCTCCACCCTTTGCTCGCTGGGCAGAGGGAAGGAAGCACCTCCCTGTATGCTCTGCA GGGGCCCCTGCAGGCCACTTTGCCTCAGAGGAAGGAGGAGAGGTCCCCTGGGAAAATGCGCACACACACACACACACACACACACACACA CGCACATGCACACACGCACGCATGCACACACACACACACATATGTGCAGGGGTGCAGCCTAAGTCAGTGGAGCCAGGACTGGCTGCGCAG CTGGGCCTGCCCCATTCTTGGAGTCCCCAGGGAATGAGCCTCAGGATCCACTTCGTCCCTTGTTTCTAAGCCAGGCTGCACGGCCGTTCC TGGGTGTGAAAGCATTCTGTCCTGTCTCCAGTCAGAGGGAACCGCCCCCAAACTAGAAAGCTAATTTCCCCCGAAGGTTCATGAAGCCAG AGGCATGCCCTTGGGGGTTGGTCGGGAGAAGGGTGAGAGGCAGTGCTGTGGAAGGGGCCTGGCCCCTGCTGCCTGCGATTCTCCCCGGGT ATGTGGCCTTCTGCCTCCTTCAGCCTCAGTTTCCTCACCTGTAACATGAGAATAATGATGCCTGCCTGCCCAGGCCACCTCCTGGGAGTG CATTGGGGACTCTAAGTGTAAAACTGTCATGAGGATGGTTTATGAAGTCCCAGGGACCACACCTGCCACTCAGGGAAGAGACCGCTGACC TCAGCCATCAGCCGTGTGCATCGTGAGGGGCAGCGAGCTCAGGACGGAGAGCTCTCCCTTGGTTCGGAGTCAGCACACCTGCACTCCCAC ACGCCCACCCAGCTGTGTGTCCTCCGCCGAGGGCGAGCCTCTCTGTGACTCTTCTGTAAAATGCACCCCATTCTGCCCACTCCTGCCACG AGACCCATCCTCTGCGATCCCGCCAGGCATGTGTGTGTGAATGCATGTGCAAAGCTCTCCATCAGAAGGGTGGTGTGGGCCCTGCAGGTC CTACCCCTCGGCCTTGAAGCTCCCTCGGGCTGCGGACTCTGCCTCCTGGGTCTGAGCATTAGAACCAGGAGAGGGGTGTCCCTGGGCAGA GCCAGGGGTGCAAACAGCCTGCAGCCATCTGGCCTTTTAAGTATAGTGTGTCGCATTTCCGGGTAGGAAGGTAGCATTTCAAGTTCAAAG AGAGGTCAAGTCATGCAACCATCTTTCCTCCAGCACTTTTGGGGTAAGGAGGACAGTTTTTGTTATGGTTTAGGGGAAATTTTCATGAAA TTTCCACCATTACCAATAGATTACTGATGTCCATGGCAAGTGATCTGTTCTTGGTATTTTGTTTTGTTTTGTTTTGGTTTTTAAATGTAA TCACCCATTGGTCAGGCCCAGGACTGGTCACCATGAGCTCTGCTAGCCACGGCCCCAACGATGCTTCCGGCTCTCATGGATTCCACAGCA AATAAAACTGTACTGACAACAGATGCACGCTGTACAGTGCCTCTGGAAGGTGCGGTGGGACTTGCTGGGTCAAAGCACGGCAGGCTCTGT GCTGGGCAAACAGGGCCCCGTGGCAGCCCAAGGGCAGCCTGCAGACGCCAGTGTGCCCGGCTTCTCCCATGTTCTGAGATGGGGGAGGGG AAGGTGTAAAAGAGAGCTGGATGGGGGCTGTGGGGACTGAGGGTCCTGGGAGGCTGGCCAGGGGCTCAGCAAGCGCCCACTTTGGGAGGC CTTGCCCTCTGGGCTGTGGTTGGACAGCACTAGGCCTGCAGGCTGGGCTGCCAGGAGGAAGAGCAGCTCCGGACCTCAGGCCCCCCCAGC AGGCGGCGATGGAGGGTGATGGGAGAATTGACGAGTGCTCGCGGGGTCTCACTCGGGAAAGGGCCGGTTAACTCTGAGCAACTGGGAGCC TCGCTCCCACCCCAGTGCCTGGAAACACAAACAGTATTTTATTGAGCCTTGGCTGCCGGGGCTCTGGTTTCAGGAAAAGCCTCAGTCCTG CATTTGTGAATTTTTAAGGAGGCAAACAAGGTGAATTTGACCTTCCTAGGAGTGGGAGTTAGGGAAAATTCAGGCTGCCCTACCAAGCCC TTGGAACCGTGCAGCAGGGGTCAGGGGAGGGATGGGCAAAGGCGGTTTGGCCCATGGAGCAGGTCTCCTGGTCCTCCCGGAGCACGGGGG GCGGGGAGCTCCAGCTCTTCATCAAGCGCCCTTTGGGGTTTCCAACTGTTTTCTGTTGTTCTCTGTGTGTCTTTTCCCCTTTTGCCTGGG GGCGGGGGCTGGAGGGGAACATACCTCTTATCTACATCACTCGGGTCTCATGTCTGAAGGGCCTGTCTCACCTGCCACCTACCTGGCCTT GGCCTCGACATCTGAGCGCCTCATCACCTCCTCTCCACATGCCCAGGGGTGCCCCTCCCAAGGCTGGCTGGGCAGGTCCCACGGACTAGG CCCCAGGAGGAGCAGTGGGCTTCCCCCAGGAAAGAGCAGAGCCAGCACTGCCTGCCTAGGGAGGGCACCCACTACCAGGCATGGCTGGTG GCTCCGTTTAAAGAAATCTTTGTCCATGTGGGAGTGGGAGGTGCTTCCCCATCCTGCCTGGAAGCCACGTCCTGGCTCTTACCGTGGACT CTGCAACAGCCGTGGTGGCCACATGAAGATGGAGGAACCTGGTGGCTCAGGGGCCCCTGATGTCACCGCCTCCAAGGCAACGGGACTGGG CAGGGCAGCTCCTCAGGAAGGATGATCCCACCTGCACGGGTCAGCCCTGCAGGGGGACAAGGCCCACCACAACCCCACCTCAGGCCACCA ACTGGACCTGTCCTGCCAAGAAGGTCCCTTCCAATTTGTCCGGCAGCTGGTTCCCTGGATACAGCCCAGACGCAGCATCATCCTCATGTT CACCTCTGCTGTGCCCTACTTCCCCCGTAAATGACAACTGTGTAGTCACCCACGCCGGGCAAGAGAACATAGGTGCCACCTGGCGGGCCA CACTTTTCAGGGGCCGTGCCTCGGGCAGGCCCGGCTCTCTGTTGTCCCGCAGCCCTCAGGGCTGGCGGTGGCTGCTCAGTGCCTAGGGAG TGCCCTTGCCACACAGCTGGGCAGCGAGCATGACTCAGCGTTTACACTCAACCAGCCGCACAGGCTCCAGAGGGGAAGTTCTCTGGGGTC CTTCTATGAGAGAGCATCCTGTCTGCAGCCCTGTTCTGGCTCCTTCCACATCATTCATGCATTCATCCCACGAGGACTATGGTACCGTGG GCTCGCTCTAGGGAAGCAAGGCAGGAGCGCACTGGGGCTACTTGGATTCCCGGGGCAGACACATATGTACCCTGCACACCCTCAAATGAG CAAAGCCACCCCACTGGCCCCTGGGTGAGGCCATGTGGACCCCCACACCTTTGGGGCCGGCTGGGCAAACCTTGAACCCCCAATTTCTGC GTGGCCTTTGGCTGCCCTCCTTCTCCAAGGCGTGACTCTTACTCCAGAGACTCAGGCGAGCACGTGTCCCTTACCTTATTTTCTCTCAAT CAAACTGAAACCCATTGTGATCCCCCATAGTCCAGTGCGGTCTCTGTTATTATTACGGGTGTCTCCCTTCCTCCCCGTGCCCAGGACAGG CCATGAGCCAGAGATACAAGGGGCCCCAGGCAAGATGCAGGGCTCTCTGCTCCTGGCTCTTTATCGTCGTCGGGACACCCTTTGTCCAAA CTCAAGGAATCCCGGGAGGTCTGGCTTTGCCGCTTTGGCTGGGACTCAGGTACCTGGGCATACCTGCATCTGTGCCCCCTCTTCGTCCCA GTGAGACGGACCCTCAGTATCCCCCCCAACATAGAACAGGAAGGTCTCAAGCCAGCCCCCTCCTCCAGTTCAATCACCCTCCCTCACATC CACCCAGAGAAGTCCAGACCTGTGGCCACTTCCATGCCCTCTCCAGACCAGGAGACACCTGCTGCTGACCTCGTGGAAAACTTAGATTTT GACATTCTGATGCTTCGGAAGTGGTGGCTCCTCCTCCTTCACCCCTCCGCCACCTGTGGGCCTCCTCTCTGCATCTCAAGAGAACAACCA GATCTTTGGGCTCCTGGGGTGTGTGCCATGCAATTTAGACGAAGTGCTTTGAAAATATGCCATTCAGTCTCTGACTAGGAAAATAAGTCT GACCTGATAGGTCCGATGTCATCAGCTCTTCAACATGAGACAAAAGAGGGGATTTTATGTTTTGAGTCATTAGAATGATATAATAATTTT CTGAATTGACATCTGGATGTTGAAATTAGGATGGTGCAAAAGGGGTCCAGGGCCTCAGGCTGGGCGCAGCAGCCAGCTCCCAATGACGCA GAAGCTGCTTCAAAACCCCCTCAACAAAGAGGGGCACATGCAAGTCACCAAAGTGGGAAGCCTTCACCAAGGCCACACCCAAAGTCTACT GATTGTCTGTCCAAAGTTCGTTGATTCCTGGCCATGAACAAGCACAATAGAAAAAGACACAGGGTCCTAGTGGCTACAAGTCAATGTGAA TTGGCACATGGTCTAGCAGTTTTAAAATCTGACAGTAGAGTATGGCAATGGGCAAGGGCCAAGAAGTCCTGAGATGGGAGGTCAGCGCTC TAACTGGGCTCAGTGGAGGTCTGTGACCAGTGTCTGGACACTAGCTACAGGGGACCGGGCAGAGGATTCTGGGCAGAAGGAAAATGTCTA >90431_90431_1_TGFB1-TBC1D16_TGFB1_chr19_41847787_ENST00000221930_TBC1D16_chr17_77926617_ENST00000310924_length(amino acids)=1026AA_BP=184 MRPQSLRRAAAAPATAGRRGRRSGRRDELVGRRGKKLLRLFRCRWEPEARGPLGATLPREEAGLGDPRPPPFAAGDACSLPAPYTASLRR PHSGPALGSRRPGLPQRLFPRPRAHPLHAAFIPGLSPEPPRILDPFSSRRRISLRPATDPLFKTTHLLVPDRAHLGYFRGILRHPRSKPP LHHCALLPEDLSFPSRPSYLLPGDPQPLQGRGLPTTPALFALSAVPGGAASPMPPSGLRLLPLLLPLLWLLVLTPGRPAAGLSTCKTIDM ELVKRKRIEAIRGQILSKLRLASPPSQGEVPPGPLPEAVLALYNSTRDRVAGESAEPEPEPEADYYAKEVTRVLMVETHNEIYDKFKQST HSIYMFFNTSELREAVPEPVLLSRAELRLLRLKLKVEQHVELYQKYSNNSWRYLSNRLLAPSDSPEWLSFDVTGVVRQWLSRGGEIEGFR LSAHCSCDSRDNTLQVDINGFTTGRRGDLATIHGMNRPFLLLMATPLERAQHLQSSRHRRALDTNYCFSPPSSSDAGLRFPDSNGLLQTP RWDEPQRVCALEQICGVFRVDLGHMRSLRLFFSDEACTSGQLVVASRESQYKVFHFHHGGLDKLSDVFQQWKYCTEMQLKDQQVAPDKTC MQFSIRRPKLPSSETHPEESMYKRLGVSAWLNHLNELGQVEEEYKLRKAIFFGGIDVSIRGEVWPFLLRYYSHESTSEEREALRLQKRKE YSEIQQKRLSMTPEEHRAFWRNVQFTVDKDVVRTDRNNQFFRGEDNPNVESMRRILLNYAVYNPAVGYSQGMSDLVAPILAEVLDESDTF WCFVGLMQNTIFVSSPRDEDMEKQLLYLRELLRLTHVRFYQHLVSLGEDGLQMLFCHRWLLLCFKREFPEAEALRIWEACWAHYQTDYFH LFICVAIVAIYGDDVIEQQLATDQMLLHFGNLAMHMNGELVLRKARSLLYQFRLLPRIPCSLHDLCKLCGSGMWDSGSMPAVECTGHHPG -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for TGFB1-TBC1D16 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for TGFB1-TBC1D16 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for TGFB1-TBC1D16 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
