
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:TMEM126B-PICALM (FusionGDB2 ID:91614) |
Fusion Gene Summary for TMEM126B-PICALM |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: TMEM126B-PICALM | Fusion gene ID: 91614 | Hgene | Tgene | Gene symbol | TMEM126B | PICALM | Gene ID | 55863 | 8301 |
| Gene name | transmembrane protein 126B | phosphatidylinositol binding clathrin assembly protein | |
| Synonyms | HT007|MC1DN29 | CALM|CLTH|LAP | |
| Cytomap | 11q14.1 | 11q14.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | complex I assembly factor TMEM126B, mitochondrial | phosphatidylinositol-binding clathrin assembly proteinclathrin assembly lymphoid myeloid leukemia protein | |
| Modification date | 20200313 | 20200322 | |
| UniProtAcc | . | Q13492 | |
| Ensembl transtripts involved in fusion gene | ENST00000358867, ENST00000393375, ENST00000534341, | ENST00000356360, ENST00000528398, ENST00000528411, ENST00000393346, ENST00000526033, ENST00000532317, | |
| Fusion gene scores | * DoF score | 3 X 1 X 3=9 | 35 X 26 X 12=10920 |
| # samples | 3 | 44 | |
| ** MAII score | log2(3/9*10)=1.73696559416621 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(44/10920*10)=-4.63332552228256 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: TMEM126B [Title/Abstract] AND PICALM [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | TMEM126B(85346822)-PICALM(85742653), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | TMEM126B-PICALM seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. TMEM126B-PICALM seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | PICALM | GO:0006897 | endocytosis | 22118466 |
| Tgene | PICALM | GO:0006898 | receptor-mediated endocytosis | 10436022 |
| Tgene | PICALM | GO:0032880 | regulation of protein localization | 10436022 |
| Tgene | PICALM | GO:0045893 | positive regulation of transcription, DNA-templated | 11425879 |
| Tgene | PICALM | GO:0048261 | negative regulation of receptor-mediated endocytosis | 10436022 |
| Tgene | PICALM | GO:1905224 | clathrin-coated pit assembly | 16262731 |
 Fusion gene breakpoints across TMEM126B (5'-gene) Fusion gene breakpoints across TMEM126B (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
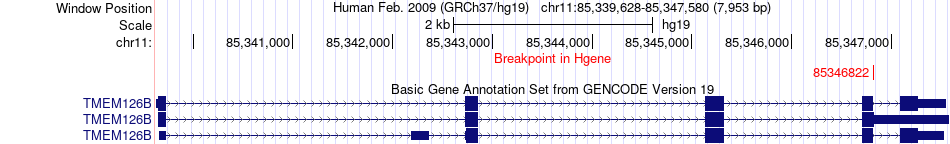 |
 Fusion gene breakpoints across PICALM (3'-gene) Fusion gene breakpoints across PICALM (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
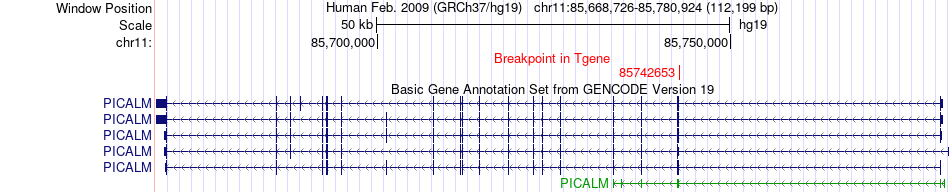 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BLCA | TCGA-FD-A3B8 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
Top |
Fusion Gene ORF analysis for TMEM126B-PICALM |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000358867 | ENST00000356360 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000358867 | ENST00000528398 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000358867 | ENST00000528411 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000393375 | ENST00000356360 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000393375 | ENST00000528398 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000393375 | ENST00000528411 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000534341 | ENST00000356360 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000534341 | ENST00000528398 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| 5CDS-5UTR | ENST00000534341 | ENST00000528411 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| Frame-shift | ENST00000358867 | ENST00000393346 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| Frame-shift | ENST00000358867 | ENST00000526033 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| Frame-shift | ENST00000358867 | ENST00000532317 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| Frame-shift | ENST00000393375 | ENST00000393346 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| Frame-shift | ENST00000393375 | ENST00000526033 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| Frame-shift | ENST00000393375 | ENST00000532317 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| In-frame | ENST00000534341 | ENST00000393346 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| In-frame | ENST00000534341 | ENST00000526033 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
| In-frame | ENST00000534341 | ENST00000532317 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000534341 | TMEM126B | chr11 | 85346822 | + | ENST00000532317 | PICALM | chr11 | 85742653 | - | 4226 | 1161 | 1148 | 2863 | 571 |
| ENST00000534341 | TMEM126B | chr11 | 85346822 | + | ENST00000526033 | PICALM | chr11 | 85742653 | - | 4327 | 1161 | 1148 | 2968 | 606 |
| ENST00000534341 | TMEM126B | chr11 | 85346822 | + | ENST00000393346 | PICALM | chr11 | 85742653 | - | 3222 | 1161 | 1148 | 2989 | 613 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000534341 | ENST00000532317 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - | 0.000473153 | 0.99952686 |
| ENST00000534341 | ENST00000526033 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - | 0.000398334 | 0.99960166 |
| ENST00000534341 | ENST00000393346 | TMEM126B | chr11 | 85346822 | + | PICALM | chr11 | 85742653 | - | 0.000753164 | 0.99924684 |
Top |
Fusion Genomic Features for TMEM126B-PICALM |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
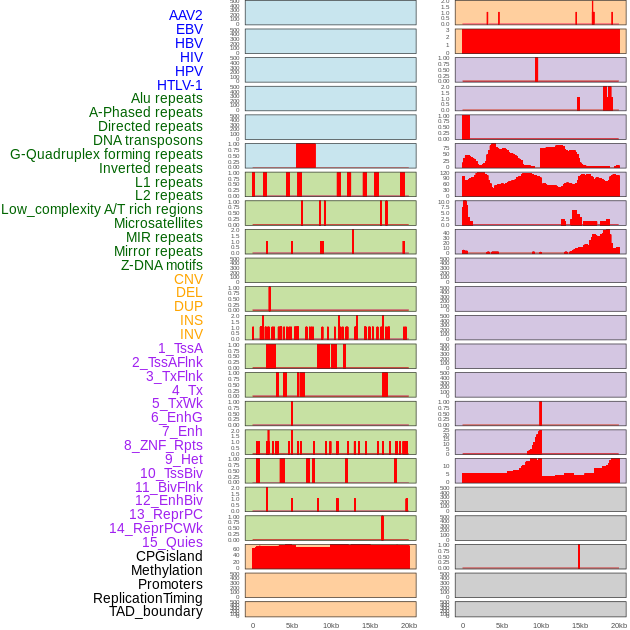 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for TMEM126B-PICALM |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:85346822/chr11:85742653) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | PICALM |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Cytoplasmic adapter protein that plays a critical role in clathrin-mediated endocytosis which is important in processes such as internalization of cell receptors, synaptic transmission or removal of apoptotic cells. Recruits AP-2 and attaches clathrin triskelions to the cytoplasmic side of plasma membrane leading to clathrin-coated vesicles (CCVs) assembly (PubMed:10436022, PubMed:16262731, PubMed:27574975). Furthermore, regulates clathrin-coated vesicle size and maturation by directly sensing and driving membrane curvature (PubMed:25898166). In addition to binding to clathrin, mediates the endocytosis of small R-SNARES (Soluble NSF Attachment Protein REceptors) between plasma membranes and endosomes including VAMP2, VAMP3, VAMP4, VAMP7 or VAMP8 (PubMed:22118466, PubMed:21808019, PubMed:23741335). In turn, PICALM-dependent SNARE endocytosis is required for the formation and maturation of autophagic precursors (PubMed:25241929). Modulates thereby autophagy and the turnover of autophagy substrates such as MAPT/TAU or amyloid precursor protein cleaved C-terminal fragment (APP-CTF) (PubMed:25241929, PubMed:24067654). {ECO:0000269|PubMed:10436022, ECO:0000269|PubMed:16262731, ECO:0000269|PubMed:21808019, ECO:0000269|PubMed:22118466, ECO:0000269|PubMed:23741335, ECO:0000269|PubMed:24067654, ECO:0000269|PubMed:25241929, ECO:0000269|PubMed:25898166, ECO:0000269|PubMed:27574975}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000358867 | + | 4 | 5 | 110_130 | 169 | 231.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000358867 | + | 4 | 5 | 141_161 | 169 | 231.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000358867 | + | 4 | 5 | 72_92 | 169 | 231.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000393375 | + | 5 | 6 | 110_130 | 139 | 201.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000393375 | + | 5 | 6 | 72_92 | 139 | 201.0 | Transmembrane | Helical |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000528398 | 0 | 19 | 14_145 | 0 | 552.0 | Domain | ENTH |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000358867 | + | 4 | 5 | 199_219 | 169 | 231.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000393375 | + | 5 | 6 | 141_161 | 139 | 201.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000393375 | + | 5 | 6 | 199_219 | 139 | 201.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000534341 | + | 1 | 4 | 110_130 | 0 | 171.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000534341 | + | 1 | 4 | 141_161 | 0 | 171.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000534341 | + | 1 | 4 | 199_219 | 0 | 171.0 | Transmembrane | Helical |
| Hgene | TMEM126B | chr11:85346822 | chr11:85742653 | ENST00000534341 | + | 1 | 4 | 72_92 | 0 | 171.0 | Transmembrane | Helical |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000356360 | 0 | 19 | 14_145 | 43 | 633.0 | Domain | ENTH | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000393346 | 0 | 20 | 14_145 | 43 | 653.0 | Domain | ENTH | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000526033 | 0 | 20 | 14_145 | 43 | 646.0 | Domain | ENTH | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000532317 | 0 | 20 | 14_145 | 43 | 611.0 | Domain | ENTH |
Top |
Fusion Gene Sequence for TMEM126B-PICALM |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >91614_91614_1_TMEM126B-PICALM_TMEM126B_chr11_85346822_ENST00000534341_PICALM_chr11_85742653_ENST00000393346_length(transcript)=3222nt_BP=1161nt ACCAAAATGGTGGTGTTCGGGTATGAGGCTGGGACTAAGCCAAGGGATTCAGGTGTGGTGCCGGTGGGAACTGAGGAAGCGCCCAAGGTT TTCAAGATGGCAGCATCTATGCATGGTCAGCCCAGTCCTTCTCTAGAAGATGCAAAACTCAGAAGACCAATGGTCATAGAAATCATAGAA AAAAATTTTGACTATCTTAGAAAAGAAATGACACAAAATATATATCAAATGGCGACATTTGGAACAACAGCTGGTTTCTCTGGAATATTC TCAAACTTCCTGTTCAGACGCTGCTTCAAGGTTAAACATGATGCTTTGAAGACATATGCATCATTGGCTACACTTCCATTTTTGTCTACT GTTGTTACTGACAAGCTTTTTGTAATTGATGCTTTGTATTCAGGTAAGTTCTTCCTTTTCCTTCTTTTTTCTTTTCTTTTCTTTTCTTTT TTTTTTTTTTTTTTAGGTTAATAAACTCTCTTTCTTCTTTAGCTGTCTATGATATATTTGAATCAAAATGTGGTAGTTTGGTATTTCAGA TAGCTAACTTTTAACTTTTTTGCTCTCTACATTATAGATCTAGAAAGGCAATTATCAATTTAGGAGAAGTCAACAAACTTCTAATAGCAT TCATATCCAGGAGAGGTGATTAACTCTTTTTTCTTTTTAGGTATCATACCGTTCCACTGCCACCAAAAGGAAGGGTTTTAATCCATTGGA TGACGCTTTGTCAAACACAAATGAAATTAATGGCGATTCCTCTAGTCTTTCAGATTATGTTTGGAATATTAAATGGTCTATACCATTATG CAGTATTTGAAGAGACACTTGAGAAAACTATACATGAAGAGTAACCAAAAAAATGAATGGTTGCTAACTTAGCAAAATGAAGTTTCTATA AAGAGGACTCAGGCATTGCTGAAAGAGTTAAAAGTAACTGTGAACAAATAATTTGTTCTGTGCCTTTTGCCTGGTATATAGCAAATACTC AAAAAGTATTCAATAATTCAATCAATAAATATAAGTTTCATCTTACACGTAAGATACAGGTCTTATCTCCTGATGGTGTGTCCATTTTGC CTGGTATATAACAGATAATAAATATCCAGTGTCAATAAATGTAACAATAAAAGTTTCATCTTTCCTCTTTGTATGTGGAAAACTTAATTC AGTGCACAAATGAGATGAATGTGAACATCCCACAGTTGGCAGACAGTTTATTTGAAAGAACTACTAATAGTAGTTGGGTGGTGGTCTTCA AATCTCTCATTACAACTCATCATTTGATGGTGTATGGAAATGAGCGTTTTATTCAGTATTTGGCTTCAAGAAACACGTTGTTTAACTTAA GCAATTTTTTGGATAAAAGTGGATTGCAAGGATATGACATGTCTACATTTATTAGGCGGTATAGTAGATATTTAAATGAGAAAGCAGTTT CATACAGACAAGTTGCATTTGATTTCACAAAAGTGAAGAGAGGGGCTGATGGAGTTATGAGAACAATGAACACAGAAAAACTCCTAAAAA CTGTACCAATTATTCAGAATCAGATGGATGCACTTCTTGATTTTAATGTTAATAGCAATGAACTTACAAATGGGGTAATAAATGCTGCCT TCATGCTCCTGTTCAAAGATGCCATTAGACTGTTTGCAGCATACAATGAAGGAATTATTAATTTGTTGGAAAAATATTTTGATATGAAAA AGAACCAATGCAAAGAAGGTCTTGACATCTATAAGAAGTTCCTAACTAGGATGACAAGAATCTCAGAGTTCCTCAAAGTTGCAGAGCAAG TTGGAATTGACAGAGGTGATATACCAGACCTTTCACAGGCCCCTAGCAGTCTTCTTGATGCTTTGGAACAACATTTAGCTTCCTTGGAAG GAAAGAAAATCAAAGATTCTACAGCTGCAAGCAGGGCAACTACACTTTCCAATGCAGTGTCTTCCCTGGCAAGCACTGGTCTATCTCTGA CCAAAGTGGATGAAAGGGAAAAGCAGGCAGCATTAGAGGAAGAACAGGCACGTTTGAAAGCTTTAAAGGAACAGCGCCTAAAAGAACTTG CAAAGAAACCTCATACCTCTTTAACAACTGCAGCCTCTCCTGTATCCACCTCAGCAGGAGGGATAATGACTGCACCAGCCATTGACATAT TTTCTACCCCTAGTTCTTCTAACAGCACATCAAAGCTGCCCAATGATCTGCTTGATTTGCAGCAGCCAACTTTTCACCCATCTGTACATC CTATGTCAACTGCTTCTCAGGTAGCAAGTACATGGGGAGATCCTTTCTCTGCTACTGTAGATGCTGTTGATGATGCCATTCCAAGCTTAA ATCCTTTCCTCACAAAAAGTAGTGGTGATGTTCACCTTTCCATTTCTTCAGATGTATCTACTTTTACTACTAGGACACCTACTCATGAAA TGTTTGTTGGATTCACTCCTTCTCCAGTTGCACAGCCACACCCTTCAGCTGGCCTTAATGTTGACTTTGAATCTGTGTTTGGAAATAAAT CTACAAATGTTATTGTAGATTCTGGGGGCTTTGATGAACTAGGTGGACTTCTCAAACCAACAGTGGCCTCTCAGAACCAGAACCTTCCTG TTGCCAAACTCCCACCTAGCAAGTTAGTATCTGATGACTTGGATTCATCTTTAGCCAACCTTGTGGGCAATCTTGGCATCGGAAATGGAA CCACTAAGAATGATGTAAATTGGAGTCAACCAGGTGAAAAGAAGTTAACTGGGGGATCTAACTGGCAACCAAAGGTTGCACCAACAACCG CTTGGAATGCTGCAACAATGGCACCCCCTGTAATGGCCTATCCTGCTACTACACCAACAGGCATGATAGGATATGGAATTCCTCCACAAA TGGGAAGTGTTCCTGTAATGACGCAACCAACCTTAATATACAGCCAGCCTGTCATGAGACCTCCAAACCCCTTTGGCCCTGTATCAGGAG CACAGATACAGTTTATGTAACTTGATGGAAGAAAATGGAATTACTCCAAAAAGACAAGTGCTCAAGCAGCAAAATCCTTACTTCCAGCAA AATCCAAACTGCTGTCTCTTAAATCTCTTAAACTCTCTTCTTCCATTAGAATGCTACAAGTAACTCAGTGAAGGCCCATGAAGGAAATTG >91614_91614_1_TMEM126B-PICALM_TMEM126B_chr11_85346822_ENST00000534341_PICALM_chr11_85742653_ENST00000393346_length(amino acids)=613AA_BP=5 MYVENLIQCTNEMNVNIPQLADSLFERTTNSSWVVVFKSLITTHHLMVYGNERFIQYLASRNTLFNLSNFLDKSGLQGYDMSTFIRRYSR YLNEKAVSYRQVAFDFTKVKRGADGVMRTMNTEKLLKTVPIIQNQMDALLDFNVNSNELTNGVINAAFMLLFKDAIRLFAAYNEGIINLL EKYFDMKKNQCKEGLDIYKKFLTRMTRISEFLKVAEQVGIDRGDIPDLSQAPSSLLDALEQHLASLEGKKIKDSTAASRATTLSNAVSSL ASTGLSLTKVDEREKQAALEEEQARLKALKEQRLKELAKKPHTSLTTAASPVSTSAGGIMTAPAIDIFSTPSSSNSTSKLPNDLLDLQQP TFHPSVHPMSTASQVASTWGDPFSATVDAVDDAIPSLNPFLTKSSGDVHLSISSDVSTFTTRTPTHEMFVGFTPSPVAQPHPSAGLNVDF ESVFGNKSTNVIVDSGGFDELGGLLKPTVASQNQNLPVAKLPPSKLVSDDLDSSLANLVGNLGIGNGTTKNDVNWSQPGEKKLTGGSNWQ -------------------------------------------------------------- >91614_91614_2_TMEM126B-PICALM_TMEM126B_chr11_85346822_ENST00000534341_PICALM_chr11_85742653_ENST00000526033_length(transcript)=4327nt_BP=1161nt ACCAAAATGGTGGTGTTCGGGTATGAGGCTGGGACTAAGCCAAGGGATTCAGGTGTGGTGCCGGTGGGAACTGAGGAAGCGCCCAAGGTT TTCAAGATGGCAGCATCTATGCATGGTCAGCCCAGTCCTTCTCTAGAAGATGCAAAACTCAGAAGACCAATGGTCATAGAAATCATAGAA AAAAATTTTGACTATCTTAGAAAAGAAATGACACAAAATATATATCAAATGGCGACATTTGGAACAACAGCTGGTTTCTCTGGAATATTC TCAAACTTCCTGTTCAGACGCTGCTTCAAGGTTAAACATGATGCTTTGAAGACATATGCATCATTGGCTACACTTCCATTTTTGTCTACT GTTGTTACTGACAAGCTTTTTGTAATTGATGCTTTGTATTCAGGTAAGTTCTTCCTTTTCCTTCTTTTTTCTTTTCTTTTCTTTTCTTTT TTTTTTTTTTTTTTAGGTTAATAAACTCTCTTTCTTCTTTAGCTGTCTATGATATATTTGAATCAAAATGTGGTAGTTTGGTATTTCAGA TAGCTAACTTTTAACTTTTTTGCTCTCTACATTATAGATCTAGAAAGGCAATTATCAATTTAGGAGAAGTCAACAAACTTCTAATAGCAT TCATATCCAGGAGAGGTGATTAACTCTTTTTTCTTTTTAGGTATCATACCGTTCCACTGCCACCAAAAGGAAGGGTTTTAATCCATTGGA TGACGCTTTGTCAAACACAAATGAAATTAATGGCGATTCCTCTAGTCTTTCAGATTATGTTTGGAATATTAAATGGTCTATACCATTATG CAGTATTTGAAGAGACACTTGAGAAAACTATACATGAAGAGTAACCAAAAAAATGAATGGTTGCTAACTTAGCAAAATGAAGTTTCTATA AAGAGGACTCAGGCATTGCTGAAAGAGTTAAAAGTAACTGTGAACAAATAATTTGTTCTGTGCCTTTTGCCTGGTATATAGCAAATACTC AAAAAGTATTCAATAATTCAATCAATAAATATAAGTTTCATCTTACACGTAAGATACAGGTCTTATCTCCTGATGGTGTGTCCATTTTGC CTGGTATATAACAGATAATAAATATCCAGTGTCAATAAATGTAACAATAAAAGTTTCATCTTTCCTCTTTGTATGTGGAAAACTTAATTC AGTGCACAAATGAGATGAATGTGAACATCCCACAGTTGGCAGACAGTTTATTTGAAAGAACTACTAATAGTAGTTGGGTGGTGGTCTTCA AATCTCTCATTACAACTCATCATTTGATGGTGTATGGAAATGAGCGTTTTATTCAGTATTTGGCTTCAAGAAACACGTTGTTTAACTTAA GCAATTTTTTGGATAAAAGTGGATTGCAAGGATATGACATGTCTACATTTATTAGGCGGTATAGTAGATATTTAAATGAGAAAGCAGTTT CATACAGACAAGTTGCATTTGATTTCACAAAAGTGAAGAGAGGGGCTGATGGAGTTATGAGAACAATGAACACAGAAAAACTCCTAAAAA CTGTACCAATTATTCAGAATCAGATGGATGCACTTCTTGATTTTAATGTTAATAGCAATGAACTTACAAATGGGGTAATAAATGCTGCCT TCATGCTCCTGTTCAAAGATGCCATTAGACTGTTTGCAGCATACAATGAAGGAATTATTAATTTGTTGGAAAAATATTTTGATATGAAAA AGAACCAATGCAAAGAAGGTCTTGACATCTATAAGAAGTTCCTAACTAGGATGACAAGAATCTCAGAGTTCCTCAAAGTTGCAGAGCAAG TTGGAATTGACAGAGGTGATATACCAGACCTTTCACAGGCCCCTAGCAGTCTTCTTGATGCTTTGGAACAACATTTAGCTTCCTTGGAAG GAAAGAAAATCAAAGATTCTACAGCTGCAAGCAGGGCAACTACACTTTCCAATGCAGTGTCTTCCCTGGCAAGCACTGGTCTATCTCTGA CCAAAGTGGATGAAAGGGAAAAGCAGGCAGCATTAGAGGAAGAACAGGCACGTTTGAAAGCTTTAAAGGAACAGCGCCTAAAAGAACTTG CAAAGAAACCTCATACCTCTTTAACAACTGCAGCCTCTCCTGTATCCACCTCAGCAGGAGGGATAATGACTGCACCAGCCATTGACATAT TTTCTACCCCTAGTTCTTCTAACAGCACATCAAAGCTGCCCAATGATCTGCTTGATTTGCAGCAGCCAACTTTTCACCCATCTGTACATC CTATGTCAACTGCTTCTCAGGTAGCAAGTACATGGGGAGATGCTGTTGATGATGCCATTCCAAGCTTAAATCCTTTCCTCACAAAAAGTA GTGGTGATGTTCACCTTTCCATTTCTTCAGATGTATCTACTTTTACTACTAGGACACCTACTCATGAAATGTTTGTTGGATTCACTCCTT CTCCAGTTGCACAGCCACACCCTTCAGCTGGCCTTAATGTTGACTTTGAATCTGTGTTTGGAAATAAATCTACAAATGTTATTGTAGATT CTGGGGGCTTTGATGAACTAGGTGGACTTCTCAAACCAACAGTGGCCTCTCAGAACCAGAACCTTCCTGTTGCCAAACTCCCACCTAGCA AGTTAGTATCTGATGACTTGGATTCATCTTTAGCCAACCTTGTGGGCAATCTTGGCATCGGAAATGGAACCACTAAGAATGATGTAAATT GGAGTCAACCAGGTGAAAAGAAGTTAACTGGGGGATCTAACTGGCAACCAAAGGTTGCACCAACAACCGCTTGGAATGCTGCAACAATGG CACCCCCTGTAATGGCCTATCCTGCTACTACACCAACAGGCATGATAGGATATGGAATTCCTCCACAAATGGGAAGTGTTCCTGTAATGA CGCAACCAACCTTAATATACAGCCAGCCTGTCATGAGACCTCCAAACCCCTTTGGCCCTGTATCAGGAGCACAGATACAGTTTATGTAAC TTGATGGAAGAAAATGGAATTACTCCAAAAAGACAAGTGCTCAAGCAGCAAAATCCTTACTTCCAGCAAAATCCAAACTGCTGTCTCTTA AATCTCTTAAACTCTCTTCTTCCATTAGAATGCTACAAGTAACTCAGTGAAGGCCCATGAAGGAAATTGGGACTAGTTTATAGGAGAACG TATCAATACAGTTTATAAAGCCAAGAATTGCTATGATTTAAGACTAAGATCTGTCTTTTTGGTGACTAACCCTTCAATTCTTTCAACTCC TGTTAATACCCATAATCAGTAACCTATCAAGAAAAGCCCTTATTTGGAAAGTGTGAAATTTGTATTTGGAAAAGCTGCCTGGAGAGAAGA ACTGTGTCCTTTACTGTATTTCAACAGGACTCTTTTGGGGGATCAAAATTAAAATTCCTAATTATGCATTATCTTTCTTTTCTCCAGTCC TCACAAATACAGAAACAATAACTGAAATTAACTTTTCTTTTTTTAAAAAAAATTATATTCAGTTTGCAGTAGACATTCCTTAAGTATTTG TATTTATTTATGATTATCAATTTTACATAACATTAATATTGTATCAGACCTCCTTATGAAAATGAGTATGGATGTGCACAGTATGTTTGA TTTTTATCCACAAGAATGAATCTGATTCAGAATGCTTTTCTCAGCTGACATACAGAGCACTAAATATTTTAAGGCAAGTCCATAGGTCTG AATCTCTTAAGAATTCTCGGCCTCTGTGGGATTTAGGGAAGCATTATAAATGCATTAATCCTTATAGTCAATTCTGTGCCTAGGATTTTG CCAGGGAACAGTTCACTGACTAGGAAAAGCACTACATTTTAAATTCAGCATTAGTGCATTGGGAAGGATCTTTACTGCTTTGTGCTTGGC ATGTCATTATTTTCCATTTGACATTAGGGCCTTTCCAAAATGAATGTGAGGAATTGCTTTCACTTCAAGACTTTCCTTCTTTTCACTAAA ACTCTAGAAGGTGTTACAAGGGGGAGGGAAGGGGGGCAAAGTCCTTGAACATTTTCTTTGGCTCGTGCCATGTTATGATCATATACCTTT TAAATAAGGGGAAATAGTATCTTTAAAGTTAATGTCTAGCCAAGAGTTTAGTAAACGAAGAATTAAACTGCACTGTTGATCGGTGCTTTG TGTAAATACATCTTTAACATTTGGGTGGAGAGGGGCCTTAAGAAGGACAGTTCATTGTAGGAAAGCAATTCTGTACATGAGTTTAAGCAT TCTTGTTGCATTGTCTCTGCAGATTCTATTTTTGTTTACAATATTAAAATGTATGTTAGCAAAATGGGTGGATTTTCAAATAAAATGCAG >91614_91614_2_TMEM126B-PICALM_TMEM126B_chr11_85346822_ENST00000534341_PICALM_chr11_85742653_ENST00000526033_length(amino acids)=606AA_BP=5 MYVENLIQCTNEMNVNIPQLADSLFERTTNSSWVVVFKSLITTHHLMVYGNERFIQYLASRNTLFNLSNFLDKSGLQGYDMSTFIRRYSR YLNEKAVSYRQVAFDFTKVKRGADGVMRTMNTEKLLKTVPIIQNQMDALLDFNVNSNELTNGVINAAFMLLFKDAIRLFAAYNEGIINLL EKYFDMKKNQCKEGLDIYKKFLTRMTRISEFLKVAEQVGIDRGDIPDLSQAPSSLLDALEQHLASLEGKKIKDSTAASRATTLSNAVSSL ASTGLSLTKVDEREKQAALEEEQARLKALKEQRLKELAKKPHTSLTTAASPVSTSAGGIMTAPAIDIFSTPSSSNSTSKLPNDLLDLQQP TFHPSVHPMSTASQVASTWGDAVDDAIPSLNPFLTKSSGDVHLSISSDVSTFTTRTPTHEMFVGFTPSPVAQPHPSAGLNVDFESVFGNK STNVIVDSGGFDELGGLLKPTVASQNQNLPVAKLPPSKLVSDDLDSSLANLVGNLGIGNGTTKNDVNWSQPGEKKLTGGSNWQPKVAPTT -------------------------------------------------------------- >91614_91614_3_TMEM126B-PICALM_TMEM126B_chr11_85346822_ENST00000534341_PICALM_chr11_85742653_ENST00000532317_length(transcript)=4226nt_BP=1161nt ACCAAAATGGTGGTGTTCGGGTATGAGGCTGGGACTAAGCCAAGGGATTCAGGTGTGGTGCCGGTGGGAACTGAGGAAGCGCCCAAGGTT TTCAAGATGGCAGCATCTATGCATGGTCAGCCCAGTCCTTCTCTAGAAGATGCAAAACTCAGAAGACCAATGGTCATAGAAATCATAGAA AAAAATTTTGACTATCTTAGAAAAGAAATGACACAAAATATATATCAAATGGCGACATTTGGAACAACAGCTGGTTTCTCTGGAATATTC TCAAACTTCCTGTTCAGACGCTGCTTCAAGGTTAAACATGATGCTTTGAAGACATATGCATCATTGGCTACACTTCCATTTTTGTCTACT GTTGTTACTGACAAGCTTTTTGTAATTGATGCTTTGTATTCAGGTAAGTTCTTCCTTTTCCTTCTTTTTTCTTTTCTTTTCTTTTCTTTT TTTTTTTTTTTTTTAGGTTAATAAACTCTCTTTCTTCTTTAGCTGTCTATGATATATTTGAATCAAAATGTGGTAGTTTGGTATTTCAGA TAGCTAACTTTTAACTTTTTTGCTCTCTACATTATAGATCTAGAAAGGCAATTATCAATTTAGGAGAAGTCAACAAACTTCTAATAGCAT TCATATCCAGGAGAGGTGATTAACTCTTTTTTCTTTTTAGGTATCATACCGTTCCACTGCCACCAAAAGGAAGGGTTTTAATCCATTGGA TGACGCTTTGTCAAACACAAATGAAATTAATGGCGATTCCTCTAGTCTTTCAGATTATGTTTGGAATATTAAATGGTCTATACCATTATG CAGTATTTGAAGAGACACTTGAGAAAACTATACATGAAGAGTAACCAAAAAAATGAATGGTTGCTAACTTAGCAAAATGAAGTTTCTATA AAGAGGACTCAGGCATTGCTGAAAGAGTTAAAAGTAACTGTGAACAAATAATTTGTTCTGTGCCTTTTGCCTGGTATATAGCAAATACTC AAAAAGTATTCAATAATTCAATCAATAAATATAAGTTTCATCTTACACGTAAGATACAGGTCTTATCTCCTGATGGTGTGTCCATTTTGC CTGGTATATAACAGATAATAAATATCCAGTGTCAATAAATGTAACAATAAAAGTTTCATCTTTCCTCTTTGTATGTGGAAAACTTAATTC AGTGCACAAATGAGATGAATGTGAACATCCCACAGTTGGCAGACAGTTTATTTGAAAGAACTACTAATAGTAGTTGGGTGGTGGTCTTCA AATCTCTCATTACAACTCATCATTTGATGGTGTATGGAAATGAGCGTTTTATTCAGTATTTGGCTTCAAGAAACACGTTGTTTAACTTAA GCAATTTTTTGGATAAAAGTGGATTGCAAGGATATGACATGTCTACATTTATTAGGCGGTATAGTAGATATTTAAATGAGAAAGCAGTTT CATACAGACAAGTTGCATTTGATTTCACAAAAGTGAAGAGAGGGGCTGATGGAGTTATGAGAACAATGAACACAGAAAAACTCCTAAAAA CTGTACCAATTATTCAGAATCAGATGGATGCACTTCTTGATTTTAATGTTAATAGCAATGAACTTACAAATGGGGTAATAAATGCTGCCT TCATGCTCCTGTTCAAAGATGCCATTAGACTGTTTGCAGCATACAATGAAGGAATTATTAATTTGTTGGAAAAATATTTTGATATGAAAA AGAACCAATGCAAAGAAGGTCTTGACATCTATAAGAAGTTCCTAACTAGGATGACAAGAATCTCAGAGTTCCTCAAAGTTGCAGAGCAAG TTGGAATTGACAGAGGTGATATACCAGACCTTTCACAGGCCCCTAGCAGTCTTCTTGATGCTTTGGAACAACATTTAGCTTCCTTGGAAG GAAAGAAAATCAAAGATTCTACAGCTGCAAGCAGGGCAACTACACTTTCCAATGCAGTGTCTTCCCTGGCAAGCACTGGTCTATCTCTGA CCAAAGTGGATGAAAGGGAAAAGCAGGCAGCATTAGAGGAAGAACAGGCACGTTTGAAAGCTTTAAAGGAACAGCGCCTAAAAGAACTTG CAAAGAAACCTCATACCTCTTTAACAACTGCAGCCTCTCCTGTATCCACCTCAGCAGGAGGGATAATGACTGCACCAGCCATTGACATAT TTTCTACCCCTAGTTCTTCTAACAGCACATCAAAGCTGCCCAATGATCTGCTTGATTTGCAGCAGCCAACTTTTCACCCATCTGTACATC CTATGTCAACTGCTTCTCAGGTAGCAAGTACATGGGGAGGATTCACTCCTTCTCCAGTTGCACAGCCACACCCTTCAGCTGGCCTTAATG TTGACTTTGAATCTGTGTTTGGAAATAAATCTACAAATGTTATTGTAGATTCTGGGGGCTTTGATGAACTAGGTGGACTTCTCAAACCAA CAGTGGCCTCTCAGAACCAGAACCTTCCTGTTGCCAAACTCCCACCTAGCAAGTTAGTATCTGATGACTTGGATTCATCTTTAGCCAACC TTGTGGGCAATCTTGGCATCGGAAATGGAACCACTAAGAATGATGTAAATTGGAGTCAACCAGGTGAAAAGAAGTTAACTGGGGGATCTA ACTGGCAACCAAAGGTTGCACCAACAACCGCTTGGAATGCTGCAACAATGAATGGCATGCATTTTCCACAATACGCACCCCCTGTAATGG CCTATCCTGCTACTACACCAACAGGCATGATAGGATATGGAATTCCTCCACAAATGGGAAGTGTTCCTGTAATGACGCAACCAACCTTAA TATACAGCCAGCCTGTCATGAGACCTCCAAACCCCTTTGGCCCTGTATCAGGAGCACAGATACAGTTTATGTAACTTGATGGAAGAAAAT GGAATTACTCCAAAAAGACAAGTGCTCAAGCAGCAAAATCCTTACTTCCAGCAAAATCCAAACTGCTGTCTCTTAAATCTCTTAAACTCT CTTCTTCCATTAGAATGCTACAAGTAACTCAGTGAAGGCCCATGAAGGAAATTGGGACTAGTTTATAGGAGAACGTATCAATACAGTTTA TAAAGCCAAGAATTGCTATGATTTAAGACTAAGATCTGTCTTTTTGGTGACTAACCCTTCAATTCTTTCAACTCCTGTTAATACCCATAA TCAGTAACCTATCAAGAAAAGCCCTTATTTGGAAAGTGTGAAATTTGTATTTGGAAAAGCTGCCTGGAGAGAAGAACTGTGTCCTTTACT GTATTTCAACAGGACTCTTTTGGGGGATCAAAATTAAAATTCCTAATTATGCATTATCTTTCTTTTCTCCAGTCCTCACAAATACAGAAA CAATAACTGAAATTAACTTTTCTTTTTTTAAAAAAAATTATATTCAGTTTGCAGTAGACATTCCTTAAGTATTTGTATTTATTTATGATT ATCAATTTTACATAACATTAATATTGTATCAGACCTCCTTATGAAAATGAGTATGGATGTGCACAGTATGTTTGATTTTTATCCACAAGA ATGAATCTGATTCAGAATGCTTTTCTCAGCTGACATACAGAGCACTAAATATTTTAAGGCAAGTCCATAGGTCTGAATCTCTTAAGAATT CTCGGCCTCTGTGGGATTTAGGGAAGCATTATAAATGCATTAATCCTTATAGTCAATTCTGTGCCTAGGATTTTGCCAGGGAACAGTTCA CTGACTAGGAAAAGCACTACATTTTAAATTCAGCATTAGTGCATTGGGAAGGATCTTTACTGCTTTGTGCTTGGCATGTCATTATTTTCC ATTTGACATTAGGGCCTTTCCAAAATGAATGTGAGGAATTGCTTTCACTTCAAGACTTTCCTTCTTTTCACTAAAACTCTAGAAGGTGTT ACAAGGGGGAGGGAAGGGGGGCAAAGTCCTTGAACATTTTCTTTGGCTCGTGCCATGTTATGATCATATACCTTTTAAATAAGGGGAAAT AGTATCTTTAAAGTTAATGTCTAGCCAAGAGTTTAGTAAACGAAGAATTAAACTGCACTGTTGATCGGTGCTTTGTGTAAATACATCTTT AACATTTGGGTGGAGAGGGGCCTTAAGAAGGACAGTTCATTGTAGGAAAGCAATTCTGTACATGAGTTTAAGCATTCTTGTTGCATTGTC >91614_91614_3_TMEM126B-PICALM_TMEM126B_chr11_85346822_ENST00000534341_PICALM_chr11_85742653_ENST00000532317_length(amino acids)=571AA_BP=5 MYVENLIQCTNEMNVNIPQLADSLFERTTNSSWVVVFKSLITTHHLMVYGNERFIQYLASRNTLFNLSNFLDKSGLQGYDMSTFIRRYSR YLNEKAVSYRQVAFDFTKVKRGADGVMRTMNTEKLLKTVPIIQNQMDALLDFNVNSNELTNGVINAAFMLLFKDAIRLFAAYNEGIINLL EKYFDMKKNQCKEGLDIYKKFLTRMTRISEFLKVAEQVGIDRGDIPDLSQAPSSLLDALEQHLASLEGKKIKDSTAASRATTLSNAVSSL ASTGLSLTKVDEREKQAALEEEQARLKALKEQRLKELAKKPHTSLTTAASPVSTSAGGIMTAPAIDIFSTPSSSNSTSKLPNDLLDLQQP TFHPSVHPMSTASQVASTWGGFTPSPVAQPHPSAGLNVDFESVFGNKSTNVIVDSGGFDELGGLLKPTVASQNQNLPVAKLPPSKLVSDD LDSSLANLVGNLGIGNGTTKNDVNWSQPGEKKLTGGSNWQPKVAPTTAWNAATMNGMHFPQYAPPVMAYPATTPTGMIGYGIPPQMGSVP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for TMEM126B-PICALM |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000356360 | 0 | 19 | 221_294 | 43.333333333333336 | 633.0 | PIMREG | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000393346 | 0 | 20 | 221_294 | 43.333333333333336 | 653.0 | PIMREG | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000526033 | 0 | 20 | 221_294 | 43.333333333333336 | 646.0 | PIMREG | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000528398 | 0 | 19 | 221_294 | 0 | 552.0 | PIMREG | |
| Tgene | PICALM | chr11:85346822 | chr11:85742653 | ENST00000532317 | 0 | 20 | 221_294 | 43.333333333333336 | 611.0 | PIMREG |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for TMEM126B-PICALM |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for TMEM126B-PICALM |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
