
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:TMEM241-PACS1 (FusionGDB2 ID:91961) |
Fusion Gene Summary for TMEM241-PACS1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: TMEM241-PACS1 | Fusion gene ID: 91961 | Hgene | Tgene | Gene symbol | TMEM241 | PACS1 | Gene ID | 85019 | 55690 |
| Gene name | transmembrane protein 241 | phosphofurin acidic cluster sorting protein 1 | |
| Synonyms | C18orf45|hVVT | MRD17|SHMS | |
| Cytomap | 18q11.2 | 11q13.1-q13.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | transmembrane protein 241putative vertebrate vrg4-like nucleotide-sugar transporter truncated variant2putative vertebrate vrg4-like nucleotide-sugar transporter variant1transmembrane protein C18orf45 | phosphofurin acidic cluster sorting protein 1PACS-1cytosolic sorting protein PACS-1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q24JQ0 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000542162, ENST00000383233, ENST00000450466, ENST00000475185, ENST00000399707, | ENST00000320580, ENST00000524815, ENST00000529757, | |
| Fusion gene scores | * DoF score | 11 X 10 X 9=990 | 21 X 18 X 12=4536 |
| # samples | 13 | 26 | |
| ** MAII score | log2(13/990*10)=-2.92891690193852 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(26/4536*10)=-4.12483711191377 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: TMEM241 [Title/Abstract] AND PACS1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | TMEM241(20889644)-PACS1(65960957), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across TMEM241 (5'-gene) Fusion gene breakpoints across TMEM241 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
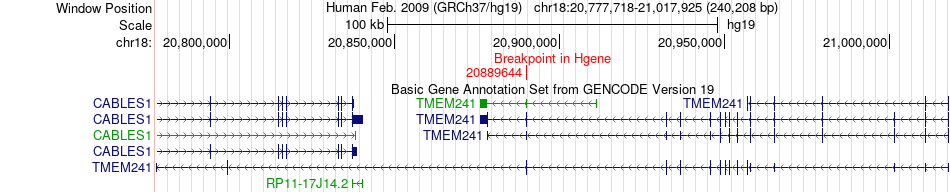 |
 Fusion gene breakpoints across PACS1 (3'-gene) Fusion gene breakpoints across PACS1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
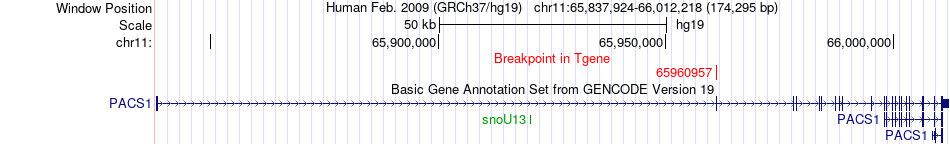 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | HNSC | TCGA-UF-A71B-01A | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
Top |
Fusion Gene ORF analysis for TMEM241-PACS1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000542162 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 3UTR-intron | ENST00000542162 | ENST00000524815 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 3UTR-intron | ENST00000542162 | ENST00000529757 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5CDS-intron | ENST00000383233 | ENST00000524815 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5CDS-intron | ENST00000383233 | ENST00000529757 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5CDS-intron | ENST00000450466 | ENST00000524815 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5CDS-intron | ENST00000450466 | ENST00000529757 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5UTR-3CDS | ENST00000475185 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5UTR-intron | ENST00000475185 | ENST00000524815 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| 5UTR-intron | ENST00000475185 | ENST00000529757 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| In-frame | ENST00000383233 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| In-frame | ENST00000450466 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| intron-3CDS | ENST00000399707 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| intron-intron | ENST00000399707 | ENST00000524815 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
| intron-intron | ENST00000399707 | ENST00000529757 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000450466 | TMEM241 | chr18 | 20889644 | - | ENST00000320580 | PACS1 | chr11 | 65960957 | + | 4946 | 943 | 476 | 3478 | 1000 |
| ENST00000383233 | TMEM241 | chr18 | 20889644 | - | ENST00000320580 | PACS1 | chr11 | 65960957 | + | 4886 | 883 | 53 | 3418 | 1121 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000450466 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + | 0.000852252 | 0.9991478 |
| ENST00000383233 | ENST00000320580 | TMEM241 | chr18 | 20889644 | - | PACS1 | chr11 | 65960957 | + | 0.000519203 | 0.9994808 |
Top |
Fusion Genomic Features for TMEM241-PACS1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| TMEM241 | chr18 | 20889643 | - | PACS1 | chr11 | 65960956 | + | 6.08E-06 | 0.9999939 |
| TMEM241 | chr18 | 20889643 | - | PACS1 | chr11 | 65960956 | + | 6.08E-06 | 0.9999939 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
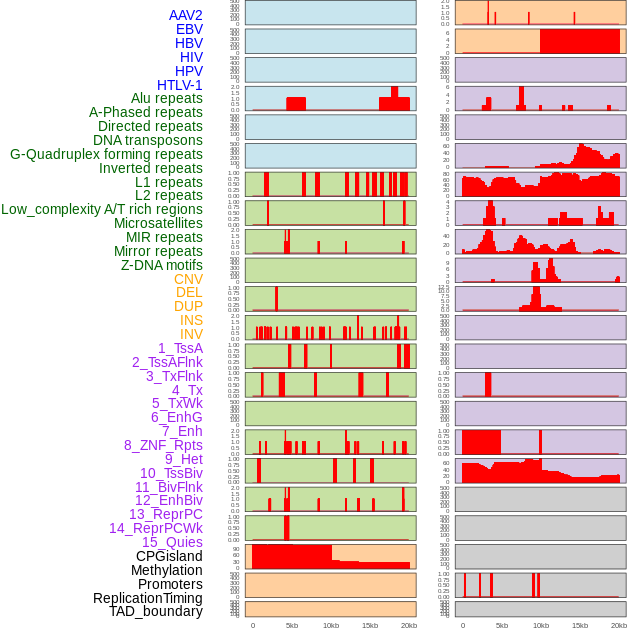 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
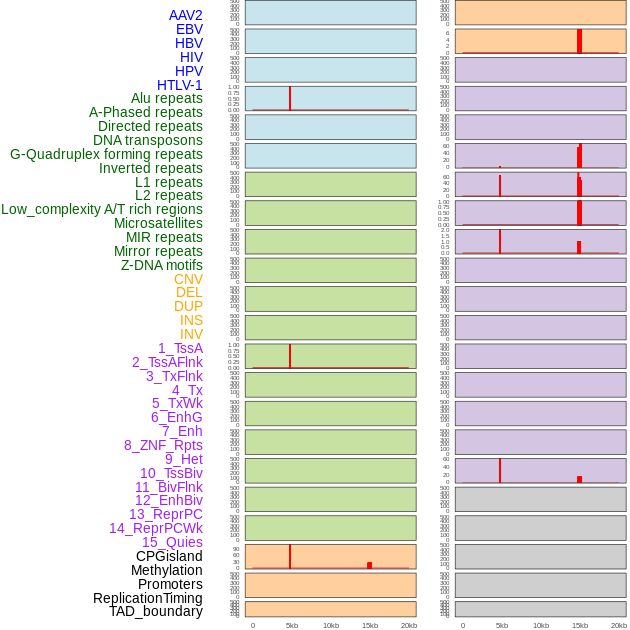 |
Top |
Fusion Protein Features for TMEM241-PACS1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr18:20889644/chr11:65960957) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| TMEM241 | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 121_141 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 146_166 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 187_207 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 211_231 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 250_270 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 32_52 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 67_87 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 7_29 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 93_113 | 276 | 297.0 | Transmembrane | Helical |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 353_377 | 118 | 964.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 279_283 | 118 | 964.0 | Compositional bias | Note=Poly-Glu | |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 168_175 | 118 | 964.0 | Region | Note=Involved in binding to AP-1 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000383233 | - | 14 | 15 | 271_291 | 276 | 297.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 121_141 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 146_166 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 187_207 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 211_231 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 250_270 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 271_291 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 32_52 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 67_87 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 7_29 | 0 | 124.0 | Transmembrane | Helical |
| Hgene | TMEM241 | chr18:20889644 | chr11:65960957 | ENST00000399707 | - | 1 | 6 | 93_113 | 0 | 124.0 | Transmembrane | Helical |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 112_115 | 118 | 964.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 35_40 | 118 | 964.0 | Compositional bias | Note=Poly-Gln | |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 55_84 | 118 | 964.0 | Compositional bias | Note=Ser-rich | |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 5_44 | 118 | 964.0 | Compositional bias | Note=Gly-rich | |
| Tgene | PACS1 | chr18:20889644 | chr11:65960957 | ENST00000320580 | 0 | 24 | 62_65 | 118 | 964.0 | Compositional bias | Note=Poly-Ala |
Top |
Fusion Gene Sequence for TMEM241-PACS1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >91961_91961_1_TMEM241-PACS1_TMEM241_chr18_20889644_ENST00000383233_PACS1_chr11_65960957_ENST00000320580_length(transcript)=4886nt_BP=883nt CCAGGGGATTTGGGCAGGCACCGTGGATCCCCGAGAAGGGGACGAGTTGACAGATGTGCGTGAGGAGGTCTCTGGTCGGCCTCACCTTTT GTACCTGCTACCTGGCTTCTTACCTCACGAACAAGTATGTGCTGTCTGTCTTGAAATTTACCTACCCTACATTATTCCAAGGGTGGCAGA CGCTCATTGGTGGACTTTTGCTTCATGTGTCCTGGAAACTGGGCTGGGTAGAGATCAACAGCAGTTCAAGATCTCATGTTCTTGTGTGGC TTCCTGCTTCAGTGCTGTTTGTGGGTATAATCTATGCTGGGTCCAGAGCATTGTCCAGACTGGCCATTCCTGTGTTTCTCACTTTGCATA ATGTAGCTGAAGTTATCATCTGTGGGTACCAGAAGTGTTTTCAGAAAGAGAAAACATCTCCTGCAAAGATCTGTAGTGCCCTCCTCCTCC TGGCCGCAGCAGGATGCCTTCCCTTCAATGACTCCCAGTTTAATCCAGATGGATATTTTTGGGCTATAATTCACTTACTCTGTGTAGGGG CTTATAAAATTCTACAGAAGTCCCAGAAACCCAGTGCATTAAGTGACATTGACCAGCAATACTTAAACTATATATTCAGTGTGGTGCTCC TGGCATTTGCATCTCATCCCACAGGTGATCTCTTCAGCGTCCTGGACTTCCCATTCCTGTACTTCTACAGATTCCATGGTAGCTGCTGTG CCAGTGGATTTTTGGGATTCTTTCTCATGTTCAGTACAGTGAAGCTAAAAAACCTTCTGGCCCCAGGGCAGTGTGCAGCCTGGATTTTCT TTGCTAAGATAATCACAGCTGGCTTATCAATATTGCTGTTTGATGCGATCCTGACCAGTGCAACCACGGGATGGCTATTCAGCTTGACCC TGAAGAAACTCGTCATGCTAAAAGAAATGGACAAAGATCTTAACTCAGTGGTCATCGCTGTGAAGCTGCAGGGTTCAAAAAGAATTCTTC GCTCCAACGAGATCGTCCTTCCAGCTAGTGGACTGGTGGAAACAGAGCTCCAATTAACCTTCTCCCTTCAGTACCCTCATTTCCTTAAGC GAGATGCCAACAAGCTGCAGATCATGCTGCAAAGGAGAAAACGTTACAAGAATCGGACCATCTTGGGCTATAAGACCTTGGCCGTGGGAC TCATCAACATGGCAGAGGTGATGCAGCATCCTAATGAAGGCGCACTGGTGCTTGGCCTACACAGCAACGTGAAGGATGTCTCTGTGCCTG TGGCAGAAATAAAGATCTACTCCCTGTCCAGCCAACCCATTGACCATGAAGGAATCAAATCCAAGCTTTCTGATCGTTCTCCTGATATTG ACAATTATTCTGAGGAAGAGGAAGAGAGTTTCTCATCAGAACAGGAAGGCAGTGATGATCCATTGCATGGGCAGGACTTGTTCTACGAAG ACGAAGATCTCCGGAAAGTGAAGAAGACCCGGAGGAAACTAACCTCAACCTCTGCCATCACAAGGCAACCTAACATCAAACAGAAGTTTG TGGCCCTCCTGAAGCGGTTTAAAGTTTCAGATGAGGTGGGCTTTGGGCTGGAGCATGTGTCCCGCGAGCAGATCCGGGAAGTGGAAGAGG ACTTGGATGAATTGTATGACAGTCTGGAGATGTACAACCCCAGCGACAGTGGCCCTGAGATGGAGGAGACAGAAAGCATCCTCAGCACGC CAAAGCCCAAGCTCAAGCCTTTCTTTGAGGGGATGTCGCAGTCCAGCTCCCAGACGGAGATTGGCAGCCTCAACAGCAAAGGCAGCCTCG GAAAAGACACCACCAGCCCTATGGAATTGGCTGCTCTAGAAAAAATTAAATCTACTTGGATTAAAAACCAAGATGACAGCTTGACTGAAA CAGACACTCTGGAAATCACTGACCAGGACATGTTTGGAGATGCCAGCACGAGTCTGGTTGTGCCGGAGAAAGTCAAAACTCCCATGAAGT CCAGTAAAACGGATCTCCAGGGCTCTGCCTCCCCCAGCAAAGTGGAGGGGGTGCACACACCCCGGCAGAAGAGGAGCACGCCCCTGAAGG AGCGGCAGCTCTCCAAGCCCCTAAGTGAGAGGACCAACAGTTCCGACAGCGAGCGCTCCCCAGATCTGGGCCACAGCACGCAGATTCCAA GAAAGGTGGTGTATGACCAGCTCAATCAGATCCTGGTGTCAGATGCAGCCCTCCCAGAAAATGTCATTCTGGTGAACACCACTGACTGGC AGGGCCAGTATGTGGCTGAGCTGCTCCAGGACCAGCGGAAGCCTGTGGTGTGCACCTGCTCCACCGTGGAGGTCCAGGCCGTGCTGTCCG CCCTGCTCACCCGGATCCAGCGCTACTGCAACTGCAACTCTTCCATGCCGAGGCCAGTGAAGGTGGCTGCTGTGGGAGGCCAGAGCTACC TGAGCTCCATCCTCAGGTTCTTTGTCAAGTCCCTGGCCAACAAGACCTCCGACTGGCTTGGCTACATGCGCTTCCTCATCATCCCCCTCG GTTCTCACCCTGTGGCCAAATACTTGGGGTCAGTCGACAGTAAATACAGTAGTTCCTTCCTGGATTCTGGTTGGAGAGATCTGTTCAGTC GCTCGGAGCCACCAGTGTCAGAGCAACTGGACGTGGCAGGGCGGGTGATGCAGTACGTCAACGGGGCAGCCACGACACACCAGCTTCCCG TGGCCGAAGCCATGCTGACTTGCCGGCATAAGTTCCCTGATGAAGACTCCTATCAGAAGTTTATTCCCTTCATTGGCGTGGTGAAGGTGG GTCTGGTTGAAGACTCTCCCTCCACAGCAGGCGATGGGGACGATTCTCCTGTGGTCAGCCTTACTGTGCCCTCCACATCACCACCCTCCA GCTCGGGCCTGAGCCGAGACGCCACGGCCACCCCTCCCTCCTCCCCATCTATGAGCAGCGCCCTGGCCATCGTGGGGAGCCCTAATAGCC CATATGGGGACGTGATTGGCCTCCAGGTGGACTACTGGCTGGGCCACCCCGGGGAGCGGAGGAGGGAAGGCGACAAGAGGGACGCCAGCT CGAAGAACACCCTCAAGAGTGTCTTCCGCTCAGTGCAGGTGTCCCGCCTGCCCCATAGTGGGGAGGCCCAGCTTTCTGGCACCATGGCCA TGACTGTGGTCACCAAAGAAAAGAACAAGAAAGTTCCCACCATCTTCCTGAGCAAGAAACCCCGAGAAAAGGAGGTGGATTCTAAGAGCC AGGTCATTGAAGGCATCAGCCGCCTCATCTGCTCAGCCAAGCAGCAGCAGACTATGCTGAGAGTGTCCATCGATGGGGTCGAGTGGAGTG ACATCAAGTTCTTCCAGCTGGCAGCCCAGTGGCCCACCCATGTCAAGCACTTTCCAGTGGGACTCTTCAGTGGCAGCAAGGCCACCTGAG GCCCTGTCTCCCAGCCACTTTCCCTCCTGGCACTGCCACCAGCCTCACCGCCTGCGGGCAGGGGGAGGCCAGCAGGCCCGGGCCCAGCAC CCCTTCCCTGGCACCAGGGTCTGCCTCTCACTCGCCCAGGTCCCGAAGGACACTGCCACAGGGACGCCTTCCCTCCCCTCCCCTCCAGCC CACCCCTGCACAGCCCCTCCTCCTTCCCGCTTTTCCCCTTCTCCCTCCTGCTCCAGGCCCAAGGCGTGTTGGTTTTGCCTTCTGGTGCCC ATAGTCCCCTGGACTGAGTCCCCCAGGCCTTCCTTCACCCGACTTCCAAACTCTTCCTTGTGGTATCAGTTTCCTTCTCGGAAATGAGAA AGCTGGAATCCTGGTCCCCAGCAGGAGAGCCTAGTCCTCCCCCAGCCCCTCCAGCCACCAGGGTGTCCTCTAGGATGCAGCTGCCAGATC CACTCACTCTGCTGCCTCCAGCAGGACCCAAGGCCACTTTCAACTCTTATGGGGTTCTCCACCTGCCCCAGAGCTTCTCAAGGGAGGGTA AGGGGGCACCCTGAGCCCACAGGACCCCTACTTCACAGCTCACAGGGGCAGGAGGCAGCTCCCCTGCCTCCAGGACCCTGTTGCTATGGT GACACAGCGTTTCTAGGACAGAGGGGCCTCCCAGTCTCCCCCCACCACCCGTGCACGACTTCCTCACCACCCCCAGGTTCCCTGCAGATG TCGTGTGTGTCCTGAGTGTTTCTTTGGTTCTTTGCACGCCAAGTCTCTTGGTTGTACCATGTGACACACCCTGTGCACTGGTCGCTGTCT TCGTGGCTTCCACCCTTGTTAATGATGCTCCTGCCTCTGCCTCCCAGCCCCTCACCCAGCACAGCTCTGCCTGGACTTGGAGAGATGGGA GGCAGACCCCCACCACCATACATGCTGTCTGTGGCCCCTCAGACATTCTGTTTCATCTCCCATTCATCTCCCTCCTCCCACCGTGTCAGT TTTTCTGCCTTTCCCTGCTCTGTTCTTCCCCCTCCTTAGGCCCCAGCCTGGGCCCAGACCCATCCTCCCAGCCAGGTTTCCCTCCAGCAG GCTCCTTCCCTCCCTGTCACCTCCCTCTCACCAACCCGGGGTCTGAGCCCCTCATTCCTGACCGTCCGTGTTCTCAGGAGTGGTTGAGGA CACAGGGCCCCAGCCCAGCCCTCTGCACCCCCCAGCCCGGCCATCTGCGCCCCACAGCCCCTTTGGAGCTTTTCTCTTGTCCTCTCACTC CTTCCCAGAAGTTTTTGCACAGAACTTCATTTTGAAAGTGTTTTTCTCATTCTCCATACCTCCCCCAAGCTCTCCTCCAGCCCTTCCCAG GGCTCAGCCCTGCTGTCCTGAGCGTCTCCTGGGCCAGAGAGAGGAGATGGGGGTGGGAGGGACTGAGTTGATGTTGGGTTTTTCATTCAA >91961_91961_1_TMEM241-PACS1_TMEM241_chr18_20889644_ENST00000383233_PACS1_chr11_65960957_ENST00000320580_length(amino acids)=1121AA_BP=277 MCVRRSLVGLTFCTCYLASYLTNKYVLSVLKFTYPTLFQGWQTLIGGLLLHVSWKLGWVEINSSSRSHVLVWLPASVLFVGIIYAGSRAL SRLAIPVFLTLHNVAEVIICGYQKCFQKEKTSPAKICSALLLLAAAGCLPFNDSQFNPDGYFWAIIHLLCVGAYKILQKSQKPSALSDID QQYLNYIFSVVLLAFASHPTGDLFSVLDFPFLYFYRFHGSCCASGFLGFFLMFSTVKLKNLLAPGQCAAWIFFAKIITAGLSILLFDAIL TSATTGWLFSLTLKKLVMLKEMDKDLNSVVIAVKLQGSKRILRSNEIVLPASGLVETELQLTFSLQYPHFLKRDANKLQIMLQRRKRYKN RTILGYKTLAVGLINMAEVMQHPNEGALVLGLHSNVKDVSVPVAEIKIYSLSSQPIDHEGIKSKLSDRSPDIDNYSEEEEESFSSEQEGS DDPLHGQDLFYEDEDLRKVKKTRRKLTSTSAITRQPNIKQKFVALLKRFKVSDEVGFGLEHVSREQIREVEEDLDELYDSLEMYNPSDSG PEMEETESILSTPKPKLKPFFEGMSQSSSQTEIGSLNSKGSLGKDTTSPMELAALEKIKSTWIKNQDDSLTETDTLEITDQDMFGDASTS LVVPEKVKTPMKSSKTDLQGSASPSKVEGVHTPRQKRSTPLKERQLSKPLSERTNSSDSERSPDLGHSTQIPRKVVYDQLNQILVSDAAL PENVILVNTTDWQGQYVAELLQDQRKPVVCTCSTVEVQAVLSALLTRIQRYCNCNSSMPRPVKVAAVGGQSYLSSILRFFVKSLANKTSD WLGYMRFLIIPLGSHPVAKYLGSVDSKYSSSFLDSGWRDLFSRSEPPVSEQLDVAGRVMQYVNGAATTHQLPVAEAMLTCRHKFPDEDSY QKFIPFIGVVKVGLVEDSPSTAGDGDDSPVVSLTVPSTSPPSSSGLSRDATATPPSSPSMSSALAIVGSPNSPYGDVIGLQVDYWLGHPG ERRREGDKRDASSKNTLKSVFRSVQVSRLPHSGEAQLSGTMAMTVVTKEKNKKVPTIFLSKKPREKEVDSKSQVIEGISRLICSAKQQQT -------------------------------------------------------------- >91961_91961_2_TMEM241-PACS1_TMEM241_chr18_20889644_ENST00000450466_PACS1_chr11_65960957_ENST00000320580_length(transcript)=4946nt_BP=943nt GCTGCTGCTGCGGGGGTCGGGCGGCGGCCAGGGGATTTGGGCAGGCACCGTGGATCCCCGAGAAGGGGACGAGTTGACAGATGTGCGTGA GGAGGTCTCTGGTCGGCCTCACCTTTTGTACCTGCTACCTGGCTTCTTACCTCACGAACAAGTATGTGCTGTCTGTCTTGAAATTTACCT ACCCTACATTATTCCAAGGGTGGCAGACGCTCATTGGTGGACTTTTGCTTCATGTGTCCTGGAAACTGGGCTGGGTAGAGATCAACAGCA GTTCAAGGCCATTCCTGTGTTTCTCACTTTGCATAATGTAGCTGAAGTTATCATCTGTGGGTACCAGAAGTGTTTTCAGAAAGAGAAAAC ATCTCCTGCAAAGATCTGTAGGTTGCACCTTGAAAGAACAAGACAAAACCAAACTTCAAGACTATCCTCCTGTTTAAAAGGAGACTAGCA GGTGTCAAAGAGAGGCGGTAAAGCTCATGATACCTGATGTAATCAGTGCCCTCCTCCTCCTGGCCGCAGCAGGATGCCTTCCCTTCAATG ACTCCCAGTTTAATCCAGATGGATATTTTTGGGCTATAATTCACTTACTCTGTGTAGGGGCTTATAAAATTCTACAGAAGTCCCAGAAAC CCAGTGCATTAAGTGACATTGACCAGCAATACTTAAACTATATATTCAGTGTGGTGCTCCTGGCATTTGCATCTCATCCCACAGGTGATC TCTTCAGCGTCCTGGACTTCCCATTCCTGTACTTCTACAGATTCCATGGTAGCTGCTGTGCCAGTGGATTTTTGGGATTCTTTCTCATGT TCAGTACAGTGAAGCTAAAAAACCTTCTGGCCCCAGGGCAGTGTGCAGCCTGGATTTTCTTTGCTAAGATAATCACAGCTGGCTTATCAA TATTGCTGTTTGATGCGATCCTGACCAGTGCAACCACGGGATGGCTATTCAGCTTGACCCTGAAGAAACTCGTCATGCTAAAAGAAATGG ACAAAGATCTTAACTCAGTGGTCATCGCTGTGAAGCTGCAGGGTTCAAAAAGAATTCTTCGCTCCAACGAGATCGTCCTTCCAGCTAGTG GACTGGTGGAAACAGAGCTCCAATTAACCTTCTCCCTTCAGTACCCTCATTTCCTTAAGCGAGATGCCAACAAGCTGCAGATCATGCTGC AAAGGAGAAAACGTTACAAGAATCGGACCATCTTGGGCTATAAGACCTTGGCCGTGGGACTCATCAACATGGCAGAGGTGATGCAGCATC CTAATGAAGGCGCACTGGTGCTTGGCCTACACAGCAACGTGAAGGATGTCTCTGTGCCTGTGGCAGAAATAAAGATCTACTCCCTGTCCA GCCAACCCATTGACCATGAAGGAATCAAATCCAAGCTTTCTGATCGTTCTCCTGATATTGACAATTATTCTGAGGAAGAGGAAGAGAGTT TCTCATCAGAACAGGAAGGCAGTGATGATCCATTGCATGGGCAGGACTTGTTCTACGAAGACGAAGATCTCCGGAAAGTGAAGAAGACCC GGAGGAAACTAACCTCAACCTCTGCCATCACAAGGCAACCTAACATCAAACAGAAGTTTGTGGCCCTCCTGAAGCGGTTTAAAGTTTCAG ATGAGGTGGGCTTTGGGCTGGAGCATGTGTCCCGCGAGCAGATCCGGGAAGTGGAAGAGGACTTGGATGAATTGTATGACAGTCTGGAGA TGTACAACCCCAGCGACAGTGGCCCTGAGATGGAGGAGACAGAAAGCATCCTCAGCACGCCAAAGCCCAAGCTCAAGCCTTTCTTTGAGG GGATGTCGCAGTCCAGCTCCCAGACGGAGATTGGCAGCCTCAACAGCAAAGGCAGCCTCGGAAAAGACACCACCAGCCCTATGGAATTGG CTGCTCTAGAAAAAATTAAATCTACTTGGATTAAAAACCAAGATGACAGCTTGACTGAAACAGACACTCTGGAAATCACTGACCAGGACA TGTTTGGAGATGCCAGCACGAGTCTGGTTGTGCCGGAGAAAGTCAAAACTCCCATGAAGTCCAGTAAAACGGATCTCCAGGGCTCTGCCT CCCCCAGCAAAGTGGAGGGGGTGCACACACCCCGGCAGAAGAGGAGCACGCCCCTGAAGGAGCGGCAGCTCTCCAAGCCCCTAAGTGAGA GGACCAACAGTTCCGACAGCGAGCGCTCCCCAGATCTGGGCCACAGCACGCAGATTCCAAGAAAGGTGGTGTATGACCAGCTCAATCAGA TCCTGGTGTCAGATGCAGCCCTCCCAGAAAATGTCATTCTGGTGAACACCACTGACTGGCAGGGCCAGTATGTGGCTGAGCTGCTCCAGG ACCAGCGGAAGCCTGTGGTGTGCACCTGCTCCACCGTGGAGGTCCAGGCCGTGCTGTCCGCCCTGCTCACCCGGATCCAGCGCTACTGCA ACTGCAACTCTTCCATGCCGAGGCCAGTGAAGGTGGCTGCTGTGGGAGGCCAGAGCTACCTGAGCTCCATCCTCAGGTTCTTTGTCAAGT CCCTGGCCAACAAGACCTCCGACTGGCTTGGCTACATGCGCTTCCTCATCATCCCCCTCGGTTCTCACCCTGTGGCCAAATACTTGGGGT CAGTCGACAGTAAATACAGTAGTTCCTTCCTGGATTCTGGTTGGAGAGATCTGTTCAGTCGCTCGGAGCCACCAGTGTCAGAGCAACTGG ACGTGGCAGGGCGGGTGATGCAGTACGTCAACGGGGCAGCCACGACACACCAGCTTCCCGTGGCCGAAGCCATGCTGACTTGCCGGCATA AGTTCCCTGATGAAGACTCCTATCAGAAGTTTATTCCCTTCATTGGCGTGGTGAAGGTGGGTCTGGTTGAAGACTCTCCCTCCACAGCAG GCGATGGGGACGATTCTCCTGTGGTCAGCCTTACTGTGCCCTCCACATCACCACCCTCCAGCTCGGGCCTGAGCCGAGACGCCACGGCCA CCCCTCCCTCCTCCCCATCTATGAGCAGCGCCCTGGCCATCGTGGGGAGCCCTAATAGCCCATATGGGGACGTGATTGGCCTCCAGGTGG ACTACTGGCTGGGCCACCCCGGGGAGCGGAGGAGGGAAGGCGACAAGAGGGACGCCAGCTCGAAGAACACCCTCAAGAGTGTCTTCCGCT CAGTGCAGGTGTCCCGCCTGCCCCATAGTGGGGAGGCCCAGCTTTCTGGCACCATGGCCATGACTGTGGTCACCAAAGAAAAGAACAAGA AAGTTCCCACCATCTTCCTGAGCAAGAAACCCCGAGAAAAGGAGGTGGATTCTAAGAGCCAGGTCATTGAAGGCATCAGCCGCCTCATCT GCTCAGCCAAGCAGCAGCAGACTATGCTGAGAGTGTCCATCGATGGGGTCGAGTGGAGTGACATCAAGTTCTTCCAGCTGGCAGCCCAGT GGCCCACCCATGTCAAGCACTTTCCAGTGGGACTCTTCAGTGGCAGCAAGGCCACCTGAGGCCCTGTCTCCCAGCCACTTTCCCTCCTGG CACTGCCACCAGCCTCACCGCCTGCGGGCAGGGGGAGGCCAGCAGGCCCGGGCCCAGCACCCCTTCCCTGGCACCAGGGTCTGCCTCTCA CTCGCCCAGGTCCCGAAGGACACTGCCACAGGGACGCCTTCCCTCCCCTCCCCTCCAGCCCACCCCTGCACAGCCCCTCCTCCTTCCCGC TTTTCCCCTTCTCCCTCCTGCTCCAGGCCCAAGGCGTGTTGGTTTTGCCTTCTGGTGCCCATAGTCCCCTGGACTGAGTCCCCCAGGCCT TCCTTCACCCGACTTCCAAACTCTTCCTTGTGGTATCAGTTTCCTTCTCGGAAATGAGAAAGCTGGAATCCTGGTCCCCAGCAGGAGAGC CTAGTCCTCCCCCAGCCCCTCCAGCCACCAGGGTGTCCTCTAGGATGCAGCTGCCAGATCCACTCACTCTGCTGCCTCCAGCAGGACCCA AGGCCACTTTCAACTCTTATGGGGTTCTCCACCTGCCCCAGAGCTTCTCAAGGGAGGGTAAGGGGGCACCCTGAGCCCACAGGACCCCTA CTTCACAGCTCACAGGGGCAGGAGGCAGCTCCCCTGCCTCCAGGACCCTGTTGCTATGGTGACACAGCGTTTCTAGGACAGAGGGGCCTC CCAGTCTCCCCCCACCACCCGTGCACGACTTCCTCACCACCCCCAGGTTCCCTGCAGATGTCGTGTGTGTCCTGAGTGTTTCTTTGGTTC TTTGCACGCCAAGTCTCTTGGTTGTACCATGTGACACACCCTGTGCACTGGTCGCTGTCTTCGTGGCTTCCACCCTTGTTAATGATGCTC CTGCCTCTGCCTCCCAGCCCCTCACCCAGCACAGCTCTGCCTGGACTTGGAGAGATGGGAGGCAGACCCCCACCACCATACATGCTGTCT GTGGCCCCTCAGACATTCTGTTTCATCTCCCATTCATCTCCCTCCTCCCACCGTGTCAGTTTTTCTGCCTTTCCCTGCTCTGTTCTTCCC CCTCCTTAGGCCCCAGCCTGGGCCCAGACCCATCCTCCCAGCCAGGTTTCCCTCCAGCAGGCTCCTTCCCTCCCTGTCACCTCCCTCTCA CCAACCCGGGGTCTGAGCCCCTCATTCCTGACCGTCCGTGTTCTCAGGAGTGGTTGAGGACACAGGGCCCCAGCCCAGCCCTCTGCACCC CCCAGCCCGGCCATCTGCGCCCCACAGCCCCTTTGGAGCTTTTCTCTTGTCCTCTCACTCCTTCCCAGAAGTTTTTGCACAGAACTTCAT TTTGAAAGTGTTTTTCTCATTCTCCATACCTCCCCCAAGCTCTCCTCCAGCCCTTCCCAGGGCTCAGCCCTGCTGTCCTGAGCGTCTCCT >91961_91961_2_TMEM241-PACS1_TMEM241_chr18_20889644_ENST00000450466_PACS1_chr11_65960957_ENST00000320580_length(amino acids)=1000AA_BP=156 MIPDVISALLLLAAAGCLPFNDSQFNPDGYFWAIIHLLCVGAYKILQKSQKPSALSDIDQQYLNYIFSVVLLAFASHPTGDLFSVLDFPF LYFYRFHGSCCASGFLGFFLMFSTVKLKNLLAPGQCAAWIFFAKIITAGLSILLFDAILTSATTGWLFSLTLKKLVMLKEMDKDLNSVVI AVKLQGSKRILRSNEIVLPASGLVETELQLTFSLQYPHFLKRDANKLQIMLQRRKRYKNRTILGYKTLAVGLINMAEVMQHPNEGALVLG LHSNVKDVSVPVAEIKIYSLSSQPIDHEGIKSKLSDRSPDIDNYSEEEEESFSSEQEGSDDPLHGQDLFYEDEDLRKVKKTRRKLTSTSA ITRQPNIKQKFVALLKRFKVSDEVGFGLEHVSREQIREVEEDLDELYDSLEMYNPSDSGPEMEETESILSTPKPKLKPFFEGMSQSSSQT EIGSLNSKGSLGKDTTSPMELAALEKIKSTWIKNQDDSLTETDTLEITDQDMFGDASTSLVVPEKVKTPMKSSKTDLQGSASPSKVEGVH TPRQKRSTPLKERQLSKPLSERTNSSDSERSPDLGHSTQIPRKVVYDQLNQILVSDAALPENVILVNTTDWQGQYVAELLQDQRKPVVCT CSTVEVQAVLSALLTRIQRYCNCNSSMPRPVKVAAVGGQSYLSSILRFFVKSLANKTSDWLGYMRFLIIPLGSHPVAKYLGSVDSKYSSS FLDSGWRDLFSRSEPPVSEQLDVAGRVMQYVNGAATTHQLPVAEAMLTCRHKFPDEDSYQKFIPFIGVVKVGLVEDSPSTAGDGDDSPVV SLTVPSTSPPSSSGLSRDATATPPSSPSMSSALAIVGSPNSPYGDVIGLQVDYWLGHPGERRREGDKRDASSKNTLKSVFRSVQVSRLPH SGEAQLSGTMAMTVVTKEKNKKVPTIFLSKKPREKEVDSKSQVIEGISRLICSAKQQQTMLRVSIDGVEWSDIKFFQLAAQWPTHVKHFP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for TMEM241-PACS1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for TMEM241-PACS1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for TMEM241-PACS1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
