
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:TNIP1-NIPAL4 (FusionGDB2 ID:92708) |
Fusion Gene Summary for TNIP1-NIPAL4 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: TNIP1-NIPAL4 | Fusion gene ID: 92708 | Hgene | Tgene | Gene symbol | TNIP1 | NIPAL4 | Gene ID | 10318 | 348938 |
| Gene name | TNFAIP3 interacting protein 1 | NIPA like domain containing 4 | |
| Synonyms | ABIN-1|NAF1|VAN|nip40-1 | ARCI6|ICHTHYIN|ICHYN | |
| Cytomap | 5q33.1 | 5q33.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | TNFAIP3-interacting protein 1A20-binding inhibitor of NF-kappa-B activation 1HIV-1 Nef-interacting proteinNef-associated factor 1 SNPvirion-associated nuclear shuttling protein | magnesium transporter NIPA4NIPA-like protein 4non-imprinted in Prader-Willi/Angelman syndrome region protein 4 | |
| Modification date | 20200327 | 20200327 | |
| UniProtAcc | Q15025 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000315050, ENST00000389378, ENST00000518977, ENST00000520931, ENST00000521591, ENST00000522226, ENST00000523200, ENST00000523338, ENST00000524280, ENST00000521423, | ENST00000521390, ENST00000311946, ENST00000435489, | |
| Fusion gene scores | * DoF score | 11 X 10 X 7=770 | 2 X 1 X 1=2 |
| # samples | 11 | 1 | |
| ** MAII score | log2(11/770*10)=-2.8073549220576 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(1/2*10)=2.32192809488736 | |
| Context | PubMed: TNIP1 [Title/Abstract] AND NIPAL4 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | TNIP1(150441688)-NIPAL4(156890102), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | TNIP1-NIPAL4 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. TNIP1-NIPAL4 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | TNIP1 | GO:0009101 | glycoprotein biosynthetic process | 9923610 |
| Hgene | TNIP1 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling | 16684768 |
| Hgene | TNIP1 | GO:0070373 | negative regulation of ERK1 and ERK2 cascade | 12220502 |
| Hgene | TNIP1 | GO:0085032 | modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade | 20010814 |
| Hgene | TNIP1 | GO:1903003 | positive regulation of protein deubiquitination | 16684768 |
 Fusion gene breakpoints across TNIP1 (5'-gene) Fusion gene breakpoints across TNIP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
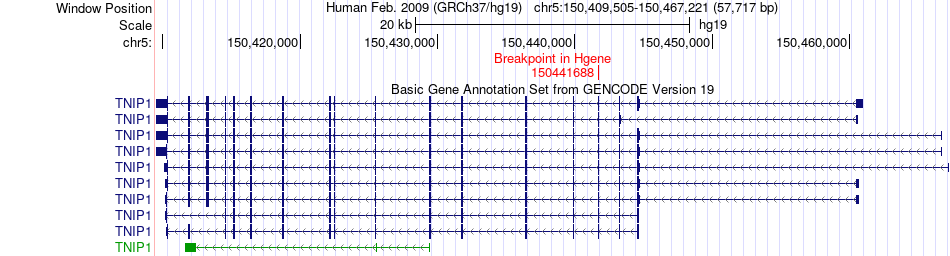 |
 Fusion gene breakpoints across NIPAL4 (3'-gene) Fusion gene breakpoints across NIPAL4 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
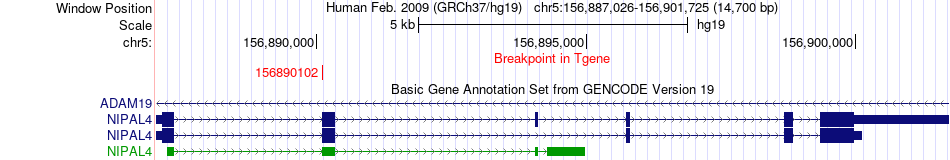 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | KIRC | TCGA-BP-4986-01A | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
Top |
Fusion Gene ORF analysis for TNIP1-NIPAL4 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000315050 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000389378 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000518977 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000520931 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000521591 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000522226 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000523200 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000523338 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| 5CDS-3UTR | ENST00000524280 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000518977 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000518977 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000521591 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000521591 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000522226 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000522226 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000523200 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000523200 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000524280 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| Frame-shift | ENST00000524280 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000315050 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000315050 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000389378 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000389378 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000520931 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000520931 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000523338 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| In-frame | ENST00000523338 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| intron-3CDS | ENST00000521423 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| intron-3CDS | ENST00000521423 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
| intron-3UTR | ENST00000521423 | ENST00000521390 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000520931 | TNIP1 | chr5 | 150441688 | - | ENST00000435489 | NIPAL4 | chr5 | 156890102 | + | 1619 | 358 | 363 | 1478 | 371 |
| ENST00000520931 | TNIP1 | chr5 | 150441688 | - | ENST00000311946 | NIPAL4 | chr5 | 156890102 | + | 3293 | 358 | 363 | 1535 | 390 |
| ENST00000389378 | TNIP1 | chr5 | 150441688 | - | ENST00000435489 | NIPAL4 | chr5 | 156890102 | + | 2207 | 946 | 951 | 2066 | 371 |
| ENST00000389378 | TNIP1 | chr5 | 150441688 | - | ENST00000311946 | NIPAL4 | chr5 | 156890102 | + | 3881 | 946 | 951 | 2123 | 390 |
| ENST00000523338 | TNIP1 | chr5 | 150441688 | - | ENST00000435489 | NIPAL4 | chr5 | 156890102 | + | 1728 | 467 | 472 | 1587 | 371 |
| ENST00000523338 | TNIP1 | chr5 | 150441688 | - | ENST00000311946 | NIPAL4 | chr5 | 156890102 | + | 3402 | 467 | 472 | 1644 | 390 |
| ENST00000315050 | TNIP1 | chr5 | 150441688 | - | ENST00000435489 | NIPAL4 | chr5 | 156890102 | + | 1728 | 467 | 472 | 1587 | 371 |
| ENST00000315050 | TNIP1 | chr5 | 150441688 | - | ENST00000311946 | NIPAL4 | chr5 | 156890102 | + | 3402 | 467 | 472 | 1644 | 390 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000520931 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.014686422 | 0.9853136 |
| ENST00000520931 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.005537761 | 0.9944622 |
| ENST00000389378 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.019336088 | 0.9806639 |
| ENST00000389378 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.005877899 | 0.9941221 |
| ENST00000523338 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.016560746 | 0.98343927 |
| ENST00000523338 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.005518195 | 0.9944818 |
| ENST00000315050 | ENST00000435489 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.016560746 | 0.98343927 |
| ENST00000315050 | ENST00000311946 | TNIP1 | chr5 | 150441688 | - | NIPAL4 | chr5 | 156890102 | + | 0.005518195 | 0.9944818 |
Top |
Fusion Genomic Features for TNIP1-NIPAL4 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| TNIP1 | chr5 | 150441687 | - | NIPAL4 | chr5 | 156890101 | + | 9.12E-05 | 0.9999088 |
| TNIP1 | chr5 | 150441687 | - | NIPAL4 | chr5 | 156890101 | + | 9.12E-05 | 0.9999088 |
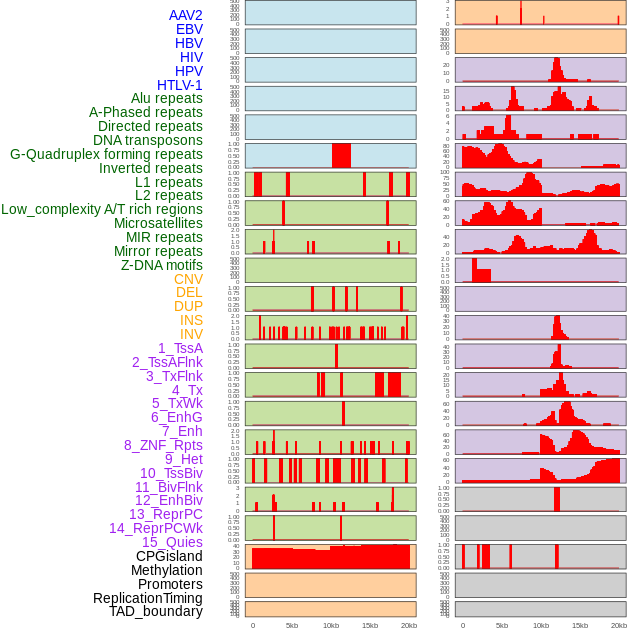
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
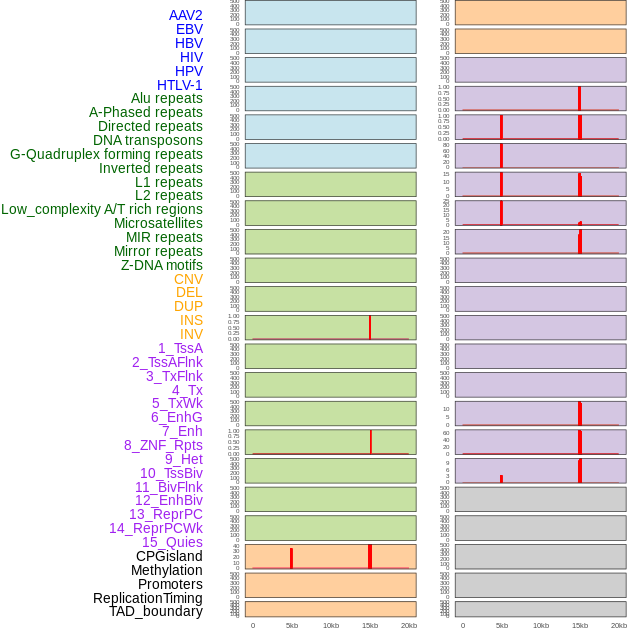
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for TNIP1-NIPAL4 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr5:150441688/chr5:156890102) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| TNIP1 | . |
| FUNCTION: Inhibits NF-kappa-B activation and TNF-induced NF-kappa-B-dependent gene expression by regulating A20/TNFAIP3-mediated deubiquitination of IKBKG; proposed to link A20/TNFAIP3 to ubiquitinated IKBKG. Involved in regulation of EGF-induced ERK1/ERK2 signaling pathway; blocks MAPK3/MAPK1 nuclear translocation and MAPK1-dependent transcription. Increases cell surface CD4(T4) antigen expression. Involved in the anti-inflammatory response of macrophages and positively regulates TLR-induced activation of CEBPB. Involved in the prevention of autoimmunity; this function implicates binding to polyubiquitin. Involved in leukocyte integrin activation during inflammation; this function is mediated by association with SELPLG and dependent on phosphorylation by SRC-family kinases. Interacts with HIV-1 matrix protein and is packaged into virions and overexpression can inhibit viral replication. May regulate matrix nuclear localization, both nuclear import of PIC (Preintegration complex) and export of GAG polyprotein and viral genomic RNA during virion production. In case of infection, promotes association of IKBKG with Shigella flexneri E3 ubiquitin-protein ligase ipah9.8 p which in turn promotes polyubiquitination of IKBKG leading to its proteasome-dependent degradation and thus is perturbing NF-kappa-B activation during bacterial infection. {ECO:0000269|PubMed:12220502, ECO:0000269|PubMed:16684768, ECO:0000269|PubMed:17016622, ECO:0000269|PubMed:17632516, ECO:0000269|PubMed:20010814}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 20_73 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 20_73 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 20_73 | 119 | 636.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 20_73 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 20_73 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 20_73 | 119 | 636.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 139_164 | 74 | 467.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 186_186 | 74 | 467.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 208_215 | 74 | 467.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 237_257 | 74 | 467.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 279_285 | 74 | 467.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 307_323 | 74 | 467.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 345_355 | 74 | 467.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 377_386 | 74 | 467.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 408_466 | 74 | 467.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 139_164 | 74 | 448.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 186_186 | 74 | 448.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 208_215 | 74 | 448.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 237_257 | 74 | 448.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 279_285 | 74 | 448.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 307_323 | 74 | 448.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 345_355 | 74 | 448.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 377_386 | 74 | 448.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 408_466 | 74 | 448.0 | Topological domain | Cytoplasmic | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 118_138 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 165_185 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 187_207 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 216_236 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 258_278 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 286_306 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 324_344 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 356_376 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 387_407 | 74 | 467.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 118_138 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 165_185 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 187_207 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 216_236 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 258_278 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 286_306 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 324_344 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 356_376 | 74 | 448.0 | Transmembrane | Helical | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 387_407 | 74 | 448.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 196_258 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 294_535 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 196_258 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 294_535 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 196_258 | 119 | 636.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 294_535 | 119 | 636.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 196_258 | 66 | 584.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 20_73 | 66 | 584.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 294_535 | 66 | 584.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 196_258 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 294_535 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 196_258 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 294_535 | 119 | 637.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 196_258 | 119 | 636.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 294_535 | 119 | 636.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 539_636 | 119 | 637.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 539_636 | 119 | 637.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 539_636 | 119 | 636.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 539_636 | 66 | 584.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 539_636 | 119 | 637.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 539_636 | 119 | 637.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 539_636 | 119 | 636.0 | Compositional bias | Note=Pro-rich |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 524_530 | 119 | 637.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 524_530 | 119 | 637.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 524_530 | 119 | 636.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 524_530 | 66 | 584.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 524_530 | 119 | 637.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 524_530 | 119 | 637.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 524_530 | 119 | 636.0 | Motif | Nuclear localization signal |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 431_588 | 119 | 637.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 452_510 | 119 | 637.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 431_588 | 119 | 637.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 452_510 | 119 | 637.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 431_588 | 119 | 636.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 452_510 | 119 | 636.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 431_588 | 66 | 584.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 452_510 | 66 | 584.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 431_588 | 119 | 637.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 452_510 | 119 | 637.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 431_588 | 119 | 637.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 452_510 | 119 | 637.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 431_588 | 119 | 636.0 | Region | Required for inhibitory activity of TNF-induced NF-kappa-B activation |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 452_510 | 119 | 636.0 | Region | Note=Ubiquitin-binding domain (UBD) |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000311946 | 0 | 6 | 1_117 | 74 | 467.0 | Topological domain | Extracellular | |
| Tgene | NIPAL4 | chr5:150441688 | chr5:156890102 | ENST00000435489 | 0 | 5 | 1_117 | 74 | 448.0 | Topological domain | Extracellular |
Top |
Fusion Gene Sequence for TNIP1-NIPAL4 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >92708_92708_1_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000315050_NIPAL4_chr5_156890102_ENST00000311946_length(transcript)=3402nt_BP=467nt AGTGAGCCGAGGAGCCTTCATAGGGACAGCCGCCCCTGGTGCACACACCCTCGTATTCTCCTGCCCTTCCCCAGCCAGGCGGGCACCACG GCAGGGGCTGAGCTACCCTCATGGAAGGGAGAGGACCGTACCGGATCTACGACCCTGGGGGCAGCGTGCCCTCAGGAGAGGCATCCGCAG CTTTTGAGCGCCTAGTGAAGGAGAATTCCCGGCTGAAGGAAAAAATGCAAGGGATAAAGATGTTAGGGGAGCTTTTGGAAGAGTCCCAGA TGGAAGCGACCAGGCTCCGGCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCCACCACCTTCTCCCTCCTTGGGCTCCTTCG ACCCCCTGGCTGAGCTCACAGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGCATGCCCCAGTGACAAGCCAGCACCAGTCC AGAAGCCTCCATCCAGTGTTCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGCCAGATTGTCAATGACCTCAGCCCTGAGGT GCCCAGCAATGCCACCTTTCACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTACATCGGCCTGGGCCTGGCATTCCTGTCTAG CTTCCTCATCGGCAGCAGCGTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACGGGAGCCACTCGAGCTGTGGATGGAGGCTT CGGCTACCTGAAAGATGCAATGTGGTGGGCTGGATTTCTCACCATGGCTGCTGGAGAAGTTGCCAACTTTGGAGCCTACGCATTTGCACC TGCAACAGTCGTCACGCCTCTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTCCTCATATTTCCTGAGGGAGAGTCTGAACCTGCT GGGGAAGCTGGGCTGTGTGATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGCTCCTGAGGAAGAGAAGGTCACTACCATCATGGA GATGGCTTCCAAGATGAAAGACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGTGTCATGCCTCATCCTCATCTTTGTCATTGCCCC ACGTTACGGGCAAAGGAATATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGCCTTCTCTGTGGCTGCTGTCAAGGGGCTGGGCAT CACCATCAAGAACTTCTTCCAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACATCCTGTCCCTCATCCTGGCACTGTCCCTCAGCAC TCAGGTCAACTTCCTCAACAGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCCCATCTACTACGTGTTCTTCACCACGGTGGTCGT TACCTCGTCCATCATCCTCTTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGCAGGCACCCTCTCGGGCTTTGTCACCATCATCTT GGGCGTGTTCATGCTGCATGCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCCCCACATGCACAAAAACCCACCCCCTTCTCCCGC CCCGGAACCCACTGTTATTAGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGAACTTGCCAGCACCTCATCACCAGAAGAGAAACC CAAAGTATTTATAATCCATTCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCCGATGGTACAGCCTGCCCTCCCAATTTCAAAACC ACCTGGTTATTTTCCAGTGCAACTGTTACCAATGGGCTCTCTTTTCTTGAGAAGTTCATTTATACCTCATCACTGTTTCCAGGAGAAAAA TCTTTACCCAAATAGCAATGGTGGCAGAACTTCCTGGAAACAGATTCAGTGACCAAATACCCAAGTTTACATCAGTGCCTGCAGGTTCCC TGGACCTTCCTTCTCATTCATTCTTTCGGTGCCATCTCTATGCCGTTGGGAAGAAGATGGAGTCTGACCCACTGAATGTAGCACAGTCCA AGGACTTCTCTAAGATATTGGTCATTGGAAGTTCCTTCACACCAATTCTCCTCCTGAGACGGAATCTCCGTTGTTGTTGTTGTTGTTGTT TTCTAGCCCAAGGATGACATAGAGCTGGCTCCCAGAGGCCCACAGAGCAATTGGCCATGCCTCCCTATCCAGAGCTGACAGGGACACAAC CAGTGTAAAATATCCTGTTGCCTTTGTCACTTCCTCTTTGGAGGCAGAAGCAAGACCTCAGCTGACCTTCTTACTGTGAAAGCCACTTGA TGTCTCAGGGAAAAATTTCAACCAGCTCATTCCCCGAGCACTCCAGCCTGGCAGTCAGCACCTCGGCATCCACCCAGTCCATCCCACCAT CACCCCTTCCCCCTCTACTTACATCCTAAGGAGTCGGTCACTGAGACATAAAGGCAGTAATCGCAGAACTGGAAACAAAACAATAATAGA GCCACAGCCAAACTCTGGTGGCCAAACCCAGTGTTGCATTTTGTCTTACTCTGAAAGAAGAACAGCAAATTCACTGCTTCAAAGTGGCCT GGCTGCCAAGCTAGAATTTGGCAGAACGCACTTTACTATTCCTCAAGGAGTCAACCAACCTATGATCTGGGGAGGTGGGAAGAGGATGAG GAGCAAAGTTGGGATTTGGCAGAAGGCAGTCCCAGGCTCTCTGGATACTAGGGGCTAACTTTTGTGTTGACTCTGGTGCTCATCTGGGAA CTTAGGAGAAACGAGCTCAGGGGTAATTTCTGGGTTGCAGCCTTAAAGGCTTGGACAGCTGTGAATCTCAATGGCCAACTGGAGGTGCAG ACTTGGCATGGGGTGCATTCTAGCTGTTGACCAGATTGCTACCGAGTCCCCTCCTCCACTGATGAGCTGCCCACACTGGGAAGCAGCATG CCCTGACTGTTCCAACACCACCTGCTATGGGGAGTACCTTTGGTCCCCTCACATTTGGCCAGAGGCATACAAAAAACCAGAGCAGCTGGA GAGGGAGATAATTACTATTCCTTCCCTTCCTATCTCCTTTCTAGCCACAAGAGTGTGGGGGTTGGAGAAGAACCATTAGAAAGGGAAATT AGTGGGCTGGTGTATCTGGAAAGAGGGAAGACTTGATCCTCAGCCCCGAGGTTGGTGCAGGGCCTCCCCTGTGTGACTCTACCTGCACTC TGTGTTTATATCCTGTGCCCTAAGTGGGCCAAGCCCAGGTAAATTCCTGCTGGCCTTGGAACTCCAAGGTTTGGCTGACCAGCAGACTGG CTCCCTGACTCTTCAGCCTCAAATCCCCAGTTTTTGATGAATGTGGATTTCTGTCTGTAATTAAAAGCAATGCAACAAGTTGGCTCTTGA >92708_92708_1_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000315050_NIPAL4_chr5_156890102_ENST00000311946_length(amino acids)=390AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVDGGFGYLKDAM WWAGFLTMAAGEVANFGAYAFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKD TGFIVFAVLLLVSCLILIFVIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNR ALDIFNTSLVFPIYYVFFTTVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIR -------------------------------------------------------------- >92708_92708_2_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000315050_NIPAL4_chr5_156890102_ENST00000435489_length(transcript)=1728nt_BP=467nt AGTGAGCCGAGGAGCCTTCATAGGGACAGCCGCCCCTGGTGCACACACCCTCGTATTCTCCTGCCCTTCCCCAGCCAGGCGGGCACCACG GCAGGGGCTGAGCTACCCTCATGGAAGGGAGAGGACCGTACCGGATCTACGACCCTGGGGGCAGCGTGCCCTCAGGAGAGGCATCCGCAG CTTTTGAGCGCCTAGTGAAGGAGAATTCCCGGCTGAAGGAAAAAATGCAAGGGATAAAGATGTTAGGGGAGCTTTTGGAAGAGTCCCAGA TGGAAGCGACCAGGCTCCGGCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCCACCACCTTCTCCCTCCTTGGGCTCCTTCG ACCCCCTGGCTGAGCTCACAGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGCATGCCCCAGTGACAAGCCAGCACCAGTCC AGAAGCCTCCATCCAGTGTTCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGCCAGATTGTCAATGACCTCAGCCCTGAGGT GCCCAGCAATGCCACCTTTCACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTACATCGGCCTGGGCCTGGCATTCCTGTCTAG CTTCCTCATCGGCAGCAGCGTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACGGGAGCCACTCGAGCTGTGGCTGCTGGAGA AGTTGCCAACTTTGGAGCCTACGCATTTGCACCTGCAACAGTCGTCACGCCTCTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTC CTCATATTTCCTGAGGGAGAGTCTGAACCTGCTGGGGAAGCTGGGCTGTGTGATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGC TCCTGAGGAAGAGAAGGTCACTACCATCATGGAGATGGCTTCCAAGATGAAAGACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGT GTCATGCCTCATCCTCATCTTTGTCATTGCCCCACGTTACGGGCAAAGGAATATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGC CTTCTCTGTGGCTGCTGTCAAGGGGCTGGGCATCACCATCAAGAACTTCTTCCAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACAT CCTGTCCCTCATCCTGGCACTGTCCCTCAGCACTCAGGTCAACTTCCTCAACAGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCC CATCTACTACGTGTTCTTCACCACGGTGGTCGTTACCTCGTCCATCATCCTCTTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGC AGGCACCCTCTCGGGCTTTGTCACCATCATCTTGGGCGTGTTCATGCTGCATGCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCC CCACATGCACAAAAACCCACCCCCTTCTCCCGCCCCGGAACCCACTGTTATTAGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGA ACTTGCCAGCACCTCATCACCAGAAGAGAAACCCAAAGTATTTATAATCCATTCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCC GATGGTACAGCCTGCCCTCCCAATTTCAAAACCACCTGGTTATTTTCCAGTGCAACTGTTACCAATGGGCTCTCTTTTCTTGAGAAGTTC >92708_92708_2_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000315050_NIPAL4_chr5_156890102_ENST00000435489_length(amino acids)=371AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVAAGEVANFGAY AFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKDTGFIVFAVLLLVSCLILIF VIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNRALDIFNTSLVFPIYYVFFT TVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIRLEDKNVLVDNIELASTSSP -------------------------------------------------------------- >92708_92708_3_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000389378_NIPAL4_chr5_156890102_ENST00000311946_length(transcript)=3881nt_BP=946nt ATGCTTACGTGCCTTTTGGCAGGTCCCCTCCCTGTTCTGGCCTGTGTTCCCCTCGGCACAAGGAAGGGGCTGGGTGGTCTCTGTGGCTCG CCTCCGCTCTGAAATTCCGGCTTTCAAGGTGGCCCCAGGCCTTCGACAGGGGCTTCCGGCGCTGGGAGTGCGGCAAGGCGCCGCGCAGGC CCGGGACGGTGGCCCCAGGGAAGTCGCTCCTAGAGGGGAAGGCCGCCGAAGGGACGAGAGACACCGCGGGCCCCCGGAACCGCAGGCAGC CGGTGTTTACCTCGCTCGGTCCCACCCCAGGCGGCCGCGGGCCCGCCCGGAGCGCGCGGCGCGATTGGGCGAGTGGCCTGCCAGTCTCCG GGGACTTTCCCAGGGGTGGGGCGGCCCGGCCAGGCCCCCGGCACTTCCTCGTCCTCGGCCCGGGTGCCCTGCCCCCGTCCAGGAGCCCTA GGAGTGCTACGGGGGGCCGGAGCCTTGCCCGGGCCGCTGCCCCGTCCCTGGATTCGGGGCTGGACGCAGCAAGCGGGGCGCTGTGTCCCC AAGCTCCCCGTCCTCGGGCGGGCACCACGGCAGGGGCTGAGCTACCCTCATGGAAGGGAGAGGACCGTACCGGATCTACGACCCTGGGGG CAGCGTGCCCTCAGGAGAGGCATCCGCAGCTTTTGAGCGCCTAGTGAAGGAGAATTCCCGGCTGAAGGAAAAAATGCAAGGGATAAAGAT GTTAGGGGAGCTTTTGGAAGAGTCCCAGATGGAAGCGACCAGGCTCCGGCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCC ACCACCTTCTCCCTCCTTGGGCTCCTTCGACCCCCTGGCTGAGCTCACAGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGC ATGCCCCAGTGACAAGCCAGCACCAGTCCAGAAGCCTCCATCCAGTGTTCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGC CAGATTGTCAATGACCTCAGCCCTGAGGTGCCCAGCAATGCCACCTTTCACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTAC ATCGGCCTGGGCCTGGCATTCCTGTCTAGCTTCCTCATCGGCAGCAGCGTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACG GGAGCCACTCGAGCTGTGGATGGAGGCTTCGGCTACCTGAAAGATGCAATGTGGTGGGCTGGATTTCTCACCATGGCTGCTGGAGAAGTT GCCAACTTTGGAGCCTACGCATTTGCACCTGCAACAGTCGTCACGCCTCTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTCCTCA TATTTCCTGAGGGAGAGTCTGAACCTGCTGGGGAAGCTGGGCTGTGTGATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGCTCCT GAGGAAGAGAAGGTCACTACCATCATGGAGATGGCTTCCAAGATGAAAGACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGTGTCA TGCCTCATCCTCATCTTTGTCATTGCCCCACGTTACGGGCAAAGGAATATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGCCTTC TCTGTGGCTGCTGTCAAGGGGCTGGGCATCACCATCAAGAACTTCTTCCAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACATCCTG TCCCTCATCCTGGCACTGTCCCTCAGCACTCAGGTCAACTTCCTCAACAGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCCCATC TACTACGTGTTCTTCACCACGGTGGTCGTTACCTCGTCCATCATCCTCTTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGCAGGC ACCCTCTCGGGCTTTGTCACCATCATCTTGGGCGTGTTCATGCTGCATGCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCCCCAC ATGCACAAAAACCCACCCCCTTCTCCCGCCCCGGAACCCACTGTTATTAGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGAACTT GCCAGCACCTCATCACCAGAAGAGAAACCCAAAGTATTTATAATCCATTCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCCGATG GTACAGCCTGCCCTCCCAATTTCAAAACCACCTGGTTATTTTCCAGTGCAACTGTTACCAATGGGCTCTCTTTTCTTGAGAAGTTCATTT ATACCTCATCACTGTTTCCAGGAGAAAAATCTTTACCCAAATAGCAATGGTGGCAGAACTTCCTGGAAACAGATTCAGTGACCAAATACC CAAGTTTACATCAGTGCCTGCAGGTTCCCTGGACCTTCCTTCTCATTCATTCTTTCGGTGCCATCTCTATGCCGTTGGGAAGAAGATGGA GTCTGACCCACTGAATGTAGCACAGTCCAAGGACTTCTCTAAGATATTGGTCATTGGAAGTTCCTTCACACCAATTCTCCTCCTGAGACG GAATCTCCGTTGTTGTTGTTGTTGTTGTTTTCTAGCCCAAGGATGACATAGAGCTGGCTCCCAGAGGCCCACAGAGCAATTGGCCATGCC TCCCTATCCAGAGCTGACAGGGACACAACCAGTGTAAAATATCCTGTTGCCTTTGTCACTTCCTCTTTGGAGGCAGAAGCAAGACCTCAG CTGACCTTCTTACTGTGAAAGCCACTTGATGTCTCAGGGAAAAATTTCAACCAGCTCATTCCCCGAGCACTCCAGCCTGGCAGTCAGCAC CTCGGCATCCACCCAGTCCATCCCACCATCACCCCTTCCCCCTCTACTTACATCCTAAGGAGTCGGTCACTGAGACATAAAGGCAGTAAT CGCAGAACTGGAAACAAAACAATAATAGAGCCACAGCCAAACTCTGGTGGCCAAACCCAGTGTTGCATTTTGTCTTACTCTGAAAGAAGA ACAGCAAATTCACTGCTTCAAAGTGGCCTGGCTGCCAAGCTAGAATTTGGCAGAACGCACTTTACTATTCCTCAAGGAGTCAACCAACCT ATGATCTGGGGAGGTGGGAAGAGGATGAGGAGCAAAGTTGGGATTTGGCAGAAGGCAGTCCCAGGCTCTCTGGATACTAGGGGCTAACTT TTGTGTTGACTCTGGTGCTCATCTGGGAACTTAGGAGAAACGAGCTCAGGGGTAATTTCTGGGTTGCAGCCTTAAAGGCTTGGACAGCTG TGAATCTCAATGGCCAACTGGAGGTGCAGACTTGGCATGGGGTGCATTCTAGCTGTTGACCAGATTGCTACCGAGTCCCCTCCTCCACTG ATGAGCTGCCCACACTGGGAAGCAGCATGCCCTGACTGTTCCAACACCACCTGCTATGGGGAGTACCTTTGGTCCCCTCACATTTGGCCA GAGGCATACAAAAAACCAGAGCAGCTGGAGAGGGAGATAATTACTATTCCTTCCCTTCCTATCTCCTTTCTAGCCACAAGAGTGTGGGGG TTGGAGAAGAACCATTAGAAAGGGAAATTAGTGGGCTGGTGTATCTGGAAAGAGGGAAGACTTGATCCTCAGCCCCGAGGTTGGTGCAGG GCCTCCCCTGTGTGACTCTACCTGCACTCTGTGTTTATATCCTGTGCCCTAAGTGGGCCAAGCCCAGGTAAATTCCTGCTGGCCTTGGAA CTCCAAGGTTTGGCTGACCAGCAGACTGGCTCCCTGACTCTTCAGCCTCAAATCCCCAGTTTTTGATGAATGTGGATTTCTGTCTGTAAT TAAAAGCAATGCAACAAGTTGGCTCTTGAGAATGGCAGTAAACTGAGGGCCCTAAGAGTGTGGTCTGCAGGGTCAAGAATAAAGATTACA >92708_92708_3_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000389378_NIPAL4_chr5_156890102_ENST00000311946_length(amino acids)=390AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVDGGFGYLKDAM WWAGFLTMAAGEVANFGAYAFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKD TGFIVFAVLLLVSCLILIFVIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNR ALDIFNTSLVFPIYYVFFTTVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIR -------------------------------------------------------------- >92708_92708_4_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000389378_NIPAL4_chr5_156890102_ENST00000435489_length(transcript)=2207nt_BP=946nt ATGCTTACGTGCCTTTTGGCAGGTCCCCTCCCTGTTCTGGCCTGTGTTCCCCTCGGCACAAGGAAGGGGCTGGGTGGTCTCTGTGGCTCG CCTCCGCTCTGAAATTCCGGCTTTCAAGGTGGCCCCAGGCCTTCGACAGGGGCTTCCGGCGCTGGGAGTGCGGCAAGGCGCCGCGCAGGC CCGGGACGGTGGCCCCAGGGAAGTCGCTCCTAGAGGGGAAGGCCGCCGAAGGGACGAGAGACACCGCGGGCCCCCGGAACCGCAGGCAGC CGGTGTTTACCTCGCTCGGTCCCACCCCAGGCGGCCGCGGGCCCGCCCGGAGCGCGCGGCGCGATTGGGCGAGTGGCCTGCCAGTCTCCG GGGACTTTCCCAGGGGTGGGGCGGCCCGGCCAGGCCCCCGGCACTTCCTCGTCCTCGGCCCGGGTGCCCTGCCCCCGTCCAGGAGCCCTA GGAGTGCTACGGGGGGCCGGAGCCTTGCCCGGGCCGCTGCCCCGTCCCTGGATTCGGGGCTGGACGCAGCAAGCGGGGCGCTGTGTCCCC AAGCTCCCCGTCCTCGGGCGGGCACCACGGCAGGGGCTGAGCTACCCTCATGGAAGGGAGAGGACCGTACCGGATCTACGACCCTGGGGG CAGCGTGCCCTCAGGAGAGGCATCCGCAGCTTTTGAGCGCCTAGTGAAGGAGAATTCCCGGCTGAAGGAAAAAATGCAAGGGATAAAGAT GTTAGGGGAGCTTTTGGAAGAGTCCCAGATGGAAGCGACCAGGCTCCGGCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCC ACCACCTTCTCCCTCCTTGGGCTCCTTCGACCCCCTGGCTGAGCTCACAGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGC ATGCCCCAGTGACAAGCCAGCACCAGTCCAGAAGCCTCCATCCAGTGTTCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGC CAGATTGTCAATGACCTCAGCCCTGAGGTGCCCAGCAATGCCACCTTTCACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTAC ATCGGCCTGGGCCTGGCATTCCTGTCTAGCTTCCTCATCGGCAGCAGCGTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACG GGAGCCACTCGAGCTGTGGCTGCTGGAGAAGTTGCCAACTTTGGAGCCTACGCATTTGCACCTGCAACAGTCGTCACGCCTCTGGGAGCG CTGAGTGTCCTCATAAGTGCCATCCTCTCCTCATATTTCCTGAGGGAGAGTCTGAACCTGCTGGGGAAGCTGGGCTGTGTGATCTGTGTG GCCGGAAGCACAGTGATGGTGATACATGCTCCTGAGGAAGAGAAGGTCACTACCATCATGGAGATGGCTTCCAAGATGAAAGACACAGGG TTCATCGTGTTTGCTGTGCTTCTGCTGGTGTCATGCCTCATCCTCATCTTTGTCATTGCCCCACGTTACGGGCAAAGGAATATCCTCATC TACATCATCATCTGCTCTGTGATCGGGGCCTTCTCTGTGGCTGCTGTCAAGGGGCTGGGCATCACCATCAAGAACTTCTTCCAGGGGCTG CCAGTTGTCCGGCACCCGCTCCCCTACATCCTGTCCCTCATCCTGGCACTGTCCCTCAGCACTCAGGTCAACTTCCTCAACAGAGCACTG GACATTTTCAACACTTCCCTGGTGTTCCCCATCTACTACGTGTTCTTCACCACGGTGGTCGTTACCTCGTCCATCATCCTCTTCAAGGAG TGGTACAGCATGTCTGCTGTGGACATTGCAGGCACCCTCTCGGGCTTTGTCACCATCATCTTGGGCGTGTTCATGCTGCATGCTTTCAAA GACCTGGACATCAGCTGCGCCAGCTTGCCCCACATGCACAAAAACCCACCCCCTTCTCCCGCCCCGGAACCCACTGTTATTAGACTGGAA GACAAGAACGTCCTTGTGGACAATATAGAACTTGCCAGCACCTCATCACCAGAAGAGAAACCCAAAGTATTTATAATCCATTCCTGAAGC TTGGAATATGTGAGTGAGAGGATGAGTCCGATGGTACAGCCTGCCCTCCCAATTTCAAAACCACCTGGTTATTTTCCAGTGCAACTGTTA >92708_92708_4_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000389378_NIPAL4_chr5_156890102_ENST00000435489_length(amino acids)=371AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVAAGEVANFGAY AFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKDTGFIVFAVLLLVSCLILIF VIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNRALDIFNTSLVFPIYYVFFT TVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIRLEDKNVLVDNIELASTSSP -------------------------------------------------------------- >92708_92708_5_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000520931_NIPAL4_chr5_156890102_ENST00000311946_length(transcript)=3293nt_BP=358nt CGGGTGCCCTGCCCCCGTCCAGGAGCCCTAGGAGTGCTACGGGGGGCCGGAGCCTTGCCCGGGCCGCTGCCCCGTCCCTGGATTCGGGGC TGGACGCAGCAAGCGGGGCGCTGTGTCCCCAAGCTCCCCGTCCTCGGGGGAGCTTTTGGAAGAGTCCCAGATGGAAGCGACCAGGCTCCG GCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCCACCACCTTCTCCCTCCTTGGGCTCCTTCGACCCCCTGGCTGAGCTCAC AGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGCATGCCCCAGTGACAAGCCAGCACCAGTCCAGAAGCCTCCATCCAGTGT TCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGCCAGATTGTCAATGACCTCAGCCCTGAGGTGCCCAGCAATGCCACCTTT CACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTACATCGGCCTGGGCCTGGCATTCCTGTCTAGCTTCCTCATCGGCAGCAGC GTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACGGGAGCCACTCGAGCTGTGGATGGAGGCTTCGGCTACCTGAAAGATGCA ATGTGGTGGGCTGGATTTCTCACCATGGCTGCTGGAGAAGTTGCCAACTTTGGAGCCTACGCATTTGCACCTGCAACAGTCGTCACGCCT CTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTCCTCATATTTCCTGAGGGAGAGTCTGAACCTGCTGGGGAAGCTGGGCTGTGTG ATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGCTCCTGAGGAAGAGAAGGTCACTACCATCATGGAGATGGCTTCCAAGATGAAA GACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGTGTCATGCCTCATCCTCATCTTTGTCATTGCCCCACGTTACGGGCAAAGGAAT ATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGCCTTCTCTGTGGCTGCTGTCAAGGGGCTGGGCATCACCATCAAGAACTTCTTC CAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACATCCTGTCCCTCATCCTGGCACTGTCCCTCAGCACTCAGGTCAACTTCCTCAAC AGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCCCATCTACTACGTGTTCTTCACCACGGTGGTCGTTACCTCGTCCATCATCCTC TTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGCAGGCACCCTCTCGGGCTTTGTCACCATCATCTTGGGCGTGTTCATGCTGCAT GCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCCCCACATGCACAAAAACCCACCCCCTTCTCCCGCCCCGGAACCCACTGTTATT AGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGAACTTGCCAGCACCTCATCACCAGAAGAGAAACCCAAAGTATTTATAATCCAT TCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCCGATGGTACAGCCTGCCCTCCCAATTTCAAAACCACCTGGTTATTTTCCAGTG CAACTGTTACCAATGGGCTCTCTTTTCTTGAGAAGTTCATTTATACCTCATCACTGTTTCCAGGAGAAAAATCTTTACCCAAATAGCAAT GGTGGCAGAACTTCCTGGAAACAGATTCAGTGACCAAATACCCAAGTTTACATCAGTGCCTGCAGGTTCCCTGGACCTTCCTTCTCATTC ATTCTTTCGGTGCCATCTCTATGCCGTTGGGAAGAAGATGGAGTCTGACCCACTGAATGTAGCACAGTCCAAGGACTTCTCTAAGATATT GGTCATTGGAAGTTCCTTCACACCAATTCTCCTCCTGAGACGGAATCTCCGTTGTTGTTGTTGTTGTTGTTTTCTAGCCCAAGGATGACA TAGAGCTGGCTCCCAGAGGCCCACAGAGCAATTGGCCATGCCTCCCTATCCAGAGCTGACAGGGACACAACCAGTGTAAAATATCCTGTT GCCTTTGTCACTTCCTCTTTGGAGGCAGAAGCAAGACCTCAGCTGACCTTCTTACTGTGAAAGCCACTTGATGTCTCAGGGAAAAATTTC AACCAGCTCATTCCCCGAGCACTCCAGCCTGGCAGTCAGCACCTCGGCATCCACCCAGTCCATCCCACCATCACCCCTTCCCCCTCTACT TACATCCTAAGGAGTCGGTCACTGAGACATAAAGGCAGTAATCGCAGAACTGGAAACAAAACAATAATAGAGCCACAGCCAAACTCTGGT GGCCAAACCCAGTGTTGCATTTTGTCTTACTCTGAAAGAAGAACAGCAAATTCACTGCTTCAAAGTGGCCTGGCTGCCAAGCTAGAATTT GGCAGAACGCACTTTACTATTCCTCAAGGAGTCAACCAACCTATGATCTGGGGAGGTGGGAAGAGGATGAGGAGCAAAGTTGGGATTTGG CAGAAGGCAGTCCCAGGCTCTCTGGATACTAGGGGCTAACTTTTGTGTTGACTCTGGTGCTCATCTGGGAACTTAGGAGAAACGAGCTCA GGGGTAATTTCTGGGTTGCAGCCTTAAAGGCTTGGACAGCTGTGAATCTCAATGGCCAACTGGAGGTGCAGACTTGGCATGGGGTGCATT CTAGCTGTTGACCAGATTGCTACCGAGTCCCCTCCTCCACTGATGAGCTGCCCACACTGGGAAGCAGCATGCCCTGACTGTTCCAACACC ACCTGCTATGGGGAGTACCTTTGGTCCCCTCACATTTGGCCAGAGGCATACAAAAAACCAGAGCAGCTGGAGAGGGAGATAATTACTATT CCTTCCCTTCCTATCTCCTTTCTAGCCACAAGAGTGTGGGGGTTGGAGAAGAACCATTAGAAAGGGAAATTAGTGGGCTGGTGTATCTGG AAAGAGGGAAGACTTGATCCTCAGCCCCGAGGTTGGTGCAGGGCCTCCCCTGTGTGACTCTACCTGCACTCTGTGTTTATATCCTGTGCC CTAAGTGGGCCAAGCCCAGGTAAATTCCTGCTGGCCTTGGAACTCCAAGGTTTGGCTGACCAGCAGACTGGCTCCCTGACTCTTCAGCCT CAAATCCCCAGTTTTTGATGAATGTGGATTTCTGTCTGTAATTAAAAGCAATGCAACAAGTTGGCTCTTGAGAATGGCAGTAAACTGAGG >92708_92708_5_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000520931_NIPAL4_chr5_156890102_ENST00000311946_length(amino acids)=390AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVDGGFGYLKDAM WWAGFLTMAAGEVANFGAYAFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKD TGFIVFAVLLLVSCLILIFVIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNR ALDIFNTSLVFPIYYVFFTTVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIR -------------------------------------------------------------- >92708_92708_6_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000520931_NIPAL4_chr5_156890102_ENST00000435489_length(transcript)=1619nt_BP=358nt CGGGTGCCCTGCCCCCGTCCAGGAGCCCTAGGAGTGCTACGGGGGGCCGGAGCCTTGCCCGGGCCGCTGCCCCGTCCCTGGATTCGGGGC TGGACGCAGCAAGCGGGGCGCTGTGTCCCCAAGCTCCCCGTCCTCGGGGGAGCTTTTGGAAGAGTCCCAGATGGAAGCGACCAGGCTCCG GCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCCACCACCTTCTCCCTCCTTGGGCTCCTTCGACCCCCTGGCTGAGCTCAC AGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGCATGCCCCAGTGACAAGCCAGCACCAGTCCAGAAGCCTCCATCCAGTGT TCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGCCAGATTGTCAATGACCTCAGCCCTGAGGTGCCCAGCAATGCCACCTTT CACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTACATCGGCCTGGGCCTGGCATTCCTGTCTAGCTTCCTCATCGGCAGCAGC GTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACGGGAGCCACTCGAGCTGTGGCTGCTGGAGAAGTTGCCAACTTTGGAGCC TACGCATTTGCACCTGCAACAGTCGTCACGCCTCTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTCCTCATATTTCCTGAGGGAG AGTCTGAACCTGCTGGGGAAGCTGGGCTGTGTGATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGCTCCTGAGGAAGAGAAGGTC ACTACCATCATGGAGATGGCTTCCAAGATGAAAGACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGTGTCATGCCTCATCCTCATC TTTGTCATTGCCCCACGTTACGGGCAAAGGAATATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGCCTTCTCTGTGGCTGCTGTC AAGGGGCTGGGCATCACCATCAAGAACTTCTTCCAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACATCCTGTCCCTCATCCTGGCA CTGTCCCTCAGCACTCAGGTCAACTTCCTCAACAGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCCCATCTACTACGTGTTCTTC ACCACGGTGGTCGTTACCTCGTCCATCATCCTCTTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGCAGGCACCCTCTCGGGCTTT GTCACCATCATCTTGGGCGTGTTCATGCTGCATGCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCCCCACATGCACAAAAACCCA CCCCCTTCTCCCGCCCCGGAACCCACTGTTATTAGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGAACTTGCCAGCACCTCATCA CCAGAAGAGAAACCCAAAGTATTTATAATCCATTCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCCGATGGTACAGCCTGCCCTC >92708_92708_6_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000520931_NIPAL4_chr5_156890102_ENST00000435489_length(amino acids)=371AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVAAGEVANFGAY AFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKDTGFIVFAVLLLVSCLILIF VIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNRALDIFNTSLVFPIYYVFFT TVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIRLEDKNVLVDNIELASTSSP -------------------------------------------------------------- >92708_92708_7_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000523338_NIPAL4_chr5_156890102_ENST00000311946_length(transcript)=3402nt_BP=467nt AGTGAGCCGAGGAGCCTTCATAGGGACAGCCGCCCCTGGTGCACACACCCTCGTATTCTCCTGCCCTTCCCCAGCCAGGCGGGCACCACG GCAGGGGCTGAGCTACCCTCATGGAAGGGAGAGGACCGTACCGGATCTACGACCCTGGGGGCAGCGTGCCCTCAGGAGAGGCATCCGCAG CTTTTGAGCGCCTAGTGAAGGAGAATTCCCGGCTGAAGGAAAAAATGCAAGGGATAAAGATGTTAGGGGAGCTTTTGGAAGAGTCCCAGA TGGAAGCGACCAGGCTCCGGCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCCACCACCTTCTCCCTCCTTGGGCTCCTTCG ACCCCCTGGCTGAGCTCACAGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGCATGCCCCAGTGACAAGCCAGCACCAGTCC AGAAGCCTCCATCCAGTGTTCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGCCAGATTGTCAATGACCTCAGCCCTGAGGT GCCCAGCAATGCCACCTTTCACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTACATCGGCCTGGGCCTGGCATTCCTGTCTAG CTTCCTCATCGGCAGCAGCGTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACGGGAGCCACTCGAGCTGTGGATGGAGGCTT CGGCTACCTGAAAGATGCAATGTGGTGGGCTGGATTTCTCACCATGGCTGCTGGAGAAGTTGCCAACTTTGGAGCCTACGCATTTGCACC TGCAACAGTCGTCACGCCTCTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTCCTCATATTTCCTGAGGGAGAGTCTGAACCTGCT GGGGAAGCTGGGCTGTGTGATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGCTCCTGAGGAAGAGAAGGTCACTACCATCATGGA GATGGCTTCCAAGATGAAAGACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGTGTCATGCCTCATCCTCATCTTTGTCATTGCCCC ACGTTACGGGCAAAGGAATATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGCCTTCTCTGTGGCTGCTGTCAAGGGGCTGGGCAT CACCATCAAGAACTTCTTCCAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACATCCTGTCCCTCATCCTGGCACTGTCCCTCAGCAC TCAGGTCAACTTCCTCAACAGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCCCATCTACTACGTGTTCTTCACCACGGTGGTCGT TACCTCGTCCATCATCCTCTTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGCAGGCACCCTCTCGGGCTTTGTCACCATCATCTT GGGCGTGTTCATGCTGCATGCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCCCCACATGCACAAAAACCCACCCCCTTCTCCCGC CCCGGAACCCACTGTTATTAGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGAACTTGCCAGCACCTCATCACCAGAAGAGAAACC CAAAGTATTTATAATCCATTCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCCGATGGTACAGCCTGCCCTCCCAATTTCAAAACC ACCTGGTTATTTTCCAGTGCAACTGTTACCAATGGGCTCTCTTTTCTTGAGAAGTTCATTTATACCTCATCACTGTTTCCAGGAGAAAAA TCTTTACCCAAATAGCAATGGTGGCAGAACTTCCTGGAAACAGATTCAGTGACCAAATACCCAAGTTTACATCAGTGCCTGCAGGTTCCC TGGACCTTCCTTCTCATTCATTCTTTCGGTGCCATCTCTATGCCGTTGGGAAGAAGATGGAGTCTGACCCACTGAATGTAGCACAGTCCA AGGACTTCTCTAAGATATTGGTCATTGGAAGTTCCTTCACACCAATTCTCCTCCTGAGACGGAATCTCCGTTGTTGTTGTTGTTGTTGTT TTCTAGCCCAAGGATGACATAGAGCTGGCTCCCAGAGGCCCACAGAGCAATTGGCCATGCCTCCCTATCCAGAGCTGACAGGGACACAAC CAGTGTAAAATATCCTGTTGCCTTTGTCACTTCCTCTTTGGAGGCAGAAGCAAGACCTCAGCTGACCTTCTTACTGTGAAAGCCACTTGA TGTCTCAGGGAAAAATTTCAACCAGCTCATTCCCCGAGCACTCCAGCCTGGCAGTCAGCACCTCGGCATCCACCCAGTCCATCCCACCAT CACCCCTTCCCCCTCTACTTACATCCTAAGGAGTCGGTCACTGAGACATAAAGGCAGTAATCGCAGAACTGGAAACAAAACAATAATAGA GCCACAGCCAAACTCTGGTGGCCAAACCCAGTGTTGCATTTTGTCTTACTCTGAAAGAAGAACAGCAAATTCACTGCTTCAAAGTGGCCT GGCTGCCAAGCTAGAATTTGGCAGAACGCACTTTACTATTCCTCAAGGAGTCAACCAACCTATGATCTGGGGAGGTGGGAAGAGGATGAG GAGCAAAGTTGGGATTTGGCAGAAGGCAGTCCCAGGCTCTCTGGATACTAGGGGCTAACTTTTGTGTTGACTCTGGTGCTCATCTGGGAA CTTAGGAGAAACGAGCTCAGGGGTAATTTCTGGGTTGCAGCCTTAAAGGCTTGGACAGCTGTGAATCTCAATGGCCAACTGGAGGTGCAG ACTTGGCATGGGGTGCATTCTAGCTGTTGACCAGATTGCTACCGAGTCCCCTCCTCCACTGATGAGCTGCCCACACTGGGAAGCAGCATG CCCTGACTGTTCCAACACCACCTGCTATGGGGAGTACCTTTGGTCCCCTCACATTTGGCCAGAGGCATACAAAAAACCAGAGCAGCTGGA GAGGGAGATAATTACTATTCCTTCCCTTCCTATCTCCTTTCTAGCCACAAGAGTGTGGGGGTTGGAGAAGAACCATTAGAAAGGGAAATT AGTGGGCTGGTGTATCTGGAAAGAGGGAAGACTTGATCCTCAGCCCCGAGGTTGGTGCAGGGCCTCCCCTGTGTGACTCTACCTGCACTC TGTGTTTATATCCTGTGCCCTAAGTGGGCCAAGCCCAGGTAAATTCCTGCTGGCCTTGGAACTCCAAGGTTTGGCTGACCAGCAGACTGG CTCCCTGACTCTTCAGCCTCAAATCCCCAGTTTTTGATGAATGTGGATTTCTGTCTGTAATTAAAAGCAATGCAACAAGTTGGCTCTTGA >92708_92708_7_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000523338_NIPAL4_chr5_156890102_ENST00000311946_length(amino acids)=390AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVDGGFGYLKDAM WWAGFLTMAAGEVANFGAYAFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKD TGFIVFAVLLLVSCLILIFVIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNR ALDIFNTSLVFPIYYVFFTTVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIR -------------------------------------------------------------- >92708_92708_8_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000523338_NIPAL4_chr5_156890102_ENST00000435489_length(transcript)=1728nt_BP=467nt AGTGAGCCGAGGAGCCTTCATAGGGACAGCCGCCCCTGGTGCACACACCCTCGTATTCTCCTGCCCTTCCCCAGCCAGGCGGGCACCACG GCAGGGGCTGAGCTACCCTCATGGAAGGGAGAGGACCGTACCGGATCTACGACCCTGGGGGCAGCGTGCCCTCAGGAGAGGCATCCGCAG CTTTTGAGCGCCTAGTGAAGGAGAATTCCCGGCTGAAGGAAAAAATGCAAGGGATAAAGATGTTAGGGGAGCTTTTGGAAGAGTCCCAGA TGGAAGCGACCAGGCTCCGGCAGAAGGCAGAGGAGCTAGTGAAGGACAACGAGCTGCTCCCACCACCTTCTCCCTCCTTGGGCTCCTTCG ACCCCCTGGCTGAGCTCACAGGAAAGGACTCAAATGTCACAGCATCTCCCACAGCCCCTGCATGCCCCAGTGACAAGCCAGCACCAGTCC AGAAGCCTCCATCCAGTGTTCCCTGCTCCACCTCTACTGCTCCTCCCAAGAAGTCCTGTGCCAGATTGTCAATGACCTCAGCCCTGAGGT GCCCAGCAATGCCACCTTTCACAGCTGGCAGGAAAGAATCAGGCAGAACTATGGCTTCTACATCGGCCTGGGCCTGGCATTCCTGTCTAG CTTCCTCATCGGCAGCAGCGTCATCCTCAAGAAGAAAGGCCTCTTGCGACTCGTGGCCACGGGAGCCACTCGAGCTGTGGCTGCTGGAGA AGTTGCCAACTTTGGAGCCTACGCATTTGCACCTGCAACAGTCGTCACGCCTCTGGGAGCGCTGAGTGTCCTCATAAGTGCCATCCTCTC CTCATATTTCCTGAGGGAGAGTCTGAACCTGCTGGGGAAGCTGGGCTGTGTGATCTGTGTGGCCGGAAGCACAGTGATGGTGATACATGC TCCTGAGGAAGAGAAGGTCACTACCATCATGGAGATGGCTTCCAAGATGAAAGACACAGGGTTCATCGTGTTTGCTGTGCTTCTGCTGGT GTCATGCCTCATCCTCATCTTTGTCATTGCCCCACGTTACGGGCAAAGGAATATCCTCATCTACATCATCATCTGCTCTGTGATCGGGGC CTTCTCTGTGGCTGCTGTCAAGGGGCTGGGCATCACCATCAAGAACTTCTTCCAGGGGCTGCCAGTTGTCCGGCACCCGCTCCCCTACAT CCTGTCCCTCATCCTGGCACTGTCCCTCAGCACTCAGGTCAACTTCCTCAACAGAGCACTGGACATTTTCAACACTTCCCTGGTGTTCCC CATCTACTACGTGTTCTTCACCACGGTGGTCGTTACCTCGTCCATCATCCTCTTCAAGGAGTGGTACAGCATGTCTGCTGTGGACATTGC AGGCACCCTCTCGGGCTTTGTCACCATCATCTTGGGCGTGTTCATGCTGCATGCTTTCAAAGACCTGGACATCAGCTGCGCCAGCTTGCC CCACATGCACAAAAACCCACCCCCTTCTCCCGCCCCGGAACCCACTGTTATTAGACTGGAAGACAAGAACGTCCTTGTGGACAATATAGA ACTTGCCAGCACCTCATCACCAGAAGAGAAACCCAAAGTATTTATAATCCATTCCTGAAGCTTGGAATATGTGAGTGAGAGGATGAGTCC GATGGTACAGCCTGCCCTCCCAATTTCAAAACCACCTGGTTATTTTCCAGTGCAACTGTTACCAATGGGCTCTCTTTTCTTGAGAAGTTC >92708_92708_8_TNIP1-NIPAL4_TNIP1_chr5_150441688_ENST00000523338_NIPAL4_chr5_156890102_ENST00000435489_length(amino acids)=371AA_BP=1 MLHLYCSSQEVLCQIVNDLSPEVPSNATFHSWQERIRQNYGFYIGLGLAFLSSFLIGSSVILKKKGLLRLVATGATRAVAAGEVANFGAY AFAPATVVTPLGALSVLISAILSSYFLRESLNLLGKLGCVICVAGSTVMVIHAPEEEKVTTIMEMASKMKDTGFIVFAVLLLVSCLILIF VIAPRYGQRNILIYIIICSVIGAFSVAAVKGLGITIKNFFQGLPVVRHPLPYILSLILALSLSTQVNFLNRALDIFNTSLVFPIYYVFFT TVVVTSSIILFKEWYSMSAVDIAGTLSGFVTIILGVFMLHAFKDLDISCASLPHMHKNPPPSPAPEPTVIRLEDKNVLVDNIELASTSSP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for TNIP1-NIPAL4 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 94_412 | 119.0 | 637.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 94_412 | 119.0 | 637.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 94_412 | 119.0 | 636.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 94_412 | 66.0 | 584.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 94_412 | 119.0 | 637.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 94_412 | 119.0 | 637.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 94_412 | 119.0 | 636.0 | Nef |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000315050 | - | 4 | 18 | 351_367 | 119.0 | 637.0 | Shigella flexneri ipah9.8 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000389378 | - | 4 | 18 | 351_367 | 119.0 | 637.0 | Shigella flexneri ipah9.8 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000518977 | - | 4 | 18 | 351_367 | 119.0 | 636.0 | Shigella flexneri ipah9.8 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000520931 | - | 3 | 17 | 351_367 | 66.0 | 584.0 | Shigella flexneri ipah9.8 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000521591 | - | 4 | 18 | 351_367 | 119.0 | 637.0 | Shigella flexneri ipah9.8 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000522226 | - | 4 | 18 | 351_367 | 119.0 | 637.0 | Shigella flexneri ipah9.8 |
| Hgene | TNIP1 | chr5:150441688 | chr5:156890102 | ENST00000523338 | - | 4 | 18 | 351_367 | 119.0 | 636.0 | Shigella flexneri ipah9.8 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for TNIP1-NIPAL4 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for TNIP1-NIPAL4 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
