
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:TRIT1-ABCA10 (FusionGDB2 ID:94265) |
Fusion Gene Summary for TRIT1-ABCA10 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: TRIT1-ABCA10 | Fusion gene ID: 94265 | Hgene | Tgene | Gene symbol | TRIT1 | ABCA10 | Gene ID | 54802 | 10349 |
| Gene name | tRNA isopentenyltransferase 1 | ATP binding cassette subfamily A member 10 | |
| Synonyms | COXPD35|GRO1|IPPT|IPT|IPTase|MOD5|hGRO1 | EST698739 | |
| Cytomap | 1p34.2 | 17q24.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | tRNA dimethylallyltransferaseIPP transferaseisopentenyl-diphosphate:tRNA isopentenyltransferasetRNA dimethylallyltransferase, mitochondrialtRNA isopentenylpyrophosphate transferase | ATP-binding cassette sub-family A member 10ATP-binding cassette A10ATP-binding cassette, sub-family A (ABC1), member 10 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q8WWZ4 | |
| Ensembl transtripts involved in fusion gene | ENST00000316891, ENST00000372818, ENST00000441669, ENST00000544981, ENST00000491865, ENST00000537223, ENST00000537440, ENST00000541099, ENST00000545233, | ENST00000423818, ENST00000519732, ENST00000269081, ENST00000416101, ENST00000432313, | |
| Fusion gene scores | * DoF score | 7 X 2 X 4=56 | 5 X 5 X 4=100 |
| # samples | 6 | 5 | |
| ** MAII score | log2(6/56*10)=0.0995356735509144 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(5/100*10)=-1 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: TRIT1 [Title/Abstract] AND ABCA10 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | TRIT1(40348990)-ABCA10(67193263), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | TRIT1-ABCA10 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. TRIT1-ABCA10 seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. TRIT1-ABCA10 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across TRIT1 (5'-gene) Fusion gene breakpoints across TRIT1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
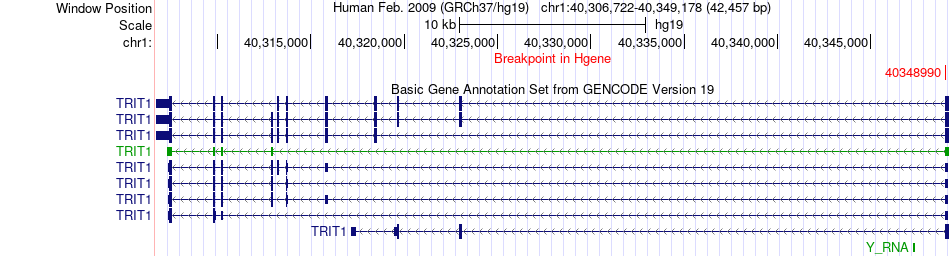 |
 Fusion gene breakpoints across ABCA10 (3'-gene) Fusion gene breakpoints across ABCA10 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
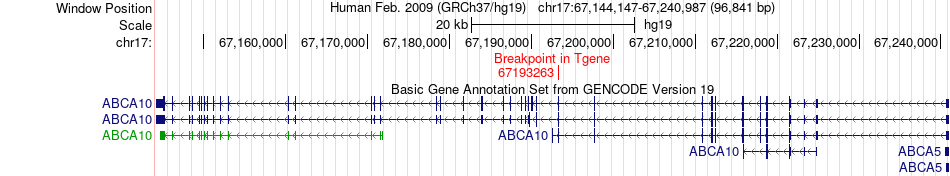 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LIHC | TCGA-PD-A5DF-01A | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
Top |
Fusion Gene ORF analysis for TRIT1-ABCA10 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000316891 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000316891 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000372818 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000372818 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000441669 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000441669 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000544981 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5CDS-intron | ENST00000544981 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000491865 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000491865 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000491865 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000537223 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000537223 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000537223 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000537440 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000537440 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000537440 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000541099 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000541099 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000541099 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000545233 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000545233 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-3CDS | ENST00000545233 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000491865 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000491865 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000537223 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000537223 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000537440 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000537440 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000541099 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000541099 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000545233 | ENST00000423818 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| 5UTR-intron | ENST00000545233 | ENST00000519732 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000316891 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000316891 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000372818 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000372818 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000441669 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000441669 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000544981 | ENST00000269081 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| Frame-shift | ENST00000544981 | ENST00000432313 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| In-frame | ENST00000316891 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| In-frame | ENST00000372818 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| In-frame | ENST00000441669 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
| In-frame | ENST00000544981 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000441669 | TRIT1 | chr1 | 40348990 | - | ENST00000416101 | ABCA10 | chr17 | 67193263 | - | 4275 | 174 | 329 | 3454 | 1041 |
| ENST00000316891 | TRIT1 | chr1 | 40348990 | - | ENST00000416101 | ABCA10 | chr17 | 67193263 | - | 4290 | 189 | 344 | 3469 | 1041 |
| ENST00000372818 | TRIT1 | chr1 | 40348990 | - | ENST00000416101 | ABCA10 | chr17 | 67193263 | - | 4290 | 189 | 344 | 3469 | 1041 |
| ENST00000544981 | TRIT1 | chr1 | 40348990 | - | ENST00000416101 | ABCA10 | chr17 | 67193263 | - | 4288 | 187 | 342 | 3467 | 1041 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000441669 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - | 0.000282532 | 0.9997174 |
| ENST00000316891 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - | 0.000295195 | 0.9997048 |
| ENST00000372818 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - | 0.000295195 | 0.9997048 |
| ENST00000544981 | ENST00000416101 | TRIT1 | chr1 | 40348990 | - | ABCA10 | chr17 | 67193263 | - | 0.000290228 | 0.9997098 |
Top |
Fusion Genomic Features for TRIT1-ABCA10 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
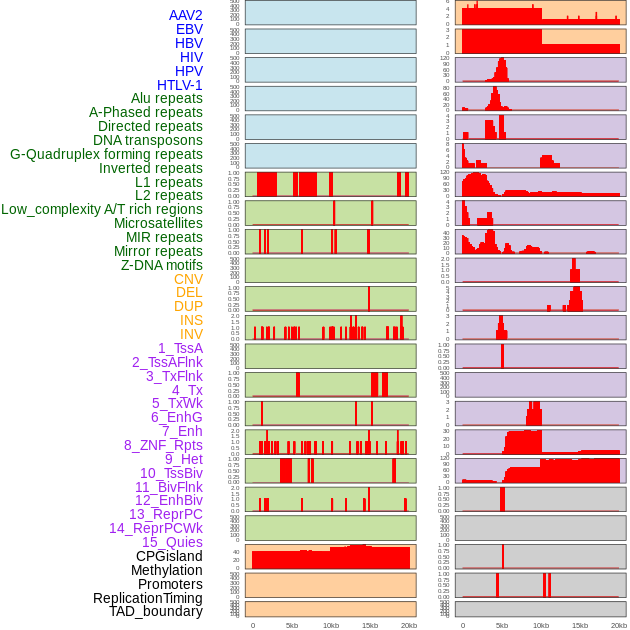 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for TRIT1-ABCA10 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr1:40348990/chr17:67193263) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | ABCA10 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Probable transporter which may play a role in macrophage lipid transport and homeostasis. {ECO:0000305|PubMed:12821155}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 32_37 | 58 | 468.0 | Region | Substrate binding |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 32_37 | 58 | 442.0 | Region | Substrate binding |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 32_37 | 58 | 386.0 | Region | Substrate binding |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 1206_1440 | 391 | 1544.0 | Domain | ABC transporter 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 391_626 | 391 | 1544.0 | Domain | ABC transporter 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 1206_1440 | 391 | 449.0 | Domain | ABC transporter 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 391_626 | 391 | 449.0 | Domain | ABC transporter 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 1206_1440 | 0 | 93.0 | Domain | ABC transporter 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 391_626 | 0 | 93.0 | Domain | ABC transporter 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 1206_1440 | 391 | 453.0 | Domain | ABC transporter 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 391_626 | 391 | 453.0 | Domain | ABC transporter 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 1239_1246 | 391 | 1544.0 | Nucleotide binding | ATP 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 427_434 | 391 | 1544.0 | Nucleotide binding | ATP 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 1239_1246 | 391 | 449.0 | Nucleotide binding | ATP 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 427_434 | 391 | 449.0 | Nucleotide binding | ATP 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 1239_1246 | 0 | 93.0 | Nucleotide binding | ATP 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 427_434 | 0 | 93.0 | Nucleotide binding | ATP 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 1239_1246 | 391 | 453.0 | Nucleotide binding | ATP 2 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 427_434 | 391 | 453.0 | Nucleotide binding | ATP 1 | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 1014_1034 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 1046_1066 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 1073_1093 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 1113_1133 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 774_794 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 890_910 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 926_946 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 985_1005 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 1014_1034 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 1046_1066 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 1073_1093 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 1113_1133 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 774_794 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 890_910 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 926_946 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 985_1005 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 1014_1034 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 1046_1066 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 1073_1093 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 1113_1133 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 135_155 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 185_205 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 210_230 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 240_260 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 264_284 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 310_330 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 774_794 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 83_103 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 890_910 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 926_946 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000423818 | 0 | 5 | 985_1005 | 0 | 93.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 1014_1034 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 1046_1066 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 1073_1093 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 1113_1133 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 774_794 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 890_910 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 926_946 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 985_1005 | 391 | 453.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 395_425 | 58 | 468.0 | Zinc finger | Note=Matrin-type |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 395_425 | 58 | 442.0 | Zinc finger | Note=Matrin-type |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 395_425 | 58 | 386.0 | Zinc finger | Note=Matrin-type |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 135_155 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 185_205 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 210_230 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 240_260 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 264_284 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 310_330 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000269081 | 10 | 40 | 83_103 | 391 | 1544.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 135_155 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 185_205 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 210_230 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 240_260 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 264_284 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 310_330 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000416101 | 10 | 39 | 83_103 | 391 | 449.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 135_155 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 185_205 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 210_230 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 240_260 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 264_284 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 310_330 | 391 | 453.0 | Transmembrane | Helical | |
| Tgene | ABCA10 | chr1:40348990 | chr17:67193263 | ENST00000432313 | 10 | 13 | 83_103 | 391 | 453.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for TRIT1-ABCA10 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >94265_94265_1_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000316891_ABCA10_chr17_67193263_ENST00000416101_length(transcript)=4290nt_BP=189nt GCAGACTGCCATAAGATGGCGTCCGTGGCGGCTGCACGAGCAGTTCCCGTGGGCAGTGGGCTCAGGGGCCTGCAACGGACCCTACCTCTT GTAGTGATTCTCGGGGCCACGGGCACCGGCAAATCCACGCTGGCGTTGCAGCTAGGCCAGCGGCTCGGCGGTGAGATCGTCAGCGCTGAC TCCATGCAGAATCAGAAATGTTATAAAAGAATATAATGGAAAGACTGGAAAAGTAGAAGCATTGCAAGGCATATTTTTTGACATATATGA AGGACAGATCACTGCAATACTTGGGCATAATGGAGCTGGTAAATCAACACTGCTAAACATTCTTAGTGGATTGTCTGTTTCTACAGAAGG TAAAAAGAATTATAATGGAATTAGACATGCAAAGCATTCAAGACATTATTGCTAAAAAATTAAGTGGTGGGCAGAAGAGAAAACTAACAC TAGGGATTGCCATCTTAGGAGATCCTCAGGTTTTGCTGCTAGATGAACCAACTGCTGGATTGGATCCCTTTTCAAGACACCGAGTGTGGA GCCTCCTGAAGGAGCATAAAGTAGACCGACTTATCCTCTTCAGTACCCAATTCATGGATGAGGCTGACATCTTGGCTGATAGGAAAGTAT TTCTGTCTAATGGGAAGTTGAAATGTGCAGGATCATCTTTGTTTCTGAAGCGAAAGTGGGGTATTGGATATCATTTAAGTTTACACAGGA ATGAAATGTGTGACACAGAAAAAATCACATCCCTTATTAAGCAGCACATTCCTGATGCCAAGTTAACAACAGAAAGTGAAGAAAAACTTG TATATAGTTTGCCTTTGGAAAAAACGAACAAATTTCCAGATCTTTACAGTGACCTTGATAAGTGTTCTGACCAGGGCATAAGGAATTATG CTGTTTCAGTGACATCTCTGAATGAAGTATTCTTGAACCTAGAAGGAAAATCAGCAATTGATGAACCAGATTTTGACATTGGGAAACAAG AGAAAATACATGTGACAAGAAATACTGGAGATGAGTCTGAAATGGAACAGGTTCTTTGTTCTCTTCCTGAAACAAGAAAGGCTGTCAGTA GTGCAGCTCTCTGGAGACGACAAATCTATGCAGTGGCAACACTTCGCTTCTTAAAGTTAAGGCGTGAAAGGAGAGCTCTTTTGTGTTTGT TACTAGTACTTGGAATTGCTTTTATCCCCATCATTCTAGAGAAGATAATGTATAAAGTAACTCGTGAAACTCATTGTTGGGAGTTTTCAC CCAGTATGTATTTCCTTTCTCTGGAACAAATCCCGAAGACGCCTCTTACCAGCCTGTTAATCGTTAATAATACAGGATCAAATATTGAAG ACCTCGTGCATTCACTGAAGTGTCAGGATATAGTTTTGGAAATAGATGACTTTAGAAACAGAAATGGCTCAGATGATCCCTCCTACAATG GAGCCATCATAGTGTCTGGTGACCAGAAGGATTACAGATTTTCTGTTGCGTGTAATACCAAGAAATTGAATTGTTTTCCTGTTCTTATGG GAATTGTTAGCAATGCCCTTATGGGAATTTTTAACTTCACGGAGCTTATTCAAATGGAGAGCACTTCATTTTCTCGTGATGACATAGTGC TGGATCTTGGTTTTATAGATGGGTCCATATTTTTGTTGTTGATCACAAACTGCGTTTCTCCTTTTATCGGCATGAGCAGCATCAGCGATT ATAAAAAAAATGTTCAATCCCAGTTATGGATTTCAGGCCTCTGGCCTTCAGCATACTGGTGTGGACAGGCTCTGGTGGACATTCCATTAT ACTTCTTGATTCTCTTTTCAATACATTTAATTTACTACTTCATATTTCTGGGATTCCAGCTTTCATGGGAACTCATGTTTGTTTTGGTGG TATGCATAATTGGTTGTGCAGTTTCTCTTATATTCCTCACATATGTGCTTTCATTCATCTTTCGCAAGTGGAGAAAAAATAATGGCTTTT GGTCTTTTGGCTTTTTTATTATCTTAATATGTGTATCCACAATTATGGTATCAACTCAATATGAAAAACTCAACTTAATTTTGTGCATGA TTTTCATACCTTCCTTCACTTTGCTGGGGTATGTCATGTTATTGATCCAGCTCGACTTTATGAGAAACTTGGACAGTCTGGACAATAGAA TAAATGAAGTCAATAAAACCATTCTTTTAACAACCTTAATACCATACCTTCAGAGTGTTATTTTCCTTTTTGTCATAAGGTGTCTGGAAA TGAAGTATGGAAATGAAATAATGAATAAAGACCCAGTTTTCAGAATCTCTCCACGGAGTAGAGAAACTCATCCCAATCCGGAAGAGCCCG AAGAAGAAGATGAAGATGTTCAAGCTGAAAGAGTCCAAGCAGCAAATGCACTCACTGCTCCAAACTTGGAGGAGGAACCAGTCATAACTG CAAGCTGTTTACACAAGGAATATTATGAGACAAAGAAAAGTTGCTTTTCAACAAGAAAGAAGAAAATAGCCATCAGAAATGTTTCCTTTT GTGTTAAAAAAGGTGAAGTTTTGGGATTACTAGGACACAATGGAGCTGGTAAAAGTACTTCCATTAAAATGATAACTGGGTGCACAAAGC CAACTGCAGGAGTGGTGGTGTTACAAGGCAGCAGAGCATCAGTAAGGCAACAGCATGACAACAGCCTCAAGTTCTTGGGGTACTGCCCTC AGGAGAACTCACTGTGGCCCAAGCTTACAATGAAAGAGCACTTGGAGTTGTATGCAGCTGTGAAAGGACTGGGCAAAGAAGATGCTGCTC TCAGTATTTCACGATTGGTGGAAGCTCTTAAGCTCCAGGAACAACTTAAGGCTCCTGTGAAAACTCTATCAGAGGGAATAAAGAGAAAGC TGTGCTTTGTGCTGAGCATCCTGGGGAACCCATCAGTGGTGCTTCTAGATGAGCCGTTCACCGGGATGGACCCCGAGGGGCAGCAGCAAA TGTGGCAGATACTTCAGGCTACCGTTAAAAACAAGGAGAGGGGCACCCTCTTGACCACCCATTACATGTCAGAGGCTGAGGCTGTGTGTG ACCGTATGGCCATGATGGTGTCAGGAACGCTAAGGTGTATTGGTTCCATTCAACATCTGAAAAACAAGTTTGGTAGAGATTATTTACTAG AAATAAAAATGAAAGAACCTACCCAGGTGGAAGCTCTCCACACAGAGATTTTGAAGCTTTTCCCACAGGCTGCTTGGCAGGAAAGATATT CCTCTTTAATGGCGTATAAGTTACCTGTGGAGGATGTCCACCCTCTATCTCGGGCCTTTTTCAAGTTAGAGGCGATGAAACAGACCTTCA ACCTGGAGGAATACAGCCTCTCTCAGGCTACCTTGGAGCAGGTATTCTTAGAACTCTGTAAAGAGCAGGAGCTGGGAAATGTTGATGATA AAATTGATACAACAGTTGAATGGAAACTTCTCCCACAGGAAGACCCTTAAAATGAAGAACCTCCTAACATTCAATTTTAGGTCCTACTAC ATTGTTAGTTTCCATAATTCTACAAGAATGTTTCCTTTTACTTCAGTTAACAAAAGAAAACATTTAATAAACATTCAATAATGATTACAG TTTTCATTTTTAAAAATTTAGGATGAAGGAAACAAGGAAATATAGGGAAAAGTAGTAGACAAAATTAACAAAATCAGACATGTTATTCAT CCCCAACATGGGTCTATTTTGTGCTTAAAAATAATTTAAAAATCATACAATATTAGGTTGGTTATCGGTTATTATCAATAAAGCTAACAC TGAGAACATTTTACAAATAAAAATATGAGGTTTTTAGCCTGAACTTCAAATGTATCAGCTATTTTTAAACATTATTTACTCGGATTCTAA TTTAATGTGACATTGACTATAAGAAGGTCTGATAAACTGATGAAATGGCACAGCATAACATTTAATTATAATGACATTCTGATTATAAAA TAAATGCATGTGAATTTTAGTACATATTGAAGTTATATGGAAGAAGATAGCCATAATCTGTAAGAAAGTACCGCAGTTAATATTTTCTTT AGCCAACTTATATTCAATGTATTTTTTATGGATCCTTTTTCAAAGGTAGTATCAGTAGGCATAGTCATTTTCTGTATCTTTTCACCTCAC AGTTCATGAACATTTCCCATGTCATTATAGCACTTTTGTGTTATATAATTGCTGCATTAATGTTATAGCTGTATTGATGTAATACCAGAG >94265_94265_1_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000316891_ABCA10_chr17_67193263_ENST00000416101_length(amino acids)=1041AA_BP=729 MFLQKVKRIIMELDMQSIQDIIAKKLSGGQKRKLTLGIAILGDPQVLLLDEPTAGLDPFSRHRVWSLLKEHKVDRLILFSTQFMDEADIL ADRKVFLSNGKLKCAGSSLFLKRKWGIGYHLSLHRNEMCDTEKITSLIKQHIPDAKLTTESEEKLVYSLPLEKTNKFPDLYSDLDKCSDQ GIRNYAVSVTSLNEVFLNLEGKSAIDEPDFDIGKQEKIHVTRNTGDESEMEQVLCSLPETRKAVSSAALWRRQIYAVATLRFLKLRRERR ALLCLLLVLGIAFIPIILEKIMYKVTRETHCWEFSPSMYFLSLEQIPKTPLTSLLIVNNTGSNIEDLVHSLKCQDIVLEIDDFRNRNGSD DPSYNGAIIVSGDQKDYRFSVACNTKKLNCFPVLMGIVSNALMGIFNFTELIQMESTSFSRDDIVLDLGFIDGSIFLLLITNCVSPFIGM SSISDYKKNVQSQLWISGLWPSAYWCGQALVDIPLYFLILFSIHLIYYFIFLGFQLSWELMFVLVVCIIGCAVSLIFLTYVLSFIFRKWR KNNGFWSFGFFIILICVSTIMVSTQYEKLNLILCMIFIPSFTLLGYVMLLIQLDFMRNLDSLDNRINEVNKTILLTTLIPYLQSVIFLFV IRCLEMKYGNEIMNKDPVFRISPRSRETHPNPEEPEEEDEDVQAERVQAANALTAPNLEEEPVITASCLHKEYYETKKSCFSTRKKKIAI RNVSFCVKKGEVLGLLGHNGAGKSTSIKMITGCTKPTAGVVVLQGSRASVRQQHDNSLKFLGYCPQENSLWPKLTMKEHLELYAAVKGLG KEDAALSISRLVEALKLQEQLKAPVKTLSEGIKRKLCFVLSILGNPSVVLLDEPFTGMDPEGQQQMWQILQATVKNKERGTLLTTHYMSE AEAVCDRMAMMVSGTLRCIGSIQHLKNKFGRDYLLEIKMKEPTQVEALHTEILKLFPQAAWQERYSSLMAYKLPVEDVHPLSRAFFKLEA -------------------------------------------------------------- >94265_94265_2_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000372818_ABCA10_chr17_67193263_ENST00000416101_length(transcript)=4290nt_BP=189nt GCAGACTGCCATAAGATGGCGTCCGTGGCGGCTGCACGAGCAGTTCCCGTGGGCAGTGGGCTCAGGGGCCTGCAACGGACCCTACCTCTT GTAGTGATTCTCGGGGCCACGGGCACCGGCAAATCCACGCTGGCGTTGCAGCTAGGCCAGCGGCTCGGCGGTGAGATCGTCAGCGCTGAC TCCATGCAGAATCAGAAATGTTATAAAAGAATATAATGGAAAGACTGGAAAAGTAGAAGCATTGCAAGGCATATTTTTTGACATATATGA AGGACAGATCACTGCAATACTTGGGCATAATGGAGCTGGTAAATCAACACTGCTAAACATTCTTAGTGGATTGTCTGTTTCTACAGAAGG TAAAAAGAATTATAATGGAATTAGACATGCAAAGCATTCAAGACATTATTGCTAAAAAATTAAGTGGTGGGCAGAAGAGAAAACTAACAC TAGGGATTGCCATCTTAGGAGATCCTCAGGTTTTGCTGCTAGATGAACCAACTGCTGGATTGGATCCCTTTTCAAGACACCGAGTGTGGA GCCTCCTGAAGGAGCATAAAGTAGACCGACTTATCCTCTTCAGTACCCAATTCATGGATGAGGCTGACATCTTGGCTGATAGGAAAGTAT TTCTGTCTAATGGGAAGTTGAAATGTGCAGGATCATCTTTGTTTCTGAAGCGAAAGTGGGGTATTGGATATCATTTAAGTTTACACAGGA ATGAAATGTGTGACACAGAAAAAATCACATCCCTTATTAAGCAGCACATTCCTGATGCCAAGTTAACAACAGAAAGTGAAGAAAAACTTG TATATAGTTTGCCTTTGGAAAAAACGAACAAATTTCCAGATCTTTACAGTGACCTTGATAAGTGTTCTGACCAGGGCATAAGGAATTATG CTGTTTCAGTGACATCTCTGAATGAAGTATTCTTGAACCTAGAAGGAAAATCAGCAATTGATGAACCAGATTTTGACATTGGGAAACAAG AGAAAATACATGTGACAAGAAATACTGGAGATGAGTCTGAAATGGAACAGGTTCTTTGTTCTCTTCCTGAAACAAGAAAGGCTGTCAGTA GTGCAGCTCTCTGGAGACGACAAATCTATGCAGTGGCAACACTTCGCTTCTTAAAGTTAAGGCGTGAAAGGAGAGCTCTTTTGTGTTTGT TACTAGTACTTGGAATTGCTTTTATCCCCATCATTCTAGAGAAGATAATGTATAAAGTAACTCGTGAAACTCATTGTTGGGAGTTTTCAC CCAGTATGTATTTCCTTTCTCTGGAACAAATCCCGAAGACGCCTCTTACCAGCCTGTTAATCGTTAATAATACAGGATCAAATATTGAAG ACCTCGTGCATTCACTGAAGTGTCAGGATATAGTTTTGGAAATAGATGACTTTAGAAACAGAAATGGCTCAGATGATCCCTCCTACAATG GAGCCATCATAGTGTCTGGTGACCAGAAGGATTACAGATTTTCTGTTGCGTGTAATACCAAGAAATTGAATTGTTTTCCTGTTCTTATGG GAATTGTTAGCAATGCCCTTATGGGAATTTTTAACTTCACGGAGCTTATTCAAATGGAGAGCACTTCATTTTCTCGTGATGACATAGTGC TGGATCTTGGTTTTATAGATGGGTCCATATTTTTGTTGTTGATCACAAACTGCGTTTCTCCTTTTATCGGCATGAGCAGCATCAGCGATT ATAAAAAAAATGTTCAATCCCAGTTATGGATTTCAGGCCTCTGGCCTTCAGCATACTGGTGTGGACAGGCTCTGGTGGACATTCCATTAT ACTTCTTGATTCTCTTTTCAATACATTTAATTTACTACTTCATATTTCTGGGATTCCAGCTTTCATGGGAACTCATGTTTGTTTTGGTGG TATGCATAATTGGTTGTGCAGTTTCTCTTATATTCCTCACATATGTGCTTTCATTCATCTTTCGCAAGTGGAGAAAAAATAATGGCTTTT GGTCTTTTGGCTTTTTTATTATCTTAATATGTGTATCCACAATTATGGTATCAACTCAATATGAAAAACTCAACTTAATTTTGTGCATGA TTTTCATACCTTCCTTCACTTTGCTGGGGTATGTCATGTTATTGATCCAGCTCGACTTTATGAGAAACTTGGACAGTCTGGACAATAGAA TAAATGAAGTCAATAAAACCATTCTTTTAACAACCTTAATACCATACCTTCAGAGTGTTATTTTCCTTTTTGTCATAAGGTGTCTGGAAA TGAAGTATGGAAATGAAATAATGAATAAAGACCCAGTTTTCAGAATCTCTCCACGGAGTAGAGAAACTCATCCCAATCCGGAAGAGCCCG AAGAAGAAGATGAAGATGTTCAAGCTGAAAGAGTCCAAGCAGCAAATGCACTCACTGCTCCAAACTTGGAGGAGGAACCAGTCATAACTG CAAGCTGTTTACACAAGGAATATTATGAGACAAAGAAAAGTTGCTTTTCAACAAGAAAGAAGAAAATAGCCATCAGAAATGTTTCCTTTT GTGTTAAAAAAGGTGAAGTTTTGGGATTACTAGGACACAATGGAGCTGGTAAAAGTACTTCCATTAAAATGATAACTGGGTGCACAAAGC CAACTGCAGGAGTGGTGGTGTTACAAGGCAGCAGAGCATCAGTAAGGCAACAGCATGACAACAGCCTCAAGTTCTTGGGGTACTGCCCTC AGGAGAACTCACTGTGGCCCAAGCTTACAATGAAAGAGCACTTGGAGTTGTATGCAGCTGTGAAAGGACTGGGCAAAGAAGATGCTGCTC TCAGTATTTCACGATTGGTGGAAGCTCTTAAGCTCCAGGAACAACTTAAGGCTCCTGTGAAAACTCTATCAGAGGGAATAAAGAGAAAGC TGTGCTTTGTGCTGAGCATCCTGGGGAACCCATCAGTGGTGCTTCTAGATGAGCCGTTCACCGGGATGGACCCCGAGGGGCAGCAGCAAA TGTGGCAGATACTTCAGGCTACCGTTAAAAACAAGGAGAGGGGCACCCTCTTGACCACCCATTACATGTCAGAGGCTGAGGCTGTGTGTG ACCGTATGGCCATGATGGTGTCAGGAACGCTAAGGTGTATTGGTTCCATTCAACATCTGAAAAACAAGTTTGGTAGAGATTATTTACTAG AAATAAAAATGAAAGAACCTACCCAGGTGGAAGCTCTCCACACAGAGATTTTGAAGCTTTTCCCACAGGCTGCTTGGCAGGAAAGATATT CCTCTTTAATGGCGTATAAGTTACCTGTGGAGGATGTCCACCCTCTATCTCGGGCCTTTTTCAAGTTAGAGGCGATGAAACAGACCTTCA ACCTGGAGGAATACAGCCTCTCTCAGGCTACCTTGGAGCAGGTATTCTTAGAACTCTGTAAAGAGCAGGAGCTGGGAAATGTTGATGATA AAATTGATACAACAGTTGAATGGAAACTTCTCCCACAGGAAGACCCTTAAAATGAAGAACCTCCTAACATTCAATTTTAGGTCCTACTAC ATTGTTAGTTTCCATAATTCTACAAGAATGTTTCCTTTTACTTCAGTTAACAAAAGAAAACATTTAATAAACATTCAATAATGATTACAG TTTTCATTTTTAAAAATTTAGGATGAAGGAAACAAGGAAATATAGGGAAAAGTAGTAGACAAAATTAACAAAATCAGACATGTTATTCAT CCCCAACATGGGTCTATTTTGTGCTTAAAAATAATTTAAAAATCATACAATATTAGGTTGGTTATCGGTTATTATCAATAAAGCTAACAC TGAGAACATTTTACAAATAAAAATATGAGGTTTTTAGCCTGAACTTCAAATGTATCAGCTATTTTTAAACATTATTTACTCGGATTCTAA TTTAATGTGACATTGACTATAAGAAGGTCTGATAAACTGATGAAATGGCACAGCATAACATTTAATTATAATGACATTCTGATTATAAAA TAAATGCATGTGAATTTTAGTACATATTGAAGTTATATGGAAGAAGATAGCCATAATCTGTAAGAAAGTACCGCAGTTAATATTTTCTTT AGCCAACTTATATTCAATGTATTTTTTATGGATCCTTTTTCAAAGGTAGTATCAGTAGGCATAGTCATTTTCTGTATCTTTTCACCTCAC AGTTCATGAACATTTCCCATGTCATTATAGCACTTTTGTGTTATATAATTGCTGCATTAATGTTATAGCTGTATTGATGTAATACCAGAG >94265_94265_2_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000372818_ABCA10_chr17_67193263_ENST00000416101_length(amino acids)=1041AA_BP=729 MFLQKVKRIIMELDMQSIQDIIAKKLSGGQKRKLTLGIAILGDPQVLLLDEPTAGLDPFSRHRVWSLLKEHKVDRLILFSTQFMDEADIL ADRKVFLSNGKLKCAGSSLFLKRKWGIGYHLSLHRNEMCDTEKITSLIKQHIPDAKLTTESEEKLVYSLPLEKTNKFPDLYSDLDKCSDQ GIRNYAVSVTSLNEVFLNLEGKSAIDEPDFDIGKQEKIHVTRNTGDESEMEQVLCSLPETRKAVSSAALWRRQIYAVATLRFLKLRRERR ALLCLLLVLGIAFIPIILEKIMYKVTRETHCWEFSPSMYFLSLEQIPKTPLTSLLIVNNTGSNIEDLVHSLKCQDIVLEIDDFRNRNGSD DPSYNGAIIVSGDQKDYRFSVACNTKKLNCFPVLMGIVSNALMGIFNFTELIQMESTSFSRDDIVLDLGFIDGSIFLLLITNCVSPFIGM SSISDYKKNVQSQLWISGLWPSAYWCGQALVDIPLYFLILFSIHLIYYFIFLGFQLSWELMFVLVVCIIGCAVSLIFLTYVLSFIFRKWR KNNGFWSFGFFIILICVSTIMVSTQYEKLNLILCMIFIPSFTLLGYVMLLIQLDFMRNLDSLDNRINEVNKTILLTTLIPYLQSVIFLFV IRCLEMKYGNEIMNKDPVFRISPRSRETHPNPEEPEEEDEDVQAERVQAANALTAPNLEEEPVITASCLHKEYYETKKSCFSTRKKKIAI RNVSFCVKKGEVLGLLGHNGAGKSTSIKMITGCTKPTAGVVVLQGSRASVRQQHDNSLKFLGYCPQENSLWPKLTMKEHLELYAAVKGLG KEDAALSISRLVEALKLQEQLKAPVKTLSEGIKRKLCFVLSILGNPSVVLLDEPFTGMDPEGQQQMWQILQATVKNKERGTLLTTHYMSE AEAVCDRMAMMVSGTLRCIGSIQHLKNKFGRDYLLEIKMKEPTQVEALHTEILKLFPQAAWQERYSSLMAYKLPVEDVHPLSRAFFKLEA -------------------------------------------------------------- >94265_94265_3_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000441669_ABCA10_chr17_67193263_ENST00000416101_length(transcript)=4275nt_BP=174nt ATGGCGTCCGTGGCGGCTGCACGAGCAGTTCCCGTGGGCAGTGGGCTCAGGGGCCTGCAACGGACCCTACCTCTTGTAGTGATTCTCGGG GCCACGGGCACCGGCAAATCCACGCTGGCGTTGCAGCTAGGCCAGCGGCTCGGCGGTGAGATCGTCAGCGCTGACTCCATGCAGAATCAG AAATGTTATAAAAGAATATAATGGAAAGACTGGAAAAGTAGAAGCATTGCAAGGCATATTTTTTGACATATATGAAGGACAGATCACTGC AATACTTGGGCATAATGGAGCTGGTAAATCAACACTGCTAAACATTCTTAGTGGATTGTCTGTTTCTACAGAAGGTAAAAAGAATTATAA TGGAATTAGACATGCAAAGCATTCAAGACATTATTGCTAAAAAATTAAGTGGTGGGCAGAAGAGAAAACTAACACTAGGGATTGCCATCT TAGGAGATCCTCAGGTTTTGCTGCTAGATGAACCAACTGCTGGATTGGATCCCTTTTCAAGACACCGAGTGTGGAGCCTCCTGAAGGAGC ATAAAGTAGACCGACTTATCCTCTTCAGTACCCAATTCATGGATGAGGCTGACATCTTGGCTGATAGGAAAGTATTTCTGTCTAATGGGA AGTTGAAATGTGCAGGATCATCTTTGTTTCTGAAGCGAAAGTGGGGTATTGGATATCATTTAAGTTTACACAGGAATGAAATGTGTGACA CAGAAAAAATCACATCCCTTATTAAGCAGCACATTCCTGATGCCAAGTTAACAACAGAAAGTGAAGAAAAACTTGTATATAGTTTGCCTT TGGAAAAAACGAACAAATTTCCAGATCTTTACAGTGACCTTGATAAGTGTTCTGACCAGGGCATAAGGAATTATGCTGTTTCAGTGACAT CTCTGAATGAAGTATTCTTGAACCTAGAAGGAAAATCAGCAATTGATGAACCAGATTTTGACATTGGGAAACAAGAGAAAATACATGTGA CAAGAAATACTGGAGATGAGTCTGAAATGGAACAGGTTCTTTGTTCTCTTCCTGAAACAAGAAAGGCTGTCAGTAGTGCAGCTCTCTGGA GACGACAAATCTATGCAGTGGCAACACTTCGCTTCTTAAAGTTAAGGCGTGAAAGGAGAGCTCTTTTGTGTTTGTTACTAGTACTTGGAA TTGCTTTTATCCCCATCATTCTAGAGAAGATAATGTATAAAGTAACTCGTGAAACTCATTGTTGGGAGTTTTCACCCAGTATGTATTTCC TTTCTCTGGAACAAATCCCGAAGACGCCTCTTACCAGCCTGTTAATCGTTAATAATACAGGATCAAATATTGAAGACCTCGTGCATTCAC TGAAGTGTCAGGATATAGTTTTGGAAATAGATGACTTTAGAAACAGAAATGGCTCAGATGATCCCTCCTACAATGGAGCCATCATAGTGT CTGGTGACCAGAAGGATTACAGATTTTCTGTTGCGTGTAATACCAAGAAATTGAATTGTTTTCCTGTTCTTATGGGAATTGTTAGCAATG CCCTTATGGGAATTTTTAACTTCACGGAGCTTATTCAAATGGAGAGCACTTCATTTTCTCGTGATGACATAGTGCTGGATCTTGGTTTTA TAGATGGGTCCATATTTTTGTTGTTGATCACAAACTGCGTTTCTCCTTTTATCGGCATGAGCAGCATCAGCGATTATAAAAAAAATGTTC AATCCCAGTTATGGATTTCAGGCCTCTGGCCTTCAGCATACTGGTGTGGACAGGCTCTGGTGGACATTCCATTATACTTCTTGATTCTCT TTTCAATACATTTAATTTACTACTTCATATTTCTGGGATTCCAGCTTTCATGGGAACTCATGTTTGTTTTGGTGGTATGCATAATTGGTT GTGCAGTTTCTCTTATATTCCTCACATATGTGCTTTCATTCATCTTTCGCAAGTGGAGAAAAAATAATGGCTTTTGGTCTTTTGGCTTTT TTATTATCTTAATATGTGTATCCACAATTATGGTATCAACTCAATATGAAAAACTCAACTTAATTTTGTGCATGATTTTCATACCTTCCT TCACTTTGCTGGGGTATGTCATGTTATTGATCCAGCTCGACTTTATGAGAAACTTGGACAGTCTGGACAATAGAATAAATGAAGTCAATA AAACCATTCTTTTAACAACCTTAATACCATACCTTCAGAGTGTTATTTTCCTTTTTGTCATAAGGTGTCTGGAAATGAAGTATGGAAATG AAATAATGAATAAAGACCCAGTTTTCAGAATCTCTCCACGGAGTAGAGAAACTCATCCCAATCCGGAAGAGCCCGAAGAAGAAGATGAAG ATGTTCAAGCTGAAAGAGTCCAAGCAGCAAATGCACTCACTGCTCCAAACTTGGAGGAGGAACCAGTCATAACTGCAAGCTGTTTACACA AGGAATATTATGAGACAAAGAAAAGTTGCTTTTCAACAAGAAAGAAGAAAATAGCCATCAGAAATGTTTCCTTTTGTGTTAAAAAAGGTG AAGTTTTGGGATTACTAGGACACAATGGAGCTGGTAAAAGTACTTCCATTAAAATGATAACTGGGTGCACAAAGCCAACTGCAGGAGTGG TGGTGTTACAAGGCAGCAGAGCATCAGTAAGGCAACAGCATGACAACAGCCTCAAGTTCTTGGGGTACTGCCCTCAGGAGAACTCACTGT GGCCCAAGCTTACAATGAAAGAGCACTTGGAGTTGTATGCAGCTGTGAAAGGACTGGGCAAAGAAGATGCTGCTCTCAGTATTTCACGAT TGGTGGAAGCTCTTAAGCTCCAGGAACAACTTAAGGCTCCTGTGAAAACTCTATCAGAGGGAATAAAGAGAAAGCTGTGCTTTGTGCTGA GCATCCTGGGGAACCCATCAGTGGTGCTTCTAGATGAGCCGTTCACCGGGATGGACCCCGAGGGGCAGCAGCAAATGTGGCAGATACTTC AGGCTACCGTTAAAAACAAGGAGAGGGGCACCCTCTTGACCACCCATTACATGTCAGAGGCTGAGGCTGTGTGTGACCGTATGGCCATGA TGGTGTCAGGAACGCTAAGGTGTATTGGTTCCATTCAACATCTGAAAAACAAGTTTGGTAGAGATTATTTACTAGAAATAAAAATGAAAG AACCTACCCAGGTGGAAGCTCTCCACACAGAGATTTTGAAGCTTTTCCCACAGGCTGCTTGGCAGGAAAGATATTCCTCTTTAATGGCGT ATAAGTTACCTGTGGAGGATGTCCACCCTCTATCTCGGGCCTTTTTCAAGTTAGAGGCGATGAAACAGACCTTCAACCTGGAGGAATACA GCCTCTCTCAGGCTACCTTGGAGCAGGTATTCTTAGAACTCTGTAAAGAGCAGGAGCTGGGAAATGTTGATGATAAAATTGATACAACAG TTGAATGGAAACTTCTCCCACAGGAAGACCCTTAAAATGAAGAACCTCCTAACATTCAATTTTAGGTCCTACTACATTGTTAGTTTCCAT AATTCTACAAGAATGTTTCCTTTTACTTCAGTTAACAAAAGAAAACATTTAATAAACATTCAATAATGATTACAGTTTTCATTTTTAAAA ATTTAGGATGAAGGAAACAAGGAAATATAGGGAAAAGTAGTAGACAAAATTAACAAAATCAGACATGTTATTCATCCCCAACATGGGTCT ATTTTGTGCTTAAAAATAATTTAAAAATCATACAATATTAGGTTGGTTATCGGTTATTATCAATAAAGCTAACACTGAGAACATTTTACA AATAAAAATATGAGGTTTTTAGCCTGAACTTCAAATGTATCAGCTATTTTTAAACATTATTTACTCGGATTCTAATTTAATGTGACATTG ACTATAAGAAGGTCTGATAAACTGATGAAATGGCACAGCATAACATTTAATTATAATGACATTCTGATTATAAAATAAATGCATGTGAAT TTTAGTACATATTGAAGTTATATGGAAGAAGATAGCCATAATCTGTAAGAAAGTACCGCAGTTAATATTTTCTTTAGCCAACTTATATTC AATGTATTTTTTATGGATCCTTTTTCAAAGGTAGTATCAGTAGGCATAGTCATTTTCTGTATCTTTTCACCTCACAGTTCATGAACATTT CCCATGTCATTATAGCACTTTTGTGTTATATAATTGCTGCATTAATGTTATAGCTGTATTGATGTAATACCAGAGTTGTTACGTTCGGGT >94265_94265_3_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000441669_ABCA10_chr17_67193263_ENST00000416101_length(amino acids)=1041AA_BP=729 MFLQKVKRIIMELDMQSIQDIIAKKLSGGQKRKLTLGIAILGDPQVLLLDEPTAGLDPFSRHRVWSLLKEHKVDRLILFSTQFMDEADIL ADRKVFLSNGKLKCAGSSLFLKRKWGIGYHLSLHRNEMCDTEKITSLIKQHIPDAKLTTESEEKLVYSLPLEKTNKFPDLYSDLDKCSDQ GIRNYAVSVTSLNEVFLNLEGKSAIDEPDFDIGKQEKIHVTRNTGDESEMEQVLCSLPETRKAVSSAALWRRQIYAVATLRFLKLRRERR ALLCLLLVLGIAFIPIILEKIMYKVTRETHCWEFSPSMYFLSLEQIPKTPLTSLLIVNNTGSNIEDLVHSLKCQDIVLEIDDFRNRNGSD DPSYNGAIIVSGDQKDYRFSVACNTKKLNCFPVLMGIVSNALMGIFNFTELIQMESTSFSRDDIVLDLGFIDGSIFLLLITNCVSPFIGM SSISDYKKNVQSQLWISGLWPSAYWCGQALVDIPLYFLILFSIHLIYYFIFLGFQLSWELMFVLVVCIIGCAVSLIFLTYVLSFIFRKWR KNNGFWSFGFFIILICVSTIMVSTQYEKLNLILCMIFIPSFTLLGYVMLLIQLDFMRNLDSLDNRINEVNKTILLTTLIPYLQSVIFLFV IRCLEMKYGNEIMNKDPVFRISPRSRETHPNPEEPEEEDEDVQAERVQAANALTAPNLEEEPVITASCLHKEYYETKKSCFSTRKKKIAI RNVSFCVKKGEVLGLLGHNGAGKSTSIKMITGCTKPTAGVVVLQGSRASVRQQHDNSLKFLGYCPQENSLWPKLTMKEHLELYAAVKGLG KEDAALSISRLVEALKLQEQLKAPVKTLSEGIKRKLCFVLSILGNPSVVLLDEPFTGMDPEGQQQMWQILQATVKNKERGTLLTTHYMSE AEAVCDRMAMMVSGTLRCIGSIQHLKNKFGRDYLLEIKMKEPTQVEALHTEILKLFPQAAWQERYSSLMAYKLPVEDVHPLSRAFFKLEA -------------------------------------------------------------- >94265_94265_4_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000544981_ABCA10_chr17_67193263_ENST00000416101_length(transcript)=4288nt_BP=187nt AGACTGCCATAAGATGGCGTCCGTGGCGGCTGCACGAGCAGTTCCCGTGGGCAGTGGGCTCAGGGGCCTGCAACGGACCCTACCTCTTGT AGTGATTCTCGGGGCCACGGGCACCGGCAAATCCACGCTGGCGTTGCAGCTAGGCCAGCGGCTCGGCGGTGAGATCGTCAGCGCTGACTC CATGCAGAATCAGAAATGTTATAAAAGAATATAATGGAAAGACTGGAAAAGTAGAAGCATTGCAAGGCATATTTTTTGACATATATGAAG GACAGATCACTGCAATACTTGGGCATAATGGAGCTGGTAAATCAACACTGCTAAACATTCTTAGTGGATTGTCTGTTTCTACAGAAGGTA AAAAGAATTATAATGGAATTAGACATGCAAAGCATTCAAGACATTATTGCTAAAAAATTAAGTGGTGGGCAGAAGAGAAAACTAACACTA GGGATTGCCATCTTAGGAGATCCTCAGGTTTTGCTGCTAGATGAACCAACTGCTGGATTGGATCCCTTTTCAAGACACCGAGTGTGGAGC CTCCTGAAGGAGCATAAAGTAGACCGACTTATCCTCTTCAGTACCCAATTCATGGATGAGGCTGACATCTTGGCTGATAGGAAAGTATTT CTGTCTAATGGGAAGTTGAAATGTGCAGGATCATCTTTGTTTCTGAAGCGAAAGTGGGGTATTGGATATCATTTAAGTTTACACAGGAAT GAAATGTGTGACACAGAAAAAATCACATCCCTTATTAAGCAGCACATTCCTGATGCCAAGTTAACAACAGAAAGTGAAGAAAAACTTGTA TATAGTTTGCCTTTGGAAAAAACGAACAAATTTCCAGATCTTTACAGTGACCTTGATAAGTGTTCTGACCAGGGCATAAGGAATTATGCT GTTTCAGTGACATCTCTGAATGAAGTATTCTTGAACCTAGAAGGAAAATCAGCAATTGATGAACCAGATTTTGACATTGGGAAACAAGAG AAAATACATGTGACAAGAAATACTGGAGATGAGTCTGAAATGGAACAGGTTCTTTGTTCTCTTCCTGAAACAAGAAAGGCTGTCAGTAGT GCAGCTCTCTGGAGACGACAAATCTATGCAGTGGCAACACTTCGCTTCTTAAAGTTAAGGCGTGAAAGGAGAGCTCTTTTGTGTTTGTTA CTAGTACTTGGAATTGCTTTTATCCCCATCATTCTAGAGAAGATAATGTATAAAGTAACTCGTGAAACTCATTGTTGGGAGTTTTCACCC AGTATGTATTTCCTTTCTCTGGAACAAATCCCGAAGACGCCTCTTACCAGCCTGTTAATCGTTAATAATACAGGATCAAATATTGAAGAC CTCGTGCATTCACTGAAGTGTCAGGATATAGTTTTGGAAATAGATGACTTTAGAAACAGAAATGGCTCAGATGATCCCTCCTACAATGGA GCCATCATAGTGTCTGGTGACCAGAAGGATTACAGATTTTCTGTTGCGTGTAATACCAAGAAATTGAATTGTTTTCCTGTTCTTATGGGA ATTGTTAGCAATGCCCTTATGGGAATTTTTAACTTCACGGAGCTTATTCAAATGGAGAGCACTTCATTTTCTCGTGATGACATAGTGCTG GATCTTGGTTTTATAGATGGGTCCATATTTTTGTTGTTGATCACAAACTGCGTTTCTCCTTTTATCGGCATGAGCAGCATCAGCGATTAT AAAAAAAATGTTCAATCCCAGTTATGGATTTCAGGCCTCTGGCCTTCAGCATACTGGTGTGGACAGGCTCTGGTGGACATTCCATTATAC TTCTTGATTCTCTTTTCAATACATTTAATTTACTACTTCATATTTCTGGGATTCCAGCTTTCATGGGAACTCATGTTTGTTTTGGTGGTA TGCATAATTGGTTGTGCAGTTTCTCTTATATTCCTCACATATGTGCTTTCATTCATCTTTCGCAAGTGGAGAAAAAATAATGGCTTTTGG TCTTTTGGCTTTTTTATTATCTTAATATGTGTATCCACAATTATGGTATCAACTCAATATGAAAAACTCAACTTAATTTTGTGCATGATT TTCATACCTTCCTTCACTTTGCTGGGGTATGTCATGTTATTGATCCAGCTCGACTTTATGAGAAACTTGGACAGTCTGGACAATAGAATA AATGAAGTCAATAAAACCATTCTTTTAACAACCTTAATACCATACCTTCAGAGTGTTATTTTCCTTTTTGTCATAAGGTGTCTGGAAATG AAGTATGGAAATGAAATAATGAATAAAGACCCAGTTTTCAGAATCTCTCCACGGAGTAGAGAAACTCATCCCAATCCGGAAGAGCCCGAA GAAGAAGATGAAGATGTTCAAGCTGAAAGAGTCCAAGCAGCAAATGCACTCACTGCTCCAAACTTGGAGGAGGAACCAGTCATAACTGCA AGCTGTTTACACAAGGAATATTATGAGACAAAGAAAAGTTGCTTTTCAACAAGAAAGAAGAAAATAGCCATCAGAAATGTTTCCTTTTGT GTTAAAAAAGGTGAAGTTTTGGGATTACTAGGACACAATGGAGCTGGTAAAAGTACTTCCATTAAAATGATAACTGGGTGCACAAAGCCA ACTGCAGGAGTGGTGGTGTTACAAGGCAGCAGAGCATCAGTAAGGCAACAGCATGACAACAGCCTCAAGTTCTTGGGGTACTGCCCTCAG GAGAACTCACTGTGGCCCAAGCTTACAATGAAAGAGCACTTGGAGTTGTATGCAGCTGTGAAAGGACTGGGCAAAGAAGATGCTGCTCTC AGTATTTCACGATTGGTGGAAGCTCTTAAGCTCCAGGAACAACTTAAGGCTCCTGTGAAAACTCTATCAGAGGGAATAAAGAGAAAGCTG TGCTTTGTGCTGAGCATCCTGGGGAACCCATCAGTGGTGCTTCTAGATGAGCCGTTCACCGGGATGGACCCCGAGGGGCAGCAGCAAATG TGGCAGATACTTCAGGCTACCGTTAAAAACAAGGAGAGGGGCACCCTCTTGACCACCCATTACATGTCAGAGGCTGAGGCTGTGTGTGAC CGTATGGCCATGATGGTGTCAGGAACGCTAAGGTGTATTGGTTCCATTCAACATCTGAAAAACAAGTTTGGTAGAGATTATTTACTAGAA ATAAAAATGAAAGAACCTACCCAGGTGGAAGCTCTCCACACAGAGATTTTGAAGCTTTTCCCACAGGCTGCTTGGCAGGAAAGATATTCC TCTTTAATGGCGTATAAGTTACCTGTGGAGGATGTCCACCCTCTATCTCGGGCCTTTTTCAAGTTAGAGGCGATGAAACAGACCTTCAAC CTGGAGGAATACAGCCTCTCTCAGGCTACCTTGGAGCAGGTATTCTTAGAACTCTGTAAAGAGCAGGAGCTGGGAAATGTTGATGATAAA ATTGATACAACAGTTGAATGGAAACTTCTCCCACAGGAAGACCCTTAAAATGAAGAACCTCCTAACATTCAATTTTAGGTCCTACTACAT TGTTAGTTTCCATAATTCTACAAGAATGTTTCCTTTTACTTCAGTTAACAAAAGAAAACATTTAATAAACATTCAATAATGATTACAGTT TTCATTTTTAAAAATTTAGGATGAAGGAAACAAGGAAATATAGGGAAAAGTAGTAGACAAAATTAACAAAATCAGACATGTTATTCATCC CCAACATGGGTCTATTTTGTGCTTAAAAATAATTTAAAAATCATACAATATTAGGTTGGTTATCGGTTATTATCAATAAAGCTAACACTG AGAACATTTTACAAATAAAAATATGAGGTTTTTAGCCTGAACTTCAAATGTATCAGCTATTTTTAAACATTATTTACTCGGATTCTAATT TAATGTGACATTGACTATAAGAAGGTCTGATAAACTGATGAAATGGCACAGCATAACATTTAATTATAATGACATTCTGATTATAAAATA AATGCATGTGAATTTTAGTACATATTGAAGTTATATGGAAGAAGATAGCCATAATCTGTAAGAAAGTACCGCAGTTAATATTTTCTTTAG CCAACTTATATTCAATGTATTTTTTATGGATCCTTTTTCAAAGGTAGTATCAGTAGGCATAGTCATTTTCTGTATCTTTTCACCTCACAG TTCATGAACATTTCCCATGTCATTATAGCACTTTTGTGTTATATAATTGCTGCATTAATGTTATAGCTGTATTGATGTAATACCAGAGTT >94265_94265_4_TRIT1-ABCA10_TRIT1_chr1_40348990_ENST00000544981_ABCA10_chr17_67193263_ENST00000416101_length(amino acids)=1041AA_BP=729 MFLQKVKRIIMELDMQSIQDIIAKKLSGGQKRKLTLGIAILGDPQVLLLDEPTAGLDPFSRHRVWSLLKEHKVDRLILFSTQFMDEADIL ADRKVFLSNGKLKCAGSSLFLKRKWGIGYHLSLHRNEMCDTEKITSLIKQHIPDAKLTTESEEKLVYSLPLEKTNKFPDLYSDLDKCSDQ GIRNYAVSVTSLNEVFLNLEGKSAIDEPDFDIGKQEKIHVTRNTGDESEMEQVLCSLPETRKAVSSAALWRRQIYAVATLRFLKLRRERR ALLCLLLVLGIAFIPIILEKIMYKVTRETHCWEFSPSMYFLSLEQIPKTPLTSLLIVNNTGSNIEDLVHSLKCQDIVLEIDDFRNRNGSD DPSYNGAIIVSGDQKDYRFSVACNTKKLNCFPVLMGIVSNALMGIFNFTELIQMESTSFSRDDIVLDLGFIDGSIFLLLITNCVSPFIGM SSISDYKKNVQSQLWISGLWPSAYWCGQALVDIPLYFLILFSIHLIYYFIFLGFQLSWELMFVLVVCIIGCAVSLIFLTYVLSFIFRKWR KNNGFWSFGFFIILICVSTIMVSTQYEKLNLILCMIFIPSFTLLGYVMLLIQLDFMRNLDSLDNRINEVNKTILLTTLIPYLQSVIFLFV IRCLEMKYGNEIMNKDPVFRISPRSRETHPNPEEPEEEDEDVQAERVQAANALTAPNLEEEPVITASCLHKEYYETKKSCFSTRKKKIAI RNVSFCVKKGEVLGLLGHNGAGKSTSIKMITGCTKPTAGVVVLQGSRASVRQQHDNSLKFLGYCPQENSLWPKLTMKEHLELYAAVKGLG KEDAALSISRLVEALKLQEQLKAPVKTLSEGIKRKLCFVLSILGNPSVVLLDEPFTGMDPEGQQQMWQILQATVKNKERGTLLTTHYMSE AEAVCDRMAMMVSGTLRCIGSIQHLKNKFGRDYLLEIKMKEPTQVEALHTEILKLFPQAAWQERYSSLMAYKLPVEDVHPLSRAFFKLEA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for TRIT1-ABCA10 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 55_58 | 58.0 | 468.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 55_58 | 58.0 | 442.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 55_58 | 58.0 | 386.0 | substrate tRNA |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 233_255 | 58.0 | 468.0 | isopentenylpyrophosphate transferase |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 233_255 | 58.0 | 442.0 | isopentenylpyrophosphate transferase |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 233_255 | 58.0 | 386.0 | isopentenylpyrophosphate transferase |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 183_187 | 58.0 | 468.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 281_283 | 58.0 | 468.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 313_331 | 58.0 | 468.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000316891 | - | 1 | 11 | 323_330 | 58.0 | 468.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 183_187 | 58.0 | 442.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 281_283 | 58.0 | 442.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 313_331 | 58.0 | 442.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000372818 | - | 1 | 10 | 323_330 | 58.0 | 442.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 183_187 | 58.0 | 386.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 281_283 | 58.0 | 386.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 313_331 | 58.0 | 386.0 | substrate tRNA |
| Hgene | TRIT1 | chr1:40348990 | chr17:67193263 | ENST00000441669 | - | 1 | 9 | 323_330 | 58.0 | 386.0 | substrate tRNA |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for TRIT1-ABCA10 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for TRIT1-ABCA10 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
